Identification of Alternaria alternata Mycotoxins by LC-SPE-NMR and Their Cytotoxic Effects to Soybean (Glycine max) Cell Suspension Culture
Abstract
:1. Introduction
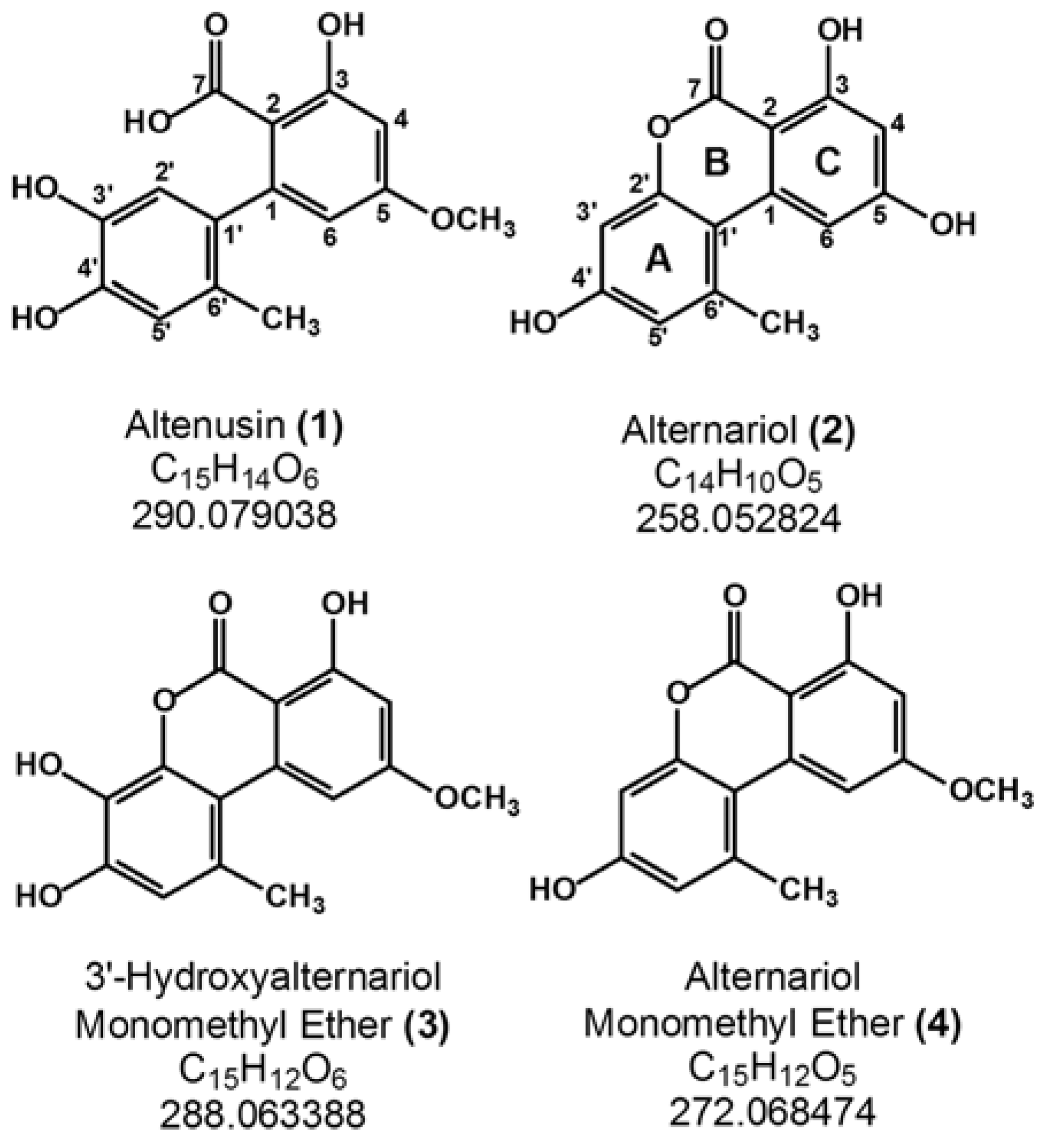
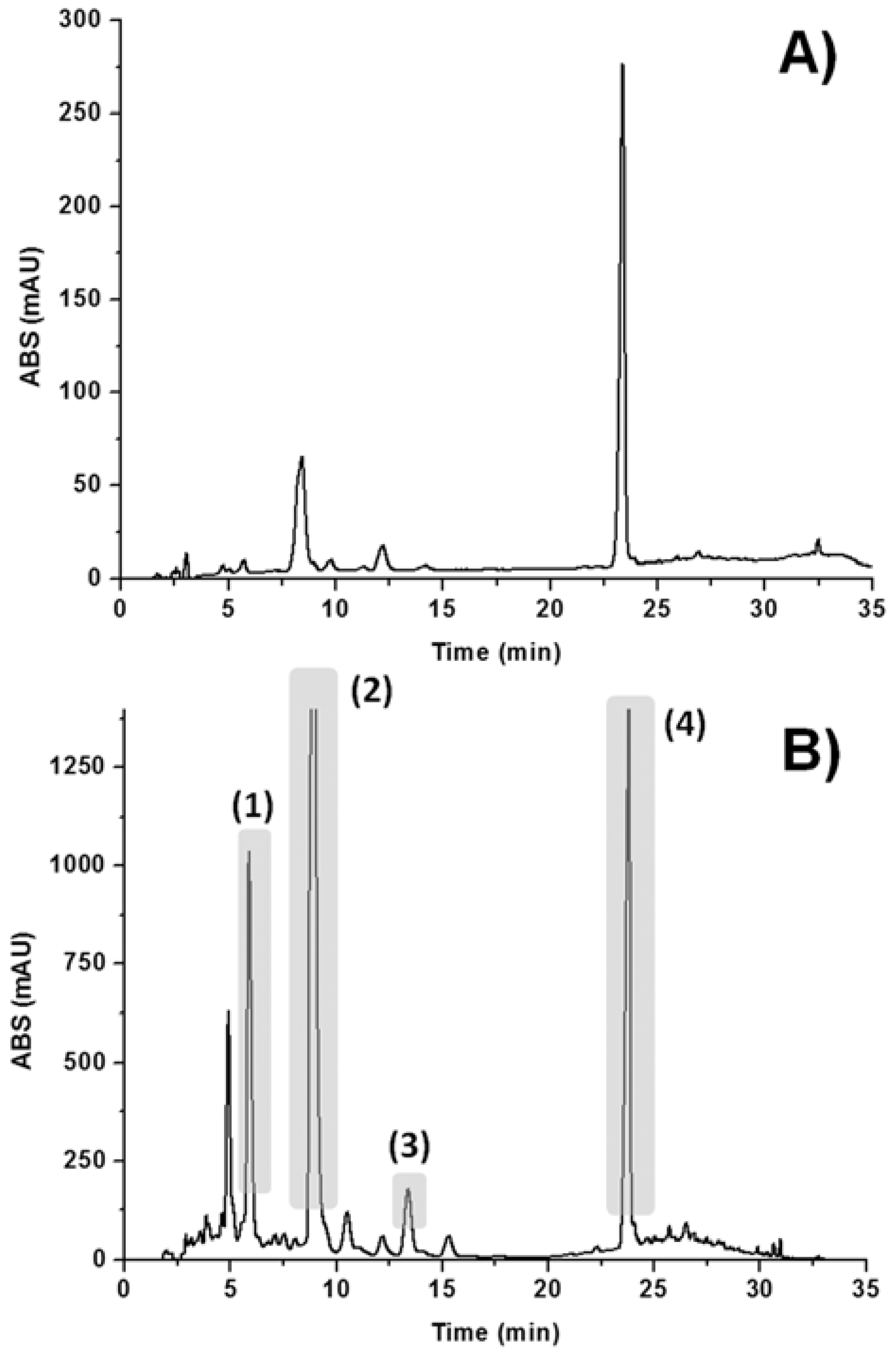
2. Results and Discussion
| Altenusin (1) | Alternariol (2) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Obtained | Reference (MeCN-d3) [26] | Obtained | Reference (CDCl3) [21] | ||||||||
| δH (J in Hz) | δC | HMBC | δH (J in Hz) | δC | δH (J in Hz) | δC | HMBC | δH (J in Hz) | δC | ||
| 1 | 147.2 | 148.3 | 139.0 | 138.1 | |||||||
| 2 | 105.3 | 106.5 | 99.6 | 97.4 | |||||||
| 3 | 165.7 | 163.0 | 164.9 | 164.1 | |||||||
| 4 | 6.47, d (2.6) | 100.6 | 2, 3, 5, 6 | 6.25, d (2.2) | 102.3 | 6.42, d (2.0) | 101.1 | 2, 3, 6 | 6.38, d (1.9) | 100.9 | |
| 5 | 165.1 | 165.3 | 165.3 | 165.5 | |||||||
| 6 | 6.19, d (2.6) | 111.4 | 1, 2, 4, 5 | 6.03, d (2.2) | 112.1 | 7.28, d (2.0) | 104.9 | 2, 4, 5, 1' | 7.27, d (2.0) | 104.4 | |
| 7 | 165.1 | 164.7 | |||||||||
| 1' | 134.7 | 135.8 | 110.2 | 109.0 | |||||||
| 2' | 6.52, s | 116.4 | 1, 1', 3', 4', 6' | 6.46, s | 116.6 | 153.9 | 152.6 | ||||
| 3' | 142.5 | 143.3 | 6.68, d (2.4) | 102.3 | 1', 2', 4', 5' | 6.32, d (2.6) | 101.6 | ||||
| 4' | 144.2 | 144.7 | 158.0 | 158.4 | |||||||
| 5' | 6.63, s | 117.1 | 1', 3', 4', 6'-CH3 | 6.55, s | 117.2 | 6.73, d (2.4) | 117.4 | 1', 3', 6'-CH3 | 6.71, d (2.6) | 117.6 | |
| 6' | 127.6 | 127.3 | 139.2 | 138.3 | |||||||
| 3-OH | 11.87, s | 11.86, s | 2, 3, 7 | ||||||||
| 5-OCH3 | 3.81, s | 56.2 | 5 | 3.87, s | 56.4 | ||||||
| 6'-CH3 | 1.90, s | 19.1 | 1', 2', 4' 5', 6' | 1.90, s | 19.3 | 2.78, s | 25.4 | 1, 1', 5', 6' | 2.77, s | 25.3 | |
| 3'-Hydroxyalternariol monomethyl ether (3) | Alternariol monomethyl ether (4) | ||||||||||
| Obtained | Reference (in DMF-d7) [26] | Obtained | Reference (in DMSO-d6) [22] | ||||||||
| δH (J in Hz) | δC | HMBC | δH (J in Hz) | δC | δH (J in Hz) | δC * | HMBC | δH (J in Hz) | δC | ||
| 1 | 138.4 | 139.5 | 138.3 | 137.8 | |||||||
| 2 | 100.0 | 99.2 | 98.1 | 98.4 | |||||||
| 3 | 165.5 | 165.4 | 165.5 | 164.0 | |||||||
| 4 | 6.52, d (1.9) | 100.3 | 2, 3, 6 | 6.63, d (1.8) | 99.7 | 6.57, d (2.0) | 101.5 | 2, 3, 5, 6 | 6.59, d (2.5) | 101.6 | |
