Bioinformatic Analysis of Differential Protein Expression in Calu-3 Cells Exposed to Carbon Nanotubes
Abstract
:1. Introduction
2. Experimental
2.1. Materials
2.2. CNT Preparation
2.3. Cell Culture and Incubation
2.4. Proteomics
2.5. Bioinformatic Analysis
3. Results and Discussion
3.1. Proteomics
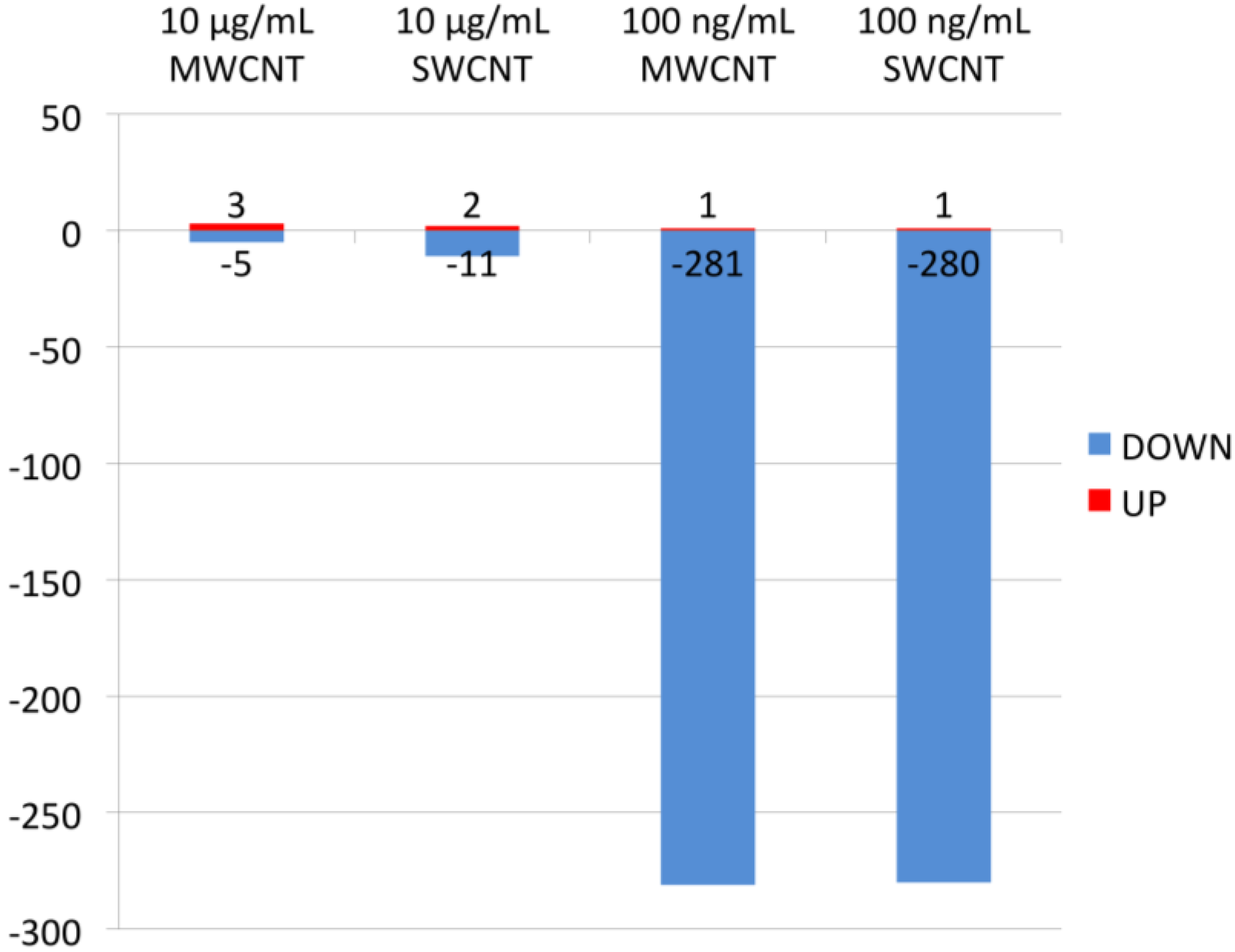
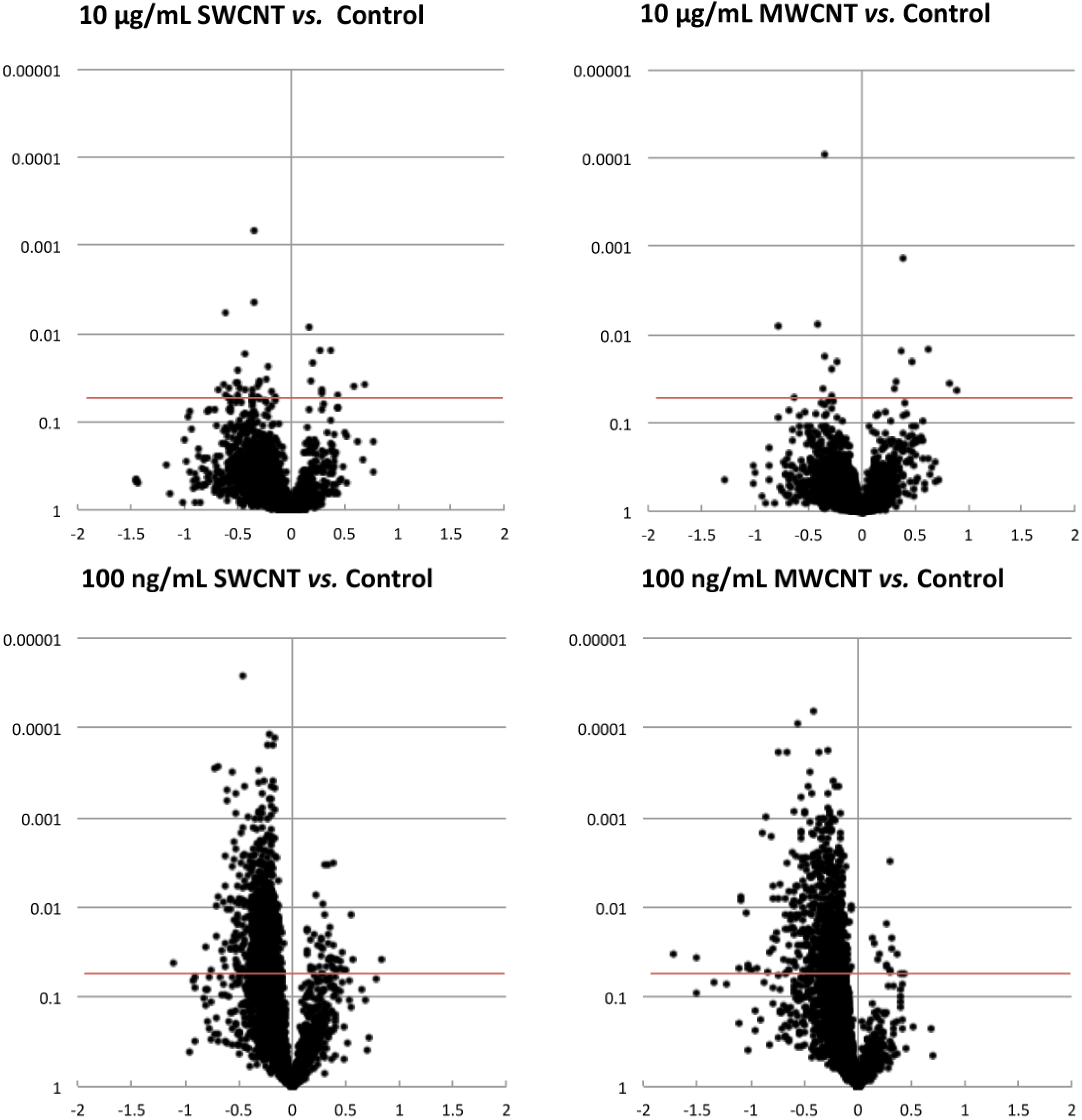
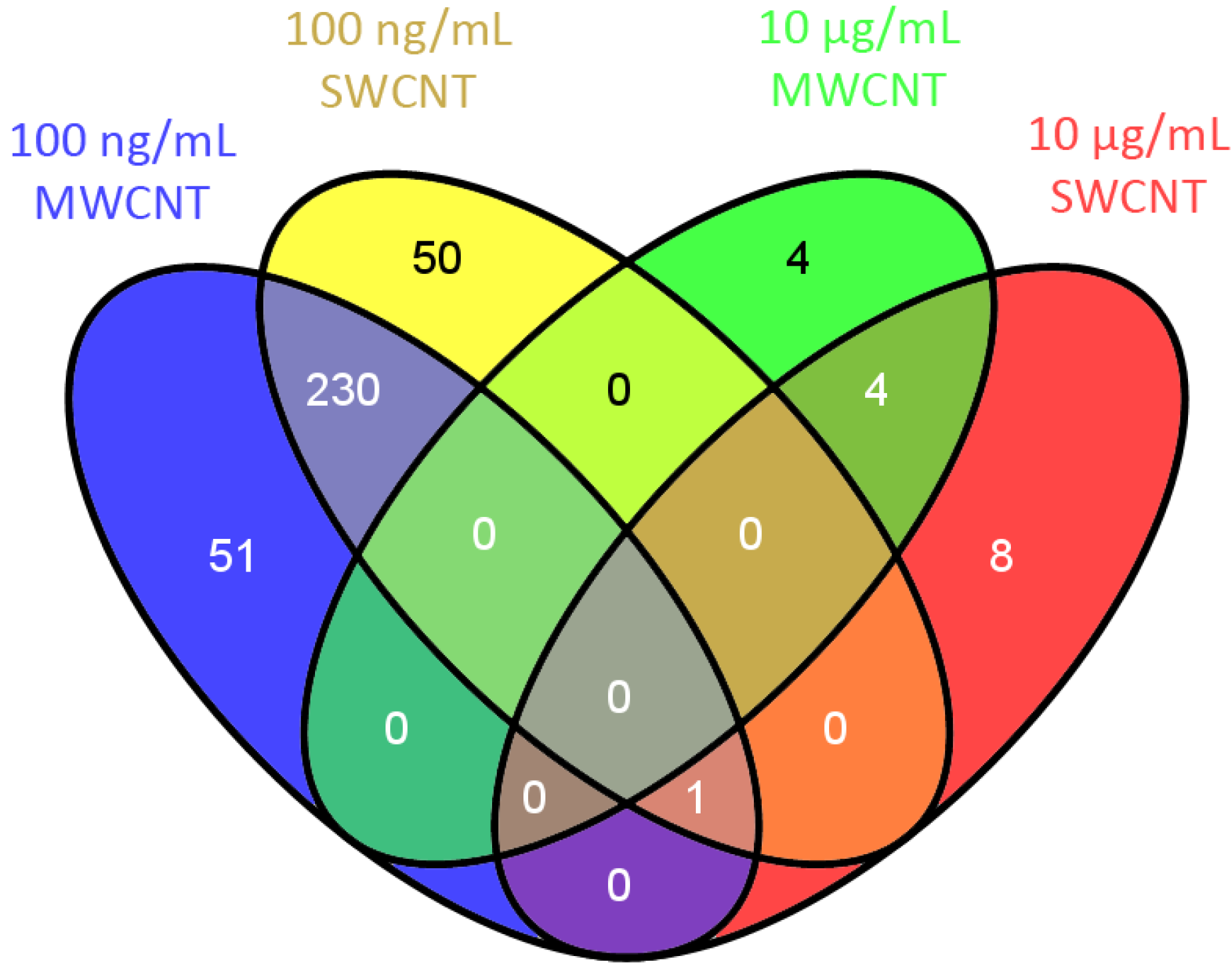
3.2. Bioinformatics
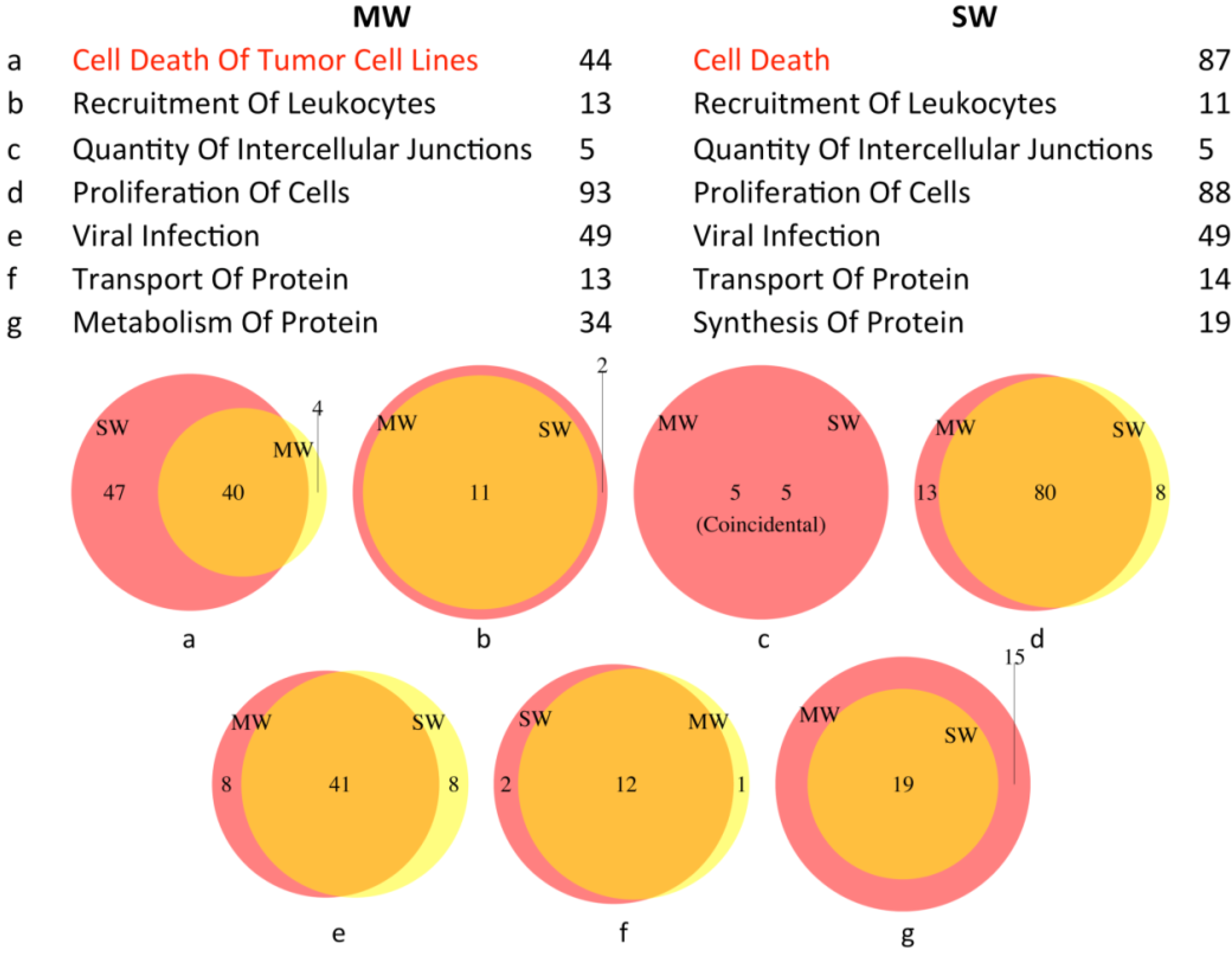
3.2.1. Upstream Regulators
3.2.2. Canonical Pathways
| Ingenuity Canonical Pathways | −log(p-value) | Ratio | Molecules | Categories | Top Functions and Diseases |
|---|---|---|---|---|---|
| Glycogen Degradation III | 5.07 | 44.40% | GAA, PYGB, TYMP, MTAP | Glycogen Degradation | Developmental Disorder; Hereditary Disorder; Metabolic Disease |
| Glycogen Degradation II | 3.63 | 37.50% | PYGB, TYMP, MTAP | Glycogen Degradation | Developmental Disorder; Hereditary Disorder; Metabolic Disease |
| RAN Signaling | 4.07 | 26.70% | KPNB1, KPNA2, XPO1, IPO5 | Cellular Growth, Proliferation and Development | Cell Signaling; DNA Replication, Recombination, and Repair; Nucleic Acid Metabolism |
