Application of DNA Aptamers and Quantum Dots to Lateral Flow Test Strips for Detection of Foodborne Pathogens with Improved Sensitivity versus Colloidal Gold
Abstract
:1. Introduction
2. Results and Discussion
2.1. Solving the Little Known Problems of Aptamer and Qdot Use in LF Test Strips
2.1.1. Immobilizing Capture Aptamers and Obtaining Intense Control Lines
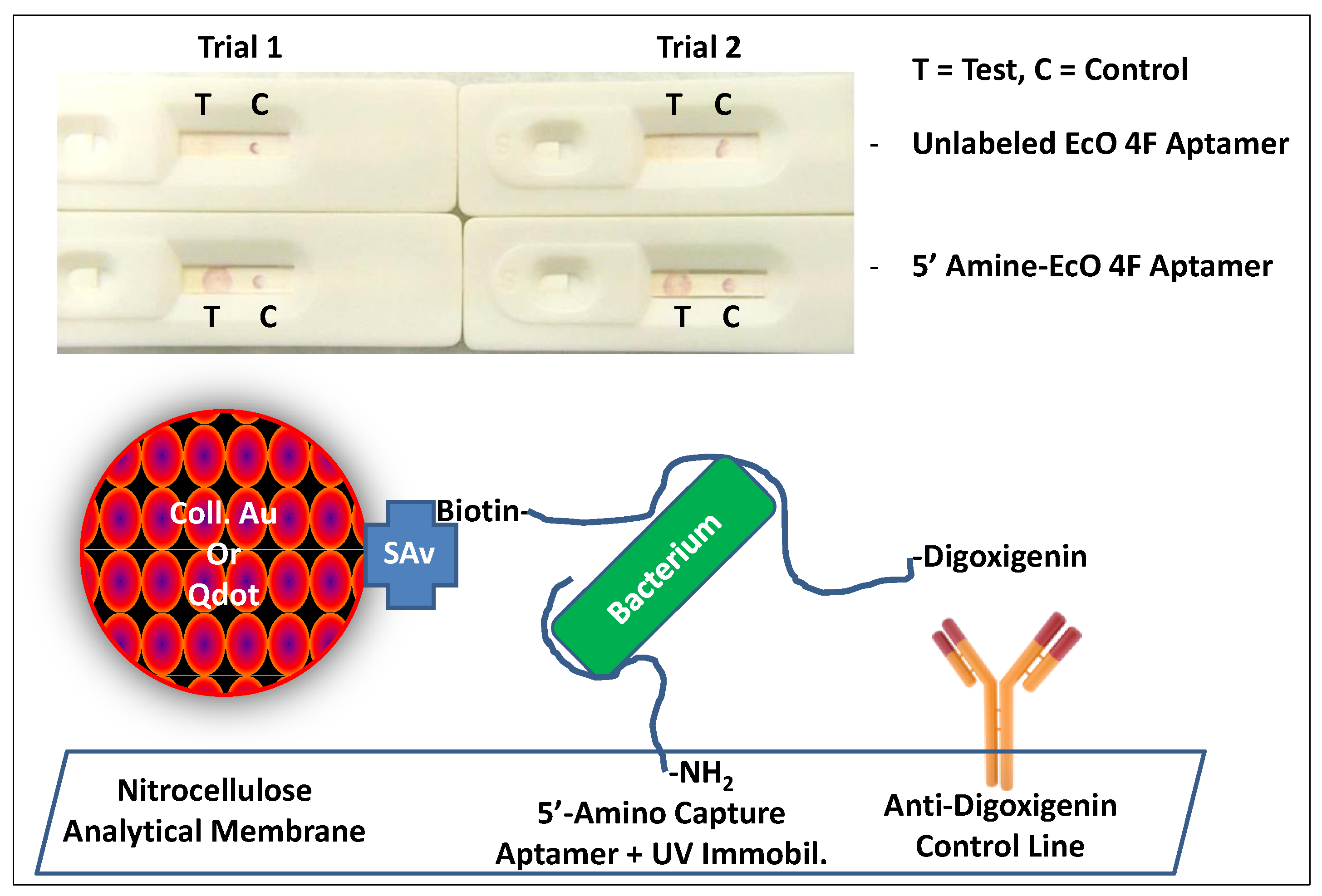
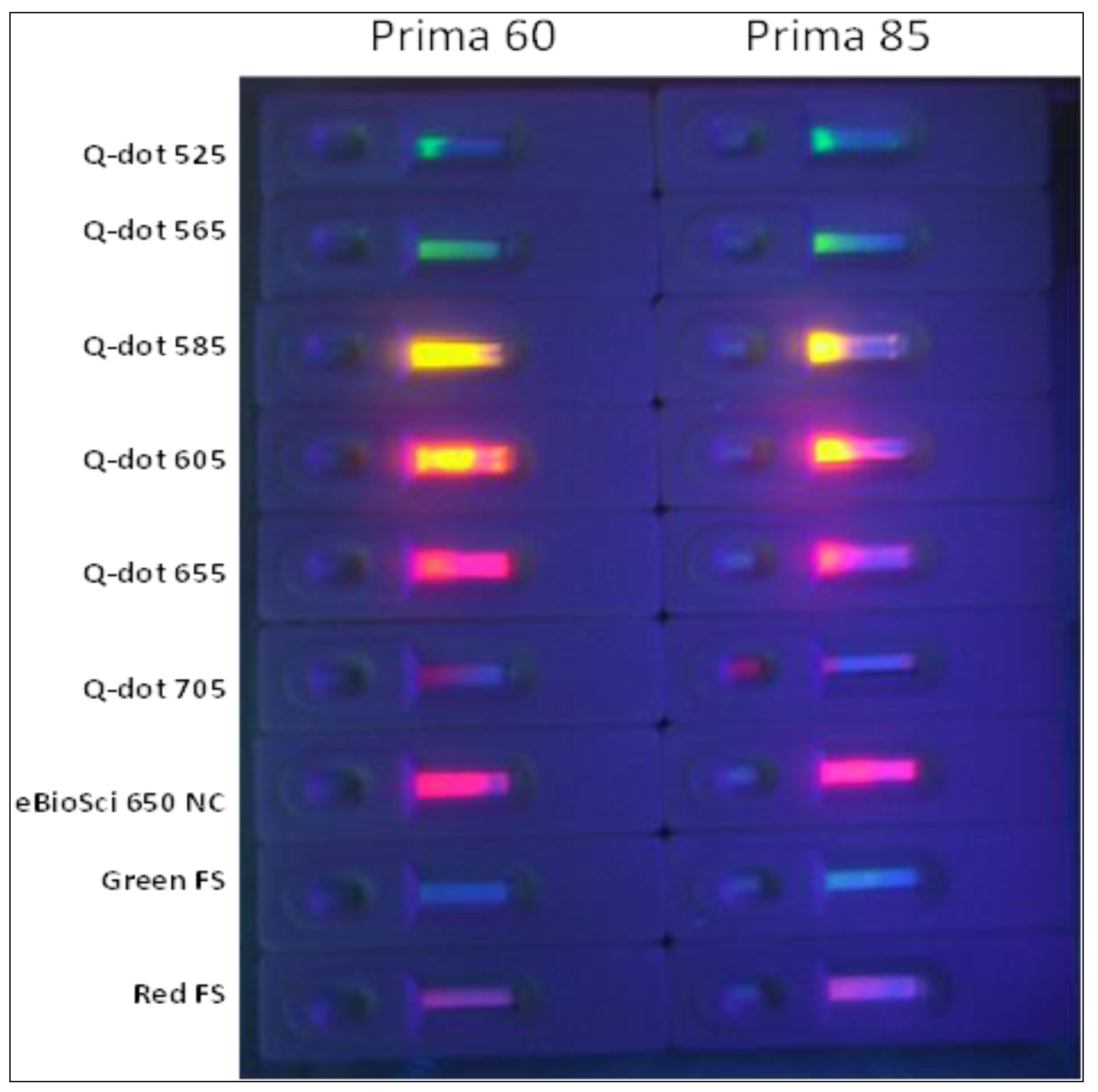
2.1.2. Sandwich Pair Screening and Choice of Optimal Analytical Membrane
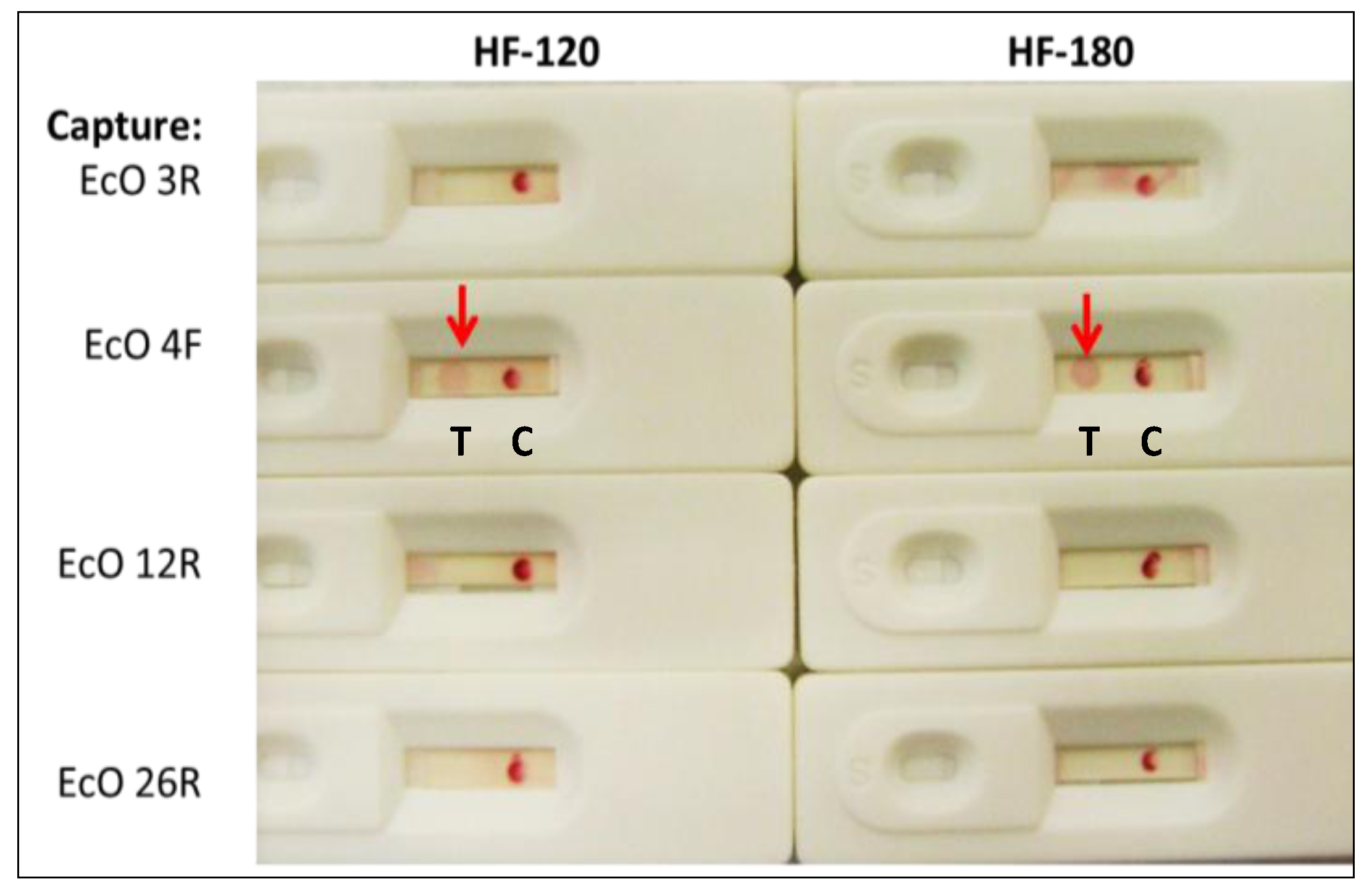

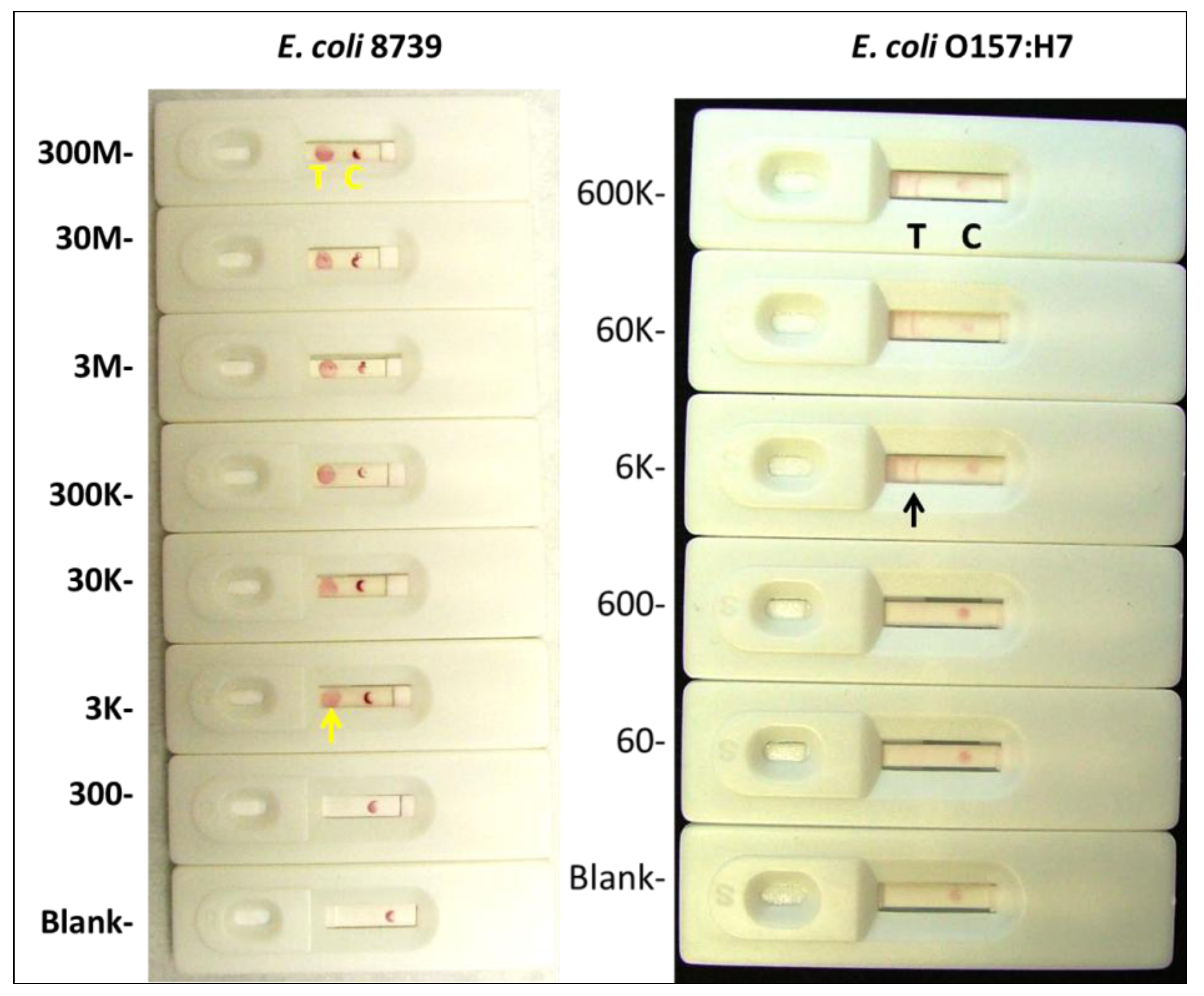
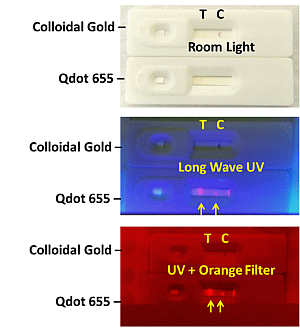
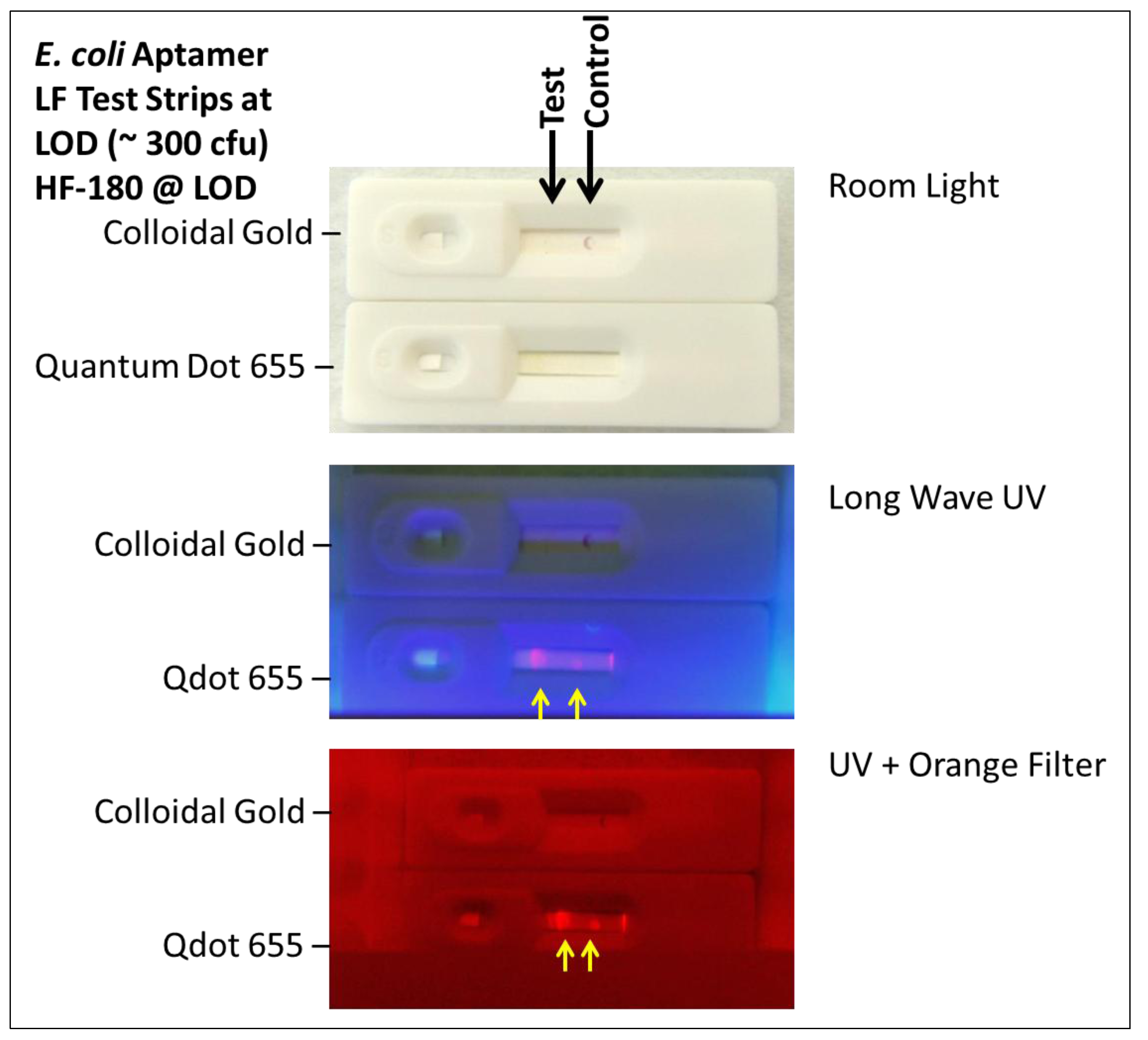
2.1.4. Visual Determination of LOD for Colloidal Gold and Qdot Versions of the E. coli LF Assay

2.1.5. Cross-Reactivity Assessment of the E. coli Aptamer LF Test
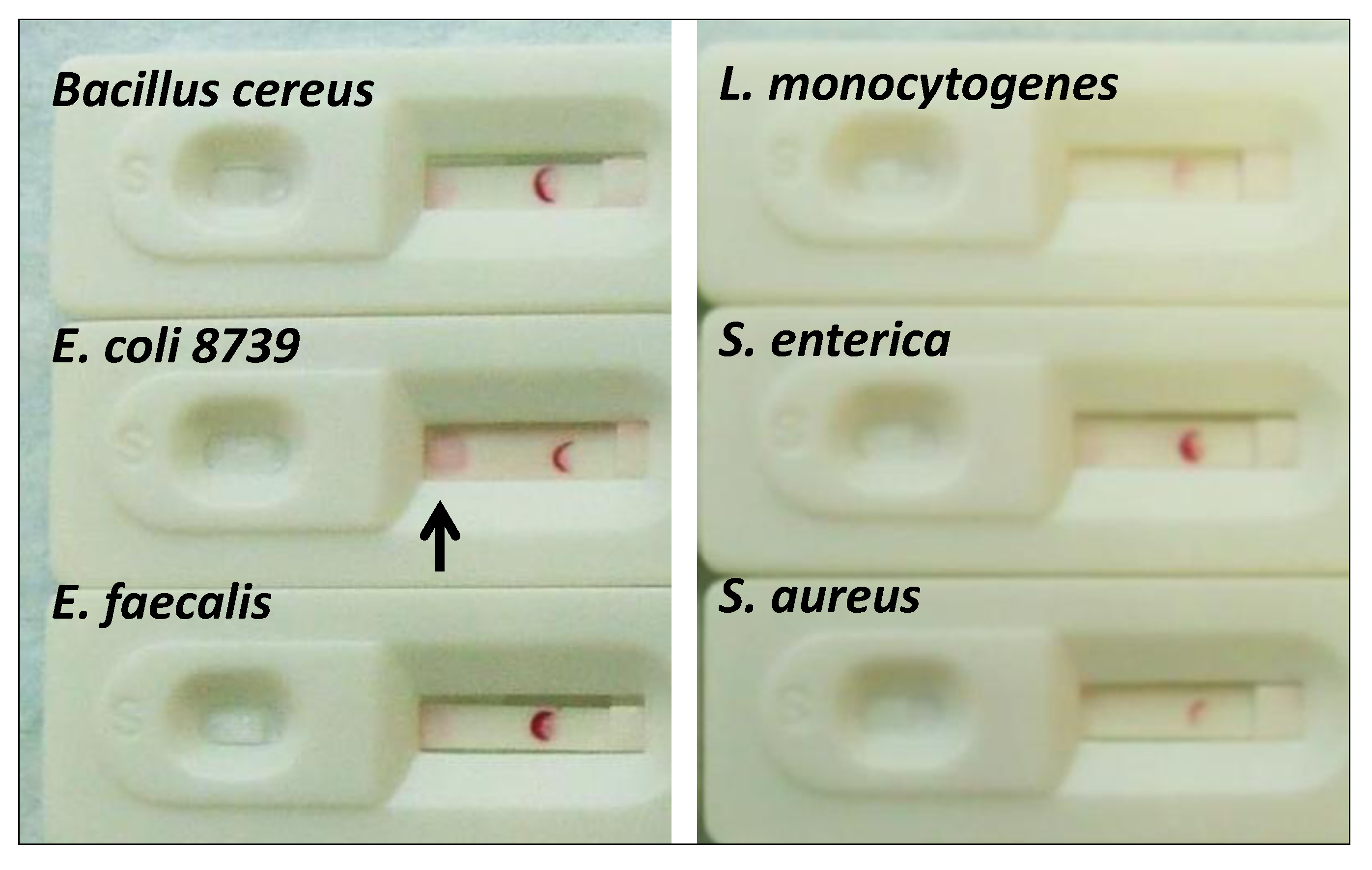
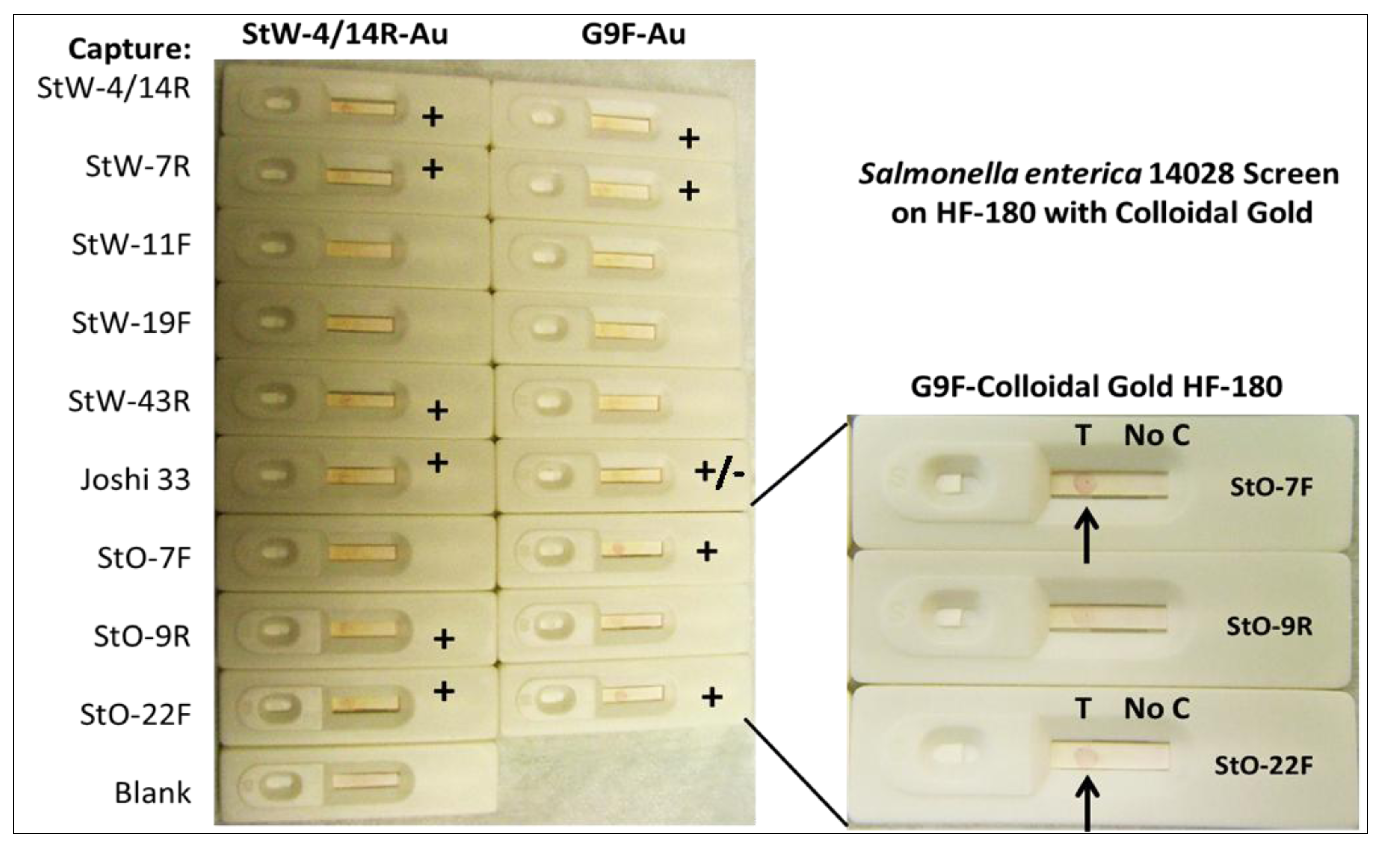
2.1.6. Screening for Other Potential Foodborne Pathogen LF Assays

3. Experimental Section
3.1. Preparation of Aptamer-Colloidal Gold and Qdot Conjugates
3.2. Assembly and Preparation of LF Test Strips
