From Cell to Beak: In-Vitro and In-Vivo Characterization of Chicken Bitter Taste Thresholds
Abstract
:1. Introduction
2. Results
2.1. All ggTas2rs Are Expressed in Transfected Cells
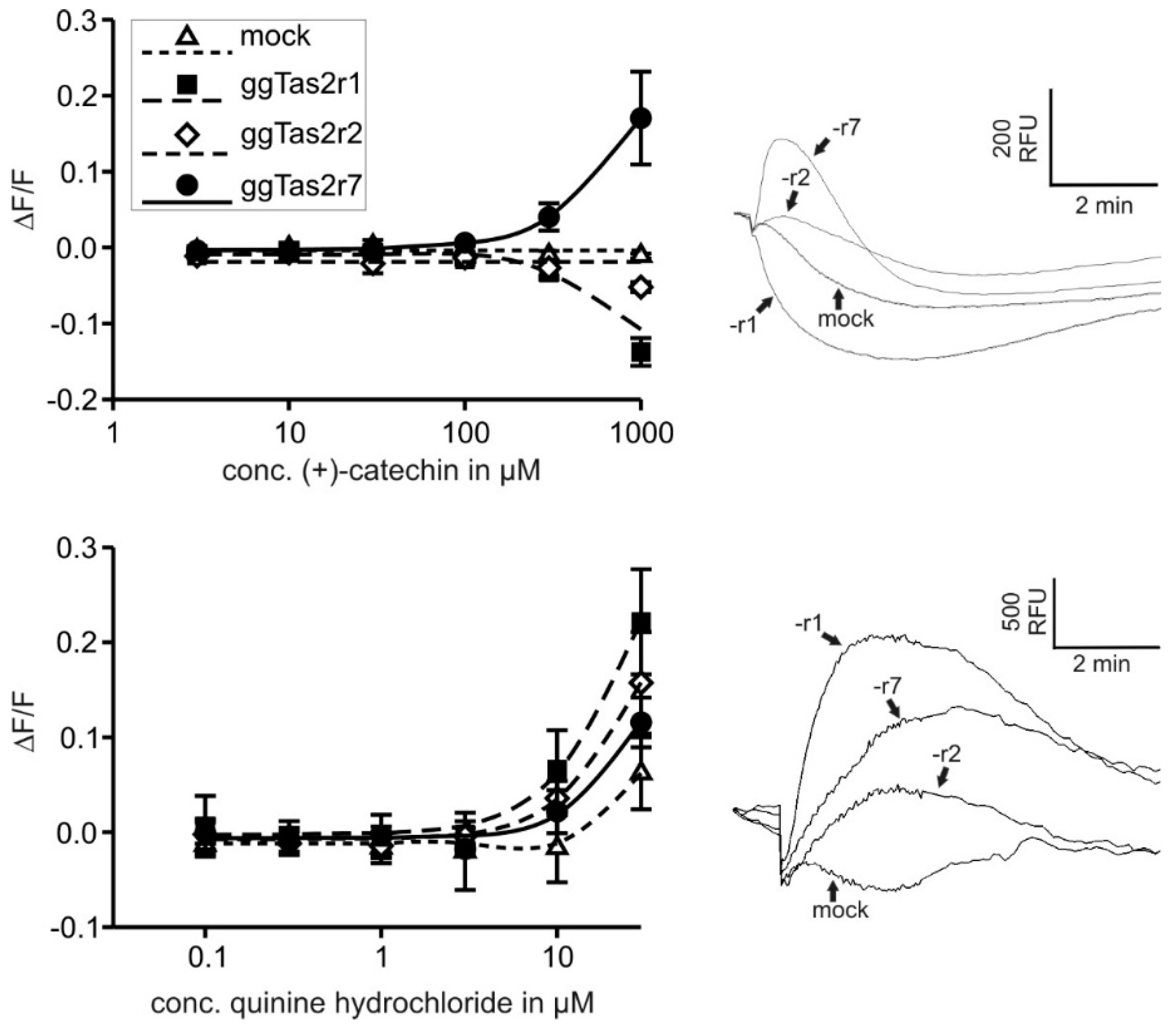
2.2. Bitter Compounds Show Different In-Vitro Detection Thresholds
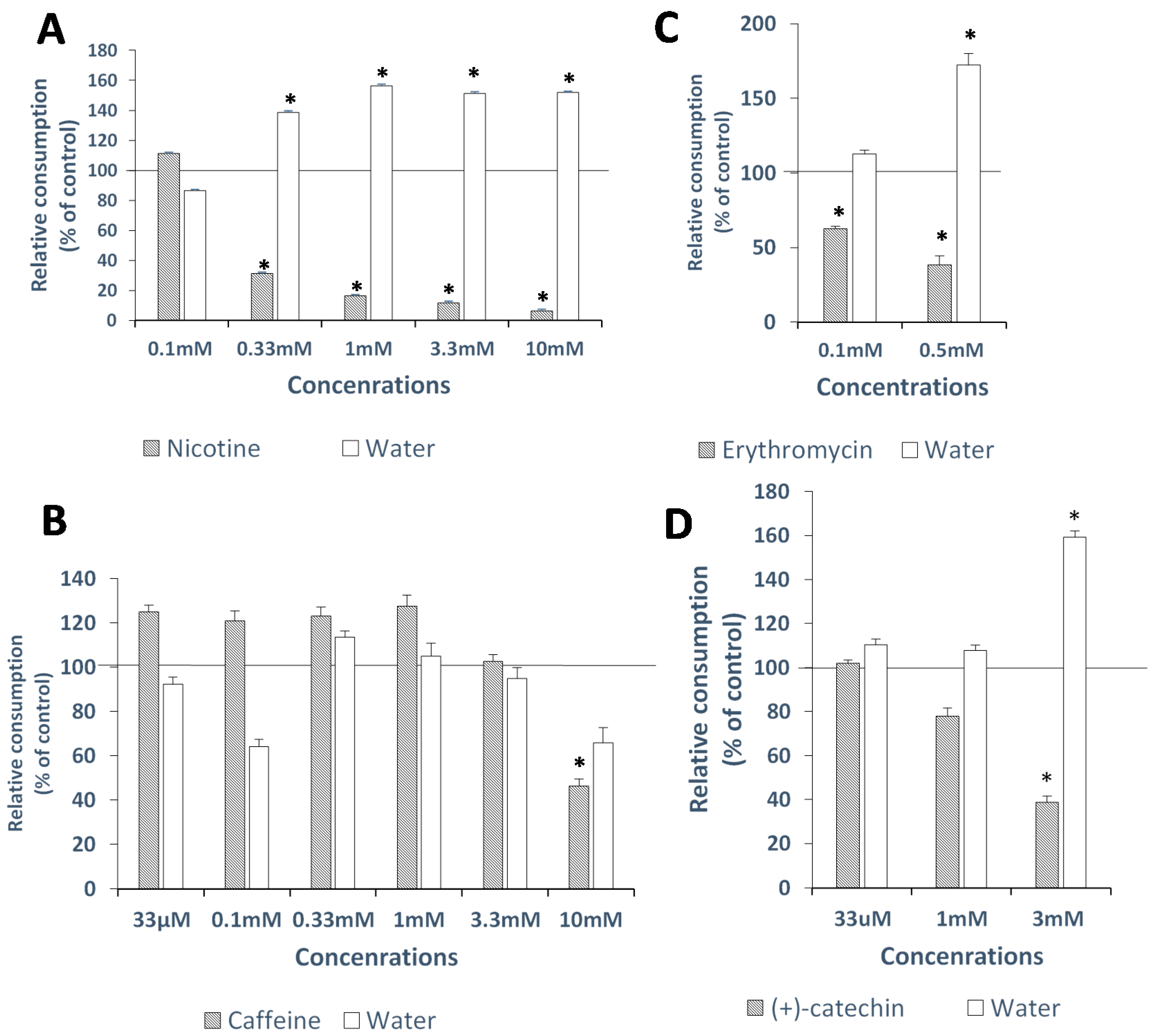
2.3. In-Vivo Avoidance Thresholds
2.3.1. Nicotine
2.3.2. Caffeine
2.3.3. Erythromycin
2.3.4. (+)-Catechin
2.4. ggTas2rs Exhibit Different Expression Levels in the Palate
2.5. Variation in In-Vivo:In-Vitro Ratios
3. Discussion
3.1. Relationships between In-Vivo and In-Vitro Thresholds
3.2. Possible Reasons for Differences between In-Vivo and In-Vitro Thresholds
3.2.1. Avoidance vs. Recognition
3.2.2. Different Expression Levels of the Cognate Receptors
3.2.3. Additional Sensory Properties
3.2.4. Test Length
4. Materials and Methords
4.1. Chemicals
4.1.1. Functional Expression of Chicken Tas2r Constructs
4.1.2. Immunocytochemistry
4.2. Taste Tests
4.2.1. Animals and Maintenance Conditions
4.2.2. In-Vivo Avoidance Threshold Detection Tests
4.2.3. mRNA Abundance (Fold Change) Calculation Using the Comparative ΔΔCt Method
4.3. Statistical Analysis
4.3.1. Taste Tests
4.3.2. Gene Expression
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lindemann, B. Taste reception. Physiol. Rev. 1996, 76, 719–766. [Google Scholar]
- Drewnowski, A.; Gomez-Carneros, C. Bitter taste, phytonutrients, and the consumer: A review. Am. J. Clin. Nutr. 2000, 72, 1424–1435. [Google Scholar] [PubMed]
- Meyerhof, W.; Batram, C.; Kuhn, C.; Brockhoff, A.; Chudoba, E.; Bufe, B.; Appendino, G.; Behrens, M. The molecular receptive ranges of human tas2r bitter taste receptors. Chem. Senses 2010, 35, 157–170. [Google Scholar] [CrossRef] [PubMed]
