Reticulitermes nelsonae, a New Species of Subterranean Termite (Rhinotermitidae) from the Southeastern United States
Abstract
:1. Introduction
| Reference citation | Caste, notes | Pages | Species from USA |
|---|---|---|---|
| Banks, N. & Snyder, T.E. (1920) | (a, s) descriptions, illustrations, key, flight times | 42–47, 148–164 | Rf, Rv, Rh, Rhp, Rt |
| Goellner, E. J. (1931) | (a, s) descriptions, illustrations, ecology | 227–234 | Rf, Rt, Ra |
| Kofoid, C.A. (1934) | (a), descriptions, flight times | 193–194 | Rf, Rv, Rh, Ra, Rhp, Rt |
| Miller, E.M. & Miller, D.B. (1943) | (a, s) descriptions, illustrations, flight times | 101–107 | Rf, Rv, Rh |
| Banks, F.A. (1946) | (a, s) descriptions, illustrations, key, flight times | 1–29 | Rf, Rv, Ra, Rt |
| Miller E.M. (1949) | (a, s) descriptions, illustrations, key, flight times | 6–7, 14–15, 20–22, 26 | Rf, Rv, Rh |
| Snyder, T.E. (1954) | (a, s) descriptions, illustrations, key, flight times | 26, 51–56 | Rf, Rv, Rh, Ra |
| Miller E. M. (1964) | (a) flight times | 5, 16 | Rf, Rv, Rh |
| Weesner, F.M. (1965) | (a) descriptions, illustrations, key, flight times | 36–44, 51 | Rf, Rv, Rh, Ra, Rhp, Rt |
