Development of a Novel, Ultra-rapid Biosensor for the Qualitative Detection of Hepatitis B Virus-associated Antigens and Anti-HBV, Based on “Membrane-engineered” Fibroblast Cells with Virus-Specific Antibodies and Antigens
Abstract
:1. Introduction
2. Experimental Section
2.1. Materials
2.2. Sensor Fabrication from Vero Cells
2.3. Sample Preparation
2.4. Assay Procedure
2.4. Fluorescence Microscopy
2.5. Experimental Design
3. Results
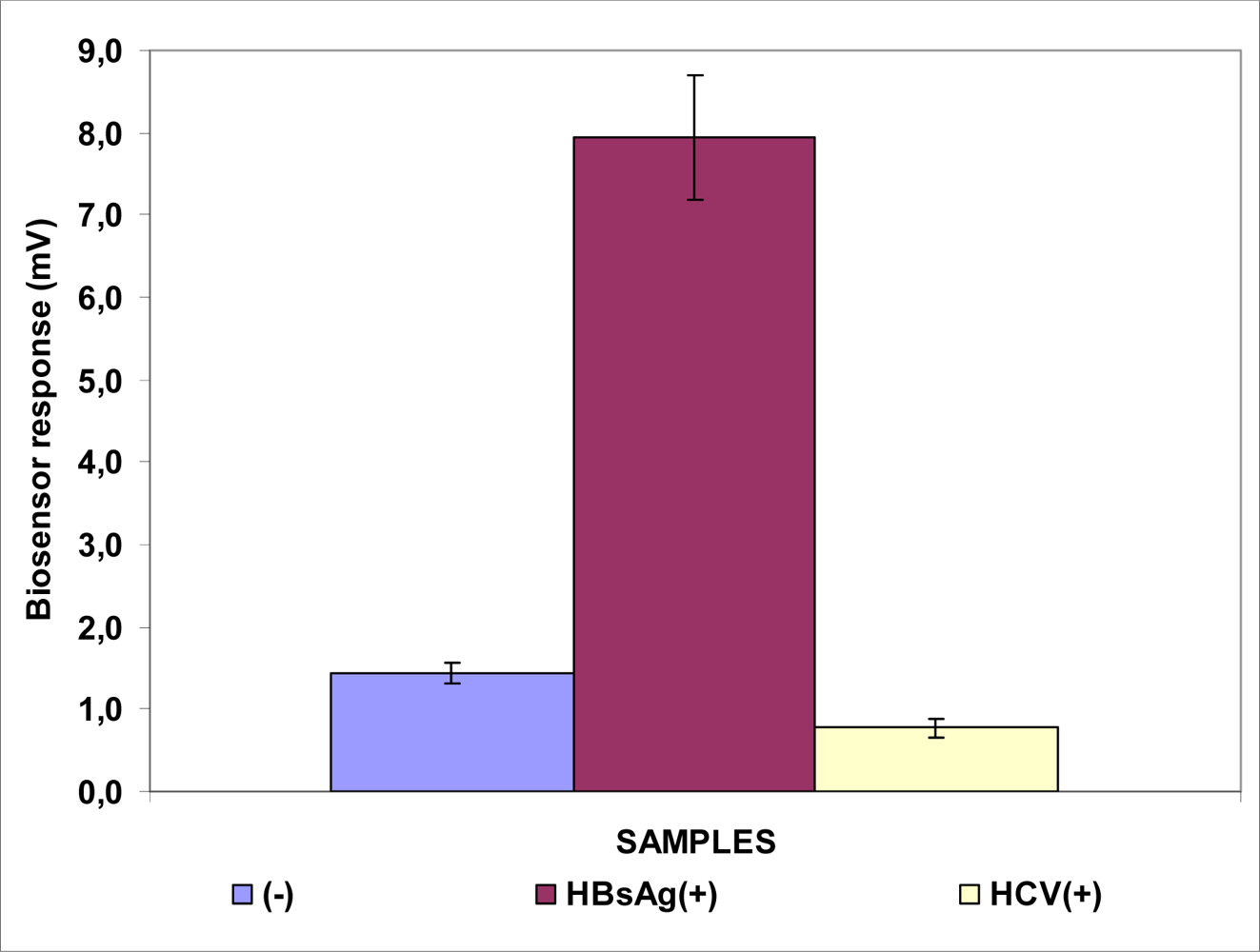
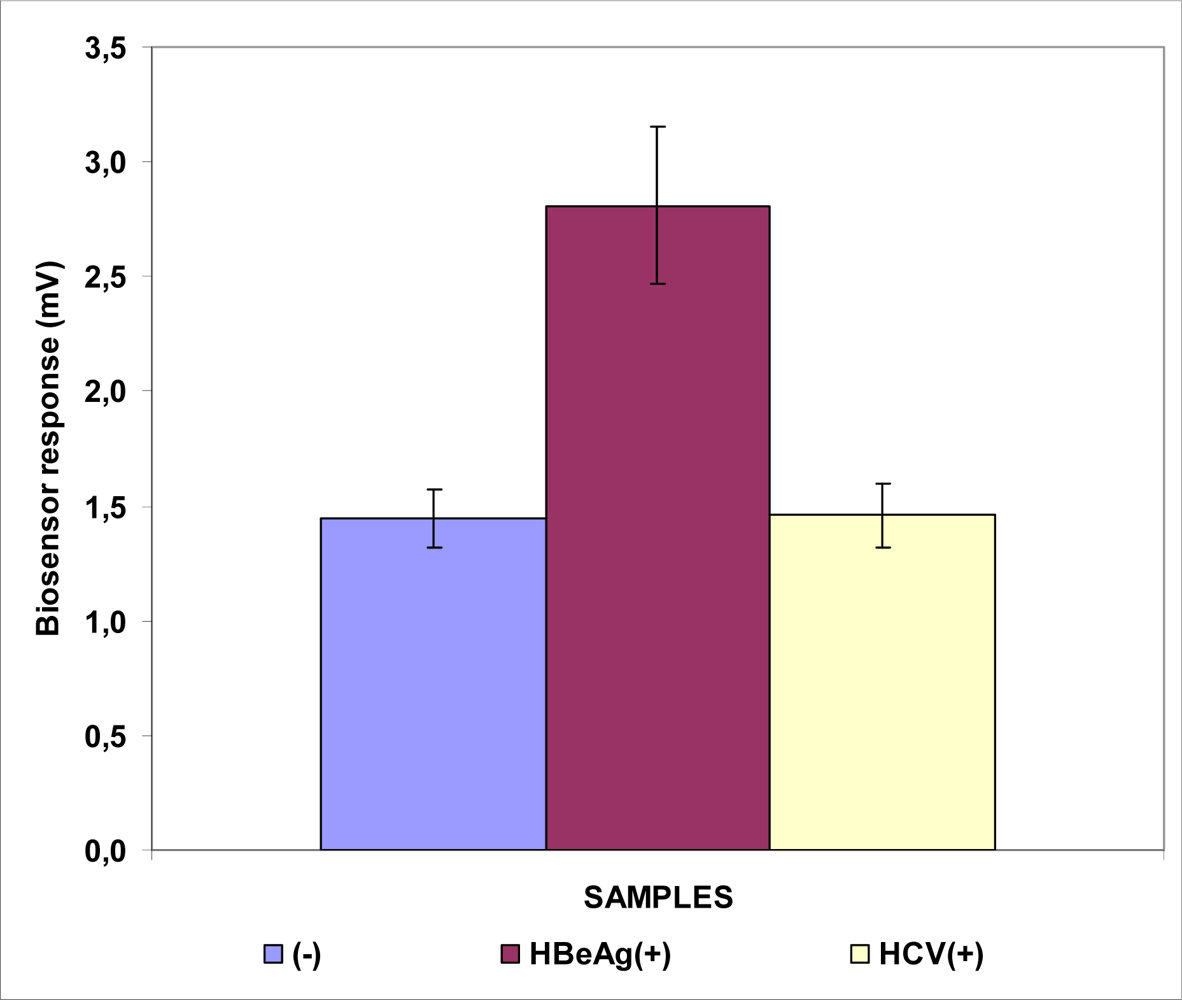
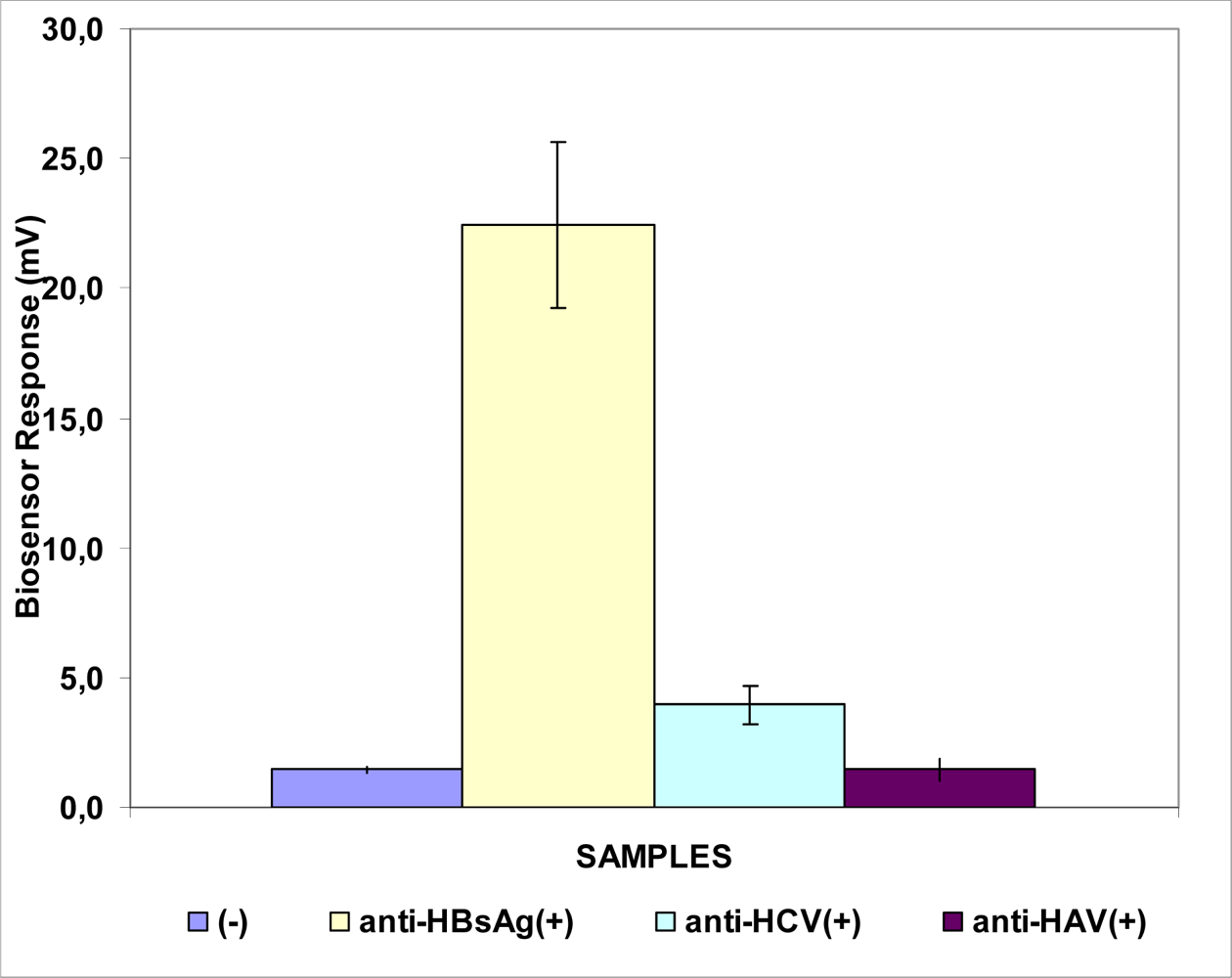
3.1. Responses of Biosensors Based on Cells Membrane-engineered with Anti-HBs, Anti-HBe or HBsAg
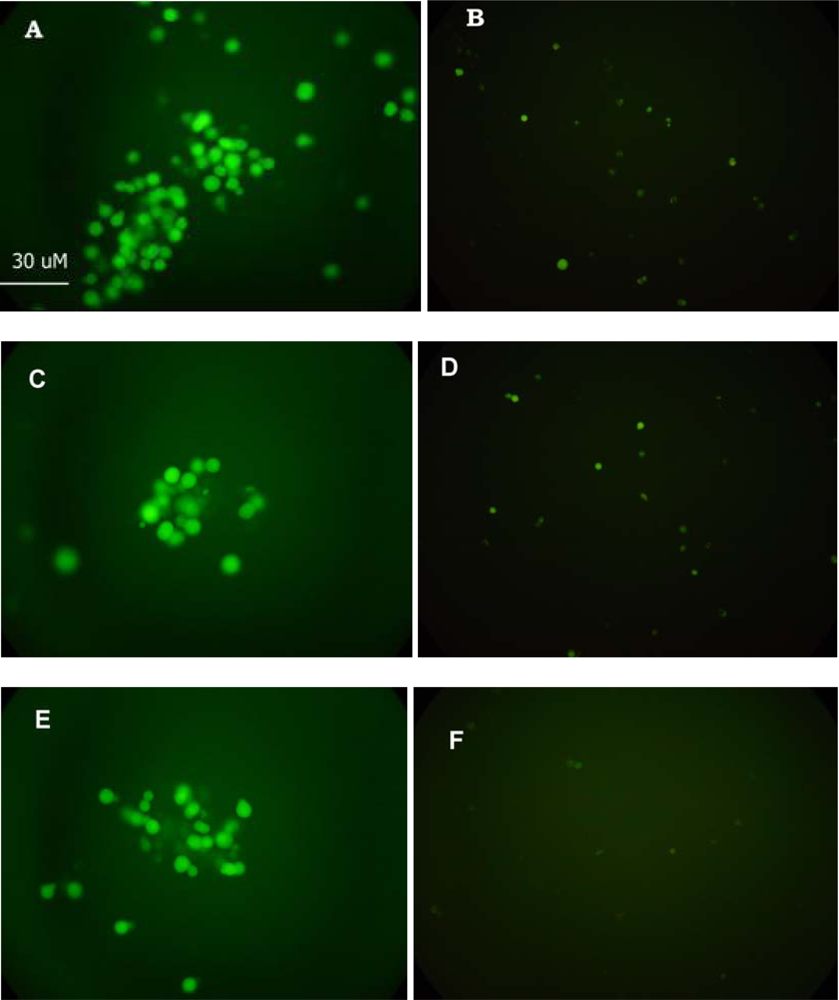
3.2. Fluorescence Microscopy Assays of Cell–virus Interactions
4. Discussion
Acknowledgments
References
- Zuckerman, A.J. Hepatitis Viruses. In Baron's Medical Microbiology, 4th Ed; Baron, S., Ed.; University of Texas Medical Branch: Galveston, TX, USA, 1996; pp. 2371–2381. [Google Scholar]
- Bonino, F.; Chiaberge, E.; Maran, E.; Piantino, P. Serological markers of HBV infectivity. Ann. Ist. Super. Sanita 1987, 24, 217–223. [Google Scholar]
- Kwon, J.A.; Yoon, S.Y.; Lee, C.K.; Lima, C.S.; Lee, K.N.; Sung, H.J.; Brennan, C.A.; Devare, S.G. Performance evaluation of three automated human immunodeficiency virus antigen–antibody combination immunoassays. J. Virol. Met 2006, 133, 20–26. [Google Scholar]
