Characterization of a Gene Encoding Clathrin Heavy Chain in Maize Up-Regulated by Salicylic Acid, Abscisic Acid and High Boron Supply
Abstract
:1. Introduction
2. Results
2.1. Cloning and Characterization of Genes ZmCHC1 and ZmCHC2 Encoding Clathrin Heavy Chain in Zea Mays
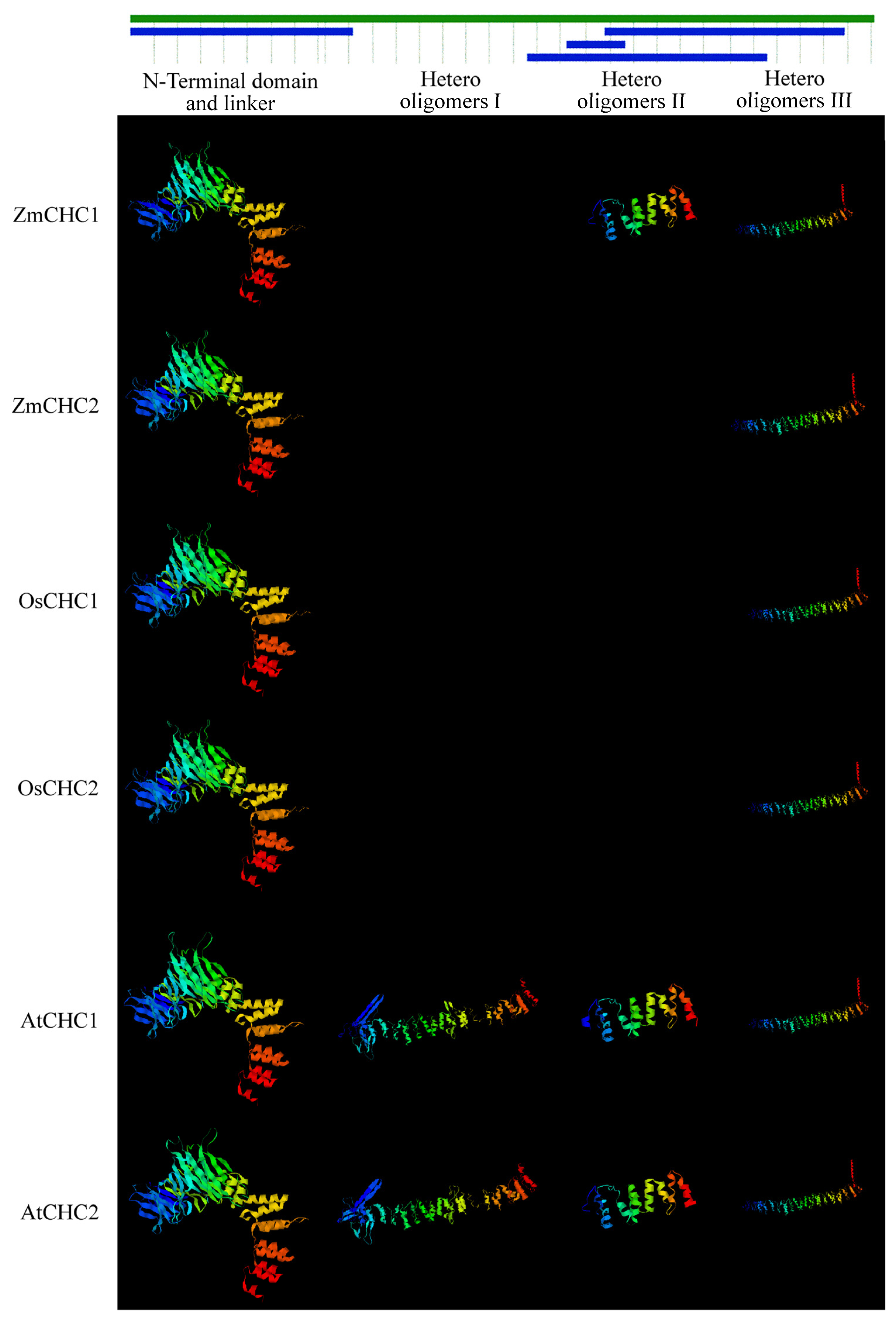
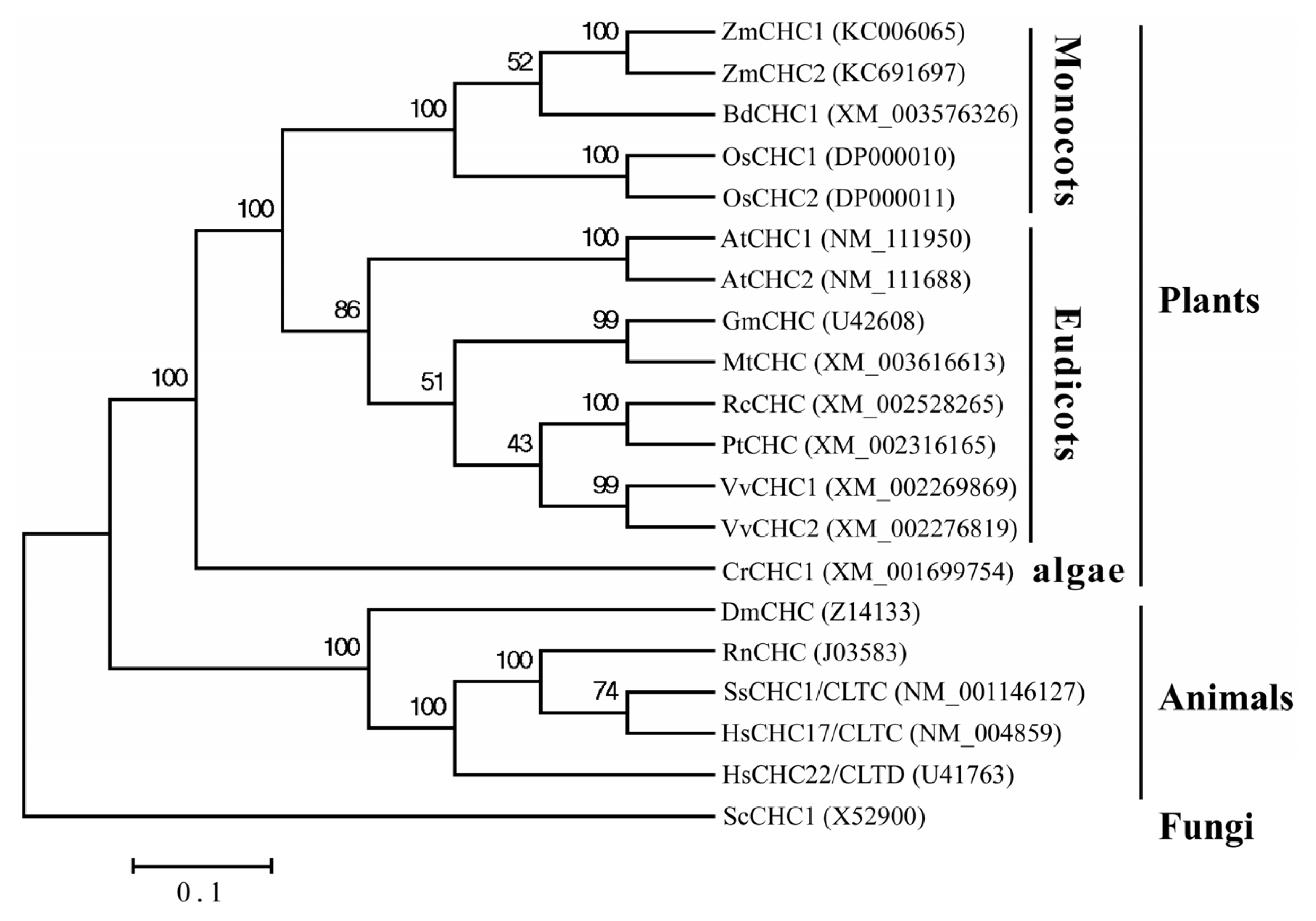
2.2. Sequence Analysis of ZmCHC1 and ZmCHC2
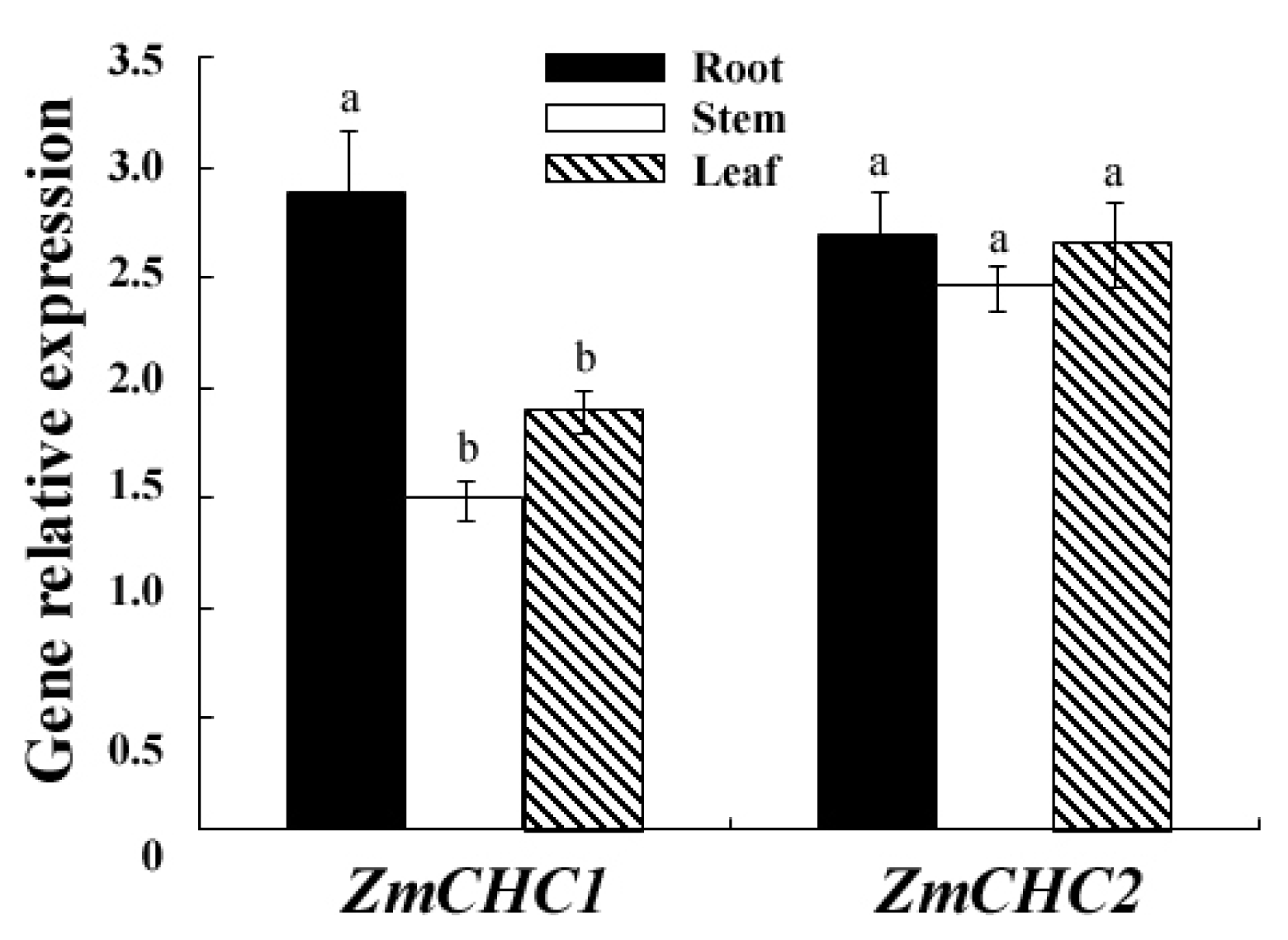
2.3. Organ Specific Expression Patterns of ZmCHC1 and ZmCHC2 in Maize Seedling
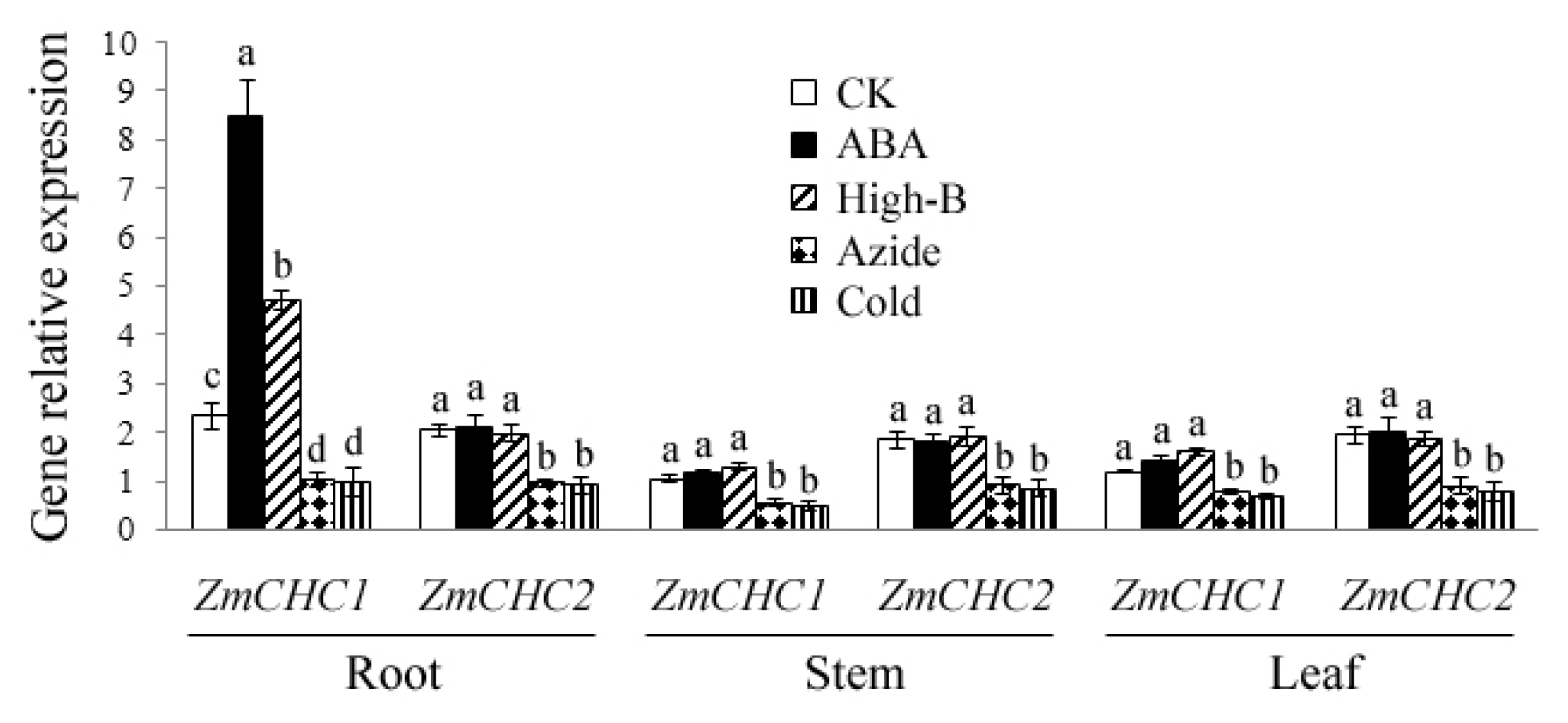
2.4. Effect of ABA, High Boron Supply, Sodium Azide or Low Temperature on Expressions of ZmCHC1 and ZmCHC2 in Maize Plants
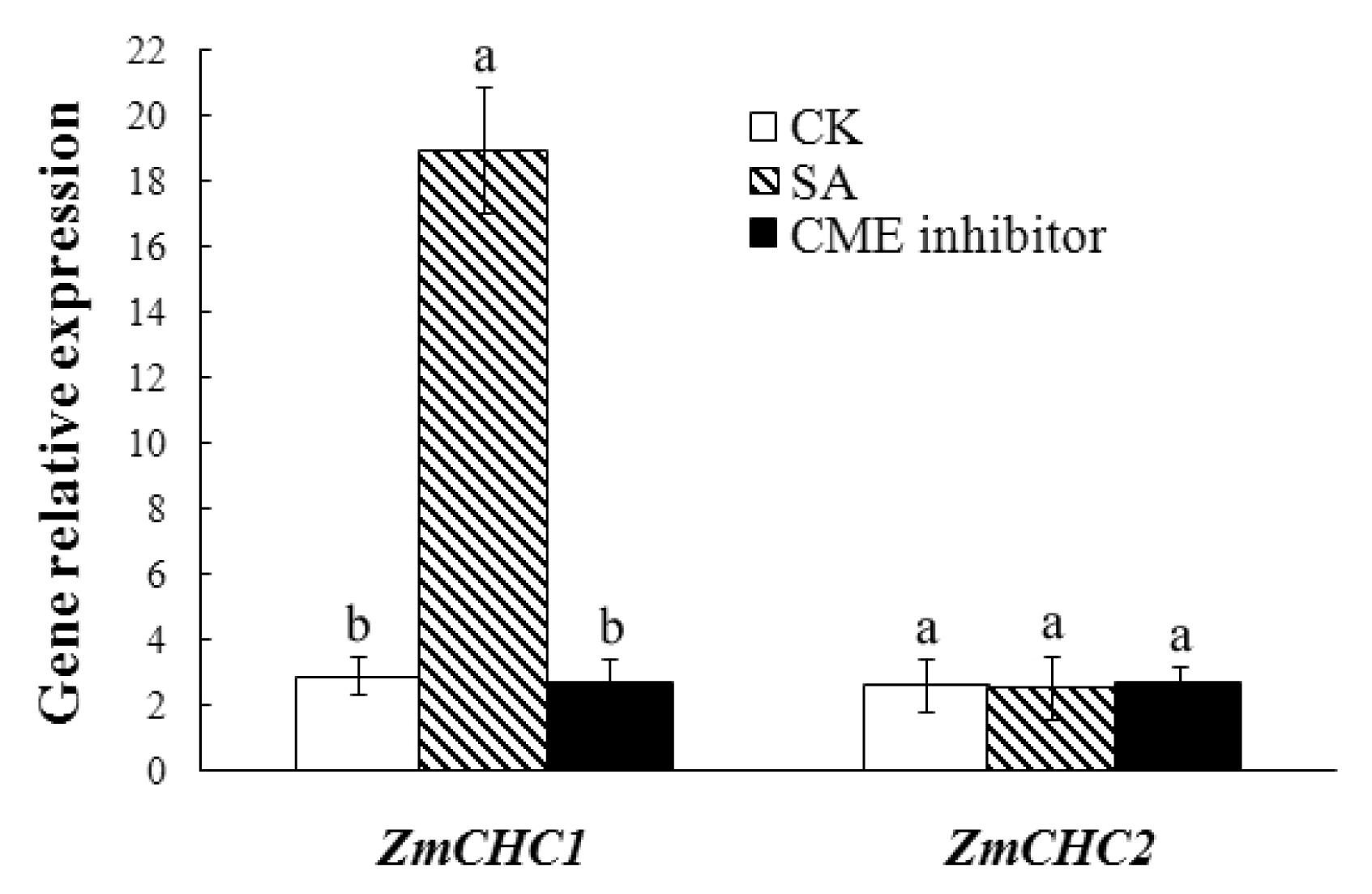
2.5. Salicylic Acid-Inducible Expression of ZmCHC1 Gene in Maize Roots
