-
 SDS Depletion from Intact Membrane Proteins by KCl Precipitation Ahead of Mass Spectrometry Analysis
SDS Depletion from Intact Membrane Proteins by KCl Precipitation Ahead of Mass Spectrometry Analysis -
 Novel Serum Glycoprotein Biomarkers Clinically Validated as a Diagnostic Test for Esophageal Adenocarcinoma
Novel Serum Glycoprotein Biomarkers Clinically Validated as a Diagnostic Test for Esophageal Adenocarcinoma -
 Deciphering Radiotherapy Resistance: A Proteomic Perspective
Deciphering Radiotherapy Resistance: A Proteomic Perspective -
 The Proteomic Signature of Oxidative Stress in Myelodysplastic Syndromes
The Proteomic Signature of Oxidative Stress in Myelodysplastic Syndromes
Journal Description
Proteomes
Proteomes
is an international, peer-reviewed, open access journal on all aspects of proteomics published quarterly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, ESCI (Web of Science), PubMed, PMC, CAPlus / SciFinder, and other databases.
- Journal Rank: JCR - Q2 (Biochemistry and Molecular Biology) / CiteScore - Q1 (Structural Biology)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 28.2 days after submission; acceptance to publication is undertaken in 4.8 days (median values for papers published in this journal in the first half of 2025).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
Impact Factor:
3.6 (2024);
5-Year Impact Factor:
3.9 (2024)
Latest Articles
Marine Bioactive Peptides in the Regulation of Inflammatory Responses: Current Trends and Future Directions
Proteomes 2025, 13(4), 53; https://doi.org/10.3390/proteomes13040053 - 13 Oct 2025
Abstract
►
Show Figures
Marine-derived bioactive peptides (MBPs) are emerging as promising natural agents for regulating inflammatory responses. MBPs, typically obtained through enzymatic hydrolysis of proteins from various marine organisms such as fish, mollusks, and algae, exhibit diverse biological activities, including antioxidant, immunomodulatory, and anti-inflammatory effects. The
[...] Read more.
Marine-derived bioactive peptides (MBPs) are emerging as promising natural agents for regulating inflammatory responses. MBPs, typically obtained through enzymatic hydrolysis of proteins from various marine organisms such as fish, mollusks, and algae, exhibit diverse biological activities, including antioxidant, immunomodulatory, and anti-inflammatory effects. The ability of MBPs to modulate key inflammatory mediators such as TNF-α, IL-6, and COX-2, primarily through pathways like NF-κB and MAPK, highlights the therapeutic potential of MBPs in managing chronic inflammatory diseases. However, most existing studies are confined to in vitro assays or animal models, with limited translation to human clinical applications. This review explores the stability, bioavailability, and metabolic rate of MBPs under physiological conditions, which remain poorly understood. In addition, a lack of standardized protocols for peptide extraction, purification, and efficacy evaluation hinders comparative analysis across studies and also different proteomics approaches for separation, purification, identification, and quantification of marine-derived peptides with therapeutic properties. The structure–function relationship of MBPs is also underexplored, limiting rational design and targeted applications in functional foods or therapeutic products. These limitations are largely due to a lack of consolidated information and integrated research efforts. To address these challenges, this review summarizes recent progress in identifying MBPs with anti-inflammatory potentials, outlines key mechanisms, and highlights current limitations. Additionally, this review also emphasizes the need to enhance mechanistic understanding, optimize delivery strategies, and advance clinical validation to fully realize the therapeutic potential of MBPs.
Full article
Open AccessArticle
Temporal and Spatial Profiling of Escherichia coli O157:H7 Surface Proteome: Insights into Intestinal Colonisation Dynamics In Vivo
by
Ricardo Monteiro, Ingrid Chafsey, Charlotte Cordonnier, Valentin Ageorges, Didier Viala, Michel Hébraud, Valérie Livrelli, Alfredo Pezzicoli, Mariagrazia Pizza and Mickaël Desvaux
Proteomes 2025, 13(4), 52; https://doi.org/10.3390/proteomes13040052 - 10 Oct 2025
Abstract
►▼
Show Figures
Background: EHEC O157:H7 causes severe gastrointestinal illness by first colonizing the large intestine. It intimately attaches to the epithelial lining, orchestrating distinctive “attaching and effacing” lesions that disrupt the host’s cellular landscape. While much is known about the well-established virulence factors, there are
[...] Read more.
Background: EHEC O157:H7 causes severe gastrointestinal illness by first colonizing the large intestine. It intimately attaches to the epithelial lining, orchestrating distinctive “attaching and effacing” lesions that disrupt the host’s cellular landscape. While much is known about the well-established virulence factors, there are much to learn about the surface proteins’ roles in a living host. Methods: This study presents the first in vivo characterisation of the surface proteome, i.e., proteosurfaceome, of Escherichia coli O157:H7 EDL933 during intestinal infection, revealing spatial and temporal adaptations critical for colonisation and survival. Using a murine ileal loop model, surface proteomic profiles were analysed at early (3 h) and late (10 h) infection stages across the ileum and colon. Results: In total, 272 proteins were identified, with only 13 shared across all conditions, reflecting substantial niche-specific adaptations. Gene ontology enrichment analyses highlighted dominant roles in metabolic, cellular, and binding functions, while subcellular localisation prediction uncovered cytoplasmic moonlighting proteins with surface activity. Comparative analyses revealed dynamic changes in protein abundance. Conclusions: These findings indicate a coordinated shift from stress adaptation and virulence to nutrient acquisition and persistence and provide a comprehensive view of EHEC O157:H7 surface proteome dynamics during infection, highlighting key adaptive proteins that may serve as targets for future therapeutic and vaccine strategies.
Full article