- Sanematsu, K.; Yoshida, R.; Shigemura, N.; Ninomiya, Y. Structure, function, and signaling of taste g-protein-coupled receptors. Curr. Pharm. Biotechnol. 2014, 15, 951–961. [Google Scholar] [CrossRef] [PubMed]
- Adler, E.; Hoon, M.A.; Mueller, K.L.; Chandrashekar, J.; Ryba, N.J.; Zuker, C.S. A novel family of mammalian taste receptors. Cell 2000, 100, 693–702. [Google Scholar] [CrossRef]
- Matsunami, H.; Montmayeur, J.-P.; Buck, L.B. A family of candidate taste receptors in human and mouse. Nature 2000, 404, 601–604. [Google Scholar] [PubMed]
- Behrens, M.; Korsching, S.I.; Meyerhof, W. Tuning properties of avian and frog bitter taste receptors dynamically fit gene repertoire sizes. Mol. Biol. Evol. 2014, 31, 3216–3227. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.V.; Chen, M.C.; Rozengurt, E. Genomic organization, expression, and function of bitter taste receptors (t2r) in mouse and rat. Physiol. Genom. 2005, 22, 139–149. [Google Scholar] [CrossRef] [PubMed]
- Jiang, P.; Josue, J.; Li, X.; Glaser, D.; Li, W.; Brand, J.G.; Margolskee, R.F.; Reed, D.R.; Beauchamp, G.K. Major taste loss in carnivorous mammals. Proc. Natl. Acad. Sci. USA 2012, 109, 4956–4961. [Google Scholar] [CrossRef] [PubMed]
- Syed, A.S.; Korsching, S.I. Positive darwinian selection in the singularly large taste receptor gene family of an ’ancient’ fish, latimeria chalumnae. BMC Genom. 2014, 15, 650. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Li, J.; Zhang, J. Molecular evidence for the loss of three basic tastes in penguins. Curr. Biol. 2015, 25, R141–R142. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Zhao, H. Birds generally carry a small repertoire of bitter taste receptor genes. Genome Biol. Evol. 2015, 7, 2705–2715. [Google Scholar] [CrossRef] [PubMed]