| 5 | 167.5 | 167.3 | 166.5 | 164.6 | |||||||
| 6 | 7.31, d (1.9) | 104.8 | 2, 4, 5, 1' | 7.30, d (1.8) | 104.2 | 7.31, d (2.0) | 103.3 | 2, 4, 5, 1' | 7.20, d (2.5) | 103.5 | |
| 7 | n.d. | 165.8 | 168.3 | 166.1 | |||||||
| 1' | 111.4 | 110.1 | 109.3 | 108.3 | |||||||
| 2' | 142.2 | 142.3 | 153.4 | 152.6 | |||||||
| 3' | 131.2 | 132.4 | 6.74, d (2.4) | 101.5 | 1', 2', 4', 5' | 6.63, d (2.0) | 99.1 | ||||
| 4' | 146.4 | 148.2 | 158.9 | 158.5 | |||||||
| 5' | 6.76, s | 117.5 | 1', 2', 3', 4', 6', 6'-CH3 | 6.82, s | 117.7 | 6.68, d (2.4) | 117.5 | 1', 3', 4', 6'-CH3 | 6.71, d (2.0) | 117.5 | |
| 6' | 128.7 | 123.3 | 138.8 | 138.4 | |||||||
| 3-OH | 11.91, s | 11.89, s | |||||||||
| 5-OCH3 | 3.92, s | 56.6 | 5 | 3.99, s | 56.3 | 3.92, s | 56.7 | 5 | 3.90, s | 55.8 | |
| 6'-CH3 | 2.70, s | 25.1 | 1, 6, 1', 2', 5', 6' | 2.74, s | 24.9 | 2.78, s | 25.7 | 1, 1', 2', 5', 6' | 2.72, s | 24.6 | |
| Compound | Detected ion (m/z) | Calculated mass (Da) | Accuracy (ppm) | Deviation (mDa) |
|---|---|---|---|---|
| Altenusin (1) (C15H14O6) | 290.07970 | 290.07904 | −2.3 | −0.7 |
| Alternariol (2) (C14H10O5) | 258.05198 | 258.05281 | 3.3 | 0.8 |
| 3'-Hydroxyalternariol monomethyl ether (3) (C15H12O6) | 288.06381 | 288.06339 | −1.5 | −0.4 |
| Alternariol monomethyl ether (4) (C15H12O5) | 272.06711 | 272.06847 | 5.0 | 1.4 |

3. Experimental
3.1. General Experimental Procedures
3.2. Fungal Cultivation and Preparation of Extracts
3.3. Separation and Trapping of Alternaria Metabolites
3.4. NMR and MS analyses
3.5. Cytotoxic (Phytotoxic) Assays
4. Conclusions
Supporting Information
Acknowledgments
References
- Strange, R.N. Phytotoxins produced by microbial plant pathogens. Nat. Prod. Rep. 2007, 24, 127–144. [Google Scholar] [CrossRef]
- Agostini, J.P.; Bushong, P.M.; Timmer, L.W. Greenhouse evaluation of products that induce host resistance for control of Scab, Melanose, and Alternaria Brown Spot of citrus. Plant Dis. 2003, 87, 69–74. [Google Scholar] [CrossRef]
- Akimitsu, K.; Peever, T.L.; Timmer, L.W. Molecular, ecological and evolutionary approaches to understanding Alternaria diseases of citrus. Mol. Plant Pathol. 2003, 4, 435–446. [Google Scholar] [CrossRef]
