Highly Sensitive Automated Method for DNA Damage Assessment: Gamma-H2AX Foci Counting and Cell Cycle Sorting
Abstract
:1. Introduction
2. Results and Discussion
2.1. Improved Automated Scoring Classifies Cells According to the Cell Cycle Phase
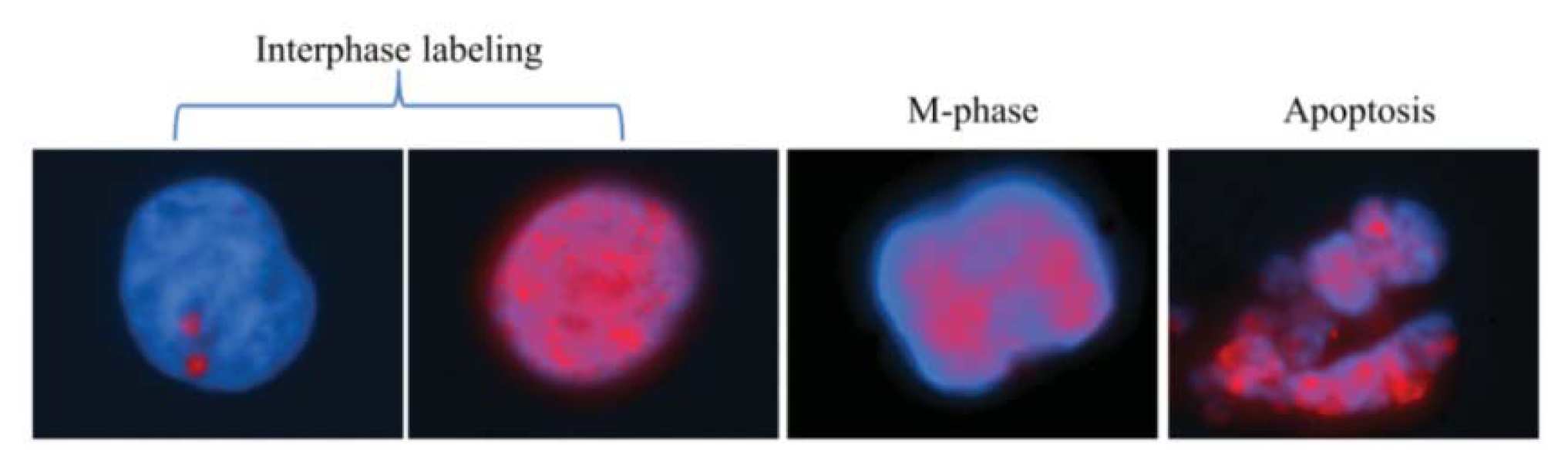
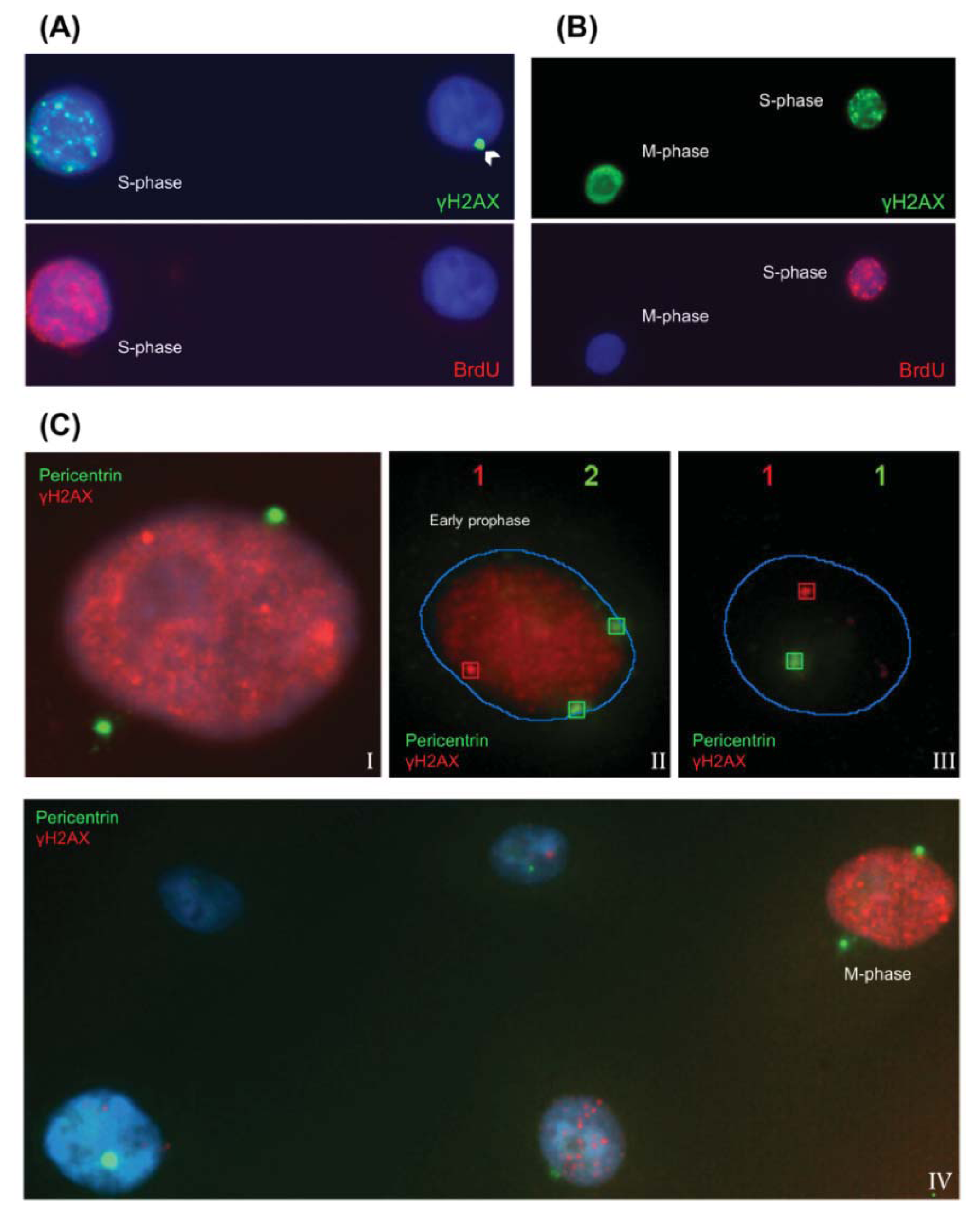
2.1.1. Characteristic γH2AX Staining Patterns of S- and M-Phase Cells
2.1.2. CENP-F to Identify G2 Cells and Define the Nuclear Area Range of Cells in G2
2.1.3. Pericentrin: The Co-Staining Marker of Choice
2.2. γH2AX Automatic Foci Counting
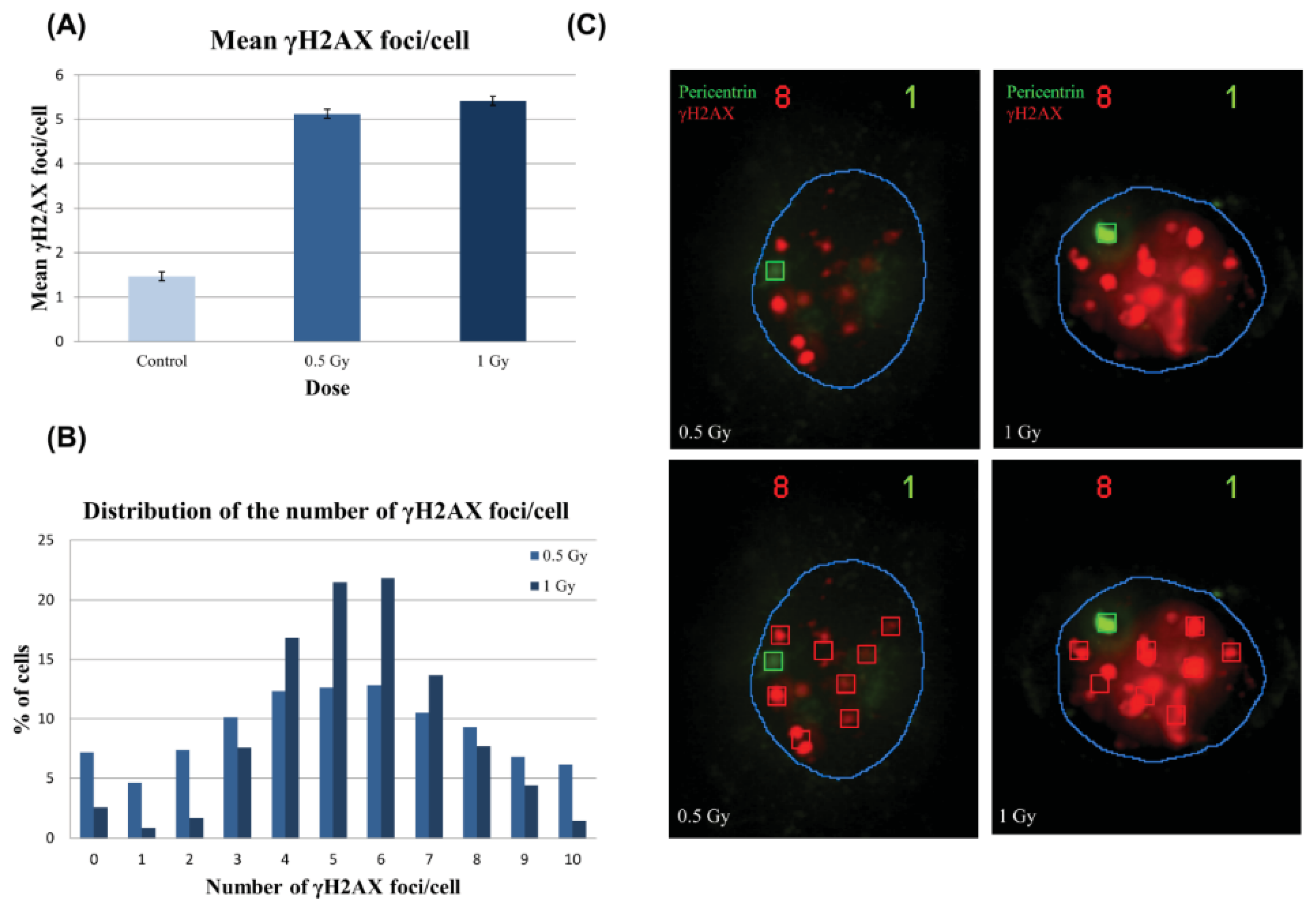
2.2.1. Foci Analysis
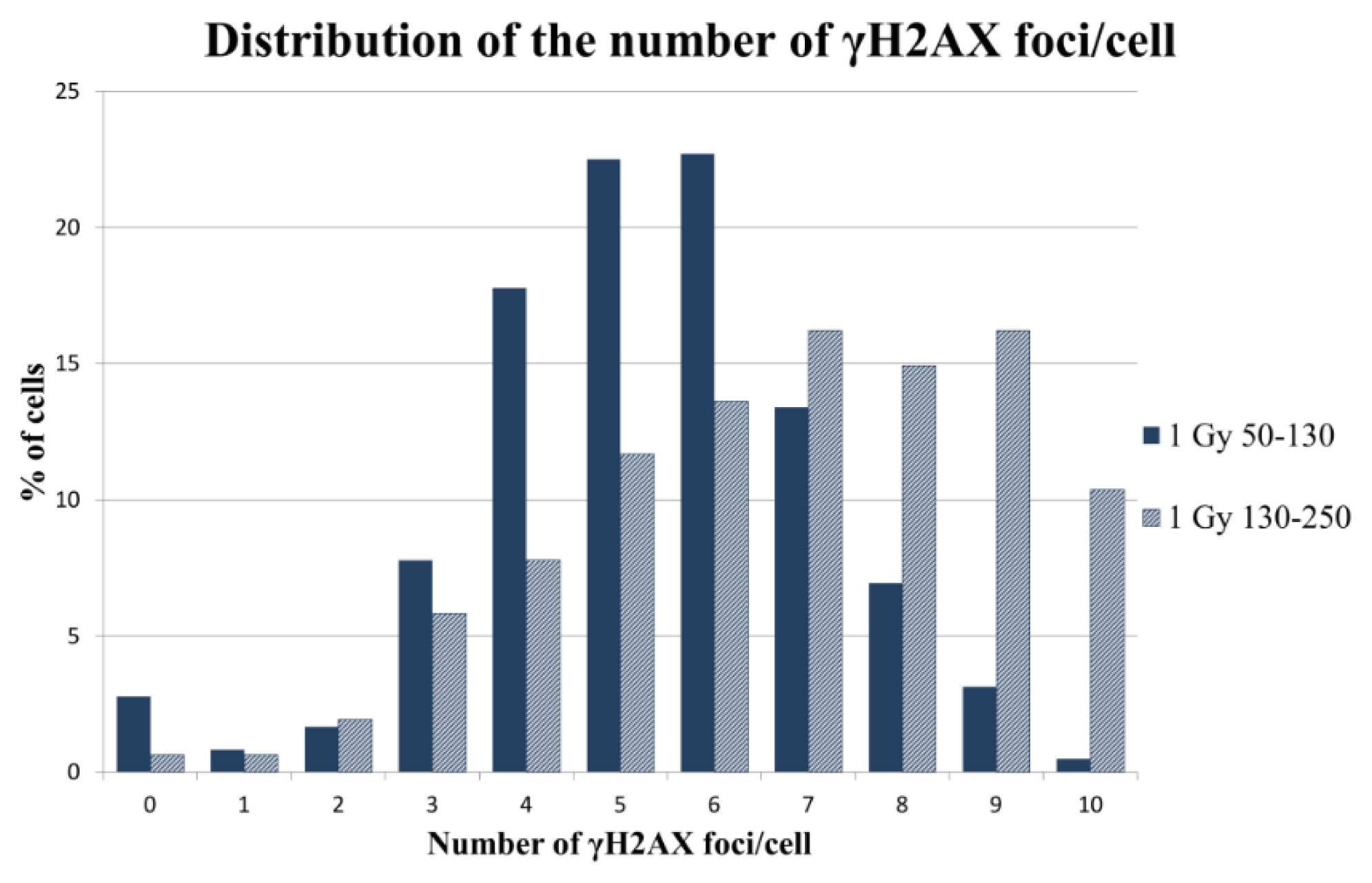
2.2.2. γH2AX Signal Saturation at High Radiation Doses in Non-Sorted Cells
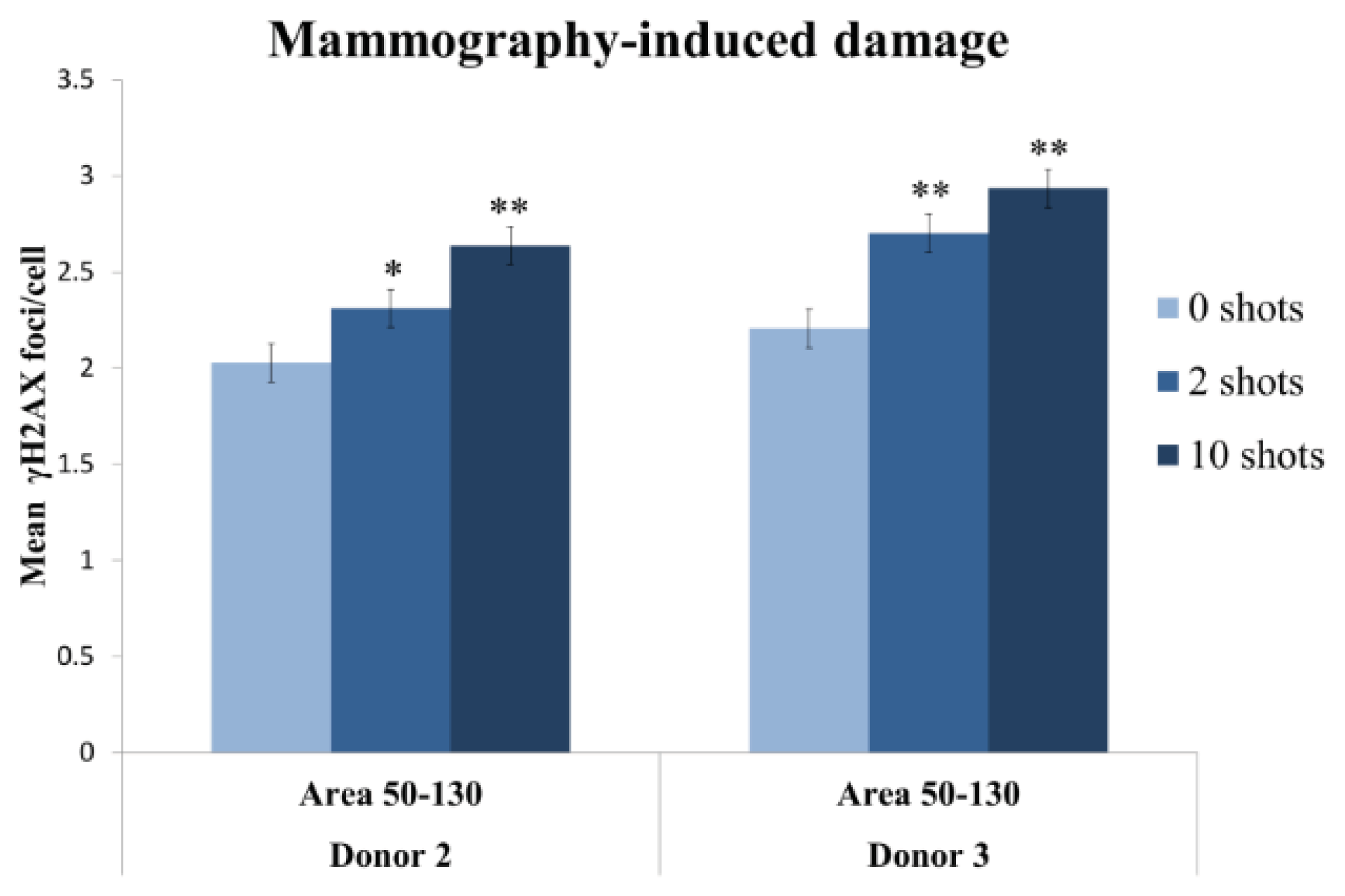
2.2.3. Two Mammography Shots Induce a Significant and Detectable Increase in the Number of Foci
2.2.4. Manual vs. Automatic γH2AX Foci Scoring: A Reliable Automated Approach
3. Discussion
4. Experimental Section
4.1. Cell Cultures
4.2. Irradiation
4.3. Immunostaining