Figure 1
Open AccessArticle
Protein-Predicted Obesity Phenotypes and Cardiovascular Events: A Secondary Analysis of UK Biobank Proteomics Data
by
Chang Liu, Bojung Seo, Qin Hui, Peter W. F. Wilson, Arshed A. Quyyumi and Yan V. Sun
Proteomes 2025, 13(4), 51; https://doi.org/10.3390/proteomes13040051 - 9 Oct 2025
Abstract
Background: Proteomic profiling may improve the understanding of obesity and cardiovascular risk prediction. This study explores the use of protein-predicted scores for body mass index (PPSBMI), body fat percentage (PPSBFP), and waist–hip ratio (PPSWHR) to estimate risk
[...] Read more.
Background: Proteomic profiling may improve the understanding of obesity and cardiovascular risk prediction. This study explores the use of protein-predicted scores for body mass index (PPSBMI), body fat percentage (PPSBFP), and waist–hip ratio (PPSWHR) to estimate risk for major adverse cardiovascular events (MACEs). Methods: We used data from the UK Biobank with proteome profiling. PPSBMI, PPSBFP, and PPSWHR were derived using the LASSO algorithm. The association between these protein scores and incident MACEs was evaluated using a competing risk model. Results: Strong to moderate correlations were observed between protein-predicted obesity phenotypes and their measured counterparts (R2: BMI = 0.78, BFP = 0.85, WHR = 0.63). Each standard deviation increment of PPSBFP and PPSWHR, but not PPSBMI, was associated with greater risk of MACEs (hazard ratio [HR] 1.25, 95% CI 1.14–1.38, p < 0.0001; HR 1.15, 95% CI 1.06–1.24, p = 0.001, respectively). For predicting MACEs, compared with the PREVENT equation (C statistic 0.694), the models adjusted for only age, sex, current smoking, and protein scores showed comparable performance (C statistics 0.684–0.688). Conclusion: Protein-predicted scores of obesity showed strong independent associations and predictive performance for MACEs, suggesting they may capture additional biological risk beyond anthropometry. These scores may complement existing risk models by providing a biologically informed approach to assessing obesity-related cardiovascular risk and improving risk stratification.
Full article
Open AccessArticle
Protein Structural Modeling Explains Rapid Oxidation in Poultry and Fish Myoglobins Compared to Livestock Myoglobins
by
Greeshma Sreejesh, Surendranath P. Suman, Gretchen G. Mafi, Morgan M. Pfeiffer and Ranjith Ramanathan
Proteomes 2025, 13(4), 50; https://doi.org/10.3390/proteomes13040050 - 8 Oct 2025
Abstract
►▼
Show Figures
Background: This study aimed to investigate rapid oxidation in poultry and fish myoglobin compared to livestock myoglobin using protein structural differences and bioinformatics tools. Methods: Myoglobins from beef (Bos taurus), bison (Bos bison), sheep (Ovis aries), goat
[...] Read more.
Background: This study aimed to investigate rapid oxidation in poultry and fish myoglobin compared to livestock myoglobin using protein structural differences and bioinformatics tools. Methods: Myoglobins from beef (Bos taurus), bison (Bos bison), sheep (Ovis aries), goat (Capra hircus), red deer (Cervus elaphus), pork (Sus scrofa), chicken (Gallus gallus), turkey (Meleagris gallopavo), yellowfin tuna (Thunnus albacares), and tilapia (Oreochromis niloticus) were analyzed to understand differences in structure and function that may influence oxidative behavior. Results: Fish and poultry had shorter or absent D-helix in their myoglobin structure than other species. Tilapia showed the largest heme cavity surface area, indicating significant internal void space, while yellowfin tuna had the largest heme cavity volume, which could affect ligand binding dynamics compared with poultry and other livestock species. However, the heme solvent-accessible area was greater in chicken and turkey than in fish and other livestock species. Tuna myoglobin contains a cysteine and fish myoglobins have fewer amino acids compared to other species. Limited knowledge is currently available on the effects of proteoform, especially post-translational modifications, on the oxidation of myoglobin from different species. Conclusions: The bioinformatics approach used in this study suggests that, in addition to physiological reasons, shorter D-helix, larger heme cavity in tilapia and yellowfin tuna, and greater solvent-accessible area in poultry contribute to increased oxidation in myoglobin from poultry and fish compared with myoglobin from livestock species.
Full article

Figure 1
Open AccessReview
Extracellular Vesicle (EV) Proteomics in Corneal Regenerative Medicine
by
Zohreh Arabpour, Hanieh Niktinat, Firouze Hatami, Amal Yaghmour, Zarife Jale Yucel, Seyyedehfatemeh Ghalibafan, Hamed Massoumi, Zahra Bibak Bejandi, Majid Salehi, Elmira Jalilian, Mahmood Ghassemi, Victor H. Guaiquil, Mark Rosenblatt and Ali R. Djalilian
Proteomes 2025, 13(4), 49; https://doi.org/10.3390/proteomes13040049 - 3 Oct 2025
Abstract
Corneal regeneration has gained growing interest in recent years, largely due to the limitations of conventional treatments and the persistent shortage of donor tissue. Among the emerging strategies, extracellular vehicles (EVs), especially those derived from mesenchymal stromal cells (MSCs), have shown great promise
[...] Read more.
Corneal regeneration has gained growing interest in recent years, largely due to the limitations of conventional treatments and the persistent shortage of donor tissue. Among the emerging strategies, extracellular vehicles (EVs), especially those derived from mesenchymal stromal cells (MSCs), have shown great promise as a cell-free therapeutic approach. These nanoscale vesicles contribute to corneal healing by modulating inflammation, supporting epithelial and stromal regeneration, and promoting nerve repair. Their therapeutic potential is largely attributed to the diverse and bioactive proteomic cargo they carry, including growth factors, cytokines, and proteins involved in extracellular matrix remodeling. This review presents a comprehensive examination of the proteomic landscape of EVs in the context of corneal regenerative medicine. We explore the biological functions of EVs in corneal epithelial repair, stromal remodeling, and neurodegeneration. In addition, we discuss advanced proteomic profiling techniques such as mass spectrometry (MS) and liquid chromatography–mass spectrometry (LC-MS/MS), which have been used to identify and characterize the protein contents of EVs. This review also compares the proteomic profiles of EVs derived from various MSC sources, including adipose tissue, bone marrow, and umbilical cord, and considers how environmental cues, such as hypoxia and inflammation, influence their protein composition. By consolidating current findings, this article aims to provide valuable insights for advancing the next generation of cell-free therapies for corneal repair and regeneration.
Full article
(This article belongs to the Topic Multi-Omics in Precision Medicine)
►▼
Show Figures