3.3. LF Screening, LOD Titration and Cross-Reactivity Experiments
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Yonekita, T.; Fujimura, T.; Morishita, N.; Matsumoto, T.; Morimatsu, F. Simple, rapid, and reliable detection of Escherichia coli O26 using immunochromatography. J. Food Prot. 2013, 76, 748–754. [Google Scholar] [CrossRef]
- Terao, Y.; Yonekita, T.; Morishita, N.; Fujimura, T.; Matsumoto, T.; Morimatsu, F. Potential rapid and simple lateral flow assay for Escherichia coli O111. J. Food Prot. 2013, 76, 755–761. [Google Scholar] [CrossRef]
- Bruno, J.G.; Carrillo, M.P.; Phillips, T. In vitro antibacterial effects of anti-lipopolysaccharide DNA aptamer-C1qrs complexes. Folia Microbiol. 2008, 53, 295–302. [Google Scholar] [CrossRef]
- Bruno, J.G.; Carrillo, M.P.; Phillips, T.; Andrews, C.J. A novel screening method for competitive FRET-aptamers applied to E. coli assay development. J. Fluoresc. 2010, 20, 1211–1223. [Google Scholar] [CrossRef]
- Bruno, J.G.; Carrillo, M.P. Development of aptamer beacons for rapid presumptive detection of Bacillus spores. J. Fluoresc. 2012, 22, 915–924. [Google Scholar] [CrossRef]