| Bile Acid Biosynthesis, Neutral Pathway | 2.15 | 25.00% | AKR1C1/AKR1C2, SCP2 | Sterol Biosynthesis | Endocrine System Development and Function; Energy Production; Lipid Metabolism |
| Methylglyoxal Degradation III | 2.04 | 22.20% | AKR7A2, AKR1C1/AKR1C2 | Aldehyde Degradation | Endocrine System Development and Function; Energy Production; Lipid Metabolism |
| Telomere Extension by Telomerase | 2.76 | 20.00% | XRCC6, HNRNPA2B1, XRCC5 | Apoptosis; Cancer | Cellular Assembly and Organization; Cellular Function and Maintenance; DNA Replication, Recombination, and Repair |
3.2.3. Biological Functions
3.3. Protein Interaction Networks
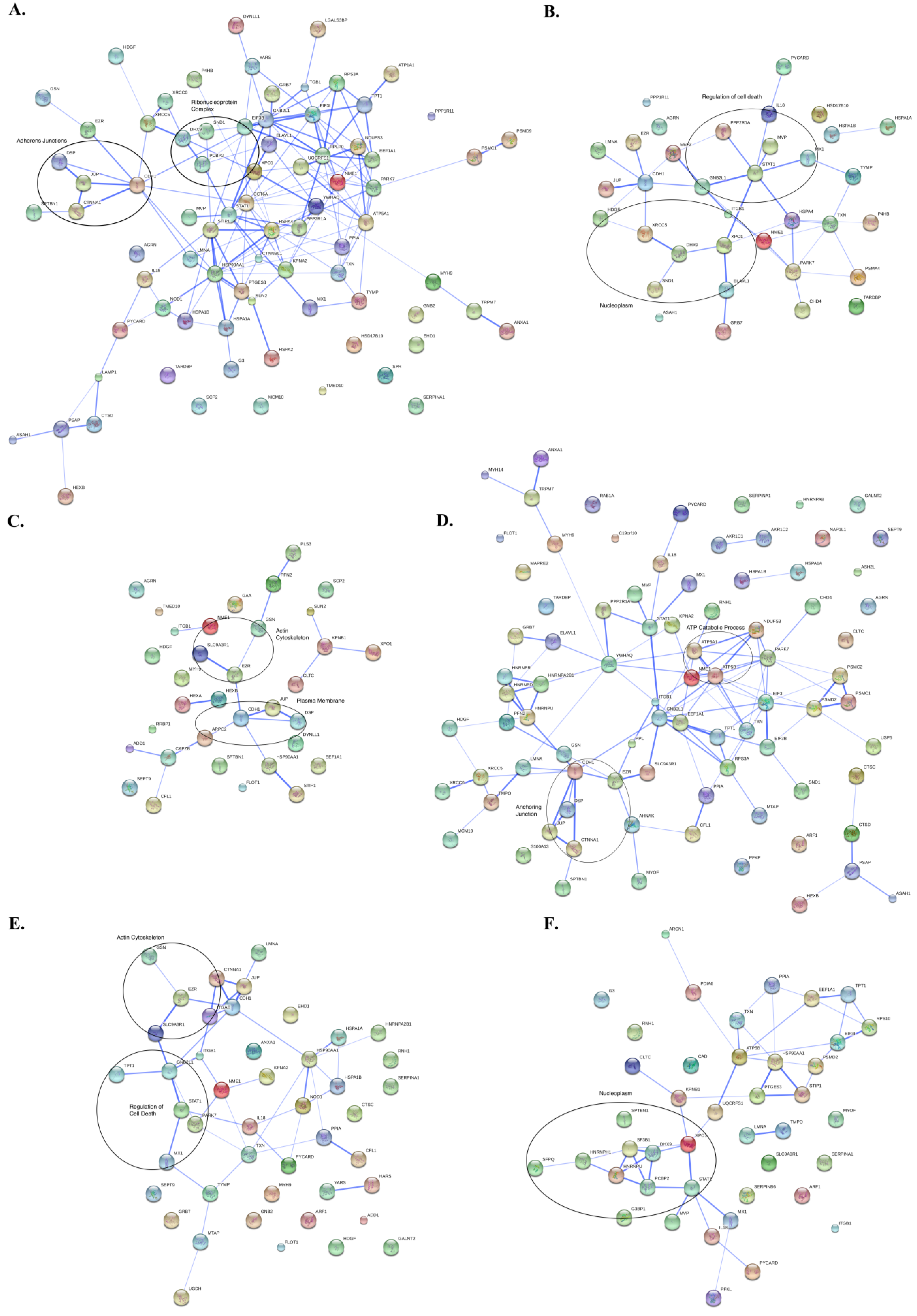
4. Discussion of Method
5. Conclusions
Supplementary Materials
Supplementary File 1Acknowledgments
Conflicts of Interest
References
- Buzea, C.; Pacheco, I.I.; Robbie, K. Nanomaterials and nanoparticles: Sources and toxicity. Biointerphases 2007, 2, MR17–MR71. [Google Scholar] [CrossRef]
- Correa-Duarte, M.A.; Wagner, N.; Rojas-Chapana, J.; Morsczeck, C.; Thie, M.; Giersig, M. Fabrication and biocompatibility of carbon nanotube-based 3D networks as scaffolds for cell seeding and growth. Nano Lett. 2004, 4, 2233–2236. [Google Scholar] [CrossRef]