3. Discussion
4. Experimental Section
4.1. Maize Plants and Growth Conditions
4.2. Salicylic Acid, Abscisic Acid, High Boron, Sodium Azide, Tyrphostin A23, or Low Temperature Treatment of Maize Plants
4.3. Molecular Cloning of CHC Homolog from Maize
4.4. Sequence Analyses and Alignments
4.5. Real-Time Quantitative RT-PCR Performance
5. Conclusions
Acknowledgments
Conflict of Interest
References
- Conner, S.D.; Schmid, S.L. Regulated portals of entry into the cell. Nature 2003, 422, 37–44. [Google Scholar]
- Murphy, A.S.; Bandyopadhyay, A.; Holstein, S.E.; Peer, W.A. Endocytotic cycling of PM proteins. Annu. Rev. Plant Biol 2005, 56, 221–251. [Google Scholar]
- Wieffer, M.; Maritzen, T.; Haucke, V. SnapShot: Endocytic trafficking. Cell 2009, 137, e381–e383. [Google Scholar]
- McMahon, H.T.; Boucrot, E. Molecular mechanism and physiological functions of clathrin-mediated endocytosis. Nat. Rev. Mol. Cell Biol 2011, 12, 517–533. [Google Scholar]
- Chen, X.; Irani, N.G.; Friml, J. Clathrin-mediated endocytosis: The gateway into plant cells. Curr. Opin. Plant Biol 2011, 14, 674–682. [Google Scholar]
- Adam, T.; Bouhidel, K.; Der, C.; Robert, F.; Najid, A.; Simon-Plas, F.; Leborgne-Castel, N. Constitutive expression of clathrin hub hinders elicitor-induced clathrin-mediated endocytosis and defense gene expression in plant cells. FEBS Lett 2012, 586, 3293–3298. [Google Scholar]
- Pearse, B.M. Clathrin: A unique protein associated with intracellular transfer of membrane by coated vesicles. Proc. Natl. Acad. Sci. USA 1976, 73, 1255–1259. [Google Scholar]