- Candotti, D.; Temple, J.; Owusu-Ofori, S.; Allain, J.P. Multiplex real-time quantitative RT-PCR assay for hepatitis B virus, hepatitis C virus, and human immunodeficiency virus type 1. J. Virol. Met 2004, 118, 39–47. [Google Scholar]
- Meric, B.; Kerman, K.; Ozkan, D.; Kara, P.; Erensoy, S.; Akarca, U.S.; Mascini, M.; Ozsoz, M. Electrochemical DNA biosensor for the detection of TT and Hepatitis B virus from PCR amplified real samples by using methylene blue. Talanta 2002, 56, 837–846. [Google Scholar]
- Ye, Y.K.; Zhao, J.H.; Yan, F.; Zhu, Y.L.; Ju, H.X. Electrochemical behaviour and detection of hepatitis B virus DNA PCR production at gold electrode. Biosens. Bioelectron 2003, 18, 1501–1508. [Google Scholar]
- Erdem, A.; Ariksoysal, D.O.; Karadeniz, H.; Kara, P.; Sengonul, A.; Sayiner, A.A.; Ozsoz, M. Electrochemical genomagnetic assay for the detection of the hepatitis B virus DNA in polymerase chain reaction amplicons by using disposable sensor technology. Electrochem. Commun 2005, 7, 815–820. [Google Scholar]
- Mandog, G.; Yanqing, L.; Hongxia, G.; Xiaoqin, W.; Lifang, F. Electrochemical detection of short sequences related to the hepatitis B virus using MB on chitosan-modified CPE. Bioelectrochem 2007, 70, 245–249. [Google Scholar]