- Roura, E.; Baldwin, M.; Klasing, K. The Avian Taste System: A review, Proceedings of the 23rd Annual, Australian Poultry Science Symposium, Sydney, Australia, 19–22 February, 2012; Poultry Research Foundation: Sydney, Australia, 2012; pp. 97–104. [Google Scholar]
- Rowland, H.M.; Rockwell Parker, M.; Jiang, P.; Reed, D.R.; Beauchamp, G.K. Comparative taste biology with special focus on birds and reptiles. In Handbook of Olfaction and Gustation; John Wiley & Sons, Inc.: Hoboken, NJ, USA; New York, NY, USA, 2015; pp. 957–982. [Google Scholar]
- Rajapaksha, P.; Wang, Z.; Venkatesan, N.; Tehrani, K.F.; Payne, J.; Swetenburg, R.L.; Kawabata, F.; Tabata, S.; Mortensen, L.J.; Stice, S.L. Labeling and analysis of chicken taste buds using molecular markers in oral epithelial sheets. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Ganchrow, D.; Ganchrow, J.R. Number and distribution of taste buds in the oral cavity of hatchling chicks. Physiol. Behav. 1985, 34, 889–894. [Google Scholar] [CrossRef]
- Cheled-Shoval, S.L.; Behrens, M.; Meyerhof, W.; Niv, M.Y.; Uni, Z. Perinatal administration of a bitter tastant influences gene expression in chicken palate and duodenum. J. Agric. Food Chem. 2014, 62, 12512–12520. [Google Scholar] [CrossRef] [PubMed]
- Cheled-Shoval, S.L.; Druyan, S.; Uni, Z. Bitter, sweet and umami taste receptors and downstream signaling effectors: Expression in embryonic and growing chicken gastrointestinal tract. Poult. Sci. 2015, 94, 1928–1941. [Google Scholar] [CrossRef] [PubMed]
- Hirose, N.; Kawabata, Y.; Kawabata, F.; Nishimura, S.; Tabata, S. Bitter taste receptor t2r1 activities were compatible with behavioral sensitivity to bitterness in chickens. Biochem. Biophys. Res. Commun. 2015, 460, 464–468. [Google Scholar] [CrossRef] [PubMed]