- Kirchhausen, T. Imaging endocytic clathrin structures in living cells. Trends Cell Biol 2009, 19, 596–605. [Google Scholar]
- Dhonukshe, P.; Aniento, F.; Hwang, I.; Robinson, D.G.; Mravec, J.; Stierhof, Y.D.; Friml, J. Clathrin-mediated constitutive endocytosis of PIN auxin efflux carriers in Arabidopsis. Curr. Biol 2007, 17, 520–527. [Google Scholar]
- Robert, S.; Kleine-Vehn, J.; Barbez, E.; Sauer, M.; Paciorek, T.; Baster, P.; Vanneste, S.; Zhang, J.; Simon, S.; Covanova, M. ABP1 mediates auxin inhibition of clathrin-dependent endocytosis in Arabidopsis. Cell 2010, 143, 111–121. [Google Scholar]
- Kitakura, S.; Vanneste, S.; Robert, S.; Lofke, C.; Teichmann, T.; Tanaka, H.; Friml, J. Clathrin mediates endocytosis and polar distribution of PIN auxin transporters in Arabidopsis. Plant Cell 2011, 23, 1920–1931. [Google Scholar]
- Blackbourn, H.D.; Jackson, A.P. Plant clathrin heavy chain: Sequence analysis and restricted localisation in growing pollen tubes. J. Cell Sci 1996, 109, 777–787. [Google Scholar]
- Fujimoto, M.; Arimura, S.; Ueda, T.; Takanashi, H.; Hayashi, Y.; Nakano, A.; Tsutsumi, N. Arabidopsis dynamin-related proteins DRP2B and DRP1A participate together in clathrin-coated vesicle formation during endocytosis. Proc. Natl. Acad. Sci. USA 2010, 107, 6094–6099. [Google Scholar]
- Food and Agriculture Organization of the United Nations. Available online: http://faostat.fao.org (on accessed 31 December 2008).