4.4. γH2AX Immunostaining Combined with a BrdU Assay
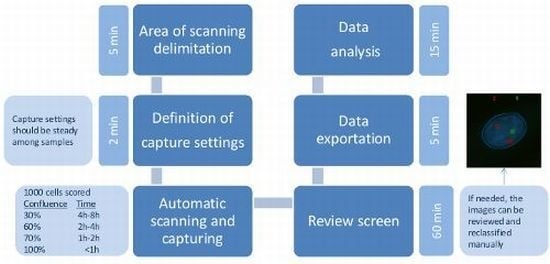
4.5. Automated Microscopy and Analysis
5. Conclusions
Acknowledgments
Conflict of Interest
References
- Celeste, A.; Difilippantonio, S.; Difilippantonio, M.J.; Fernandez-Capetillo, O.; Pilch, D.R.; Sedelnikova, O.A.; Eckhaus, M.; Ried, T.; Bonner, W.M.; Nussenzweig, A. H2AX haploinsufficiency modifies genomic stability and tumor susceptibility. Cell 2003, 114, 371–383. [Google Scholar]
- Sedelnikova, O.A.; Pilch, D.R.; Redon, C.; Bonner, W.M. Histone H2AX in DNA damage and repair. Cancer Biol. Ther 2003, 2, 233–235. [Google Scholar]
- Rogakou, E.P.; Pilch, D.R.; Orr, A.H.; Ivanova, V.S.; Bonner, W.M. DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J. Biol. Chem 1998, 273, 5858–5868. [Google Scholar]
- Paull, T.T.; Rogakou, E.P.; Yamazaki, V.; Kirchgessner, C.U.; Gellert, M.; Bonner, W.M. A critical role for histone H2AX in recruitment of repair factors to nuclear foci after DNA damage. Curr. Biol 2000, 10, 886–895. [Google Scholar]