Figure 1
Open AccessArticle
Proteomic Characterization of Primary Human Pancreatic Cancer Cell Lines Following Long-Term Exposure to Gemcitabine
by
Manoj Amrutkar, Yuchuan Li, Anette Vefferstad Finstadsveen, Caroline S. Verbeke and Ivar P. Gladhaug
Proteomes 2025, 13(4), 48; https://doi.org/10.3390/proteomes13040048 - 1 Oct 2025
Abstract
Background: Gemcitabine (GEM) remains a cornerstone in the treatment of pancreatic cancer. Upon exposure to GEM, pancreatic cancer cells (PCCs) tend to adapt quickly to outcompete drug-induced cytotoxicity, thereby contributing to treatment failure. Thus, understanding GEM-induced molecular changes in PCCs is important. Methods:
[...] Read more.
Background: Gemcitabine (GEM) remains a cornerstone in the treatment of pancreatic cancer. Upon exposure to GEM, pancreatic cancer cells (PCCs) tend to adapt quickly to outcompete drug-induced cytotoxicity, thereby contributing to treatment failure. Thus, understanding GEM-induced molecular changes in PCCs is important. Methods: Three primary PCC lines (PCC-1, PCC-2, PCC-7) and Mia PaCa-2 cultured for 40 passages (p) in the absence (control) or presence of GEM (GemR) were assessed for phenotypic changes. Proteome profiles for all PCCs at p10, p20, p25, p30, p35, and p40 were obtained using mass spectrometry (MS). Protein expression was determined using immunoblotting. Differentially abundant proteins (DAPs) were evaluated for enrichment of functional and biological attributes and protein–protein interactions. Results: GEM sensitivity and growth were both reduced in GemR versus paired controls for all four PCC lines. MS mapped > 7000 proteins in each PCC line, and the abundance of 70–83% of these was found to be significantly altered when comparing all sample groups. Proteomic changes in GemR versus paired controls differed remarkably among the PCCs and were affected by passaging and treatment duration. DAPs at p40 were mostly related to metabolic pathways, including nucleotide metabolism and diverse cell growth processes. Several closely related DAPs and multiple hub proteins in each PCC line were identified. Conclusions: Overall, this study revealed cell-line-specific, heterogeneous changes in proteome profiles of PCCs following their long-term exposure to GEM, and these were likely affected by treatment duration, dosage, and passaging.
Full article
(This article belongs to the Special Issue Proteomics in Chronic Diseases: Issues and Challenges)
►▼
Show Figures

Graphical abstract
Open AccessReview
In Search of Ideal Solutions for Cancer Diagnosis: From Conventional Methods to Protein Biomarkers in Liquid Biopsy
by
Anca-Narcisa Neagu, Pathea S. Bruno, Claudiu-Laurentiu Josan, Natalie Waterman, Hailey Morrissiey, Victor T. Njoku and Costel C. Darie
Proteomes 2025, 13(4), 47; https://doi.org/10.3390/proteomes13040047 - 23 Sep 2025
Cited by 1
Abstract
►▼
Show Figures
Cancer detection has made significant progress, moving from conventional methods to innovative, non-invasive or minimally invasive approaches aimed at improving early diagnosis, precision, and treatment outcomes. This review examines current and emerging diagnostic technologies, including liquid biopsy (LB), molecular biomarkers, and artificial intelligence
[...] Read more.
Cancer detection has made significant progress, moving from conventional methods to innovative, non-invasive or minimally invasive approaches aimed at improving early diagnosis, precision, and treatment outcomes. This review examines current and emerging diagnostic technologies, including liquid biopsy (LB), molecular biomarkers, and artificial intelligence (AI). LB analyzes biomarkers in bodily fluids, showing promise in detecting tumors at molecular levels, monitoring cancer progression, and predicting treatment responses. The assignment of specific proteoforms, often linked to tumor subtype, stage, and therapy resistance, adds another layer of diagnostic precision, offering valuable insights for personalized oncology. However, the clinical application of LB faces challenges related to sensitivity, specificity, tumor heterogeneity, and a lack of standardized protocols. Relatively high costs, complex result interpretation, and privacy concerns also hinder its widespread adoption in clinical practice. Despite these challenges, advancements in AI, nanotechnology, and multi-omics strategies offer opportunities to enhance cancer diagnostic accuracy. Future developments, including wearable biosensors and lab-on-a-chip technologies, could lead to personalized, real-time cancer detection with improved patient outcomes, potentially redefining cancer care and fostering a more proactive, patient-centered healthcare approach.
Full article