| Clément, J.L., Howard, R., Blum, M. & Lloyd, H. (1986) | (a, s) descriptions, life history | 67–70 | Rf, Rv, Rh, Rm |
| Nutting, W.L. (1990) | (a, s) descriptions, illustrations, key, flight times | 997–1030 | Rf, Rv, Rh, Ra, Rhp, Rt |
| Scheffrahn RH, Su N-Y. (1994) | (a, s) descriptions, illustrations, key, flight times | 465–473 | Rf, Rv, Rh |
| Hostettler, N.C., Hall, D.W. & Scheffrahn, R.H. (1995) | (s) descriptions, photographs | 119–129 | Rf, Rv, Rh |
| Ye, W., Lee, C.-Y., Scheffrahn, R.H., Aleong, J.M., Su, N.-Y., Bennett, G.W., Scharf, M.E., (2004) | (s) descriptions, photographs | 815–822 | Rf, Rv, Rh, Ra |
| Brown, K., Kard, B. & Payton, M. (2005) | (a, s) descriptions, photograph | 277–284 | Rf, Rv, Rh |
| Austin, J.W., Bagnères, A.-G., Szalanski, A.L., Scheffrahn, R.H., Heintschel, B.P., Messenger, M.T., Clement, J.-L. & Gold, R.E. (2007) | (a, s) descriptions, illustrations, photographs, flight times | 1–26 | Rf, Rv, Rh, Rm, Ra, Rhp, Rt |
| Wang, C., Zhou, X., Li, S., Schwinghammer, M., Scharf, M., Buczkowski, G. & Bennett, G. (2009) | (a, s) descriptions, key, photographs | 1029–1036 | Rf, Rv, Rh, Ra, Rt |
2. Materials and Methods
2.1. Specimens
| Soldier | |||