- MacPhail, S.H.; Banath, J.P.; Yu, T.Y.; Chu, E.H.M.; Lambur, H.; Olive, P.L. Expression of phosphorylated histone H2AX in cultured cell lines following exposure to X-rays. Int. J. Radiat. Biol 2003, 79, 351–358. [Google Scholar]
- Banáth, J.P.; MacPhail, S.H.; Olive, P.L. Radiation sensitivity, H2AX phosphorylation, and kinetics of repair of DNA strand breaks in irradiated cervical cancer cell lines. Cancer Res 2004, 64, 7144–7149. [Google Scholar]
- Klokov, D.; MacPhail, S.M.; Banáth, J.P.; Byrne, J.P.; Olive, P.L. Phosphorylated histone H2AX in relation to cell survival in tumor cells and xenografts exposed to single and fractionated doses of X-rays. Radiother Oncol 2006, 80, 223–229. [Google Scholar]
- Böcker, W.; Iliakis, G. Computational methods for analysis of foci: Validation for radiation-induced γ-H2AX foci in human cells. Radiat. Res 2006, 165, 113–124. [Google Scholar]
- Runge, R.; Hiemann, R.; Wendisch, M.; Kasten-Pisula, U.; Storch, K.; Zoephel, K.; Kotzerke, J. Fully automated interpretation of ionizing radiation-induced γH2AX foci by the novel pattern recognition system AKLIDES®. Int. J. Radiat. Biol 2012, 88, 439–447. [Google Scholar]