- Zhou, X.; Liu, L.; Hu, M.; Wang, L.; Hu, J. Detection of hepatitis B virus by piezoelectric biosensor. J. Pharm. Biomed. Anal 2002, 27, 341–345. [Google Scholar]
- Li, X-M.; Ju, H.Q.; Ding, C.F.; Zhang, S.S. Nucleic acid biosensor for detection of hepatitis B virus using 2,9-dimethyl-1,10-phenanthroline copper complex as electrochemical indicator. Anal. Chim. Acta 2007, 582, 158–163. [Google Scholar]
- Ly, S.Y.; Cho, N.S. Diagnosis of human hepatitis B virus in non-treated blood by the bovine IgC DNA-linked carbon nanotube biosensor. J. Clin. Virol 2009, 44, 43–47. [Google Scholar]
- Yao, C.; Zhu, T.; Tang, J.; Wu, R.; Chen, Q.; Chen, M.; Zhang, B.; Huang, J.; Fu, W. Hybridization assay of hepatiis B virus by QCM peptide nucleic acid biosensor. Biosens. Bioelectron 2008, 23, 879–885. [Google Scholar]
- Hassen, W.M.; Chaix, C.; Abdelghani, A.; Bessueille, F.; Leonard, D.; Jaffrezic-Renault, N. An impedemetric DNA sensor based on functionalized magnetic nanoparticles for HIV and HBV detection. Sens. Actuat. B Chem 2008, 134, 755–760. [Google Scholar]
- Chung, J.W.; Kim, S.D.; Bernhardt, R.; Pyun, J.C. Application of SPR biosensor for medical diagnostics of human hepatitis B virus (hHBV). Sens. Actuat. B Chem 2005, 111–112, 416–422. [Google Scholar]
