Recent Advances in Multi-Drug Resistance (MDR) Efflux Pump Inhibitors of Gram-Positive Bacteria S. aureus
Abstract
:1. Introduction
2. Efflux Pumps in Gram Positive Bacteria
| Bacterial strain | Transport protein family | Efflux pump | Substrates |
|---|---|---|---|
| Staphylococcus aureus | MFS [9,41,42,43,44] | NorA | NOR, CPX |
| NorB | NOR, CPX, SPX | ||
| MdeA | Macrolides | ||
| MATE [45] | Tet38 | tetracyclines | |
| MepA | FQ, glycylcyclines | ||
| SMR [39,60,61] | Smr, QacG, QacH | - | |
| Staphylococcus spp | MFS [9,46,47] | Mef(A) | Macrolides |
| ABC [47,48,49] | MsrA | Macrolides, type B streptogramins | |
| Staphylococcus haemolyticus | MFS [42] | MdeA | Macrolides, lincosamides type A streptogramins |
| Staphylococcus lentus | [54] | FexA | - |
| Streptomyces coelicolor | MFS [38] | CmlR1, CmlR2 | Chloramphenicol |
| Streptomyces spp | MFS [50] | Cml, Cmlv, Cmr, Cmx, CmA | - |
| Streptococcus spp | MFS [9,45,47] | Mef(A) | Macrolides |
| Streptococcus pneumoniae | ABC [51] | Msr(D) | Macrolides, ketolides |
| MFS [52] | PmrA | NOR, CPX | |
| Clostridium difficile | MFS [53] | Cme | Erythromycin |
| MATE [45] | CdeA | FQ | |
| Bacillus subtilis | MFS [9,46] | LmrB | Lincosamides |
| Bmr, Bmr3, Blt | FQ | ||
| Bacillus glutamicum | MFS [9,46] | LmrB | Lincosamides |
3. Current Approaches in Search for EPIs in Gram Positive Bacteria
3.1. Plant-Derived NorA EPIs and Their Chemical Modifications

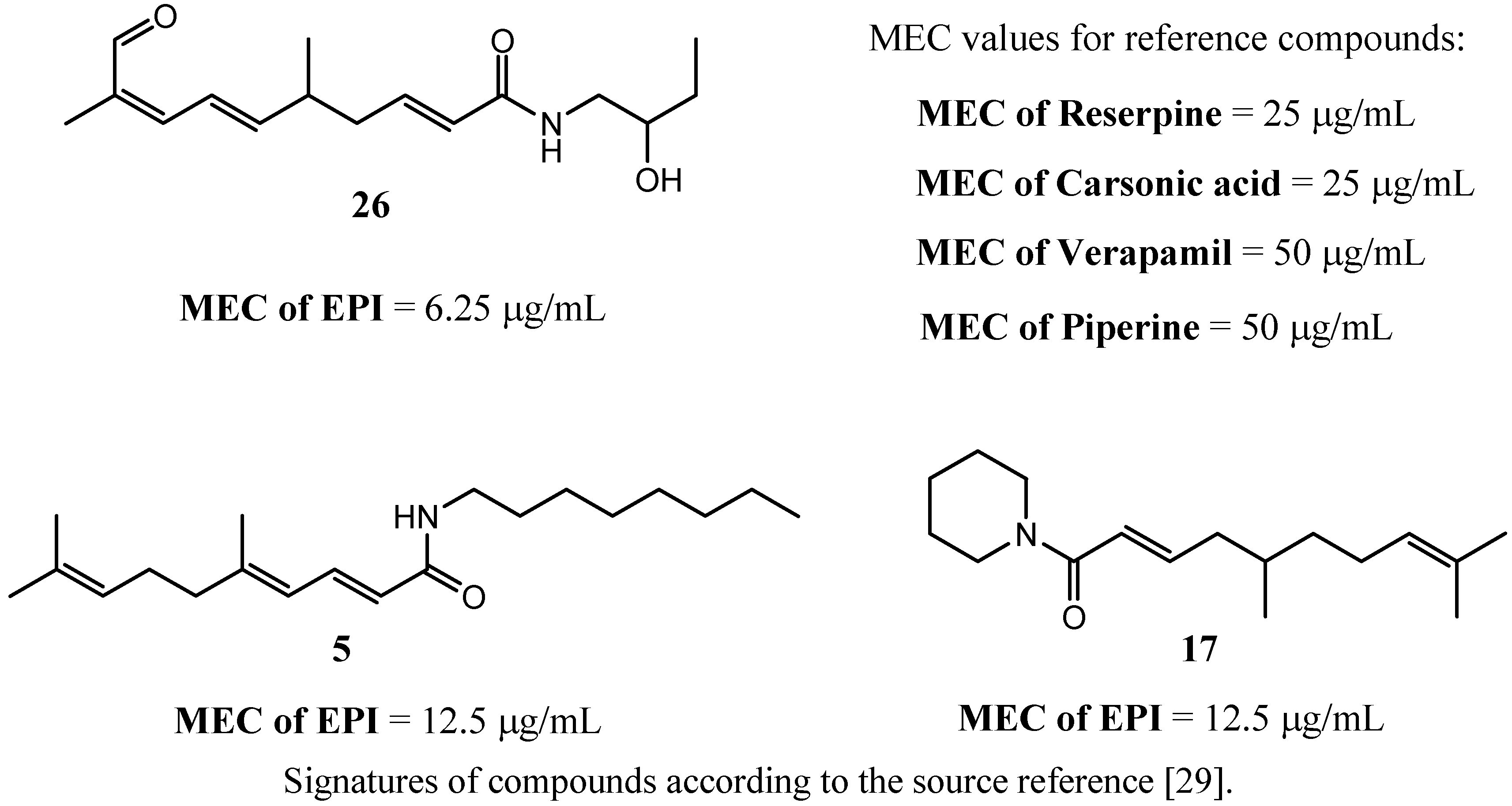
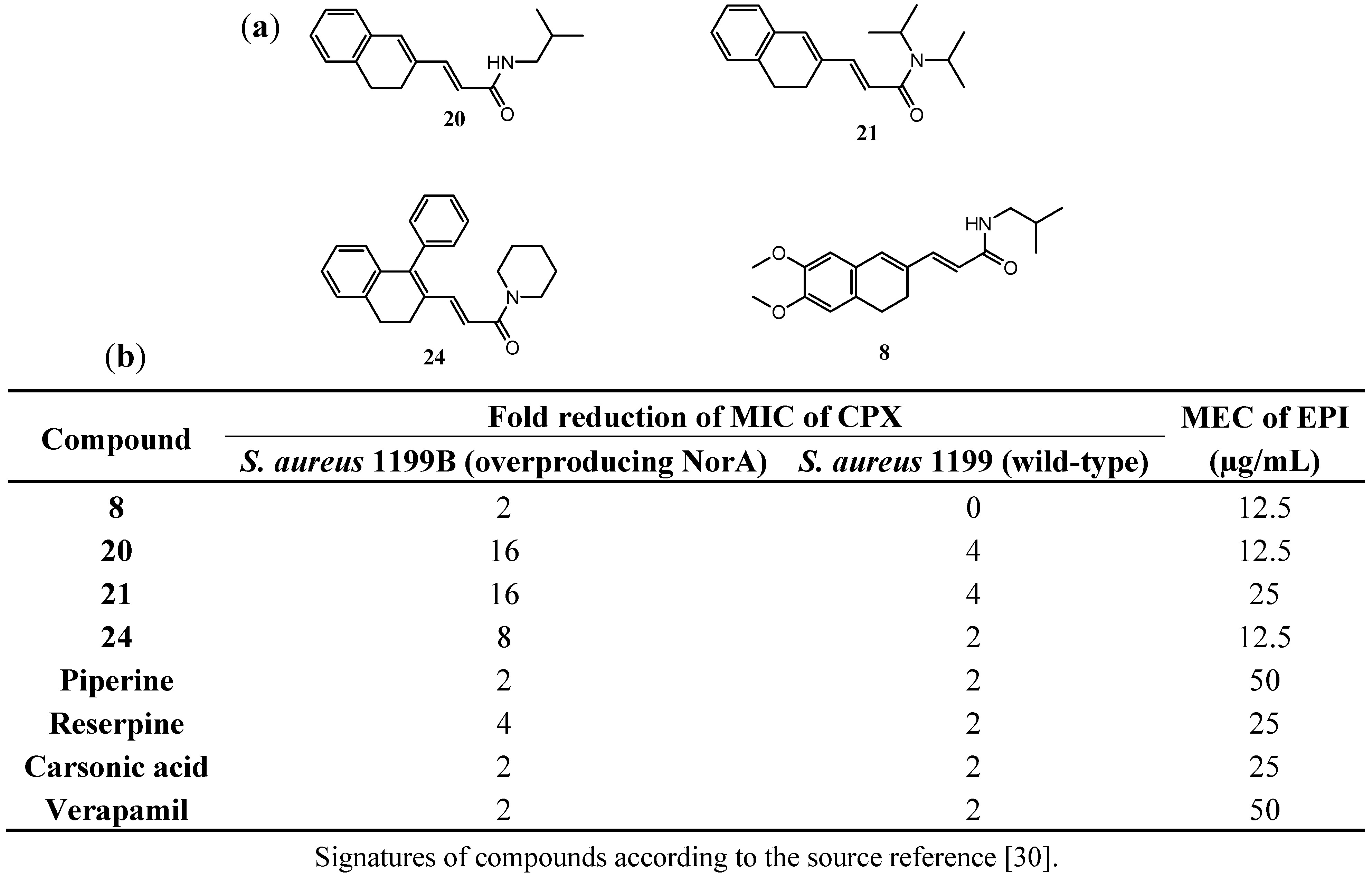
3.2. Chalcones and Alkenamide Inhibitors of the NorA in S. aureus

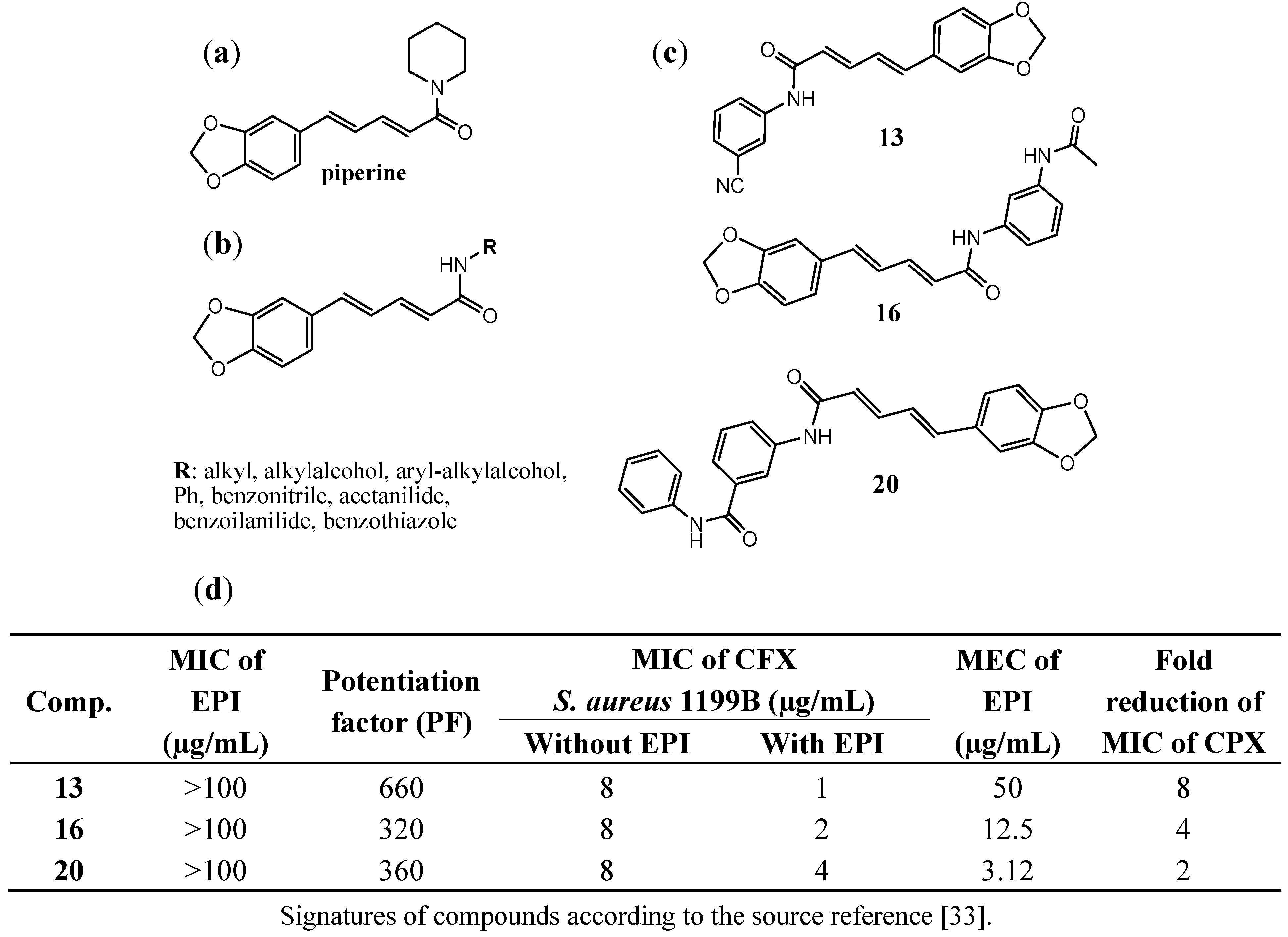
3.3. Piperine EPIs against NorA in S. aureus
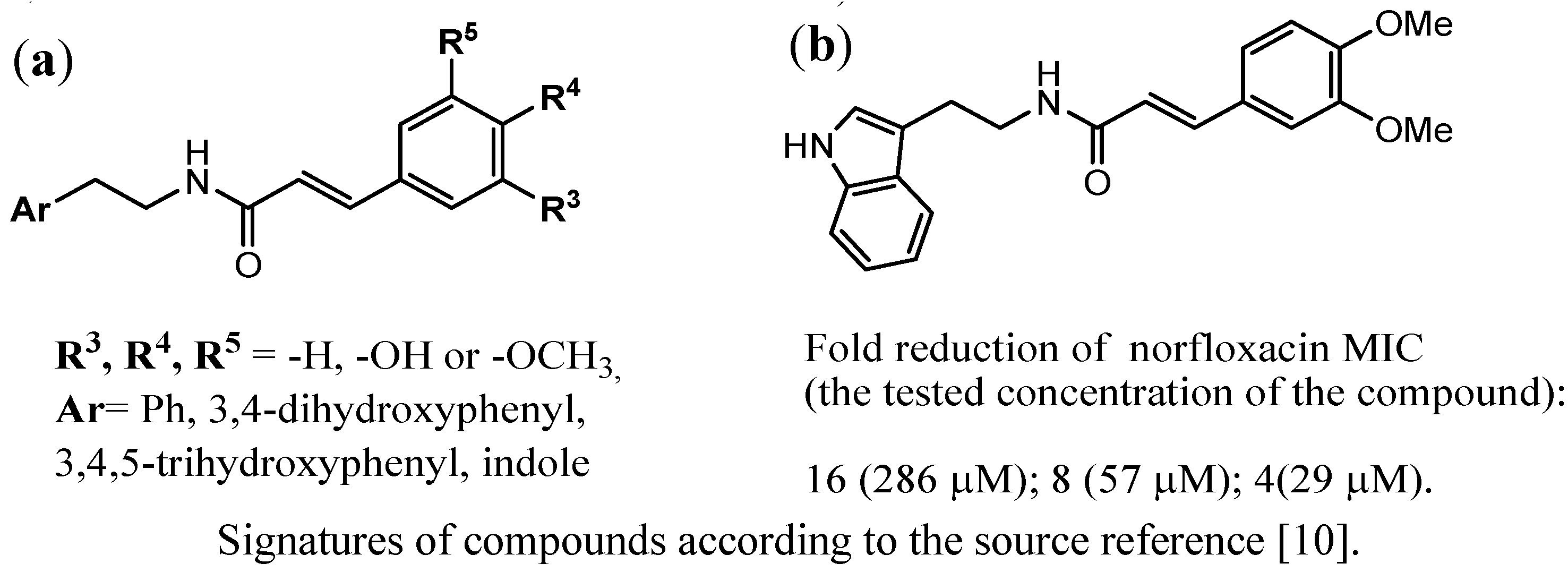
3.4. Indoles as Inhibitors of the NorA of S. aureus
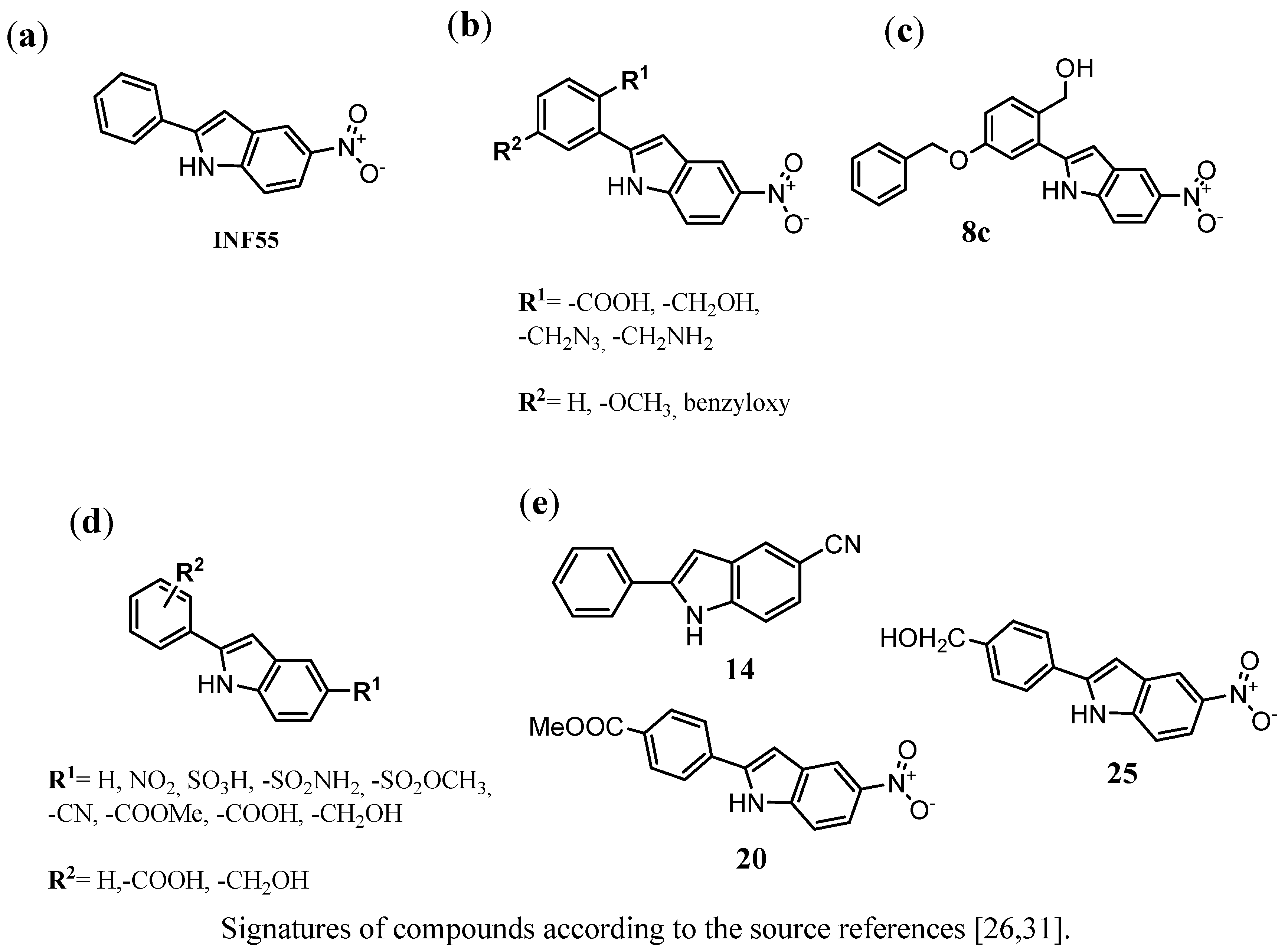
3.5. Other Chemical Groups of NorA EPIs
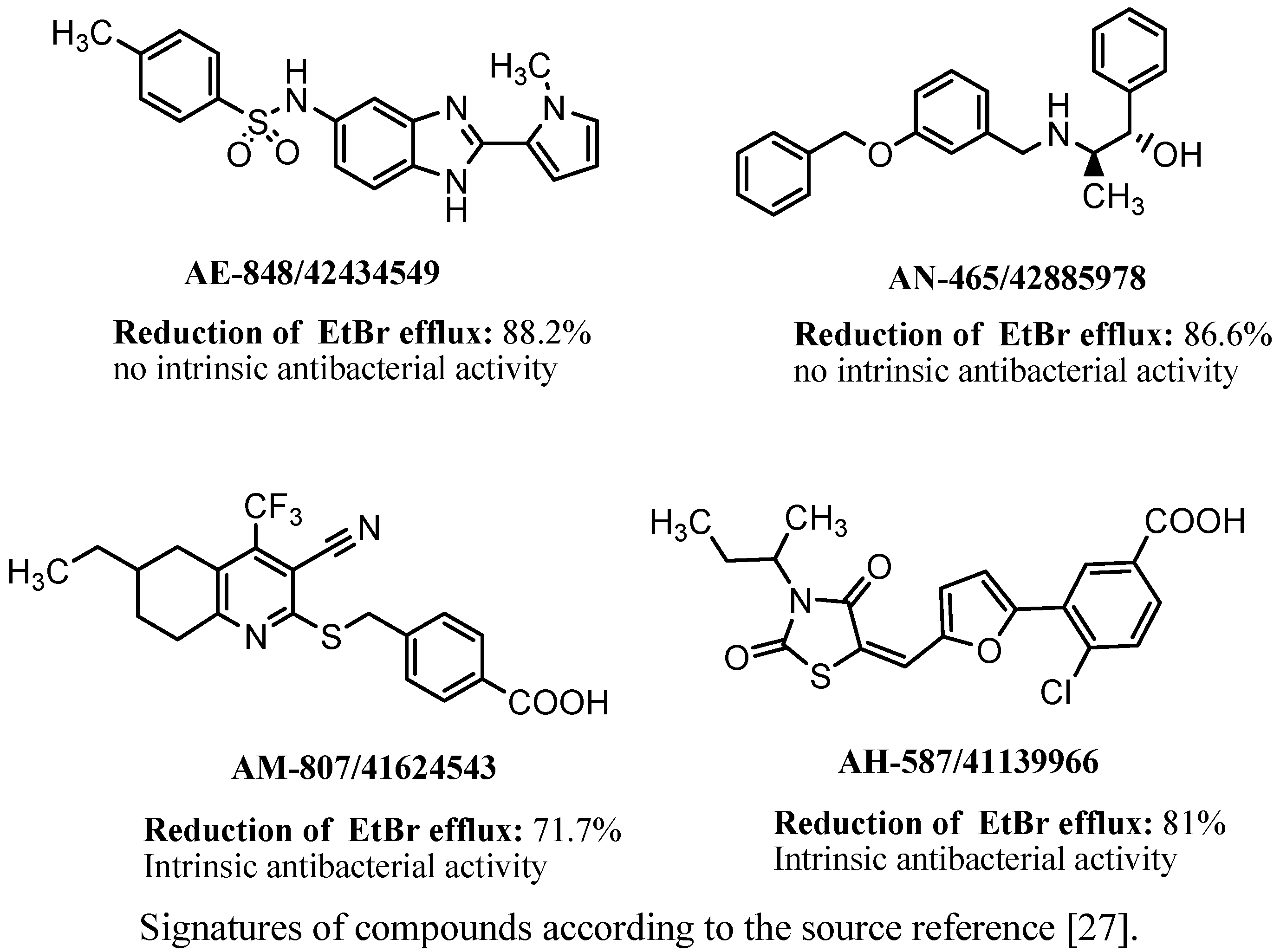
3.6. PSSRI-Based EPIs of NorA and MepA in S. aureus
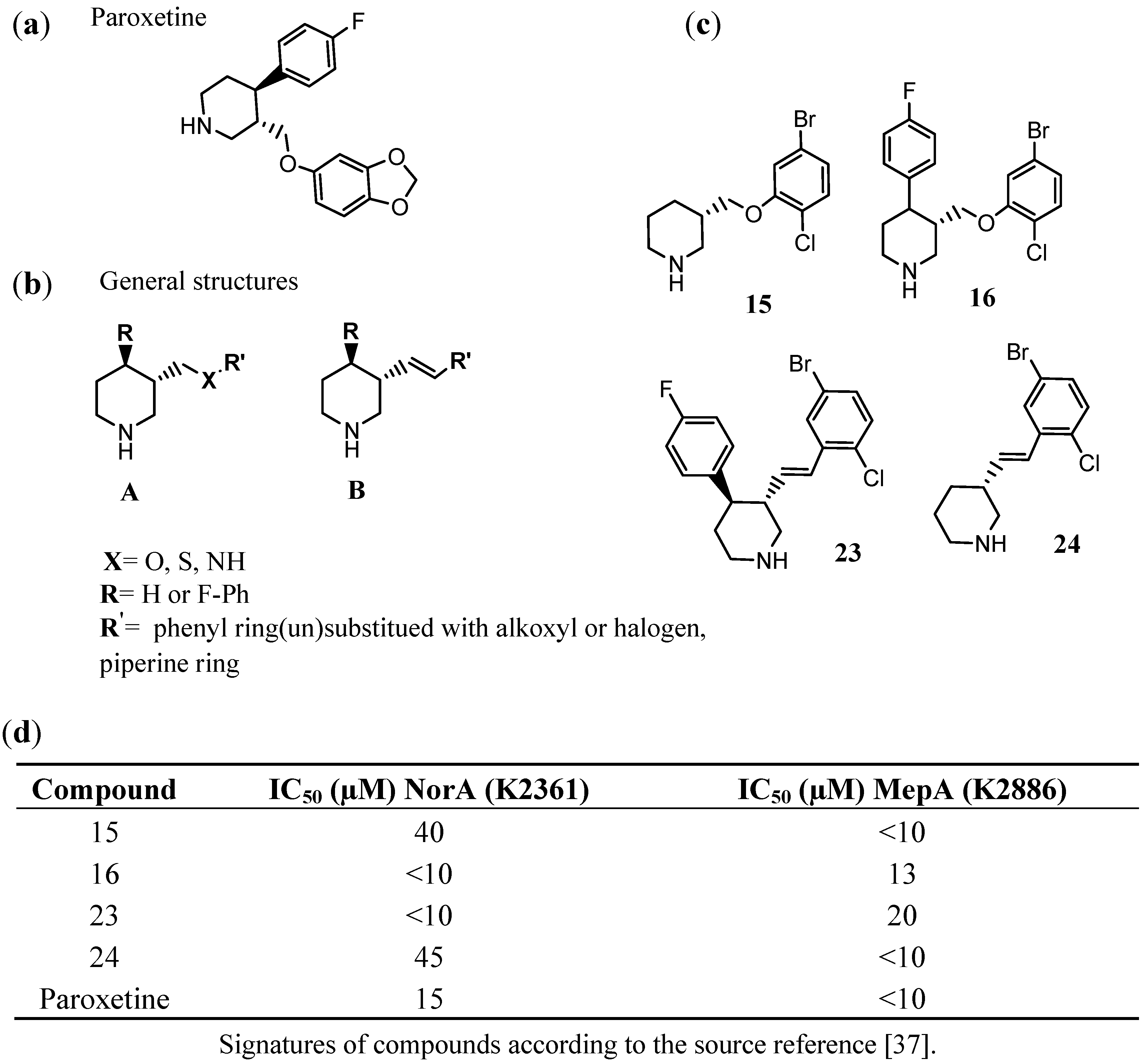
3.7. Search for Inhibitors of Other Efflux Pumps of Gram-Positive Bacteria
4. Conclusions
Acknowledgments
Conflict of Interest
References and Notes
- Fournier Dit Chabert, J.; Marquez, B.; Neville, L.; Joucla, L.; Broussous, S.; Bouhours, P.; David, E.; Pellet-Rostaing, S.; Marquet, B.; Moreau, N.; et al. Synthesis and evaluation of new arylbenzo[b]thiophene and diarylthiophene derivatives as inhibitors of the NorA multidrug transporter of Staphylococcus aureus. Bioorg. Med. Chem. 2007, 15, 4482–4497. [Google Scholar]