| Site ID | County | Sample Size | Collection Date |
| Site 1s | McIntosh Co. | 7 | Nov 2007 |
| Site 2s | McIntosh Co. | 16 | Feb 2007 |
| Site 3s | McIntosh Co. | 9 | Feb 2007 |
| Site 4s | McIntosh Co. | 2 | Feb 2007 |
| Site 5s | McIntosh Co. | 3 | Feb 2007 |
| Site 6s | McIntosh Co. | 4 | Feb 2007 |
| Site 7s | McIntosh Co. | 11 | Feb 2007 |
| Site 8s | McIntosh Co. | 2 | Jul 2009 |
| Site 9s | McIntosh Co. | 1 | Jul 2009 |
| Site 10s | McIntosh Co. | 2 | Jul 2009 |
| Site 11s | McIntosh Co. | 2 | Jul 2009 |
| Site 12s | Thomas Co. | 1 | Nov 2009 |
| Site 13s | Thomas Co. | 2 | Nov 2009 |
| Site 14s | Thomas Co. | 4 | Nov 2009 |
| Site 15s | Thomas Co. | 1 | Nov 2009 |
| Site 16s | Thomas Co. | 2 | Nov 2009 |
| Site 17s | Thomas Co. | 1 | Nov 2009 |
| Site 18s | Thomas Co. | 2 | Nov 2009 |
| Site 19s | Thomas Co. | 1 | Nov 2009 |
| Site 20s | Thomas Co. | 2 | Nov 2009 |
| Site 21s | Thomas Co. | 2 | Nov 2009 |
| Site 22s | Thomas Co. | 1 | Nov 2009 |
| Site 23s | Thomas Co. | 1 | Nov 2009 |
| Site 24s | Thomas Co. | 1 | Nov 2009 |
| Site 25s | Thomas Co. | 1 | Nov 2009 |
| Site 26s | Thomas Co. | 1 | Nov 2009 |
| Site 27s | Thomas Co. | 2 | Nov 2009 |
| Site 28s | Thomas Co. | 3 | Nov 2009 |
