Functionalization and Self-Assembly of DNA Bidimensional Arrays
Abstract
:1. Introduction
2. Results and Discussion
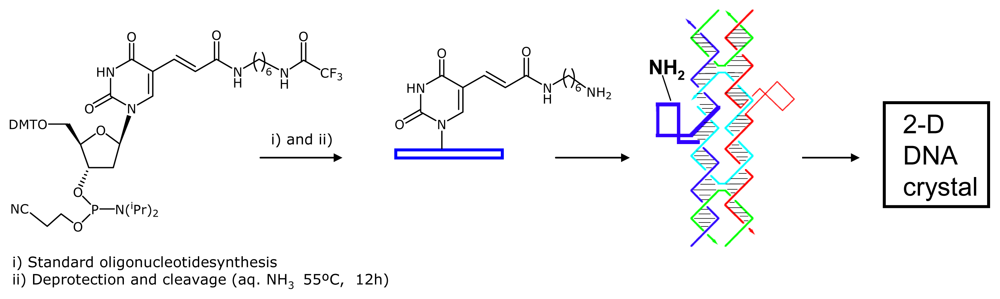
2.1. Amino-Modified DNA Arrays
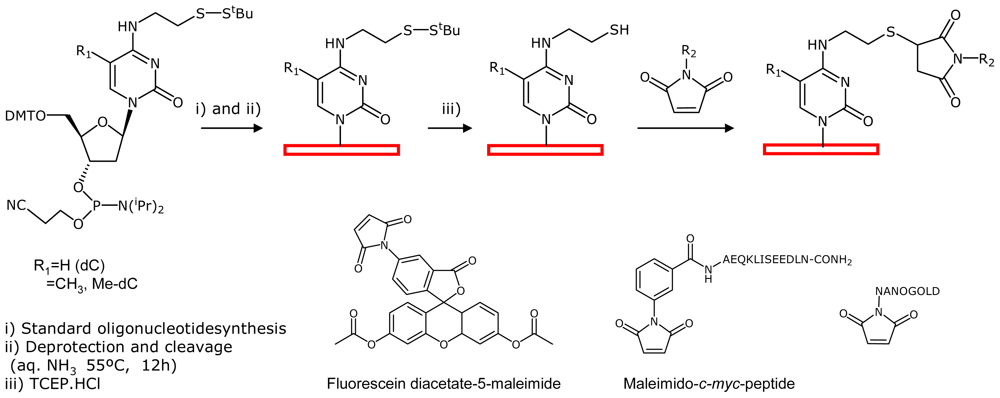
2.2. Fluorescein-, c-myc Peptide- and Nanogold-Modified DNA Arrays
3. Experimental Section
3.1. Oligonucleotide Synthesis
3.2. Preparation of DNA-Conjugates
3.3. Oligonucleotide Purification
3.4. Formation of Hydrogen-Bonded Complexes and DNA Arrays
3.5. Atomic Force Microscopy (AFM) Imaging
4. Conclusions
Supplementary Information
ijms-12-05641-s001.pdfAcknowledgments
References
- Aldaye, FA; Palmer, AL; Sleiman, HF. Assembling materials with DNA as the guide. Science 2008, 321, 1795–1799. [Google Scholar]
- Gothelf, KV; LaBean, TH. DNA-programmed assembly of nanostructures. Org Biomol Chem 2005, 3, 4023–4037. [Google Scholar]
- Seeman, NC; Lukeman, PS. Nucleic acid nanostructures: bottom-up control of geometry on the nanoscale. Rep Prog Phys 2005, 68, 237–270. [Google Scholar]
- Lin, C; Liu, Y; Yan, H. Designer DNA nanoarchitectures. Biochemistry 2009, 48, 1663–1680. [Google Scholar]
- Eritja, R. Solid-phase synthesis of modified oligonucleotides. Int J Pept Res Ther 2007, 13, 53–68. [Google Scholar]
- Winfree, E; Liu, F; Wenzler, LA; Seeman, NC. Design and self-assembly of two-dimensional DNA crystals. Nature 1998, 394, 539–544. [Google Scholar]
- Sharma, J; Ke, Y; Lin, C; Chhabra, R; Wang, Q; Nangreave, J; Liu, Y; Yan, H. DNA-tile-directed self-assembly of quantum dots into two-dimensional nanopatterns. Angew Chem Int Ed 2008, 47, 5157–5159. [Google Scholar]
- Seeman, NC. At the crossroads of chemistry, biology and materials: structural DNA nanotechnology. Chem Biol 2003, 10, 1151–1159. [Google Scholar]
- Yan, H; Park, SH; Finkelstein, G; Reif, JH; LaBean, TH. DNA-templated self-assembly of protein arrays and highly conductive nanowires. Science 2003, 301, 1882–1884. [Google Scholar]
- Li, H; Park, SH; Reif, JH; LaBean, TH; Yan, H. DNA-templated self-assembly of protein and nanoparticle linear arrays. J Am Chem Soc 2004, 126, 418–419. [Google Scholar]
- Rothemund, PWK. Folding DNA to create nanoscale shapes and patterns. Nature 2006, 440, 297–302. [Google Scholar]