- Zhang, Y.; Zhang, Z.; Yang, F. A sensitive immunoassay for determination of hepatitis B surface antigen and antibody in human serum using capillary electrophoresis with chemiluminescence detection. J. Chromatogr 2007, 857, 100–107. [Google Scholar]
- Kintzios, S. Cell-based sensors in clinical chemistry. Mini Rev. Clin. Chem 2007, 7, 1019–1026. [Google Scholar]
- Kintzios, S.; Pistola, E.; Panagiotopoulos, P.; Bomsel, M.; Alexandropoulos, N.; Bem, F.; Varveri, C.; Ekonomou, G.; Biselis, J.; Levin, R. Bioelectric Recognition Assay (BERA). Biosens. Bioelectron 2001, 16, 325–336. [Google Scholar]
- Moschopoulou, G.; Vitsa, K.; Bem, F.; Vassilakos, N.; Perdikaris, A.; Blouhos, P.; Yialouris, C.; Frosyniotis, D.; Anthopoulos, I.; Mangana, O.; Nomikou, K.; Rodeva, V.; Kostova, D.; Grozeva, S.; Michaelides, A.; Simonian, A.; Kintzios, S. Engineering of the membrane of fibroblast cells with virus-specific antibodies: A novel biosensor tool for virus detection. Biosens. Bioelectron 2008, 24, 1027–1030. [Google Scholar]
- Mavrikou, S.; Flampouri, K.; Moschopoulou, G.; Mangana, O.; Michaelides, A.; Kintzios, S. Assessment of organophosphate and carbamate pesticide residues in cigarette tobacco with a novel cell biosensor. Sensors 2008, 8, 2818–2832. [Google Scholar]