- Jucha, A.; Wegierek-Ciuk, A.; Koza, Z.; Lisowska, H.; Wojcik, A.; Wojewodzka, M.; Lankoff, A. FociCounter: A freely available PC programme for quantitative and qualitative analysis of gamma-H2AX foci. Mutat. Res 2010, 696, 16–20. [Google Scholar]
- González, J.E.; Lee, M.; Barquinero, J.F.; Valente, M.; Roch-Lefèvre, S.; García, O. Quantitative image analysis of gamma-H2AX foci induced by ionizing radiation applying open source programs. Anal. Quant. Cytol. Histol 2012, 34, 66–71. [Google Scholar]
- MacPhail, S.H.; Banáth, J.P.; Yu, Y.; Chu, E.; Olive, P.L. Cell cycle-dependent expression of phosphorylated histone H2AX: Reduced expression in unirradiated but not X-irradiated G1-phase cells. Radiat. Res 2003, 159, 759–767. [Google Scholar]
- Yoshida, K.; Yoshida, S.H.; Shimoda, C.; Morita, T. Expression and radiation-induced phosphorylation of histone H2AX in mammalian cells. J. Radiat. Res 2003, 44, 47–51. [Google Scholar]
- Löbrich, M.; Shibata, A.; Beucher, A.; Fisher, A.; Ensminger, M.; Goodarzi, A.A.; Jeggo, P.A. γH2AX foci analysis for monitoring DNA double-strand break repair: Strengths, limitations and optimization. Cell Cycle 2010, 9, 662–669. [Google Scholar]
- Huang, X.; Darzynkiewicz, Z. Cytometric assessment of histone H2AX phosphorylation: A reporter of DNA damage. Methods Mol. Biol 2006, 314, 73–80. [Google Scholar]
- Hernández, L.; Terradas, M.; Martín, M.; Feijoo, P.; Soler, D.; Tusell, L.; Genescà, A. Increased mammogram-induced DNA damage in mammary epithelial cells aged in vitro. PLoS One 2013, 8, e63052. [Google Scholar]
- Ward, I.M.; Chen, J. Histone H2AX is phosphorylated in an ATR-dependent manner in response to replicational stress. J. Biol. Chem 2001, 276, 47759–47762. [Google Scholar]
- Mühlhans, J.; Brandstätter, J.H.; Gießl, A. The centrosomal protein pericentrin identified at the basal body complex of the connecting cilium in mouse photoreceptors. PLoS One 2011, 6, e26496. [Google Scholar]