- Bruno, J.G.; Phillips, T.; Carrillo, M.P.; Crowell, R. Plastic-adherent DNA aptamer-magnetic bead and quantum dot sandwich assay for Campylobacter detection. J. Fluoresc. 2009, 19, 427–435. [Google Scholar] [CrossRef]
- Bruno, J.G.; Richarte, A.M.; Carrillo, M.P.; Edge, A. An aptamer beacon responsive to botulinum toxins. Biosens. Bioelectron. 2012, 31, 240–243. [Google Scholar] [CrossRef]
- Bruno, J.G.; Sivils, J.C.; Carrillo, M.P.; Richarte, A.M.; Phillips, T.; Andrews, C.; Garcia, A.; Montez, T.; Edge, A. Chapter 2: Novel aptamer-based diagnostic test formats. In Biomedical Applications of Aptamers; Bruno, J.G., Ed.; Nova Science Publishers: Hauppauge, NY, USA, 2013; pp. 15–32. [Google Scholar]
- Bruno, J.G.; Carrillo, M.P.; Phillips, T.; Savage, A.; Koke, J.R. Chapter 3: Aptamer-based detection and therapeutics to prevent and treat E. coli infections. In E. coli Infections: Causes, Treatment and Prevention; Rogers, M.C., Peterson, N.D., Eds.; Nova Science Publishers: Hauppauge, NY, USA, 2011; pp. 91–112. [Google Scholar]