- Tester, M.; Langridge, P. Breeding technologies to increase crop production in a changing world. Science 2010, 327, 818–822. [Google Scholar]
- Lescot, M.; Ddhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; van de Peer, Y.; Rouzd, P.; Rombauts, S. PlantCARE: A database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 2002, 30, 325–327. [Google Scholar]
- Kirchhausen, T.; Harrison, S.C.; Chow, E.P.; Mattaliano, R.J.; Ramachandran, K.L.; Smart, J.; Brosius, J. Clathrin heavy chain: Molecular cloning and complete primary structure. Proc. Natl. Acad. Sci. USA 1987, 84, 8805–8809. [Google Scholar]
- Lemmon, S.K.; Pellicena-Palle, A.; Conley, K.; Freund, C.L. Sequence of the clathrin heavy chain from Saccharomyces cerevisiae and requirement of the COOH terminus for clathrin function. J. Cell Biol 1991, 112, 65–80. [Google Scholar]
- Doge, G.R.; Kovalszky, I.; McBride, O.W.; Yi, H.F.; Chu, M.; Saitta, B.; Stokes, D.; Iozzo, R.V. Human clathrin heavy chain (CLTL): Partial molecular cloning, expression, and mapping of the gene to human chromosome 17q11-qter. Genomics 1991, 11, 174–178. [Google Scholar]
- Sirotkin, H.; Morrow, B.; DasGupta, R.; Goldberg, R.; Patanjali, S.R.; Shi, G.; Cannizzaro, L.; Shprintzen, R.; Weissman, S.M.; Kucherlapati, R. Isolation of a new clathrin heavy chain gene with muscle-specific expression from the region commonly deleted in velo-cardio-facial syndrome. Hum. Mol. Genet. 1996, 5, 617–624. [Google Scholar]
- Bazinet, C.; Katzen, A.L.; Morgan, M.; Mahowald, A.P.; Lemmon, S.K. The Drosophila clathrin heavy chain gene: Clathrin function is essential in a multicellular organism. Genetics 1993, 134, 1119–1134. [Google Scholar]
- Wingen, C.; Stumpges, B.; Hoch, M.; Behr, M. Expression and localization of clathrin heavy chain in Drosophila melanogaster. Gene Expr. Patterns 2009, 9, 549–554. [Google Scholar]
- Hölzenspies, J.J.; Roelen, B.A.J.; Colenbrander, B.; Romijn, R.A.P.; Hemrika, W.; Stoorvogel, W.; van Haeften, T. Clathrin is essential for meiotic spindle function in oocytes. Reproduction 2010, 140, 223–233. [Google Scholar]
- Nathke, I.S.; Heuser, J.; Lupas, A.; Stockton, J.; Turck, C.W.; Brodsky, F.M. Folding and trimerisation of clathrin subunits at the triskelion hub. Cell 1992, 68, 899–910. [Google Scholar]
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol. Biol. Evol 2007, 24, 1596–1599. [Google Scholar]
- Bannai, H.; Tamada, Y.; Maruyama, O.; Nakai, K.; Miyano, S. Extensive feature detection of N-terminal protein sorting signals. Bioinformatics 2002, 18, 298–305. [Google Scholar]
- Bari, R.; Jones, J.D. Role of plant hormones in plant defence responses. Plant Mol. Biol 2009, 69, 473–488. [Google Scholar]
- An, C.; Mou, Z. Salicylic acid and its function in plant immunity. J. Integr. Plant Biol 2011, 53, 412–428. [Google Scholar]
- Vernooij, B.; Friedrich, L.; Morse, A.; Rest, R.; Kolditz-Jawahar, R.; Ward, E.; Uknef, S.; Kessmann, H.; Ryalf, J. Salicylic acid is not the translocated signal responsible for inducing systemic acquired resistance but is required for signal transduction. Plant Cell 1994, 6, 959–965. [Google Scholar]
- Pérez-Gómez, J.; Moore, I. Plant endocytosis: It is clathrin after all. Curr. Biol. 2007, 17, R217–R219. [Google Scholar]
- Geldner, N.; Jurgens, G. Endocytosis in signalling and development. Curr. Opin. Plant Biol 2006, 9, 589–594. [Google Scholar]
- Holstein, S.E. Clathrin and plant endocytosis. Traffic 2002, 3, 614–620. [Google Scholar]