- Moghimi, S.M.; Hunter, A.C.; Murray, J.C. Nanomedicine: Current status and future prospects. FASEB J. 2005, 19, 311–330. [Google Scholar] [CrossRef]
- Wong, S.K.N.; Dai, H. Single walled carbon nanotubes for transport and delivery of biological cargos. Physica Status Solidi B 2006, 243, 3561–3566. [Google Scholar] [CrossRef]
- Cui, H.F.; Vashist, S.K.; Al-Rubeaan, K.; Luong, J.H.; Sheu, F.S. Interfacing carbon nanotubes with living mammalian cells and cytotoxicity issues. Chem. Res. Toxicol. 2010, 23, 1131–1147. [Google Scholar] [CrossRef]
- Muller, J.; Huaux, F.; Moreau, N.; Misson, P.; Heilier, J.F.; Delos, M.; Arras, M.; Fonseca, A.; Nagy, J.B.; Lison, D. Respiratory toxicity of multi-wall carbon nanotubes. Toxicol. Appl. Pharmacol. 2005, 207, 221–231. [Google Scholar] [CrossRef]
- Zhu, Y.; Chidekel, A.; Shaffer, T.H. Cultured human airway epithelial cells (calu-3): A model of human respiratory function, structure, and inflammatory responses. Crit. Care Res. Pract. 2010. [Google Scholar] [CrossRef]