- Kudo, K.; Kawabata, F.; Nomura, T.; Aridome, A.; Nishimura, S.; Tabata, S. Isolation of chicken taste buds for real-time Ca2+ imaging. Anim. Sci. J. 2014, 85, 904–909. [Google Scholar] [CrossRef] [PubMed]
- Kudo, K.; Shiraishi, J.; Nishimura, S.; Bungo, T.; Tabata, S. The number of taste buds is related to bitter taste sensitivity in layer and broiler chickens. Anim. Sci. J. 2010, 81, 240–244. [Google Scholar] [CrossRef] [PubMed]
- Kudo, K.; Wakamatsu, K.; Nishimura, S.; Tabata, S. Gustducin is expressed in the taste buds of the chicken. Anim. Sci. J. 2010, 81, 666–672. [Google Scholar] [CrossRef] [PubMed]
- Mazzoni, M.; Bombardi, C.; Vallorani, C.; Sirri, F.; De Giorgio, R.; Caio, G.; Grandis, A.; Sternini, C.; Clavenzani, P. Distribution of alpha-transducin and alpha-gustducin immunoreactive cells in the chicken (gallus domesticus) gastrointestinal tract. Poult. Sci. 2016, 95, 1624–1630. [Google Scholar] [CrossRef] [PubMed]
- Roura, E.; Baldwin, M.W.; Klasing, K. The avian taste system: Potential implications in poultry nutrition. Anim. Feed Sci. Technol. 2013, 180, 1–9. [Google Scholar] [CrossRef]
- Go, Y. Lineage-specific expansions and contractions of the bitter taste receptor gene repertoire in vertebrates. Mol. Biol. Evol. 2006, 23, 964–972. [Google Scholar] [CrossRef] [PubMed]
- Cheled-Shoval, S.L.; Reicher, N.; Niv, M.Y.; Uni, Z. Detecting thresholds for bitter, umami and sweet tastants in broiler chicken using a two choice test method. Poult. Sci. 2017. [Google Scholar] [CrossRef] [PubMed]
- Wiener, A.; Shudler, M.; Levit, A.; Niv, M.Y. Bitterdb: A database of bitter compounds. Nucleic Acid Res. 2012, 40, D413–D419. [Google Scholar] [CrossRef] [PubMed]