- Dhonukshe, P.; Tanaka, H.; Goh, T.; Ebine, K.; Mähönen, A.P.; Prasad, K.; Blilou, I.; Geldner, N.; Xu, J.; Uemura, T.; et al. Generation of cell polarity in plants links endocytosis, auxin distribution and cell fate decisions. Nature 2008, 456, 962–966. [Google Scholar]
- Boutté, Y.; Frescatada-Rosa, M.; Men, S.; Chow, C.M.; Ebine, K.; Gustavsson, A.; Johansson, L.; Ueda, T.; Moore, I.; Jurgens, G.; et al. Endocytosis restricts Arabidopsis KNOLLE syntaxin to the cell division plane during late cytokinesis. EMBO J 2010, 29, 546–558. [Google Scholar]
- Takeda, S.; Gapper, C.; Kaya, H.; Bell, E.; Kuchitsu, K.; Dolan, L. Local positive feedback regulation determines cell shape in root hair cells. Science 2008, 319, 1241–1244. [Google Scholar]
- Zhao, Y.; Yan, A.; Feijó, J.A.; Furutani, M.; Takenawa, T.; Hwang, I.; Fu, Y.; Yang, Z. Phosphoinositides regulate clathrindependent endocytosis at the tip of pollen tubes in Arabidopsis and tobacco. Plant Cell 2010, 22, 4031–4044. [Google Scholar]
- Ivanov, R.; Gaude, T. Endocytosis and endosomal regulation of the S-receptor kinase during the self-incompatibility response in Brassica oleracea. Plant Cell 2009, 21, 2107–2117. [Google Scholar]
- Robatzek, S.; Chinchilla, D.; Boller, T. Ligand-induced endocytosis of the pattern recognition receptor FLS2 in Arabidopsis. Genes Dev 2006, 20, 537–542. [Google Scholar]
- Sutter, J.U.; Sieben, C.; Hartel, A.; Eisenach, C.; Thiel, G.; Blatt, M.R. Abscisic acid triggers the endocytosis of the Arabidopsis KAT1 K+ channel and its recycling to the plasma membrane. Curr. Biol 2007, 17, 1396–1402. [Google Scholar]
- Takano, J.; Miwa, K.; Yuan, L.; von Wirén, N.; Fujiwara, T. Endocytosis and degradation of BOR1, a boron transporter of Arabidopsis thaliana, regulated by boron availability. Proc. Natl. Acad. Sci. USA 2005, 102, 12276–12281. [Google Scholar]
- Takano, J.; Tanaka, M.; Toyoda, A.; Miwa, K.; Kasai, K.; Fuji, K.; Onouchi, H.; Naito, S.; Fujiwara, T. Polar localization and degradation of Arabidopsis boron transporters through distinct trafficking pathways. Proc. Natl. Acad. Sci. USA 2010, 107, 5220–5225. [Google Scholar]
- Onelli, E.; Prescianotto-Baschong, C.; Caccianiga, M.; Moscatelli, A. Clathrin-dependent and independent endocytic pathways in tobacco protoplasts revealed by labeling with charged nanogold. J. Exp. Bot 2008, 59, 3051–3068. [Google Scholar]
- Prescianotto-Baschong, C.; Riezman, H. Morphology of the yeast endocytic pathway. Mol. Biol. Cell 1998, 9, 173–189. [Google Scholar]
- Baluska, F.; Hlavacka, A.; Samaj, J.; Palme, K.; Robinson, D.G.; Matoh, T.; McCurdy, D.; Menzel, D.; Volkmann, D. F-actin dependent endocytosis of cell wall pectins in meristematic root cells. Insights from Brefeldin A-induced compartments. Plant Physiol 2002, 130, 422–431. [Google Scholar]
- Koornnef, A.; Pieterse, C.M. Cross talk in defence signaling. Plant Physiol 2008, 146, 839–844. [Google Scholar]
- Mika, A.; Boenisch, M.J.; Hopff, D.; Luthje, S. Membrane-bound guaiacol peroxidases from maize (Zea mays L.) roots are regulated by methyl jasmonate, salicylic acid, and pathogen elicitors. J. Exp. Bot. 2010, 61, 831–841. [Google Scholar]
- Leborgne-Castel, N.; Adam, T.; Bouhidel, K. Endocytosis in plant-microbe interactions. Protoplasma 2010, 247, 177–193. [Google Scholar]
- Leborgne-Castel, N.; Lherminier, J.; Der, C.; Fromentin, J.; Houot, V.; Simon-Plas, F. The plant defense elicitor cryptogein stimulates clathrin-mediated endocytosis correlated with reactive oxygen species production in Bright Yellow-2 tobacco cells. Plant Physiol 2008, 146, 1255–1266. [Google Scholar]