- Bush, K.; Miller, G.H. Bacterial enzymatic resistance: Beta-lactamases and aminoglycoside-modifying enzymes. Curr. Opin. Microbiol. 1998, 1, 509–515. [Google Scholar] [CrossRef]
- Sabatini, S.; Gosetto, F.; Serritella, S.; Manfroni, G.; Tabarrini, O.; Iraci, N.; Brincat, J.P.; Carosati, E.; Villarini, M.; Kaatz, G.W.; et al. Pyrazolo[4,3-c][1,2]benzothiazines 5,5-dioxide: A promising new class of Staphylococcus aureus NorA efflux pump inhibitors. J. Med. Chem. 2012, 55, 3568–3572. [Google Scholar] [CrossRef]
- Ruiz, J. Mechanisms of resistance to quinolones: Target alterations, decreased accumulation and DNA gyrase protection. J. Antimicrob. Chemother. 2003, 51, 1109–1117. [Google Scholar] [CrossRef]
- Nikaido, H. Molecular basis of bacterial outer membrane permeability revisited. Microbiol. Mol. Biol. Rev. 2003, 67, 593–656. [Google Scholar] [CrossRef]
- Li, X.Z.; Nikaido, H. Efflux-mediated drug resistance in bacteria: An update. Drugs 2009, 69, 1555–1623. [Google Scholar] [CrossRef]
- Kohler, T.; Pechere, J.C.; Plesiat, P. Bacterial antibiotic efflux systems of medical importance. Cell Mol. Life Sci. 1999, 56, 771–778. [Google Scholar] [CrossRef]
- Mahamoud, A.; Chevalier, J.; Alibert-Franco, S.; Kern, W.V.; Pages, J.M. Antibiotic efflux pumps in Gram-negative bacteria: The inhibitor response strategy. J. Antimicrob. Chemother. 2007, 59, 1223–1229. [Google Scholar] [CrossRef]
- Poole, K.; Lomovskaya, O. Can efflux inhibitors really counter resistance? Drug Discov. Today 2006, 3, 145–152. [Google Scholar] [CrossRef]
- Michalet, S.; Cartier, G.; David, B.; Mariotte, A.M.; Dijoux-franca, M.G.; Kaatz, G.W.; Stavri, M.; Gibbons, S. N-Caffeoylphenalkylamide derivatives as bacterial efflux pump inhibitors. Bioorg. Med. Chem. Lett. 2007, 17, 1755–1758. [Google Scholar] [CrossRef]
- Mahamoud, A.; Chevalier, J.; Davin-Regli, A.; Barbe, J.; Pages, J.M. Quinoline derivatives as promising inhibitors of antibiotic efflux pump in multidrug resistant Enterobacter aerogenes isolates. Curr. Drug Targets 2006, 7, 843–847. [Google Scholar] [CrossRef]
- Pages, J.M.; Amaral, L.; Fanning, S. An original deal for new molecule: Reversal of efflux pump activity, a rational strategy to combat gram-negative resistant bacteria. Curr. Med. Chem. 2011, 18, 2969–2980. [Google Scholar] [CrossRef]