- Molina, Ò.; Sarrate, Z.; Vidal, F.; Blanco, J. FISH on sperm: Spot-counting to stop counting? Not yet. Fertil. Steril. 2009, 92, 1474–1480. [Google Scholar]





© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Hernández, L.; Terradas, M.; Martín, M.; Tusell, L.; Genescà, A. Highly Sensitive Automated Method for DNA Damage Assessment: Gamma-H2AX Foci Counting and Cell Cycle Sorting. Int. J. Mol. Sci. 2013, 14, 15810-15826. https://doi.org/10.3390/ijms140815810
Hernández L, Terradas M, Martín M, Tusell L, Genescà A. Highly Sensitive Automated Method for DNA Damage Assessment: Gamma-H2AX Foci Counting and Cell Cycle Sorting. International Journal of Molecular Sciences. 2013; 14(8):15810-15826. https://doi.org/10.3390/ijms140815810
Chicago/Turabian StyleHernández, Laia, Mariona Terradas, Marta Martín, Laura Tusell, and Anna Genescà. 2013. "Highly Sensitive Automated Method for DNA Damage Assessment: Gamma-H2AX Foci Counting and Cell Cycle Sorting" International Journal of Molecular Sciences 14, no. 8: 15810-15826. https://doi.org/10.3390/ijms140815810