- Banbury, D.N.; Oakley, J.D.; Sessions, R.B.; Banting, G. Tyrphostin A23 inhibits internalization of the transferrin receptor by perturbing the interaction between tyrosine motifs and the medium chain subunit of the AP-2 adaptor complex. J. Biol. Chem 2003, 278, 12022–12028. [Google Scholar]
- Murashige, T.; Skoog, F. A revised medium for rapid growth and bioassay with tobacco tissue cultures. Physiol. Plant 1962, 15, 473–497. [Google Scholar]
- Wan, X.R.; Li, L. Molecular cloning and characterization of a dehydration-inducible cDNA encoding a putative 9-cis-epoxycarotenoid dioxygenase in Arachis hypogaea L. DNA Seq 2005, 16, 217–223. [Google Scholar]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol 1990, 215, 403–410. [Google Scholar]
- Schwede, T.; Kopp, J.; Guex, N.; Peitsch, M.C. SWISS-MODEL: An automated protein homology-modeling server. Nucleic Acids Res 2003, 31, 3381–3385. [Google Scholar]
- Arnold, K.; Bordoli, L.; Kopp, J.; Schwede, T. The SWISS-MODEL workspace: A web-based environment for protein structure homology modeling. Bioinformatics 2006, 22, 195–201. [Google Scholar]
- Muller, P.Y.; Janovjak, H.; Miserez, A.R.; Dobbie, Z. Processing of gene expression data generated by quantitative real-time RT-PCR. Biotechniques 2002, 32, 1372–1379. [Google Scholar]
- Jahn, R.; Lang, T.; Sudhof, T.C. Membrane fusion. Cell 2003, 112, 519–533. [Google Scholar]
- Surpin, M.; Raikhel, N. Traffic jams affect plant development and signal transduction. Nat. Rev. Mol. Cell Biol 2004, 5, 100–109. [Google Scholar]



| cis-Acting elements | Position (upstream of the start codon of ZmCHC1 or ZmCHC2) | Core sequence | Function | ||
|---|---|---|---|---|---|
| ZmCHC1 | ZmCHC2 | ZmCHC1 | ZmCHC2 | ||
| TCA-element | −403 to −412 | GAGAAGAAGA | Salicylic acid responsiveness | ||
| ABRE | −152 to −143 | −880 to −875 | CCGACGTGGC | CACGTG | Abscisic acid responsiveness |
| −1163 to −1158 | CGTGGA | ||||
| −1189 to −1180 | TGGGGGTGGC | ||||
| −1441 to −1436 | TACGTG | ||||
| Box-W1 | −1470 to −1475 | −1279 to −1274 | TTGACC | TTGACC | Fungal elicitor responsiveness |
| TGACG-motif | −66 to −70 −847 to −843 | −945 to −949 | TGACG TGACG | TGACG | Methyl jasmonate responsiveness |
| −1065 to −1069 | TGACG | ||||
| −1245 to −1241 | TGACG | ||||
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Zeng, M.-H.; Liu, S.-H.; Yang, M.-X.; Zhang, Y.-J.; Liang, J.-Y.; Wan, X.-R.; Liang, H. Characterization of a Gene Encoding Clathrin Heavy Chain in Maize Up-Regulated by Salicylic Acid, Abscisic Acid and High Boron Supply. Int. J. Mol. Sci. 2013, 14, 15179-15198. https://doi.org/10.3390/ijms140715179
Zeng M-H, Liu S-H, Yang M-X, Zhang Y-J, Liang J-Y, Wan X-R, Liang H. Characterization of a Gene Encoding Clathrin Heavy Chain in Maize Up-Regulated by Salicylic Acid, Abscisic Acid and High Boron Supply. International Journal of Molecular Sciences. 2013; 14(7):15179-15198. https://doi.org/10.3390/ijms140715179
Chicago/Turabian StyleZeng, Mu-Heng, Sheng-Hong Liu, Miao-Xian Yang, Ya-Jun Zhang, Jia-Yong Liang, Xiao-Rong Wan, and Hong Liang. 2013. "Characterization of a Gene Encoding Clathrin Heavy Chain in Maize Up-Regulated by Salicylic Acid, Abscisic Acid and High Boron Supply" International Journal of Molecular Sciences 14, no. 7: 15179-15198. https://doi.org/10.3390/ijms140715179
APA StyleZeng, M.-H., Liu, S.-H., Yang, M.-X., Zhang, Y.-J., Liang, J.-Y., Wan, X.-R., & Liang, H. (2013). Characterization of a Gene Encoding Clathrin Heavy Chain in Maize Up-Regulated by Salicylic Acid, Abscisic Acid and High Boron Supply. International Journal of Molecular Sciences, 14(7), 15179-15198. https://doi.org/10.3390/ijms140715179