- Di Pizio, A.; Niv, M.Y. Promiscuity and selectivity of bitter molecules and their receptors. Bioorg. Med. Chem. Lett. 2015, 23, 4082–4091. [Google Scholar] [CrossRef] [PubMed]
- Shi, P.; Zhang, J. Contrasting modes of evolution between vertebrate sweet/umami receptor genes and bitter receptor genes. Mol. Biol. Evol. 2006, 23, 292–300. [Google Scholar] [CrossRef] [PubMed]
- Baldwin, M.W.; Toda, Y.; Nakagita, T.; O’Connell, M.J.; Klasing, K.C.; Misaka, T.; Edwards, S.V.; Liberles, S.D. Evolution of sweet taste perception in hummingbirds by transformation of the ancestral umami receptor. Science 2014, 345, 929–933. [Google Scholar] [CrossRef] [PubMed]
- Dey, B.; Kawabata, F.; Kawabata, Y.; Yoshida, Y.; Nishimura, S.; Tabata, S. Identification of functional bitter taste receptors and their antagonist in chickens. Biochem. Biophys. Res. Commun. 2016, 482, 693–699. [Google Scholar] [CrossRef] [PubMed]
- Hyde, R.J.; Feller, R.P. Age and sex effects on taste of sucrose, nacl, citric acid and caffeine. Neurobiol. Aging 1981, 2, 315–318. [Google Scholar] [CrossRef]
- Purves, D.; Augustine, G.J.; Fitzpatrick, D.; Katz, L.C.; LaMantia, A.-S.; McNamara, J.O.; Williams, S.M. The chemical senses. In Neuroscience, 2nd ed.; Sinauer Associates Inc.: Sunderland, MA, USA, 2001. [Google Scholar]
- McMahon, D.B.; Shikata, H.; Breslin, P.A. Are human taste thresholds similar on the right and left sides of the tongue? Chem. Senses 2001, 26, 875–883. [Google Scholar] [CrossRef] [PubMed]
- Bufe, B.; Breslin, P.A.; Kuhn, C.; Reed, D.R.; Tharp, C.D.; Slack, J.P.; Kim, U.K.; Drayna, D.; Meyerhof, W. The molecular basis of individual differences in phenylthiocarbamide and propylthiouracil bitterness perception. Curr. Biol. 2005, 15, 322–327. [Google Scholar] [CrossRef] [PubMed]