- Joshi, R.; Janagama, H.; Dwivedi, H.P.; Senthil Kumar, T.M.; Jaykus, L.A.; Schefers, J.; Sreevatsan, S. Selection, characterization, and application of DNA aptamers for the capture and detection of Salmonella enterica serovars. Mol. Cell Probes 2009, 23, 20–28. [Google Scholar] [CrossRef]
- Moon, J.; Kim, G.; Lee, S.; Park, S. Identification of Salmonella Typhimurium-specific DNA aptamers developed using whole-cell SELEX and FACS analysis. J. Microbiol. Methods 2013, 95, 162–166. [Google Scholar] [CrossRef]
- Jayasena, S.D. Aptamers: An emerging class of molecules that rival antibodies in diagnostics. Clin. Chem. 1999, 45, 1628–1650. [Google Scholar]
- Marx, V. Finding the right antibody for the job. Nat. Methods 2013, 10, 703–703. [Google Scholar] [CrossRef]
- Bordeaux, J.; Welsh, A.W.; Agarwal, S.; Killiam, E.; Baquero, M.T.; Hanna, J.A.; Anagnostou, V.K.; Rimm, D.L. Antibody validation. BioTechniques 2010, 48, 197–209. [Google Scholar] [CrossRef]
- Vivekananda, J.; Kiel, J.L. Anti-Francisella tularensis DNA aptamers detect tularemia antigen from different subspecies by Aptamer-Linked Immobilized Sorbent Assay. Lab Invest. 2006, 86, 610–618. [Google Scholar]