- Pages, J.M.; Amaral, L. Mechanisms of drug efflux and strategies to combat them: Challenging the efflux pump of Gram-negative bacteria. Biochim. Biophys. Acta 2009, 1794, 826–833. [Google Scholar] [CrossRef]
- Martins, M.; Dastidar, S.G.; Fanning, S.; Kristiansen, J.E.; Molnar, J.; Pages, J.M.; Schelz, Z.; Spengler, G.; Viveiros, M.; Amaral, L. Potential role of non-antibiotics (helper compounds) in the treatment of multidrug-resistant Gram-negative infections: Mechanisms for their direct and indirect activities. Int. J. Antimicrob. Agents 2008, 31, 198–208. [Google Scholar] [CrossRef]
- Nikaido, H.; Pages, J.M. Broad-specificity efflux pumps and their role in multidrug resistance of Gram-negative bacteria. FEMS Microbiol. Rev. 2012, 36, 340–363. [Google Scholar] [CrossRef]
- Bolla, J.M.; Alibert-Franco, S.; Handzlik, J.; Chevalier, J.; Mahamoud, A.; Boyer, G.; Kiec-Kononowicz, K.; Pages, J.M. Strategies for bypassing the membrane barrier in multidrug resistant Gram-negative bacteria. FEBS Lett. 2011, 585, 1682–1690. [Google Scholar] [CrossRef]
- Handzlik, J.; Szymanska, E.; Chevalier, J.; Otrebska, E.; Kiec-Kononowicz, K.; Pages, J.M.; Alibert, S. Amine-alkyl derivatives of hydantoin: New tool to combat resistant bacteria. Eur. J. Med. Chem. 2011, 46, 5807–5816. [Google Scholar] [CrossRef]
- Handzlik, J.; Szymanska, E.; Alibert, S.; Chevalier, J.; Otrebska, E.; Pekala, E.; Pages, J.M.; Kiec-Kononowicz, K. Search for new tools to combat Gram-negative resistant bacteria among amine derivatives of 5-arylidenehydantoin. Bioorg. Med. Chem. 2013, 21, 135–145. [Google Scholar] [CrossRef]
- Pradel, E.; Pages, J.M. The AcrAB-TolC efflux pump contributes to multidrug resistance in the nosocomial pathogen Enterobacter aerogenes. Antimicrob. Agents Chemother. 2002, 46, 2640–2643. [Google Scholar] [CrossRef]
- Ghisalberti, D.; Masi, M.; Pages, J.M.; Chevalier, J. Chloramphenicol and expression of multidrug efflux pump in Enterobacter aerogenes. Biochem. Biophys. Res. Commun. 2005, 328, 1113–1118. [Google Scholar] [CrossRef]
- Lavigne, J.P.; Sotto, A.; Nicolas-Chanoine, M.H.; Bouziges, N.; Bourg, G.; Davin-Regli, A.; Pages, J.M. Membrane permeability, a pivotal function involved in antibiotic resistance and virulence in Enterobacter aerogenes clinical isolates. Clin. Microbiol. Infect. 2012, 18, 539–545. [Google Scholar] [CrossRef]