- Peres, N.A.; Timmer, L.W. Evaluation of the Alter-Rater model for spray timing for control of Alternaria Brown Spot on murcott tangor in Brazil. Crop Protec. 2006, 25, 454–460. [Google Scholar] [CrossRef]
- Magnani, R.F.; de Souza, G.D.; Rodrigues-Filho, E. Analysis of alternariol and alternariol monomethyl ether on flavedo and albedo tissues of tangerines (Citrus reticulata) with symptoms of Alternaria Brown Spot. J. Agric. Food Chem. 2007, 55, 4980–4986. [Google Scholar]
- Asam, S.; Konitzer, K.; Schieberle, P.; Rychlik, M. Stable isotope dilution assays of alternariol and alternariol monomethyl ether in beverages. J. Agric. Food Chem. 2009, 57, 5152–5160. [Google Scholar]
- Ruberta, J.; Dzumanb, Z.; Vaclavikovab, M.; Zachariasovab, M.; Solera, C.; Hajslova, J. Analysis of mycotoxins in barley using ultra high liquid chromatography high resolution mass spectrometry: Comparison of efficiency and efficacy of different extraction procedures. Talanta 2012, 99, 712–719. [Google Scholar] [CrossRef]
- Abia, W.A.; Warth, B.; Sulyok, M.; Krska, R.; Tchana, A.N.; Njobeh, P.B.; Dutton, M.F.; Moundipa, P.F. Determination of multi-mycotoxin occurrence in cereals, nuts and their products in Cameroon by liquid chromatography tandem mass spectrometry (LC-MS/MS). Food Control. 2013, 31, 438–453. [Google Scholar] [CrossRef]
- Cirlini, M.; Dall’Asta, C.; Galaverna, G. Hyphenated chromatographic techniques for structural characterization and determination of masked mycotoxins. J. Chromatogr. A 2012, 1255, 145–152. [Google Scholar] [CrossRef]
- Dong, Z.G.; Liu, G.T.; Dong, Z.M.; Qian, Y.Z.; An, Y.H.; Miao, J.A.; Zhen, Y.Z. Induction of mutagenesis and transformation by the extract of Alternaria alternata isolated from grains in Linxian, China. Carcinogenesis 1987, 8, 989–991. [Google Scholar] [CrossRef]
- An, Y.-H.; Zhao, T.-Z.; Miao, J.; Liu, G.-T.; Zheng, Y.-Z.; Xu, Y.-M.; van Etten, R.L. Isolation, identification, and mutagenicity of alternariol monomethyl ether. J. Agric. Food Chem. 1989, 37, 1341–1343. [Google Scholar] [CrossRef]
- Pavia, D.L.; Lampman, G.M.; Kriz, G.S. Introduction to Spectroscopy, 2nd ed; Saunders College Publishing: Philadelphia, PA, USA, 1996; pp. 303–371. [Google Scholar]