- Kershner, RJ; Bozano, LD; Micheel, CM; Hung, AM; Fornof, AR; Cha, JN; Rettner, CT; Bersani, M; Frommer, J; Rothemund, PWK; Wallraff, GM. Placement and orientation of individual DNA shapes on lithographically patterned surfaces. Nat Nanotechnol 2009, 4, 557–561. [Google Scholar]
- Xiao, SJ; Liu, F; Rosen, AE; Hainfeld, JF; Seeman, NC; Musier-Forsyth, K; Kiehl, RA. Selfassembly of metallic nanoparticle arrays by DNA scaffolding. J Nanopart Res 2002, 4, 313–317. [Google Scholar]
- Le, JD; Pinto, Y; Seeman, NC; Musier-Forsyth, K; Taton, TA; Kiehl, RA. DNA-templated self-assembly of metallic nanocomponent arrays on a surface. Nano Lett 2004, 4, 2343–2347. [Google Scholar]
- Williams, BAR; Lund, K; Liu, Y; Yan, H; Chaput, JC. Self-assembled peptide nanoarrays: An approach to studying protein-protein interactions. Angew Chem Int Ed 2007, 46, 3051–3054. [Google Scholar]
- Garibotti, AV; Sisquella, X; Martínez, E; Eritja, R. Assembly of two-dimensional DNA crystals carrying N4-[2-(tert-butyldisulfanyl)ethyl]-cytosine residues. Helv Chim Acta 2009, 92, 1466–1472. [Google Scholar]
- Gottschling, D; Seliger, H; Tarrasón, G; Piulats, J; Eritja, R. Synthesis of oligodeoxynucleotides containing N4-mercaptoethylcytosine and their use in the preparation of oligonucleotide-peptide conjugates carrying c-myc tag sequence. Bioconjugate Chem 1998, 9, 831–837. [Google Scholar]
- Pérez-Rentero, S; Garibotti, AV; Eritja, R. Solid-phase synthesis of oligodeoxynucleotides carrying N4-[2-t-butyldisulfanyl)ethyl]-5-methylcytosine. Molecules 2010, 15, 5692–5707. [Google Scholar]
- Manning, B; Pérez-Rentero, S; Garibotti, AV; Ramos, R; Eritja, R. Modified oligonucleotides for biosensing applications. Sensor Lett 2009, 7, 774–781. [Google Scholar]
- Aviñó, A; Grijalvo, S; Pérez-Rentero, S; Garibotti, A; Terrazas, M; Eritja, R. Synthesis of oligonucleotide-peptide conjugates for biomedical and technological applications. Meth Mol Biol 2011, 751, 223–238. [Google Scholar]
- De la Torre, BG; Morales, JC; Aviñó, A; Iacopino, D; Ongaro, A; Fitzmaurice, D; Murphy, D; Doyle, H; Redmond, G; Eritja, R. Synthesis of oligonucleotides carrying anchoring groups and their use in the preparation of oligonucleotide-gold conjugates. Helv Chim Acta 2002, 85, 2594–2607. [Google Scholar]