- Askoura, M.; Mottawea, W.; Abujamel, T.; Taher, I. Efflux pump inhibitors (EPIs) as new antimicrobial agents against Pseudomonas aeruginosa. Libyan J. Med. 2011, 6. [Google Scholar] [CrossRef]
- Drew, D.; Klepsch, M.M.; Newstead, S.; Flaig, R.; De Gier, J.W.; Iwata, S.; Beis, K. The structure of the efflux pump AcrB in complex with bile acid. Mol. Membr. Biol. 2008, 25, 677–682. [Google Scholar] [CrossRef]
- Sennhauser, G.; Bukowska, M.A.; Briand, C.; Grutter, M.G. Crystal structure of the multidrug exporter MexB from Pseudomonas aeruginosa. J. Mol. Biol. 2009, 389, 134–145. [Google Scholar] [CrossRef]
- Pages, J.M.; Sandrine, A.F.; Mahamoud, A.; Bolla, J.M.; Davin-Regli, A.; Chevalier, J.; Garnotel, E. Efflux pumps of gram-negative bacteria, a new target for new molecules. Curr. Top. Med. Chem. 2010, 10, 1848–1857. [Google Scholar] [CrossRef]
- Ambrus, J.I.; Kelso, M.J.; Bremner, J.B.; Ball, A.R.; Casadei, G.; Lewis, K. Structure-activity relationships of 2-aryl-1H-indole inhibitors of the NorA efflux pump in Staphylococcus aureus. Bioorg. Med. Chem. Lett. 2008, 18, 4294–4297. [Google Scholar] [CrossRef]
- Brincat, J.P.; Carosati, E.; Sabatini, S.; Manfroni, G.; Fravolini, A.; Raygada, J.L.; Patel, D.; Kaatz, G.W.; Cruciani, G. Discovery of novel inhibitors of the NorA multidrug transporter of Staphylococcus aureus. J. Med. Chem. 2011, 54, 354–365. [Google Scholar] [CrossRef]
- Holler, J.G.; Slotved, H.C.; Molgaard, P.; Olsen, C.E.; Christensen, S.B. Chalcone inhibitors of the NorA efflux pump in Staphylococcus aureus whole cells and enriched everted membrane vesicles. Bioorg. Med. Chem. 2012, 20, 4514–4521. [Google Scholar] [CrossRef]
- Thota, N.; Koul, S.; Reddy, M.V.; Sangwan, P.L.; Khan, I.A.; Kumar, A.; Raja, A.F.; Andotra, S.S.; Qazi, G.N. Citral derived amides as potent bacterial NorA efflux pump inhibitors. Bioorg. Med. Chem. 2008, 16, 6535–6543. [Google Scholar] [CrossRef]
- Thota, N.; Reddy, M.V.; Kumar, A.; Khan, I.A.; Sangwan, P.L.; Kalia, N.P.; Koul, J.L.; Koul, S. Substituted dihydronaphthalenes as efflux pump inhibitors of Staphylococcus aureus. Eur. J. Med. Chem. 2010, 45, 3607–3616. [Google Scholar] [CrossRef]
- Samosorn, S.; Bremner, J.B.; Ball, A.; Lewis, K. Synthesis of functionalized 2-aryl-5-nitro-1H-indoles and their activity as bacterial NorA efflux pump inhibitors. Bioorg. Med. Chem. 2006, 14, 857–865. [Google Scholar] [CrossRef]