- Bonenberger, J.; Doumanas, M. Overcoming sensitivity limitations of lateral-flow immunoassays with a novel labeling technique. IVD Technol. 2006, 5, 41–46. [Google Scholar]
- Juntunen, E.; Myyryläinen, T.; Salminen, T.; Soukka, T.; Pettersson, K. Performance of fluorescent europium(III) nanoparticles and colloidal gold reporters in lateral flow bioaffinity assay. Anal. Biochem. 2012, 428, 31–38. [Google Scholar] [CrossRef]
- Bai, Y.; Tian, C.; Wei, X.; Wang, Y.; Wang, D.; Shi, X. A sensitive lateral flow test strip based on silica nanoparticle/CdTe quantum dot composite reporter probes. RSC Adv. 2012, 2, 1778–1781. [Google Scholar] [CrossRef]
- Li, X.; Lu, D.; Sheng, Z.; Chen, K.; Guo, X.; Jin, M.; Han, H. A fast and sensitive immunoassay of avian influenza virus based on label-free quantum dot probe and lateral flow test strip. Talanta 2012, 100, 1–6. [Google Scholar] [CrossRef]
- Yang, H.; Li, D.; He, R.; Guo, Q.; Wang, K.; Zhang, X.; Huang, P.; Cui, D. A novel quantum dots-based point of care test for syphilis. Nanoscale Res. Lett. 2010, 5, 875–881. [Google Scholar] [CrossRef]