| # | Name | Sequence (5′-3′) |
|---|---|---|
| 1 | A1 | GATGGCGACATCCTGCCGCTATGATTACACAGCCTGAGCATTGACAC |
| 2 | A2 | GTAGCGCCGTTAGTGGATGTC |
| 3 | A3 | TGTAGTATCGTGGCTGTGTAATCATAGCGGCACCAACTGGCA |
| 4 | A4 | GACTGCGTGTCAATGCTCACCGATGCAACCAG |
| 5 | A5 | CTGACGCTGGTTGCATCGGACGATACTACATGCCAGTTGGACTAACGG |
| 6 | B1 | CGCTACCGTGCATCATGGACTAACCAGTGCTCGCTGATTTTTCAGCGAGTTACCGCATCGGACTCGGACAGCAGC |
| 7 | B2 | CGTCAGGCTGCTGTGCTCGTGC |
| 8 | B3 | AGTACAACGCCACCGATGCGGTCACTGGTTAGTGGATTGCGT |
| 9 | B4 | GCCATCCGTCGATACGGCACCATGATGCACG |
| 10 | B5 | GCAGTCGCACGACCTGGCGTCTGTTGGCTTTTGCCAACAGTTTGTACTACGCAATCCTGCCGTATCGACG |
| 11 | B1-amino | CGCTACCGTGCATCATGGACTAACCAGTGCTCGCTGATTXTTCAGCGAGTTACCGCATCGGACAGCAGC; X = amino-dT |
| 12 | B1-thiol | CGCTACCGTGCATCATGGACTAACCAGTGCTCGCTGATTYTTCAGCGAGTTACCGCATCGGACAGCAGC, Y = t-butyldithio-ethyl-dC, t-butyldithio-ethyl-5-methyl-dC |
| 13 | B5-thiol | GCAGTCGCACGACCTGGCGTCTGTTGGCTTYTGCCAACAGTTTGTACTACGCAATCCTGCCGTATCGACG, Y = t-butyldithio-ethyl-dC, t-butyldithio-ethyl-5-methyl-dC |
| 14 | B5-fluorescein | GCAGTCGCACGACCTGGCGTCTGTTGGCTTZTGCCAACAGTTTGTACTACGCAATCCTGCCGTATCGACG; Z = N-(fluorescein-maleimido-S-ethyl)-5-methyl-dC |
| 15 | B5-c-myc peptide | GCAGTCGCACGACCTGGCGTCTGTTGGCTTZTGCCAACAGTTTGTACTACGCAATCCTGCCGTATCGACG; Z = N-(c-myc-peptide-maleimido-S-ethyl)-5-methyl-dC; c-myc peptide sequence: Maleimido-Ala-Glu-Gln-Lys-Leu-Ile-Ser-Glu-Glu-Asp-Leu-Asn-CONH2 |
| 16 | B1-Nanogold | CGCTACCGTGCATCATGGACTAACCAGTGCTCGCTGATTZTTCAGCGAGTTACCGCATCGGACAGCAGC, Z = N-(Nanogold-maleimido-S-ethyl)-dC |
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Garibotti, A.V.; Pérez-Rentero, S.; Eritja, R. Functionalization and Self-Assembly of DNA Bidimensional Arrays. Int. J. Mol. Sci. 2011, 12, 5641-5651. https://doi.org/10.3390/ijms12095641
Garibotti AV, Pérez-Rentero S, Eritja R. Functionalization and Self-Assembly of DNA Bidimensional Arrays. International Journal of Molecular Sciences. 2011; 12(9):5641-5651. https://doi.org/10.3390/ijms12095641
Chicago/Turabian StyleGaribotti, Alejandra V., Sónia Pérez-Rentero, and Ramon Eritja. 2011. "Functionalization and Self-Assembly of DNA Bidimensional Arrays" International Journal of Molecular Sciences 12, no. 9: 5641-5651. https://doi.org/10.3390/ijms12095641