- Powell, D.W. Barrier function of epithelia. Am. J. Physiol. 1981, 241, G275–G288. [Google Scholar]
- Banga, A.; Witzmann, F.A.; Petrache, H.I.; Blazer-Yost, B.L. Functional effects of nanoparticle exposure on calu-3 airway epithelial cells. Cell. Physiol. Biochem. 2012, 29, 197–212. [Google Scholar] [CrossRef]
- Blazer-Yost, B.L.; Banga, A.; Amos, A.; Chernoff, E.; Lai, X.; Li, C.; Mitra, S.; Witzmann, F.A. Effect of carbon nanoparticles on renal epithelial cell structure, barrier function, and protein expression. Nanotoxicology 2011, 5, 354–371. [Google Scholar] [CrossRef]
- Prince, K.L.; Colvin, S.C.; Park, S.; Lai, X.; Witzmann, F.A.; Rhodes, S.J. Developmental analysis and influence of genetic background on the lhx3 w227ter mouse model of combined pituitary hormone deficiency disease. Endocrinology 2013, 154, 738–748. [Google Scholar] [CrossRef]
- Lai, X.; Blazer-Yost, B.L.; Clack, J.W.; Fears, S.L.; Mitra, S.; Ntim, S.A.; Ringham, H.N.; Witzmann, F.A. Protein expression profiles of intestinal epithelial co-cultures: Effect of functionalised carbon nanotube exposure. Int. J. Biomed. Nanosci. Nanotechnol. 2013, 3, 127–162. [Google Scholar] [CrossRef]