Figure 1
Open AccessSystematic Review
Neurogranin as a Synaptic Biomarker in Mild Traumatic Brain Injury: A Systematic Review of Diagnostic and Pathophysiological Evidence
by
Ioannis Mavroudis, Foivos Petridis, Eleni Karantali and Dimitrios Kazis
Proteomes 2025, 13(3), 46; https://doi.org/10.3390/proteomes13030046 - 19 Sep 2025
Abstract
►▼
Show Figures
Neurogranin (NRGN), a synaptic protein essential for plasticity and memory function, is gaining recognition as a promising biomarker for mild traumatic brain injury (mTBI). This systematic review brings together findings from six studies that measured neurogranin levels in biofluids—including serum, cerebrospinal fluid (CSF),
[...] Read more.
Neurogranin (NRGN), a synaptic protein essential for plasticity and memory function, is gaining recognition as a promising biomarker for mild traumatic brain injury (mTBI). This systematic review brings together findings from six studies that measured neurogranin levels in biofluids—including serum, cerebrospinal fluid (CSF), plasma, and exosomes—during both the acute and chronic phases following injury. In the acute phase of mTBI, elevated levels of neurogranin were consistently observed in serum samples, suggesting its potential as a diagnostic marker. These increases appear to reflect immediate synaptic disturbances caused by injury. In contrast, studies focusing on the chronic phase reported a decrease in exosomal neurogranin levels, pointing to ongoing synaptic dysfunction well after the initial trauma. This temporal shift in neurogranin expression highlights its dual utility—both as an early indicator of injury and as a longer-term marker of synaptic integrity. However, interpreting these findings is not straightforward. The studies varied considerably in terms of sample type, timing of measurements, and control for potential confounding factors such as physical activity. Such variability makes direct comparisons difficult and may influence the outcomes observed. Additionally, none of the studies included proteoform-specific analyses of neurogranin, an omission that limits our understanding of the molecular changes underlying mTBI-related synaptic alterations. Due to heterogeneity across study designs and outcome measures, a meta-analysis could not be performed. Instead, a narrative synthesis was conducted, revealing consistent patterns in neurogranin dynamics over time and underscoring the influence of biofluid selection on measured outcomes. Overall, the current evidence supports neurogranin’s potential as both a diagnostic and mechanistic biomarker for mTBI. Yet, to fully realize its clinical utility, future research must prioritize standardized protocols, the inclusion of proteoform profiling, and rigorous longitudinal validation studies.
Full article

Figure 1
Open AccessArticle
Comparative Analysis of Plasma Extracellular Vesicle Isolation Methods for Purity Assessment and Biomarker Discovery
by
Alexandra T. Star, Melissa Hewitt, Amanpreet Badhwar, Wen Ding, Tammy-Lynn Tremblay, Jennifer J. Hill, William G. Willmore, Jagdeep K. Sandhu and Arsalan S. Haqqani
Proteomes 2025, 13(3), 45; https://doi.org/10.3390/proteomes13030045 - 18 Sep 2025
Abstract
Background: Extracellular vesicles (EVs) are an important source of blood biomarkers and are emerging as next-generation therapeutics. Demonstrating the purity of isolated EVs is essential for applications ranging from proteomics-based biomarker discovery to biomanufacturing. In this study, we systematically evaluated multiple EV isolation
[...] Read more.
Background: Extracellular vesicles (EVs) are an important source of blood biomarkers and are emerging as next-generation therapeutics. Demonstrating the purity of isolated EVs is essential for applications ranging from proteomics-based biomarker discovery to biomanufacturing. In this study, we systematically evaluated multiple EV isolation methods for plasma and developed a scoring method to identify the approach best suited for proteomics. Methods: Commonly used enrichment techniques, including size-exclusion chromatography (SEC) and precipitation-based methods, were compared against the starting plasma in terms of particle yield and size, proteomic overlap, depletion of abundant plasma proteins, and enrichment of EV markers and unique proteins. To enable rigorous purity assessment, we established a targeted parallel reaction monitoring (PRM) mass spectrometry assay that quantified key EV markers and contaminant proteins across preparations. Results: Among the methods tested, SEC showed the greatest enrichment of EV markers and unique proteins, with the lowest level of contaminants, resulting in the highest overall purity scores. SEC also allowed for the detection of EV-free proteins. Other methods, by contrast, performed sub-optimally and were less reliable for proteomics-driven biomarker discovery. Conclusions: SEC provides the most EV-enriched plasma isolates for proteomics information, with minimal contamination from plasma proteins. The PRM-based purity scoring offers an objective means of benchmarking EV preparations and may help standardize EV isolation quality for both biomarker discovery and therapeutic manufacturing.
Full article
(This article belongs to the Topic Liquid Biopsy: A Modern Method Transforming Biomedicine)
►▼
Show Figures
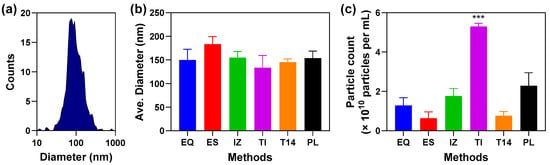
Figure 1
Open AccessArticle
Proteomic Analysis of Mechanical Injury Effects in Papaya Fruit at Two Maturity Stages
by
Francisco Antonio Reyes-Soria, Eliel Ruiz-May, Enrique Castaño, Miguel Ángel Herrera-Alamillo, José Miguel Elizalde-Contreras, Samuel David Gamboa-Tuz, Lidia F. E. Huerta-Nuñez, Jesús Alejandro Zamora-Briseño and Luis Carlos Rodríguez-Zapata
Proteomes 2025, 13(3), 44; https://doi.org/10.3390/proteomes13030044 - 18 Sep 2025
Abstract
Background: Mechanical damage to fruit during harvesting is nearly inevitable, with certain species, such as papaya, being particularly prone to spoilage. Postharvest handling can induce mechanical injuries that impair ripening and reduce shelf life, leading to significant economic losses. Although several studies have
[...] Read more.
Background: Mechanical damage to fruit during harvesting is nearly inevitable, with certain species, such as papaya, being particularly prone to spoilage. Postharvest handling can induce mechanical injuries that impair ripening and reduce shelf life, leading to significant economic losses. Although several studies have shed light on the molecular bases of mechanical damage, other aspects remain to be described (plant hormone inter-talk, physiological changes, and regulatory networks). Methods: In this study, we investigated proteomic changes in papaya fruit at two distinct ripening stages following mechanical damage. A total of 3230 proteins were identified, representing the most comprehensive proteomic analysis of papaya to date and the first assessment of proteins regulated by mechanical stress. Results: Proteins involved in ethylene biosynthesis were up-regulated on Day 2 but down-regulated on Day 12, with a similar trend observed for proteins in the abscisic acid synthesis pathway. Enzymes associated with photosynthesis, carbon fixation, primary metabolism, and carotenoid synthesis were down-regulated at both stages. In contrast, those related to plasmodesmata, calcium signaling, kinases, pathogenesis, cell wall remodeling, and proteases were up-regulated. Conclusions: These findings are thoroughly discussed, and a general model of the events triggered by mechanical impact in papaya is proposed. Our results provide a comprehensive framework for understanding papaya’s response to mechanical damage.
Full article
(This article belongs to the Section Proteome Bioinformatics)
►▼
Show Figures