- Vitazkova, S.K.; Long, E.; Paul, A.; Glendinning, J.I. Mice suppress malaria infection by sampling a ‘bitter’chemotherapy agent. Anim. Behav. 2001, 61, 887–894. [Google Scholar] [CrossRef]
- Ganchrow, J.R.; Ganchrow, D. Taste bud development in chickens (gallus gallus domesticus). Anat. Rec. 1987, 218, 88–93. [Google Scholar] [CrossRef] [PubMed]
- Gyekis, J.P.; Dingman, M.A.; Revitsky, A.R.; Bryant, B.P.; Vandenbergh, D.J.; Frank, M.E.; Blizard, D.A. Gustatory, trigeminal, and olfactory aspects of nicotine intake in three mouse strains. Behav. Genet. 2012, 42, 820–829. [Google Scholar] [CrossRef] [PubMed]
- Stevens, J.C. Detection of tastes in mixture with other tastes: Issues of masking and aging. Chem. Senses 1996, 21, 211–221. [Google Scholar] [CrossRef] [PubMed]
- Drewnowski, A. The science and complexity of bitter taste. Nutr. Rev. 2001, 59, 163–169. [Google Scholar] [CrossRef] [PubMed]
- Breslin, P.A.; Beauchamp, G.K. Suppression of bitterness by sodium: Variation among bitter taste stimuli. Chem. Senses 1995, 20, 609–623. [Google Scholar] [CrossRef] [PubMed]
- Breslin, P.A. Interactions among salty, sour and bitter compounds. Trends Food Sci. Technol. 1996, 7, 390–399. [Google Scholar] [CrossRef]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time rt–pcr. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef] [PubMed]
- Druyan, S.; Levi, E. Reduced O2 concentration during cam development–its effect on angiogenesis and gene expression in the broiler embryo cam. Gene Expr. Patterns 2012, 12, 236–244. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: All compounds used in this study are commercially available. |

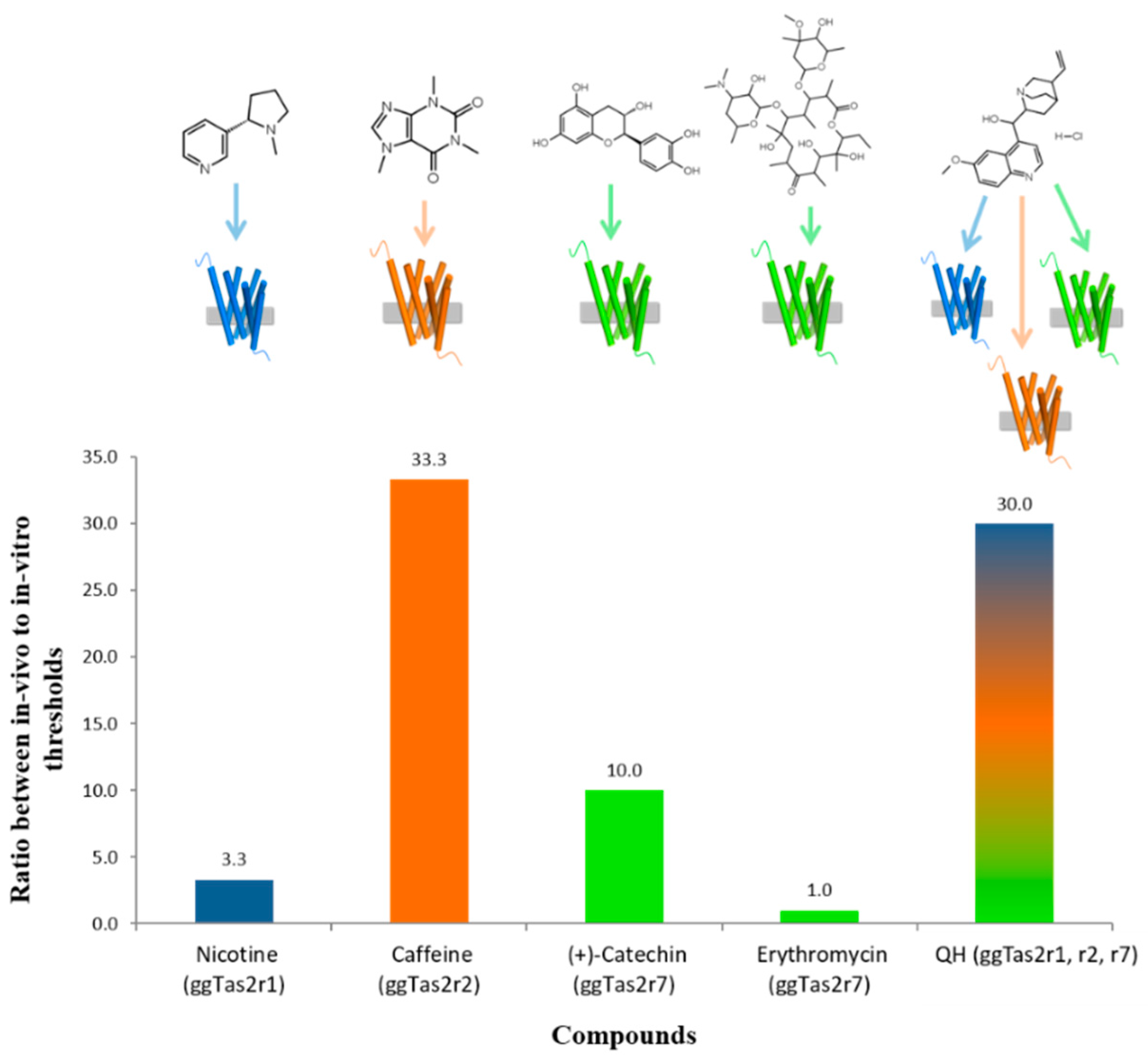