- Sangwan, P.L.; Koul, J.L.; Koul, S.; Reddy, M.V.; Thota, N.; Khan, I.A.; Kumar, A.; Kalia, N.P.; Qazi, G.N. Piperine analogs as potent Staphylococcus aureus NorA efflux pump inhibitors. Bioorg. Med. Chem. 2008, 16, 9847–9857. [Google Scholar]
- Nargotra, A.; Sharma, S.; Koul, J.L.; Sangwan, P.L.; Khan, I.A.; Kumar, A.; Taneja, S.C.; Koul, S. Quantitative structure activity relationship (QSAR) of piperine analogs for bacterial NorA efflux pump inhibitors. Eur. J. Med. Chem. 2009, 44, 4128–4135. [Google Scholar] [CrossRef]
- Sabatini, S.; Gosetto, F.; Manfroni, G.; Tabarrini, O.; Kaatz, G.W.; Patel, D.; Cecchetti, V. Evolution from a natural flavones nucleus to obtain 2-(4-Propoxyphenyl)quinoline derivatives as potent inhibitors of the S. aureus NorA efflux pump. J. Med. Chem. 2011, 54, 5722–5736. [Google Scholar] [CrossRef]
- Wei, P.; Kaatz, G.W.; Kerns, R.J. Structural differences between paroxetine and femoxetine responsible for differential inhibition of Staphylococcus aureus efflux pumps. Bioorg. Med. Chem. Lett. 2004, 14, 3093–3097. [Google Scholar] [CrossRef]
- Abulrob, A.N.; Suller, M.T.; Gumbleton, M.; Simons, C.; Russell, A.D. Identification and biological evaluation of grapefruit oil components as potential novel efflux pump modulators in methicillin-resistant Staphylococcus aureus bacterial strains. Phytochemistry 2004, 65, 3021–3027. [Google Scholar] [CrossRef]
- German, N.; Kaatz, G.W.; Kerns, R.J. Synthesis and evaluation of PSSRI-based inhibitors of Staphylococcus aureus multidrug efflux pumps. Bioorg. Med. Chem. Lett. 2008, 18, 1368–1373. [Google Scholar] [CrossRef]
- Okandeji, B.O.; Greenwald, D.M.; Wroten, J.; Sello, J.K. Synthesis and evaluation of inhibitors of bacterial drug efflux pumps of the major facilitator superfamily. Bioorg. Med. Chem. 2011, 19, 7679–7689. [Google Scholar] [CrossRef]
- Jarmula, A.; Oblak, E.; Wawrzycka, D.; Gutowicz, J. Efflux-mediated antimicrobial multidrug resistance. Postepy Hig. Med. Dosw. (Online) 2011, 65, 216–227. [Google Scholar]
- Li, X.Z.; Nikaido, H. Efflux-mediated drug resistance in bacteria. Drugs 2004, 64, 159–204. [Google Scholar] [CrossRef]