Figure 1
Open AccessArticle
Proteomic Analysis of Sputum from Patients with Active Tuberculosis
by
Endrei Marcantonio, Amy M. Woron, A. Christian Whelen and Sladjana Prisic
Proteomes 2025, 13(3), 43; https://doi.org/10.3390/proteomes13030043 - 12 Sep 2025
Abstract
►▼
Show Figures
Background: Patients with pulmonary tuberculosis (TB) typically produce sputa, which are used to identify the pathogen. Sputum also contains host proteins that may aid in diagnosis. We hypothesized that sputa from TB patients will have unique proteomes when compared to other lung diseases.
[...] Read more.
Background: Patients with pulmonary tuberculosis (TB) typically produce sputa, which are used to identify the pathogen. Sputum also contains host proteins that may aid in diagnosis. We hypothesized that sputa from TB patients will have unique proteomes when compared to other lung diseases. Methods: Sputa were collected from 219 patients with suspected TB. Neutrophil-derived protein calprotectin (CP), which was used as a marker for lung damage, was quantified and compared between TB and non-TB groups. Three sputa with high or low CP from each group were selected and analyzed using label-free proteomics. Results: There was no difference in CP amounts between TB and non-TB groups. However, TB samples had other differentially abundant neutrophil-associated proteins. Compared to low CP, samples with high CP had much smaller number of proteins that could differentiate between TB and non-TB groups. Only two proteins, MUC5AC and MMP8, were more abundant in TB samples, regardless of CP levels. Conclusions: Our findings suggest that TB sputa may have unique proteomes that depend on CP levels, which should be further validated due to the small sample size. Therefore, controlled and more advanced TB may need a different set of biomarkers to reliably distinguish TB from other lung diseases.
Full article

Figure 1
Open AccessArticle
Proteomic Analysis of the Periodontal Ligament During Orthodontic Movement: A Study in Rats
by
Camila Chierici Marcantonio, Maria Eduarda Scordamaia Lopes, Lélio Fernando Ferreira Soares, Cristiane Ribeiro Salmon, Francisco Humberto Nociti Junior, James Deschner, Andressa Vilas Boas Nogueira and Joni Augusto Cirelli
Proteomes 2025, 13(3), 42; https://doi.org/10.3390/proteomes13030042 - 11 Sep 2025
Abstract
►▼
Show Figures
The periodontal ligament (PDL) is a dynamic connective tissue that absorbs and transmits mechanical forces, playing a critical role during orthodontic tooth movement (OTM). This study aimed to characterize the proteomic profile of rat PDLs subjected to OTM. Ten Holtzman rats were allocated
[...] Read more.
The periodontal ligament (PDL) is a dynamic connective tissue that absorbs and transmits mechanical forces, playing a critical role during orthodontic tooth movement (OTM). This study aimed to characterize the proteomic profile of rat PDLs subjected to OTM. Ten Holtzman rats were allocated into Control and OTM groups. After 15 days of force application, hemimaxillae were harvested, and PDL tissues from the first maxillary molars were isolated via laser capture microdissection. Protein extracts were analyzed using liquid chromatography–tandem mass spectrometry (LC-MS/MS), followed by quantitative and enrichment analyses. Immunohistochemistry was performed to validate selected proteins. The full proteomic datasets supporting these findings are available in the PRIDE repository under the identifiers PXD055817 and PXD033647. A total of 1121 proteins were identified; 101 were exclusive to the OTM group, 324 to the control, and 696 shared. Among the 335 proteins with differential abundance, 334 were downregulated and one (Prelp) was upregulated in the OTM group. Enrichment analysis revealed that differentially abundant proteins were associated with molecular functions such as protein binding, and cellular components including extracellular exosomes, focal adhesions, and the extracellular matrix. Immunohistochemical analysis confirmed the presence of Prelp, Rbm3, and Cirbp in PDL tissues. These findings demonstrate that OTM significantly alters the proteomic landscape of the PDL and identify key proteins potentially involved in periodontal remodeling.
Full article

Figure 1
Open AccessArticle
Bidirectional Interaction Between PGE2-Preconditioned Mesenchymal Stem Cells and Myofibroblasts Mediates Anti-Fibrotic Effects: A Proteomic Investigation into Equine Endometrial Fibrosis Reversal
by
Lidice Méndez-Pérez, Yat Sen Wong, Belén O. Ibáñez, Ioanna Martinez-Hormaza, Lleretny Rodríguez-Álvarez and Fidel Ovidio Castro
Proteomes 2025, 13(3), 41; https://doi.org/10.3390/proteomes13030041 - 8 Sep 2025
Abstract
►▼
Show Figures
Background: Endometrosis is a prevalent fibrotic condition in mares that impairs reproductive efficiency by inducing transdifferentiation of endometrial stromal cells into myofibroblasts, leading to excessive ECM deposition. Methods: To elucidate the molecular mechanisms underlying fibrosis resolution, this study employed comprehensive proteomic techniques, including
[...] Read more.
Background: Endometrosis is a prevalent fibrotic condition in mares that impairs reproductive efficiency by inducing transdifferentiation of endometrial stromal cells into myofibroblasts, leading to excessive ECM deposition. Methods: To elucidate the molecular mechanisms underlying fibrosis resolution, this study employed comprehensive proteomic techniques, including LC-MS/MS and SILAC, to analyze the interaction between myofibroblasts and mesenchymal stem cells derived from the endometrium (ET-eMSCs) preconditioned with PGE2. An in vitro co-culture system was used, with samples collected at baseline and after 48 h. Results: Proteomic analysis identified significant alterations in proteins associated with ECM remodeling, immune regulation, and cellular stress response. Notably, proteins involved in collagen degradation, antioxidant defense, and growth factor signaling pathways were differentially abundant. Network analyses demonstrated robust interactions among these proteins, suggesting coordinated modulatory effects. The data indicate that PGE2-primed ET-eMSCs induce a shift in myofibroblast secretory profiles, promoting a reduction in ECM stiffness, tissue reorganization, and activation of resolution pathways. Data are available via ProteomeXchange with identifier PXD067551. Conclusions: These findings reinforce the therapeutic potential of mesenchymal stem cell-based interventions for fibrotic diseases of the endometrium, opening avenues for regenerative strategies to restore reproductive function in mares.
Full article