- Berlina, A.N.; Taranova, N.A.; Zherdev, A.V.; Vengerov, Y.Y.; Dzantiev, B.B. Quantum dot-based lateral flow immunoassay for detection of chloramphenicol in milk. Anal. Bioanal. Chem. 2013, 405, 4997–5000. [Google Scholar] [CrossRef]
- Berlina, A.N.; Taranova, N.A.; Zherdev, A.V.; Sankov, M.N.; Andreev, I.V.; Martynov, A.I.; Dzantiev, B.B. Quantum-dot-based immunochromatographic assay for total IgE in human serum. PLoS One 2013, 8, e77485. [Google Scholar] [CrossRef]
- Hampl, J.; Hall, M.; Mufti, N.A.; Yao, Y.M.; MacQueen, D.B.; Wright, W.H.; Cooper, D.E. Upconverting phosphor reporters in immunochromatographic assays. Anal. Biochem. 2001, 288, 176–187. [Google Scholar] [CrossRef]
- Queiros, R.B.; de-los-Santos-Álvarez, N.; Noronha, J.P.; Sales, M.G.F. A label-free DNA aptamer-based impedance biosensor for the detection of E. coli outer membrane proteins. Sens. Actuators B: Chem. 2013, 181, 766–772. [Google Scholar] [CrossRef]
- Smiley, S.; Derosa, M.; Blais, B. Immobilization of DNA aptamers on polyester cloth for antigen detection by dot blot immunoenzymatic assay (aptablot). J. Nucleic Acids 2013. [Google Scholar] [CrossRef]