| Site 29s | Thomas Co. | 2 | Nov 2009 |
| Site 30s | Thomas Co. | 2 | Nov 2009 |
| Site 31s | Thomas Co. | 1 | Nov 2009 |
| Site 32s | Thomas Co. | 1 | Nov 2009 |
| Site 33s | Thomas Co. | 1 | Jan 2010 |
| Site 34s | Thomas Co. | 1 | Jan 2010 |
| Site 35s | Thomas Co. | 2 | Mar 2010 |
| Total = 35 | N = 96 | ||
| Alate | |||
| Site ID | County | Sample Size | Collection Date |
| Site 4a | McIntosh Co. | 14 | Feb 2007 |
| Site 6a | McIntosh Co. | 6 | Feb 2007 |
| Site 4a | McIntosh Co. | 19 | Mar 2007 |
| Site 36a | McIntosh Co. | 19 | Mar 2007 |
| Site 3a | McIntosh Co. | 5 | Feb 2007 |
| Site 2a | McIntosh Co. | 14 | Feb 2007 |
| Site 5a | McIntosh Co. | 1 | Feb 2007 |
| Site 7a | McIntosh Co. | 13 | Feb 2007 |
| Site 37a | McIntosh Co. | 50 | May 2005 |
| Total = 9 | N = 141 | ||
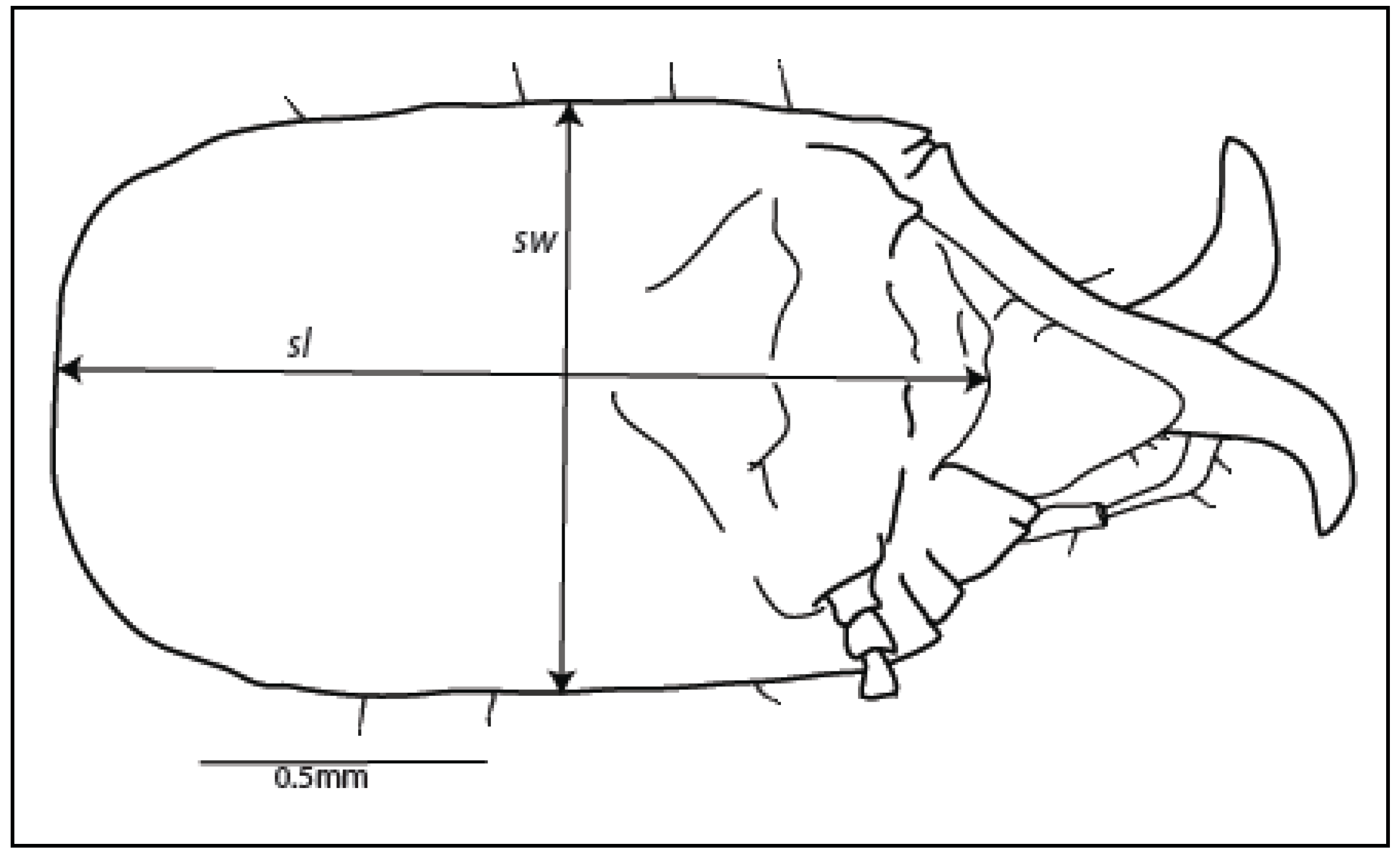
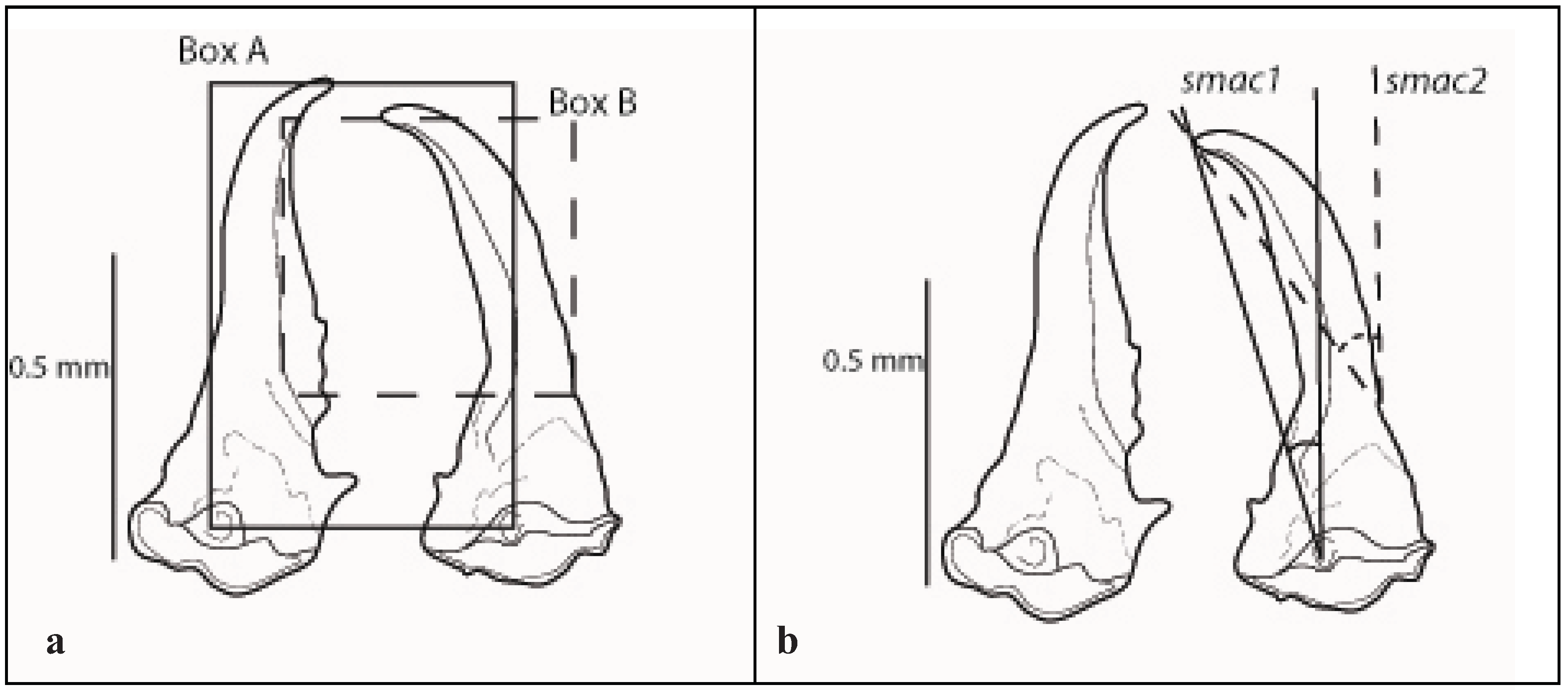
2.2. Soldier
2.3. Alate
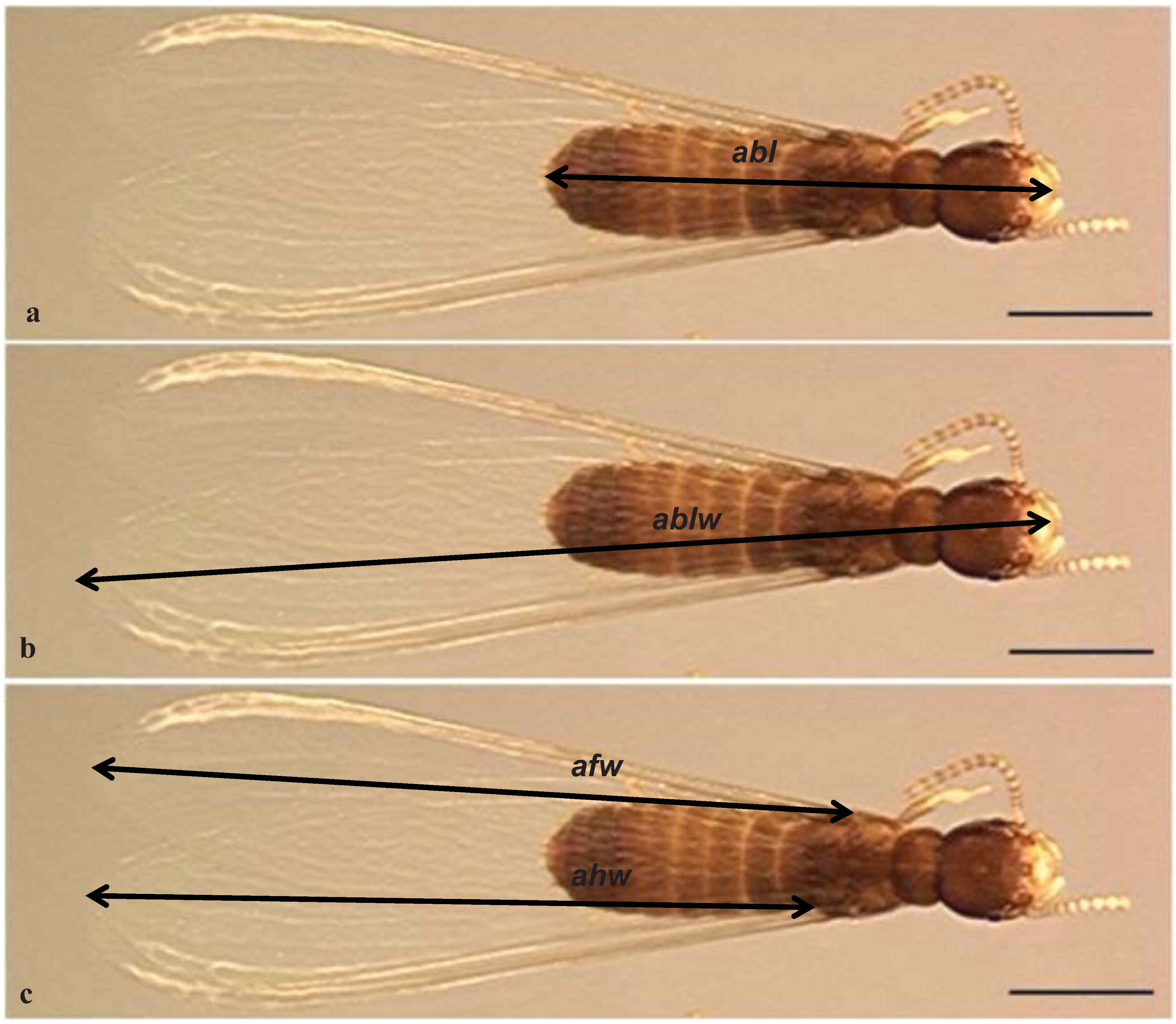

2.4. Morphometrics, Imaging and Statistics
2.5. Behavior
2.6. Dichotomous Key
2.7. Molecular Data
| Gene of interest | Primer pair (forward and reverse) | Sequences (5’→3’) |
|---|---|---|
| COII (685 bp) | TL2-J-3037 | ATG GCA GAT TAG TGC AAT GG |