- Whelan, R.J.; Zare, R.N. Single-cell immunosensors for protein detection. Biosens. Bioelectron 2008, 18, 1065–1072. [Google Scholar]
- Moschopoulou, G.; Kintzios, S. Application of “membrane-engineering” to bioelectric recognition cell sensors for the detection of picomole concentrations of superoxide radical: a novel biosensor principle. Anal. Chim. Acta 2006, 573–574, 90–96. [Google Scholar]
- Mowat, W.P.; Dawson, S. Detection and identification of plant viruses by ELISA using crude sap extracts and unfractionated antisera. J. Virol. Meth 1987, 15, 233–247. [Google Scholar]
- Guo, M.; Yanqing, L.; Hongxia, G.; Xiaoqin, W.; Lifang, F. Electrochemical detection of short sequences related to the hepatitis B virus using MB on chitosan-modified CPE. Bioelectrochem 2007, 70, 245–249. [Google Scholar]
- Taylor, P.; Pickard, G.; Gammie, A.; Atkins, M. Comparison of the ADVIA centaur® and Abbott AxSYM immunoassay systems for a routine diagnostic virology laboratory. J. Clin. Virol 2004, 30, S11–S15. [Google Scholar]
- Katsanakis, N.; Katsivelis, A.; Kintzios, S. Immobilization of electroporated cells: physiological effects of the shape of calcium alginate matrices and foetal calf serum. Sensors 2009, 9, 378–385. [Google Scholar]
| Sample type | Number of samples |
|---|---|
| HBsAg(+), anti-HBsAg(+) | 35 |