- Kind, T.; Fiehn, O. Advances in structure elucidation of small molecules using mass spectrometry. Bioanal. Rev. 2010, 2, 23–60. [Google Scholar] [CrossRef]
- Pan, Z.; Raftery, D. Comparing and combining NMR spectroscopy and mass spectrometry in metabolomics. Anal. Bioanal. Chem. 2007, 387, 525–527. [Google Scholar] [CrossRef]
- Pusecker, K.; Schewitz, J.; Gfrörer, P.; Tseng, L.-H.; Albert, K.; Bayer, E. On-Line coupling of capillary electrochromatography, capillary electrophoresis, and capillary HPLC with nuclear magnetic resonance spectroscopy. Anal. Chem. 1998, 70, 3280–3285. [Google Scholar]
- Simpson, A.J.; Tseng, L.-H.; Simpson, M.J.; Spraul, M.; Braumann, U.; Kingery, W.L.; Kelleher, B.P.; Hayes, M.H.B. The application of LC-NMR and LC-SPE-NMR to compositional studies of natural organic matter. Analyst 2004, 129, 1216–1222. [Google Scholar] [CrossRef]
- Jaroszewski, J.W. Hyphenated NMR methods in natural products research, Part 2: HPLC-SPE-NMR and other new trends in NMR hyphenation. Planta Med. 2005, 71, 795–802. [Google Scholar] [CrossRef]
- Daolio, C.; Schneider, B. Coupling Liquid Chromatography and Other Separation Techniques to Nuclear Magnetic Resonance Spectroscopy. In Hyphenated and Alternative Methods of Detection in Chromatography; Shalliker, R.A., Ed.; Taylor & Francis–CRC Press: Oxford, UK, 2012; Volume 104, pp. 61–98, Chromatographic Science Series. [Google Scholar]
- Koskela, H.; Ervasti, M.; Björk, H.; Vanninen, P. On-Flow Pulsed field gradient heteronuclear correlation spectrometry in off-Line LC-SPE-NMR analysis of chemicals related to the chemical weapons convention. Anal. Chem. 2009, 81, 1262–1269. [Google Scholar]
- Seger, C.; Godejohann, M.; Spraul, M.; Stuppner, H.; Hadacek, F.J. Reaction product analysis by high-performance liquid chromatography-solid-phase extraction-nuclear magnetic resonance: Application to the absolute configuration determination of naturally occurring polyyne alcohols. J. Chromatogr. A 2006, 1136, 82–88. [Google Scholar] [CrossRef]
- Koch, K.; Podlech, J.; Pfeiffer, E.; Metzler, M. Total synthesis of alternariol. J. Org. Chem. 2005, 70, 3275–3276. [Google Scholar] [CrossRef]