- Lai, X.; Agarwal, M.; Lvov, Y.M.; Pachpande, C.; Varahramyan, K.; Witzmann, F.A. Proteomic profiling of halloysite clay nanotube exposure in intestinal cell co-culture. J. Appl. Toxicol. 2013, 33, 1316–1329. [Google Scholar]
- Vidanapathirana, A.K.; Lai, X.; Hilderbrand, S.C.; Pitzer, J.E.; Podila, R.; Sumner, S.J.; Fennell, T.R.; Wingard, C.J.; Witzmann, F.A.; Brown, J.M. Multi-walled carbon nanotube directed gene and protein expression in cultured human aortic endothelial cells is influenced by suspension medium. Toxicology 2012, 302, 114–122. [Google Scholar] [CrossRef]
- Lai, X.; Bacallao, R.L.; Blazer-Yost, B.L.; Hong, D.; Mason, S.B.; Witzmann, F.A. Characterization of the renal cyst fluid proteome in autosomal dominant polycystic kidney disease (adpkd) patients. Proteomics Clin. Appl. 2008, 2, 1140–1152. [Google Scholar] [CrossRef]
- Keller, A.; Nesvizhskii, A.I.; Kolker, E.; Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by ms/ms and database search. Anal. Chem. 2002, 74, 5383–5392. [Google Scholar] [CrossRef]
- Nesvizhskii, A.I.; Keller, A.; Kolker, E.; Aebersold, R. A statistical model for identifying proteins by tandem mass spectrometry. Anal. Chem. 2003, 75, 4646–4658. [Google Scholar] [CrossRef]
- Seattle Proteome Center (SPC). The Trans-Proteomic Pipeline (TPP, v. 3.3.0). Available online: http://tools.proteomecenter.org/software.php (accessed on 25 July 2013).
- Lai, X.; Wang, L.; Tang, H.; Witzmann, F.A. A novel alignment method and multiple filters for exclusion of unqualified peptides to enhance label-free quantification using peptide intensity in LC-MS/MS. J. Proteome Res. 2011, 10, 4799–4812. [Google Scholar] [CrossRef]
- Monroe, M.E.; Shaw, J.L.; Daly, D.S.; Adkins, J.N.; Smith, R.D. Masic: A software program for fast quantitation and flexible visualization of chromatographic profiles from detected LC-MS(/MS) features. Comput. Biol. Chem. 2008, 32, 215–217. [Google Scholar] [CrossRef]
- Storey, J.D. A direct approach to false discovery rates. J. R. Stat. Soc. Series B Stat. Methodol. 2002, 64, 479–498. [Google Scholar] [CrossRef]
- Ingenuity Systems. The Ingenuity Pathway Analysis (IPA) web based software. Available online: http://www.ingenuity.com (accessed on 23 September 2013).
- STRING. Available online: http://string-db.org/ (accessed on 3 March 2013).