- Seal, J.; Braven, H.; Wallace, P. Point-of-care nucleic acid lateral-flow tests. IVD Technol. 2006, 12, 41–51. [Google Scholar]
- Posthuma-Trumpie, G.A.; Korf, J.; van Amerongen, A. Lateral flow (immuno)assay: Its strengths, weaknesses, opportunities and threats. A literature survey. Anal. Bioanal. Chem. 2009, 393, 569–582. [Google Scholar] [CrossRef]
- Bruno, J.G.; Carrillo, M.P.; Richarte, A.M.; Phillips, T.; Andrews, C.; Lee, J.S. Development, screening, and analysis of a small DNA aptamer library potentially useful for diagnosis and passive immunity of arboviruses. BMC Res. Notes 2012, 5, 633. [Google Scholar] [CrossRef]
- Swift, J.L.; Cramb, D.T. Nanoparticles as fluorescence labels: Is size all that matters? Biophys. J. 2008, 95, 865–876. [Google Scholar] [CrossRef]
- McNamara, J.O.; Kolonias, D.; Pastor, F.; Mittler, R.S.; Chen, L.; Giangrande, P.H.; Sullenger, B.; Gilboa, E. Multivalent 4–1BB binding aptamers costimulate CD8+ T cells and inhibit tumor growth in mice. Clin. Invest. 2008, 118, 376–386. [Google Scholar] [CrossRef]