- Tan, N.; Tao, Y.; Pan, J.; Wang, S.; Xu, F.; She, Z.; Lin, Y.; Jones, E.B.G. Isolation, structure elucidation, and mutagenicity of four alternariol derivatives produced by the mangrove endophytic fungus N. 2240. Chem. Nat. Comp. 2008, 44, 296–300. [Google Scholar] [CrossRef]
- Bradburn, N.; Coker, R.D.; Blunden, G.; Turner, C.H.; Crabb, T.A. 5'-Epialtenuene and neoaltenuene, dibenzo-α-pyrones from Alternaria alternata cultured on rice. Phytochemistry 1994, 35, 665–669. [Google Scholar] [CrossRef]
- Andersen, B.; Kroger, E.; Roberts, R.G. Chemical and morphological segregation of Alternaria arborescens, A. infectoria and A. tenuissima species-groups. Mycol. Res. 2002, 106, 170–182. [Google Scholar] [CrossRef]
- Meng, X.; Mao, Z.; Lou, J.; Xu, L.; Zhong, L.; Peng, Y.; Zhou, L.; Wang, M. Benzopyranones from the endophytic fungus Hyalodendriella sp. Ponipodef12 and their bioactivities. Molecules 2012, 17, 11303–11314. [Google Scholar] [CrossRef]
- Brugger, E.-M.; Wagner, J.; Schumacher, D.M.; Koch, K.; Podlech, J.; Metzler, M.; Lehmann, L. Mutagenicity of the mycotoxin alternariol in cultured mammalian cells. Toxicol. Lett. 2006, 164, 221–230. [Google Scholar] [CrossRef]
- Aly, A.H.; Edrada-Ebel, R.; Indriani, I.D.; Wray, V.; Müller, W.E.G.; Totzke, F.; Zirrgiebel, U.; Schächtele, C.; Kubbutat, M.H.G.; Lin, W.H.; et al. Cytotoxic metabolites from the fungal endophyte Alternaria sp. and their subsequent detection in its host plant Polygonum senegalense. J. Nat. Prod. 2008, 71, 972–980. [Google Scholar] [CrossRef]
- Nakanishi, S.; Toki, S.; Saytoh, Y.; Tsukuda, E.; Kawahara, K.; Ando, K.; Matsuda, Y. Isolation of myosin light chain kinase inhibitors from microorganisms: Dehydroaltenusin, altenusin, atrovenetinone, and cyclooctasulfur. Biosci. Biotech. Biochem. 1995, 59, 1333–1335. [Google Scholar] [CrossRef]
- Robbana-Barnat, S.; Lafarge-Frayssinet, C.; Frayssinet, C. Use of cell cultures for predicting the biological effects of mycotoxins. Cell Biol. Toxicol. 1989, 5, 217–226. [Google Scholar] [CrossRef]
- Kuramata, M.; Fujioka, S.; Shimada, A.; Kawano, T.; Kimura, Y. Citrinolactones A, B and C, and sclerotinin C, plant growth regulators from Penicillium citrinum. Biosci. Biotech. Biochem. 2007, 71, 499–503. [Google Scholar] [CrossRef]