- Szklarczyk, D.; Franceschini, A.; Kuhn, M.; Simonovic, M.; Roth, A.; Minguez, P.; Doerks, T.; Stark, M.; Muller, J.; Bork, P.; et al. The string database in 2011: Functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res. 2011, 39, D561–D568. [Google Scholar] [CrossRef]
- Oberdorster, G.; Oberdorster, E.; Oberdorster, J. Nanotoxicology: An emerging discipline evolving from studies of ultrafine particles. Environ. Health Perspect. 2005, 113, 823–839. [Google Scholar] [CrossRef]
- Ge, C.; Du, J.; Zhao, L.; Wang, L.; Liu, Y.; Li, D.; Yang, Y.; Zhou, R.; Zhao, Y.; Chai, Z.; et al. Binding of blood proteins to carbon nanotubes reduces cytotoxicity. Proc. Natl. Acad. Sci. USA 2011, 108, 16968–16973. [Google Scholar]
- Shannahan, J.H.; Brown, J.M.; Chen, R.; Ke, P.C.; Lai, X.; Mitra, S.; Witzmann, F.A. Comparison of nanotube-protein corona composition in cell culture media. Small 2013, 9, 2171–2181. [Google Scholar] [CrossRef]
- Monopoli, M.P.; Åberg, C.; Salvati, A.; Dawson, K.A. Biomolecular coronas provide the biological identity of nanosized materials. Nat. Nanotechnol. 2012, 7, 779–786. [Google Scholar] [CrossRef]
- Kam, N.W.; Dai, H. Carbon nanotubes as intracellular protein transporters: Generality and biological functionality. J. Am. Chem. Soc. 2005, 127, 6021–6026. [Google Scholar] [CrossRef]
- Rollins, B.M.; Burn, M.; Coakley, R.D.; Chambers, L.A.; Hirsh, A.J.; Clunes, M.T.; Lethem, M.I.; Donaldson, S.H.; Tarran, R. A2b adenosine receptors regulate the mucus clearance component of the lung’s innate defense system. Am. J. Resp. Cell Mol. 2008, 39, 190–197. [Google Scholar] [CrossRef]
- Kawkitinarong, K.; Linz-McGillem, L.; Birukov, K.G.; Garcia, J.G. Differential regulation of human lung epithelial and endothelial barrier function by thrombin. Am. J. Resp. Cell Mol. 2004, 31, 517–527. [Google Scholar] [CrossRef]
- Weis, K. Importins and exportins: How to get in and out of the nucleus. Trends Biochem. Sci. 1998, 23, 185–189. [Google Scholar] [CrossRef]
- Lee, J.C.; Jong, H.S.; Yoo, C.G.; Han, S.K.; Shim, Y.S.; Kim, Y.W. Telomerase activity in lung cancer cell lines and tissues. Lung Cancer 1998, 21, 99–103. [Google Scholar] [CrossRef]
- Chen, Y.; Qu, K.; Zhao, C.; Wu, L.; Ren, J.; Wang, J.; Qu, X. Insights into the biomedical effects of carboxylated single-wall carbon nanotubes on telomerase and telomeres. Nat. Commun. 2012, 3, e1074. [Google Scholar] [CrossRef]
- Kultz, D. Molecular and evolutionary basis of the cellular stress response. Annu. Rev. Physiol. 2005, 67, 225–257. [Google Scholar] [CrossRef]
- Holt, B.D.; Short, P.A.; Rape, A.D.; Wang, Y.L.; Islam, M.F.; Dahl, K.N. Carbon nanotubes reorganize actin structures in cells and ex vivo. ACS Nano 2010, 4, 4872–4878. [Google Scholar] [CrossRef]
- Zahm, J.M.; Kaplan, H.; Herard, A.L.; Doriot, F.; Pierrot, D.; Somelette, P.; Puchelle, E. Cell migration and proliferation during the in vitro wound repair of the respiratory epithelium. Cell Motil. Cytoskel. 1997, 37, 33–43. [Google Scholar] [CrossRef]
- Chun, J.; Prince, A. Tlr2-induced calpain cleavage of epithelial junctional proteins facilitates leukocyte transmigration. Cell Host Microbe 2009, 5, 47–58. [Google Scholar] [CrossRef]
- Chin, J.E.; Hatfield, C.A.; Winterrowd, G.E.; Brashler, J.R.; Vonderfecht, S.L.; Fidler, S.F.; Griffin, R.L.; Kolbasa, K.P.; Krzesicki, R.F.; Sly, L.M.; et al. Airway recruitment of leukocytes in mice is dependent on alpha4-integrins and vascular cell adhesion molecule-1. Am. J. Physiol. 1997, 272, L219–L229. [Google Scholar]
- Hudder, A.; Nathanson, L.; Deutscher, M.P. Organization of mammalian cytoplasm. Mol. Cell. Biol. 2003, 23, 9318–9326. [Google Scholar] [CrossRef]