| Compound | Threshold Concentrations (mM) | Receptor Activated In-Vitro | |
|---|---|---|---|
| In-Vitro | In-Vivo | ggTas2r1 | |
| Nicotine | 0.1 (Behrens et al. [7]) | 0.33 | ggTas2r2 |
| Caffeine | 0.3 (Behrens et al. [7]) | 10 | ggTas2r7 |
| Erythromycin | 0.1 (Behrens et al. [7]) * | ≤0.1 ** | ggTas2r7 |
| (+)-Catechin | 0.3 | 3 | ggTas2r1, ggTas2r2, ggTas2r7 |
| QH | 0.01 | 0.3 (Cheled-Shoval et al. [26]) | ggTas2r1 |
| Tastant | Concentrations Tested (In-Vivo) | Concentrations Tested (In-Vitro) |
|---|---|---|
| Nicotine | 0.1 mM, 0.33 mM, 1 mM, 3.3 mM, 10 mM | Behrens et al. [7] |
| Caffeine | 33 µM, 0.1 mM, 0.33 mM, 1 mM, 3.3 mM, 10 mM | Behrens et al. [7] |
| Erythromycin | 0.1 mM, 0.5 mM | Behrens et al. [7] |
| (+)-Catechin | 33 µM, 3 mM, 1 mM | 3 µM, 10 µM, 30 µM, 100 µM, 300 µM, 1000 µM |
| QH | Cheled-Shoval et al. [26] | 0.1 µM, 0.3 µM, 1 µM, 3 µM, 10 µM, 30 µM |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cheled-Shoval, S.; Behrens, M.; Korb, A.; Di Pizio, A.; Meyerhof, W.; Uni, Z.; Niv, M.Y. From Cell to Beak: In-Vitro and In-Vivo Characterization of Chicken Bitter Taste Thresholds. Molecules 2017, 22, 821. https://doi.org/10.3390/molecules22050821
Cheled-Shoval S, Behrens M, Korb A, Di Pizio A, Meyerhof W, Uni Z, Niv MY. From Cell to Beak: In-Vitro and In-Vivo Characterization of Chicken Bitter Taste Thresholds. Molecules. 2017; 22(5):821. https://doi.org/10.3390/molecules22050821
Chicago/Turabian StyleCheled-Shoval, Shira, Maik Behrens, Ayelet Korb, Antonella Di Pizio, Wolfgang Meyerhof, Zehava Uni, and Masha Y. Niv. 2017. "From Cell to Beak: In-Vitro and In-Vivo Characterization of Chicken Bitter Taste Thresholds" Molecules 22, no. 5: 821. https://doi.org/10.3390/molecules22050821