Figure 1
Open AccessArticle
The Biological Variation in Serum ACE and CPN/CPB2 Activity in Healthy Individuals as Measured by the Degradation of Dabsylated Bradykinin—Reference Data and the Importance of Pre-Analytical Standardization
by
Malte Bayer, Michael Snyder and Simone König
Proteomes 2025, 13(3), 40; https://doi.org/10.3390/proteomes13030040 - 27 Aug 2025
Abstract
Background: Bradykinin (BK) is an inflammatory mediator. The degradation of labeled synthetic BK in biofluids can be used to report on the activity of angiotensin-converting enzyme (ACE) and basic carboxypeptidases N and CBP2, for which the neuropeptide is a substrate. Clinical studies have
[...] Read more.
Background: Bradykinin (BK) is an inflammatory mediator. The degradation of labeled synthetic BK in biofluids can be used to report on the activity of angiotensin-converting enzyme (ACE) and basic carboxypeptidases N and CBP2, for which the neuropeptide is a substrate. Clinical studies have shown significant changes in the serum activity of these enzymes in patients with inflammatory diseases. Methods: Here, we investigated variation in the cleavage of dabsylated synthetic BK (DBK) in serum and the formation of the major enzymatic fragments using a thin-layer chromatography-based neuropeptide reporter assay (NRA) in a large cohort of healthy volunteers from the international human Personal Omics Profiling consortium based at Stanford University. Results: Four major outcomes were reported. First, a set of NRA reference data for the healthy population was delivered, which is important for future investigations of patient sera. Second, it was shown that the measured serum degradation capacity for DBK was significantly higher in males than in females. There was no significant correlation of the NRA results with ethnicity, body mass index or overnight fasting. Third, a batch effect was noted among sampling sites (HUPO conferences). Thus, we used subcohorts rather than the entire collection for data mining. Fourth, as the low-cost and robust NRA is sensitive to enzyme activity, it provides such a necessary quick test to eliminate degraded and/or otherwise questionable samples. Conclusions: The results reiterate the critical importance of a high level of standardization in pre-analytical sample collection and processing—most notably, sample quality should be evaluated before conducting any large and expensive omics analyses.
Full article
(This article belongs to the Section Proteomics Technology and Methodology Development)
►▼
Show Figures

Figure 1
Open AccessArticle
Adiposome Proteomics Uncover Molecular Signatures of Cardiometabolic Risk in Obese Individuals
by
Mohamed Saad Rakab, Monica C. Asada, Imaduddin Mirza, Mohammed H. Morsy, Amro Mostafa, Francesco M. Bianco, Mohamed M. Ali, Chandra Hassan, Mario A. Masrur, Brian T. Layden and Abeer M. Mahmoud
Proteomes 2025, 13(3), 39; https://doi.org/10.3390/proteomes13030039 - 26 Aug 2025
Abstract
Background: Adipose-derived extracellular vesicles (adiposomes) are emerging as key mediators of inter-organ communication, yet their molecular composition and role in obesity-related pathophysiology remain underexplored. This study integrates clinical phenotyping with proteomic analysis of visceral adipose-derived adiposomes to identify obesity-linked molecular disruptions. Methods: Seventy-five
[...] Read more.
Background: Adipose-derived extracellular vesicles (adiposomes) are emerging as key mediators of inter-organ communication, yet their molecular composition and role in obesity-related pathophysiology remain underexplored. This study integrates clinical phenotyping with proteomic analysis of visceral adipose-derived adiposomes to identify obesity-linked molecular disruptions. Methods: Seventy-five obese and forty-seven lean adults were extensively profiled for metabolic, inflammatory, hepatic, and vascular parameters. Adiposomes isolated from visceral fat underwent mass spectrometry-based proteomic analysis, followed by differential abundance, pathway enrichment, regulatory network modeling, and clinical association testing. Results: Obese individuals exhibited widespread cardiometabolic dysfunction. Proteomics revealed 64 adiposomal proteins with differential abundance. Upregulated proteins (e.g., CRP, C9, APOC1) correlated with visceral adiposity, systemic inflammation, and endothelial dysfunction. In contrast, downregulated proteins (e.g., ADIPOQ, APOD, TTR, FGB, FGG) were associated with enhanced nitric oxide bioavailability and vascular protection, suggesting loss of homeostatic signaling. Network analyses identified TNF and IL1 as key upstream regulators driving inflammatory and oxidative stress pathways. Decision tree and random forest models accurately classified obesity, hypertension, diabetes, dyslipidemia, and hepatic steatosis (AUC = 0.908–0.994), identifying predictive protein signatures related to complement activation, inflammation, and lipid transport. Conclusion: Obesity alters adiposome proteomic cargo, reflecting and potentially mediating systemic inflammation, metabolic dysregulation, and vascular impairment.
Full article
(This article belongs to the Special Issue Proteomics in Chronic Diseases: Issues and Challenges)
►▼
Show Figures