- Mallikaratchy, P.R.; Ruggiero, A.; Gardner, J.R.; Kuryavyi, V.; Maguire, W.F.; Heaney, M.L.; McDevitt, M.R.; Patel, D.J.; Scheinberg, D.A. A multivalent DNA aptamer specific for the B-cell receptor on human lymphoma and leukemia. Nucleic Acids Res. 2011, 39, 2458–2469. [Google Scholar] [CrossRef]
- Zhao, X.; Lis, J.T.; Shi, H. A systematic study of the features critical for designing a high avidity multivalent aptamer. Nucleic Acid Ther. 2013, 23, 238–242. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Bruno, J.G. Application of DNA Aptamers and Quantum Dots to Lateral Flow Test Strips for Detection of Foodborne Pathogens with Improved Sensitivity versus Colloidal Gold. Pathogens 2014, 3, 341-355. https://doi.org/10.3390/pathogens3020341
Bruno JG. Application of DNA Aptamers and Quantum Dots to Lateral Flow Test Strips for Detection of Foodborne Pathogens with Improved Sensitivity versus Colloidal Gold. Pathogens. 2014; 3(2):341-355. https://doi.org/10.3390/pathogens3020341
Chicago/Turabian StyleBruno, John G. 2014. "Application of DNA Aptamers and Quantum Dots to Lateral Flow Test Strips for Detection of Foodborne Pathogens with Improved Sensitivity versus Colloidal Gold" Pathogens 3, no. 2: 341-355. https://doi.org/10.3390/pathogens3020341