- Tseng, T.T.; Gratwick, K.S.; Kollman, J.; Park, D.; Nies, D.H.; Goffeau, A.; Saier, M.H., Jr. The RND permease superfamily: An ancient, ubiquitous and diverse family that includes human disease and development proteins. J. Mol. Microbiol. Biotechnol. 1999, 1, 107–125. [Google Scholar]
- Truong-Bolduc, Q.C.; Dunman, P.M.; Strahilevitz, J.; Projan, S.J.; Hooper, D.C. MgrA is a multiple regulator of two new efflux pumps in Staphylococcus aureus. J. Bacteriol. 2005, 187, 2395–2405. [Google Scholar] [CrossRef]
- DeMarco, C.E.; Cushing, L.A.; Frempong-Manso, E.; Seo, S.M.; Jaravaza, T.A.; Kaatz, G.W. Efflux-related resistance to norfloxacin, dyes, and biocides in bloodstream isolates of Staphylococcus aureus. Antimicrob. Agents Chemother. 2007, 51, 3235–3239. [Google Scholar] [CrossRef]
- Huang, J.; O'Toole, P.W.; Shen, W.; Amrine-Madsen, H.; Jiang, X.; Lobo, N.; Palmer, L.M.; Voelker, L.; Fan, F.; Gwynn, M.N.; et al. Novel chromosomally encoded multidrug efflux transporter MdeA in Staphylococcus aureus. Antimicrob. Agents Chemother. 2004, 48, 909–917. [Google Scholar] [CrossRef]
- Kaatz, G.W.; McAleese, F.; Seo, S.M. Multidrug resistance in Staphylococcus aureus due to overexpression of a novel multidrug and toxin extrusion (MATE) transport protein. Antimicrob. Agents Chemother. 2005, 49, 1857–1864. [Google Scholar] [CrossRef]
- Piddock, L.J. Clinically relevant chromosomally encoded multidrug resistance efflux pumps in bacteria. Clin. Microbiol. Rev. 2006, 19, 382–402. [Google Scholar] [CrossRef]
- Pozzi, G.; Iannelli, F.; Oggioni, M.R.; Santagati, M.; Stefani, S. Genetic elements carrying macrolide efflux genes in streptococci. Curr. Drug Targets Infect. Disord. 2004, 4, 203–206. [Google Scholar] [CrossRef]
- Reynolds, E.; Ross, J.I.; Cove, J.H. Msr(A) and related macrolide/streptogramin resistance determinants: incomplete transporters? Int. J. Antimicrob. Agents 2003, 22, 228–236. [Google Scholar] [CrossRef]
- Portillo, A.; Ruiz-Larrea, F.; Zarazaga, M.; Alonso, A.; Martinez, J.L.; Torres, C. Macrolide resistance genes in Enterococcus spp. Antimicrob. Agents Chemother. 2000, 44, 967–971. [Google Scholar] [CrossRef]
- Schwarz, S.; Kehrenberg, C.; Doublet, B.; Cloeckaert, A. Molecular basis of bacterial resistance to chloramphenicol and florfenicol. FEMS Microbiol. Rev. 2004, 28, 519–542. [Google Scholar] [CrossRef]
- Robertson, G.T.; Doyle, T.B.; Lynch, A.S. Use of an efflux-deficient streptococcus pneumoniae strain panel to identify ABC-class multidrug transporters involved in intrinsic resistance to antimicrobial agents. Antimicrob. Agents Chemother. 2005, 49, 4781–4783. [Google Scholar] [CrossRef]