| TK-N-3785 | GTT TAA GAG ACC AGT ACT TG | |
| COI (~800 bp) (partial sequence) | C1-J-2195 | TTG ATT CTT TTG GTC ACT CCA TGA AGT |
| TL2-N-3014 | TCC TAA TTG CAC TTA ATC TGC CAT ATT |
3. Results and Discussion
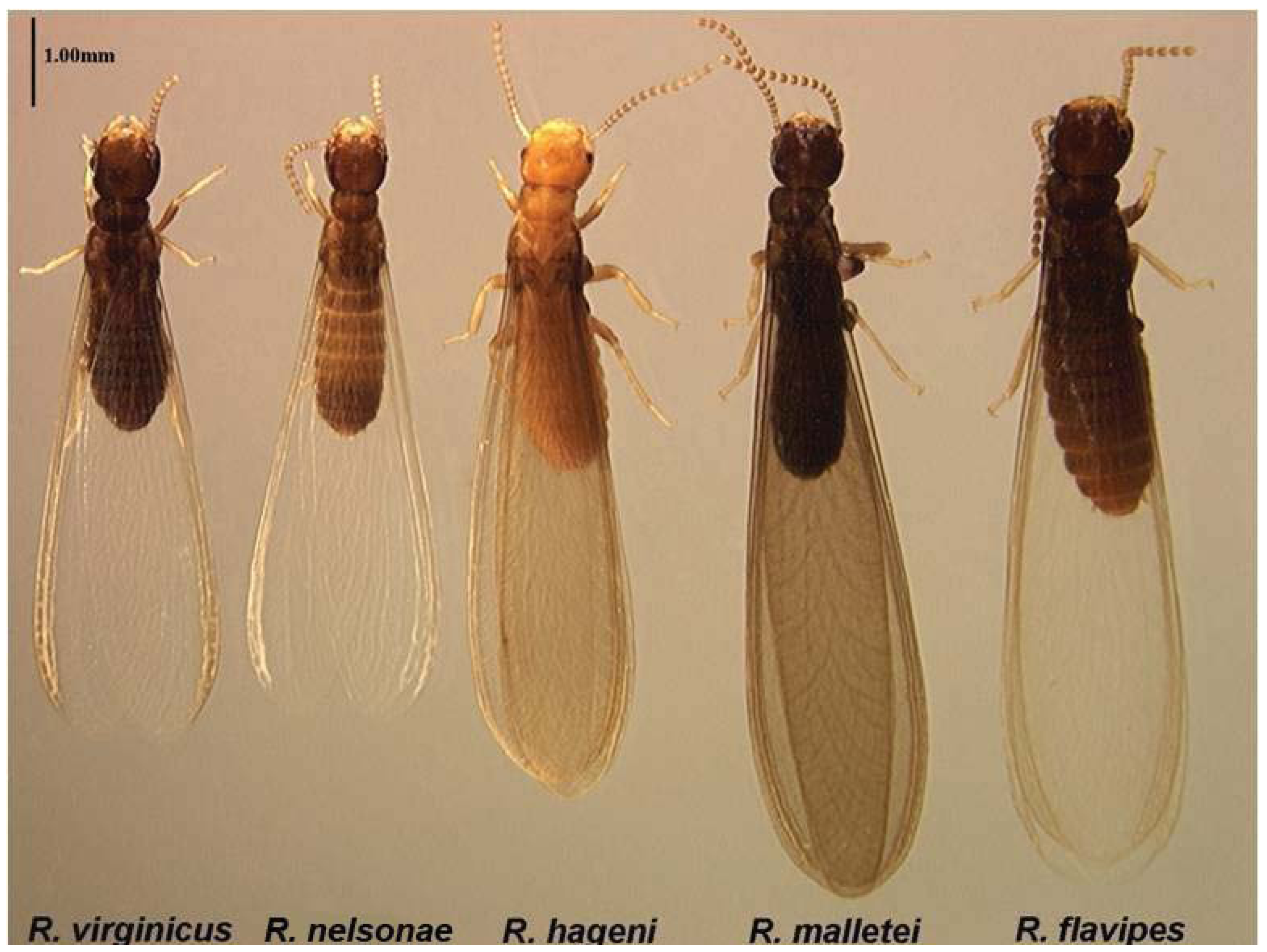
3.1. Morphological Characters
3.2. Behavior
| Soldier head capsule | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Species | Length (sl), mm | Width (sw), mm | Ratio of length: width (sl:sw) | ||||||||||||
| Mean | Std. dev. | Min. a | Max. b | Range of mean ± 1 | Mean | Std. dev. | Min. a | Max. b | Range of mean ± 1 | Mean | Std. dev. | Min. a | Max. b | Range of mean ± 1 | |
| R. flavipes | 1.693 | 0.119 | 1.21 | 1.91 | 1.57–1.81 | 1.044 | 0.074 | 0.73 | 1.17 | 0.97–1.12 | 1.625 | 0.084 | 1.43 | 1.83 | 1.54–1.71 |
| R. virginicus | 1.625 | 0.068 | 1.37 | 1.84 | 1.56–1.69 | 0.920 | 0.039 | 0.76 | 1.01 | 0.88–0.96 | 1.767 | 0.063 | 1.56 | 1.95 | 1.70–1.83 |
| R. hageni | 1.434 | 0.092 | 1.16 | 1.59 | 1.34–1.53 | 0.862 | 0.030 | 0.75 | 0.91 | 0.82–0.90 | 1.656 | 0.073 | 1.44 | 1.78 | 1.58–1.73 |
| R. malletei | 1.490 | 0.058 | 1.33 | 1.64 | 1.43–1.55 | 0.879 | 0.029 | 0.78 | 0.95 | 0.85–0.91 | 1.695 | 0.066 | 1.52 | 1.87 | 1.63–1.76 |