Figure 1
Open AccessArticle
Identification of Cancer-Associated Proteins in Colorectal Cancer Using Mass Spectrometry
by
Naoyuki Toyota, Ryo Konno, Shuhei Iwata, Shin Fujita, Yoshio Kodera, Rei Noguchi, Tadashi Kondo, Yusuke Kawashima and Yuki Yoshimatsu
Proteomes 2025, 13(3), 38; https://doi.org/10.3390/proteomes13030038 - 12 Aug 2025
Abstract
►▼
Show Figures
Background: Colorectal cancer (CRC) is a leading cause of cancer-related mortality worldwide, with a multifactorial etiology involving genetic and environmental factors. Advanced proteomics offers valuable insights into the molecular mechanisms of cancer, identifying proteins that function as mediators in tumor biology. Methods: In
[...] Read more.
Background: Colorectal cancer (CRC) is a leading cause of cancer-related mortality worldwide, with a multifactorial etiology involving genetic and environmental factors. Advanced proteomics offers valuable insights into the molecular mechanisms of cancer, identifying proteins that function as mediators in tumor biology. Methods: In this study, we used mass spectrometry-based data-independent acquisition (DIA) to analyze the proteomic landscape of CRC. We compared protein abundance in normal and tumor tissues from 16 patients with CRC to identify cancer-associated proteins and examine their roles in disease progression. Results: The analysis identified 10,329 proteins, including 531 cancer-associated proteins from the Catalogue Of Somatic Mutations In Cancer (COSMIC) database, and 48 proteins specifically linked to CRC. Notably, clusters of proteins showed consistent increases or decreases in abundance across disease stages, suggesting their roles in tumorigenesis and progression. Conclusions: Our findings suggest that proteome abundance trends may contribute to the identification of biomarker candidates and therapeutic targets in colorectal cancer. However, given the limited sample size and lack of subtype stratification, further studies using larger, statistically powered cohorts are warranted to establish clinical relevance. These proteins may provide insights into drug resistance and tumor heterogeneity. Limitations of the study include the inability to detect low-abundance proteins and reliance on protein abundance rather than functional activity. Future complementary approaches, such as affinity proteomics, are suggested to address these limitations.
Full article

Figure 1
Open AccessReview
Uncovering Enzyme-Specific Post-Translational Modifications: An Overview of Current Methods
by
Nashira H. Ridgeway and Kyle K. Biggar
Proteomes 2025, 13(3), 37; https://doi.org/10.3390/proteomes13030037 - 11 Aug 2025
Abstract
►▼
Show Figures
Post-translational modifications (PTMs) govern a multitude of protein functions within the cell, surpassing the basic function(s) encoded directly within the amino acid sequence. Despite the historical discovery of PTMs dating back over a century, recent technological advancements have facilitated the rapid expansion of
[...] Read more.
Post-translational modifications (PTMs) govern a multitude of protein functions within the cell, surpassing the basic function(s) encoded directly within the amino acid sequence. Despite the historical discovery of PTMs dating back over a century, recent technological advancements have facilitated the rapid expansion of the known PTM landscape. However, the elucidation of enzyme–substrate relationships responsible for PTMs, particularly for those less studied, remains a challenging endeavor. This review provides an extensive overview of methods employed in the discovery of enzyme-specific substrates for PTM catalysis. Beginning with traditional experimental approaches rooted in chemistry, biochemistry and cell biology, this review progresses to recently developed computational strategies tailored for identifying enzyme–substrate interactions. The analysis reflects on the remarkable progress achieved in PTM research to date, underscoring the increasing role of computational and high-throughput techniques in expediting enzyme–substrate discovery. Furthermore, it highlights the potential of artificial intelligence to revolutionize PTM research and emphasizes the importance of unbiased high-throughput analysis in advancing our understanding of PTM networks. Ultimately, the review advocates for the integration of sophisticated computational strategies with experimental techniques to unravel the complex enzyme–substrate networks governing PTM-mediated cellular processes.
Full article

Figure 1
Open AccessArticle
Cell Surface Proteomics Reveals Hypoxia-Regulated Pathways in Cervical and Bladder Cancer
by
Faris Alanazi, Ammar Sharif, Melissa Kidd, Emma-Jayne Keevill, Vanesa Biolatti, Richard D. Unwin, Peter Hoskin, Ananya Choudhury, Tim A. D. Smith and Conrado G. Quiles
Proteomes 2025, 13(3), 36; https://doi.org/10.3390/proteomes13030036 - 5 Aug 2025
Abstract
Background Plasma membrane proteins (PMPs) play key roles in cell signalling, adhesion, and trafficking, and are attractive therapeutic targets in cancer due to their surface accessibility. However, their typically low abundance limits detection by conventional proteomic approaches. Methods: To improve PMP detection, we
[...] Read more.
Background Plasma membrane proteins (PMPs) play key roles in cell signalling, adhesion, and trafficking, and are attractive therapeutic targets in cancer due to their surface accessibility. However, their typically low abundance limits detection by conventional proteomic approaches. Methods: To improve PMP detection, we employed a surface proteomics workflow combining cell surface biotinylation and affinity purification prior to LC-MS/MS analysis in cervical (SiHa) and bladder (UMUC3) cancer cell lines cultured under normoxic (21% O2) or hypoxic (0.1% O2) conditions. Results: In SiHa cells, 43 hypoxia-upregulated proteins were identified exclusively in the biotin-enriched fraction, including ITGB2, ITGA7, AXL, MET, JAG2, and CAV1/CAV2. In UMUC3 cells, 32 unique upregulated PMPs were detected, including CD55, ADGRB1, SLC9A1, NECTIN3, and ACTG1. These proteins were not observed in corresponding whole-cell lysates and are associated with extracellular matrix remodelling, immune modulation, and ion transport. Biotinylation enhanced the detection of membrane-associated pathways such as ECM organisation, integrin signalling, and PI3K–Akt activation. Protein–protein interaction analysis revealed links between membrane receptors and intracellular stress regulators, including mitochondrial proteins. Conclusions: These findings demonstrate that surface biotinylation improves the sensitivity and selectivity of plasma membrane proteomics under hypoxia, revealing hypoxia-responsive proteins and pathways not captured by standard whole-cell analysis.
Full article
(This article belongs to the Section Proteomics of Human Diseases and Their Treatments)
►▼
Show Figures