- Yoshida, H.; Bogaki, M.; Nakamura, S.; Ubukata, K.; Konno, M. Nucleotide sequence and characterization of the Staphylococcus aureus norA gene, which confers resistance to quinolones. J. Bacteriol. 1990, 172, 6942–6949. [Google Scholar]
- Pumbwe, L.; Piddock, L.J. Identification and molecular characterisation of CmeB, a Campylobacter jejuni multidrug efflux pump. FEMS Microbiol. Lett. 2002, 206, 185–189. [Google Scholar] [CrossRef]
- Kehrenberg, C.; Schwarz, S. fexA, a novel Staphylococcus lentus gene encoding resistance to florfenicol and chloramphenicol. Antimicrob. Agents Chemother. 2004, 48, 615–618. [Google Scholar] [CrossRef]
- Borges-Walmsley, M.I.; McKeegan, K.S.; Walmsley, A.R. Structure and function of efflux pumps that confer resistance to drugs. Biochem. J. 2003, 376, 313–338. [Google Scholar] [CrossRef]
- Borges-Walmsley, M.I.; Walmsley, A.R. The structure and function of drug pumps. Trends Microbiol. 2001, 9, 71–79. [Google Scholar] [CrossRef]
- Paulsen, I.T.; Brown, M.H.; Skurray, R.A. Proton-dependent multidrug efflux systems. Microbiol. Rev. 1996, 60, 575–608. [Google Scholar]
- Poole, K. Efflux-mediated antimicrobial resistance. J. Antimicrob. Chemother. 2005, 56, 20–51. [Google Scholar] [CrossRef]
- Markham, P.N.; Neyfakh, A.A. Efflux-mediated drug resistance in Gram-positive bacteria. Curr. Opin. Microbiol. 2001, 4, 509–514. [Google Scholar] [CrossRef]
- Kikukawa, T.; Nara, T.; Araiso, T.; Miyauchi, S.; Kamo, N. Two-component bacterial multidrug transporter, EbrAB: Mutations making each component solely functional. Biochim. Biophys. Acta 2006, 1758, 673–679. [Google Scholar] [CrossRef]
- Putman, M.; van Veen, H.W.; Konings, W.N. Molecular properties of bacterial multidrug transporters. Microbiol. Mol. Biol. Rev. 2000, 64, 672–693. [Google Scholar] [CrossRef]
- Wasaznik, A.; Grinholc, M.; Bielawski, K.P. Active efflux as the multidrug resistance mechanism. Postepy Hig. Med. Dosw. (Online) 2009, 63, 123–133. [Google Scholar]
- Omote, H.; Hiasa, M.; Matsumoto, T.; Otsuka, M.; Moriyama, Y. The MATE proteins as fundamental transporters of metabolic and xenobiotic organic cations. Trends Pharmacol. Sci. 2006, 27, 587–593. [Google Scholar] [CrossRef]