| R. nelsonae | 1.407 | 0.127 | 1.14 | 1.72 | 1.28–1.42 | 0.784 | 0.054 | 0.70 | 0.99 | 0.73–0.84 | 1.793 | 0.092 | 1.52 | 1.98 | 1.70–1.89 |
| Soldier mandible angle of curvature | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Species | smac1, ° | smac2, ° | ||||||||
| Mean | Std. dev. | Min. a | Max. b | Range of Mean ± 1 | Mean | Std. dev. | Min. a | Max. b | Range of Mean ± 1 | |
| R. flavipes | 10.61 | 2.089 | 7.3 | 15.1 | 8.0–13.0 | 27.29 | 2.730 | 22.1 | 31.4 | 25.0–30.0 |
| R. virginicus | 13.62 | 1.719 | 11.7 | 18.1 | 12.0–15.0 | 32.60 | 2.003 | 29.4 | 36.7 | 31.0–35.0 |
| R. hageni | 8.39 | 1.199 | 6.9 | 11.2 | 7.0–10.0 | 23.39 | 1.307 | 21.5 | 27.3 | 22.0–25.0 |
| R. malletei | 10.51 | 1.366 | 8.1 | 13.2 | 9.0–12.0 | 25.85 | 2.282 | 22.4 | 32.9 | 24.0–28.0 |
| R. nelsonae | 10.66 | 2.206 | 7.2 | 14.6 | 8.0–13.0 | 27.27 | 2.654 | 24.1 | 34.0 | 25.0–30.0 |
| Species | Body (abl), mm | Body-wing (ablw), mm | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Mean | Std. dev. | Min. a | Max. b | Range of mean ± 1 | Mean | Std. dev. | Min. a | Max. b | Range of mean ± 1 | |
| R. flavipes | 4.783 | 0.383 | 3.77 | 5.83 | 4.40–5.17 | 8.973 | 0.402 | 8.05 | 9.94 | 8.57–9.38 |