- Naganuma, M.; Nishida, M.; Kuramochi, K.; Sugawara, F.; Yoshida, H.; Mizushina, Y. 1-Deoxyrubralactone, a novel specific inhibitor of families X and Y of eukaryotic DNA polymerases from a fungal strain derived from sea algae. Bioorg. Med. Chem. 2008, 16, 2939–2944. [Google Scholar]
- Lehmann, L.; Wagner, J.; Metzler, M. Estrogenic and clastogenic potential of the mycotoxin alternariol in cultured mammalian cells. Food Chem. Toxicol. 2006, 44, 398–408. [Google Scholar] [CrossRef]
- Fliegmann, J.; Schüler, G.; Boland, W.; Ebel, J.; Mithöfer, A. The role of octadecanoids and functional mimics in soybean defence response. Biol. Chem. 2003, 384, 437–446. [Google Scholar]
- Sample Availability: Samples of the compounds alternariol and alternariol monomethyl ether are available from the authors.
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
De Souza, G.D.; Mithöfer, A.; Daolio, C.; Schneider, B.; Rodrigues-Filho, E. Identification of Alternaria alternata Mycotoxins by LC-SPE-NMR and Their Cytotoxic Effects to Soybean (Glycine max) Cell Suspension Culture. Molecules 2013, 18, 2528-2538. https://doi.org/10.3390/molecules18032528
De Souza GD, Mithöfer A, Daolio C, Schneider B, Rodrigues-Filho E. Identification of Alternaria alternata Mycotoxins by LC-SPE-NMR and Their Cytotoxic Effects to Soybean (Glycine max) Cell Suspension Culture. Molecules. 2013; 18(3):2528-2538. https://doi.org/10.3390/molecules18032528
Chicago/Turabian StyleDe Souza, Gezimar D., Axel Mithöfer, Cristina Daolio, Bernd Schneider, and Edson Rodrigues-Filho. 2013. "Identification of Alternaria alternata Mycotoxins by LC-SPE-NMR and Their Cytotoxic Effects to Soybean (Glycine max) Cell Suspension Culture" Molecules 18, no. 3: 2528-2538. https://doi.org/10.3390/molecules18032528
APA StyleDe Souza, G. D., Mithöfer, A., Daolio, C., Schneider, B., & Rodrigues-Filho, E. (2013). Identification of Alternaria alternata Mycotoxins by LC-SPE-NMR and Their Cytotoxic Effects to Soybean (Glycine max) Cell Suspension Culture. Molecules, 18(3), 2528-2538. https://doi.org/10.3390/molecules18032528