| HBeAg(+) | 25 |
| HCV(+), anti-HCV(+) | 32 |
| anti-HAV IgM(+) | 6 |
| Total positive | 98 |
| Negative | 35 |
| Total | 133 |
© 2009 by the authors; licensee MDPI, Basel, Switzerland This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Perdikaris, A.; Alexandropoulos, N.; Kintzios, S. Development of a Novel, Ultra-rapid Biosensor for the Qualitative Detection of Hepatitis B Virus-associated Antigens and Anti-HBV, Based on “Membrane-engineered” Fibroblast Cells with Virus-Specific Antibodies and Antigens. Sensors 2009, 9, 2176-2186. https://doi.org/10.3390/s90302176
Perdikaris A, Alexandropoulos N, Kintzios S. Development of a Novel, Ultra-rapid Biosensor for the Qualitative Detection of Hepatitis B Virus-associated Antigens and Anti-HBV, Based on “Membrane-engineered” Fibroblast Cells with Virus-Specific Antibodies and Antigens. Sensors. 2009; 9(3):2176-2186. https://doi.org/10.3390/s90302176
Chicago/Turabian StylePerdikaris, Antonios, Nikos Alexandropoulos, and Spiridon Kintzios. 2009. "Development of a Novel, Ultra-rapid Biosensor for the Qualitative Detection of Hepatitis B Virus-associated Antigens and Anti-HBV, Based on “Membrane-engineered” Fibroblast Cells with Virus-Specific Antibodies and Antigens" Sensors 9, no. 3: 2176-2186. https://doi.org/10.3390/s90302176