- Barabási, A.-L.; Bonabeau, E. Scale-free networks. Sci. Am. 2003, 288, 50–59. [Google Scholar]
- Delprato, A. Topological and functional properties of the small gtpases protein interaction network. PLoS One 2012, 7, e44882. [Google Scholar] [CrossRef]
- Fievet, B.; Louvard, D.; Arpin, M. Erm proteins in epithelial cell organization and functions. Biochim. Biophys. Acta 2007, 1773, 653–660. [Google Scholar] [CrossRef]
- Srinivasula, S.M.; Poyet, J.L.; Razmara, M.; Datta, P.; Zhang, Z.; Alnemri, E.S. The pyrin-card protein asc is an activating adaptor for caspase-1. J. Biol. Chem. 2002, 277, 21119–21122. [Google Scholar]
- Maier, T.; Güell, M.; Serrano, L. Correlation of mrna and protein in complex biological samples. FEBS Lett. 2009, 583, 3966–3973. [Google Scholar] [CrossRef]
- Yeung, E.S. Genome‐wide correlation between mrna and protein in a single cell. Angew. Chem. Int. Ed. Engl. 2011, 50, 583–585. [Google Scholar] [CrossRef]
- Tilton, S.C.; Karin, N.; Tolic, A.; Xie, Y.; Lai, X.; Hamilton, R.F.; Waters, K.; Holian, A.; Witzmann, F.A.; Orr, G. Three human cell types respond to multi-walled carbon nanotubes and titanium dioxide nanobelts with cell-specific transcriptomic and proteomic expression patterns. Nanotoxicology 2013. [Google Scholar] [CrossRef] [Green Version]
- Aebersold, R.; Burlingame, A.L.; Bradshaw, R.A. Western blots vs. Srm assays: Time to turn the tables? Mol. Cell. Proteomics 2013, 12, 2381–2382. [Google Scholar] [CrossRef]
- Shannahan, J.H.; Lai, X.; Ke, P.C.; Podila, R.; Brown, J.M.; Witzmann, F.A. Silver nanoparticle protein corona composition in cell culture media. PLoS One 2013, 8, e74001. [Google Scholar]
- Dewey, S.; Lai, X.; Witzmann, F.A.; Sohal, M.; Gomes, A.V. Proteomic analysis of hearts from akita mice suggests that increases in soluble epoxide hydrolase and antioxidative programming are key changes in early stages of diabetic cardiomyopathy. J. Proteome Res. 2013, 12, 3920–3933. [Google Scholar] [CrossRef]
- Lai, X.; Wang, L.; Witzmann, F.A. Issues and applications in label-free quantitative mass spectrometry. Int. J. Proteomics 2013. [Google Scholar] [CrossRef]
- Owen, M.K.; Witzmann, F.A.; McKenney, M.L.; Lai, X.; Berwick, Z.C.; Moberly, S.P.; Alloosh, M.; Sturek, M.; Tune, J.D. Perivascular adipose tissue potentiates contraction of coronary vascular smooth muscle: Influence of obesity. Circulation 2013, 128, 9–18. [Google Scholar]
- Koehler, G.; Wilson, R.C.; Goodpaster, J.V.; Sonsteby, A.; Lai, X.; Witzmann, F.A.; You, J.S.; Rohloff, J.; Randall, S.K.; Alsheikh, M. Proteomic study of low-temperature responses in strawberry cultivars (fragaria × ananassa) that differ in cold tolerance. Plant Physiol. 2012, 159, 1787–1805. [Google Scholar] [CrossRef] [Green Version]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Li, P.; Lai, X.; Witzmann, F.A.; Blazer-Yost, B.L. Bioinformatic Analysis of Differential Protein Expression in Calu-3 Cells Exposed to Carbon Nanotubes. Proteomes 2013, 1, 219-239. https://doi.org/10.3390/proteomes1030219
Li P, Lai X, Witzmann FA, Blazer-Yost BL. Bioinformatic Analysis of Differential Protein Expression in Calu-3 Cells Exposed to Carbon Nanotubes. Proteomes. 2013; 1(3):219-239. https://doi.org/10.3390/proteomes1030219
Chicago/Turabian StyleLi, Pin, Xianyin Lai, Frank A. Witzmann, and Bonnie L. Blazer-Yost. 2013. "Bioinformatic Analysis of Differential Protein Expression in Calu-3 Cells Exposed to Carbon Nanotubes" Proteomes 1, no. 3: 219-239. https://doi.org/10.3390/proteomes1030219
APA StyleLi, P., Lai, X., Witzmann, F. A., & Blazer-Yost, B. L. (2013). Bioinformatic Analysis of Differential Protein Expression in Calu-3 Cells Exposed to Carbon Nanotubes. Proteomes, 1(3), 219-239. https://doi.org/10.3390/proteomes1030219