- Van Bambeke, F.; Balzi, E.; Tulkens, P.M. Antibiotic efflux pumps. Biochem. Pharmacol. 2000, 60, 457–470. [Google Scholar] [CrossRef]
- Kumar, A.; Schweizer, H.P. Bacterial resistance to antibiotics: Active efflux and reduced uptake. Adv. Drug Deliv. Rev. 2005, 57, 1486–1513. [Google Scholar] [CrossRef]
- Lubelski, J.; Konings, W.N.; Driessen, A.J. Distribution and physiology of ABC-type transporters contributing to multidrug resistance in bacteria. Microbiol. Mol. Biol. Rev. 2007, 71, 463–476. [Google Scholar] [CrossRef]
- Zgurskaya, H.I.; Nikaido, H. Multidrug resistance mechanisms: Drug efflux across two membranes. Mol. Microbiol. 2000, 37, 219–225. [Google Scholar] [CrossRef]
- Wang, K.; Pei, H.; Huang, B.; Zhu, X.; Zhang, J.; Zhou, B.; Zhu, L.; Zhang, Y.; Zhou, F.F. The Expression of ABC Efflux Pump, Rv1217c-Rv1218c, and Its Association with Multidrug Resistance of Mycobacterium tuberculosis in China. Curr. Microbiol. 2013, 66, 222–226. [Google Scholar] [CrossRef]
- Zechini, B.; Versace, I. Inhibitors of multidrug resistant efflux systems in bacteria. Recent. Pat. Antiinfect. Drug Discov. 2009, 4, 37–50. [Google Scholar] [CrossRef]
- Holler, J.G.; Christensen, S.B.; Slotved, H.C.; Rasmussen, H.B.; Guzman, A.; Olsen, C.E.; Petersen, B.; Molgaard, P. Novel inhibitory activity of the Staphylococcus aureus NorA efflux pump by a kaempferol rhamnoside isolated from Persea lingue Nees. J. Antimicrob. Chemother. 2012, 67, 1138–1144. [Google Scholar] [CrossRef]
- ClinicalTrials.gov. A service of the US National Institutes of Health. Available online: www.clinicaltrials.gov (accessed on 31 January 2013).
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Handzlik, J.; Matys, A.; Kieć-Kononowicz, K. Recent Advances in Multi-Drug Resistance (MDR) Efflux Pump Inhibitors of Gram-Positive Bacteria S. aureus. Antibiotics 2013, 2, 28-45. https://doi.org/10.3390/antibiotics2010028
Handzlik J, Matys A, Kieć-Kononowicz K. Recent Advances in Multi-Drug Resistance (MDR) Efflux Pump Inhibitors of Gram-Positive Bacteria S. aureus. Antibiotics. 2013; 2(1):28-45. https://doi.org/10.3390/antibiotics2010028
Chicago/Turabian StyleHandzlik, Jadwiga, Anna Matys, and Katarzyna Kieć-Kononowicz. 2013. "Recent Advances in Multi-Drug Resistance (MDR) Efflux Pump Inhibitors of Gram-Positive Bacteria S. aureus" Antibiotics 2, no. 1: 28-45. https://doi.org/10.3390/antibiotics2010028