Figure 1
Open AccessArticle
Multiplex Targeted Proteomic Analysis of Cytokine Ratios for ICU Mortality in Severe COVID-19
by
Rúben Araújo, Cristiana P. Von Rekowski, Tiago A. H. Fonseca, Cecília R. C. Calado, Luís Ramalhete and Luís Bento
Proteomes 2025, 13(3), 35; https://doi.org/10.3390/proteomes13030035 - 2 Aug 2025
Abstract
►▼
Show Figures
Background: Accurate and timely prediction of mortality in intensive care unit (ICU) patients, particularly those with COVID-19, remains clinically challenging due to complex immune responses. Proteomic cytokine profiling holds promise for refining mortality risk assessment. Methods: Serum samples from 89 ICU patients (55
[...] Read more.
Background: Accurate and timely prediction of mortality in intensive care unit (ICU) patients, particularly those with COVID-19, remains clinically challenging due to complex immune responses. Proteomic cytokine profiling holds promise for refining mortality risk assessment. Methods: Serum samples from 89 ICU patients (55 discharged, 34 deceased) were analyzed using a multiplex 21-cytokine panel. Samples were stratified into three groups based on time from collection to outcome: ≤48 h (Group 1: Early), >48 h to ≤7 days (Group 2: Intermediate), and >7 days to ≤14 days (Group 3: Late). Cytokine levels, simple cytokine ratios, and previously unexplored complex ratios between pro- and anti-inflammatory cytokines were evaluated. Machine learning-based feature selection identified the most predictive ratios, with performance evaluated by area under the curve (AUC), sensitivity, and specificity. Results: Complex cytokine ratios demonstrated superior predictive accuracy compared to traditional severity markers (APACHE II, SAPS II, SOFA), individual cytokines, and simple ratios, effectively distinguishing discharged from deceased patients across all groups (AUC: 0.918–1.000; sensitivity: 0.826–1.000; specificity: 0.775–0.900). Conclusions: Multiplex cytokine profiling enhanced by computationally derived complex ratios may offer robust predictive capabilities for ICU mortality risk stratification, serving as a valuable tool for personalized prognosis in critical care.
Full article

Figure 1
Open AccessArticle
Molecular Remodeling of the Sperm Proteome Following Varicocele Sclero-Embolization: Implications for Semen Quality Improvement
by
Domenico Milardi, Edoardo Vergani, Francesca Mancini, Fiorella Di Nicuolo, Emanuela Teveroni, Emanuele Pierpaolo Vodola, Alessandro Oliva, Giuseppe Grande, Alessandro Cina, Roberto Iezzi, Michela Cicchinelli, Federica Iavarone, Silvia Baroni, Alberto Ferlin, Andrea Urbani and Alfredo Pontecorvi
Proteomes 2025, 13(3), 34; https://doi.org/10.3390/proteomes13030034 - 15 Jul 2025
Abstract
Background: Varicocele is a common condition involving the dilation of veins in the scrotum, often linked to male infertility and testicular dysfunction. This study aimed to elucidate the molecular effects of successful varicocele treatment on sperm proteomes following percutaneous sclero-embolization. Methods: High-resolution tandem
[...] Read more.
Background: Varicocele is a common condition involving the dilation of veins in the scrotum, often linked to male infertility and testicular dysfunction. This study aimed to elucidate the molecular effects of successful varicocele treatment on sperm proteomes following percutaneous sclero-embolization. Methods: High-resolution tandem mass spectrometry was performed for proteomic profiling of pooled sperm lysates from five patients exhibiting improved semen parameters before and after (3 and 6 months) varicocele sclero-embolization. Data were validated by Western blot analysis. Results: Seven proteins were found exclusively in varicocele patients before surgery—such as stathmin, IFT20, selenide, and ADAM21—linked to inflammation and oxidative stress. After sclero-embolization, 55 new proteins emerged, including antioxidant enzymes like selenoprotein P and GPX3. Thioredoxin (TXN) and peroxiredoxin (PRDX3) were upregulated, indicating restoration of key antioxidant pathways. Additionally, the downregulation of some histones and the autophagy-related protein ATG9A suggests a shift toward an improved chromatin organization and a healthier cellular environment post-treatment. Conclusions: Varicocele treatment that improves sperm quality and fertility parameters leads to significant proteome modulation. These changes include reduced oxidative stress and broadly restored sperm maturation. Despite the limited patient cohort analyzed, these preliminary findings provide valuable insights into how varicocele treatment might enhance male fertility and suggest potential biomarkers for improved male infertility treatment strategies.
Full article
(This article belongs to the Section Proteomics of Human Diseases and Their Treatments)
►▼
Show Figures

Graphical abstract
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Biomedicines, Metabolites, Proteomes, Genes, J
Multi-Omics in Precision Medicine
Topic Editors: Michele Costanzo, Armando CeveniniDeadline: 31 December 2025
Topic in
IJMS, Metabolites, Molecules, Proteomes, Biomedicines, IJTM
Liquid Biopsy: A Modern Method Transforming Biomedicine
Topic Editors: Michele Costanzo, Marianna CaterinoDeadline: 31 March 2026

Special Issues
Special Issue in
Proteomes
Proteomics in Diabetes: From Mechanisms to Biomarkers
Guest Editor: Sindre Lee-ØdegårdDeadline: 15 November 2025
Special Issue in
Proteomes
Plant Genomics and Proteomics
Guest Editors: Setsuko Komatsu, Pingfang YangDeadline: 31 March 2026