| R. virginicus | 4.021 | 0.214 | 3.56 | 4.44 | 3.81–4.24 | 7.414 | 0.213 | 6.89 | 7.90 | 7.20–7.63 |
| R. hageni | 4.083 | 0.323 | 3.41 | 5.35 | 3.76–4.41 | 7.810 | 0.318 | 7.25 | 8.64 | 7.49–8.13 |
| R. malletei | 4.023 | 0.302 | 3.53 | 4.99 | 3.72–4.33 | 8.238 | 0.394 | 6.91 | 9.28 | 7.84–8.63 |
| R. nelsonae | 3.928 | 0.236 | 3.26 | 4.63 | 3.69–4.16 | 7.080 | 0.291 | 6.53 | 7.88 | 6.79–7.37 |
| Species | Forewing (afw), mm | Hind wing (ahw), mm | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Mean | Std. dev. | Min. a | Max. b | Range of mean ± 1 | Mean | Std. dev. | Min. a | Max. b | Range of mean ± 1 | |
| R. flavipes | 6.810 | 0.331 | 5.97 | 7.74 | 6.48–7.14 | 6.550 | 0.332 | 5.70 | 7.44 | 1.29–1.35 |
| R. virginicus | 5.532 | 0.193 | 5.15 | 6.05 | 5.34–5.73 | 5.418 | 0.193 | 4.78 | 5.78 | 1.32–1.37 |
| R. hageni | 5.965 | 0.263 | 5.47 | 6.52 | 5.70–6.23 | 5.739 | 0.257 | 5.24 | 6.19 | 1.28–1.34 |
| R. malletei | 6.375 | 0.328 | 5.13 | 7.31 | 6.04–6.70 | 6.100 | 0.339 | 4.92 | 6.95 | 1.27–1.32 |
| R. nelsonae | 5.430 | 0.212 | 4.94 | 5.98 | 5.22–5.64 | 5.315 | 0.297 | 4.81 | 6.21 | 1.27–1.31 |
3.3. Dichotomous Key
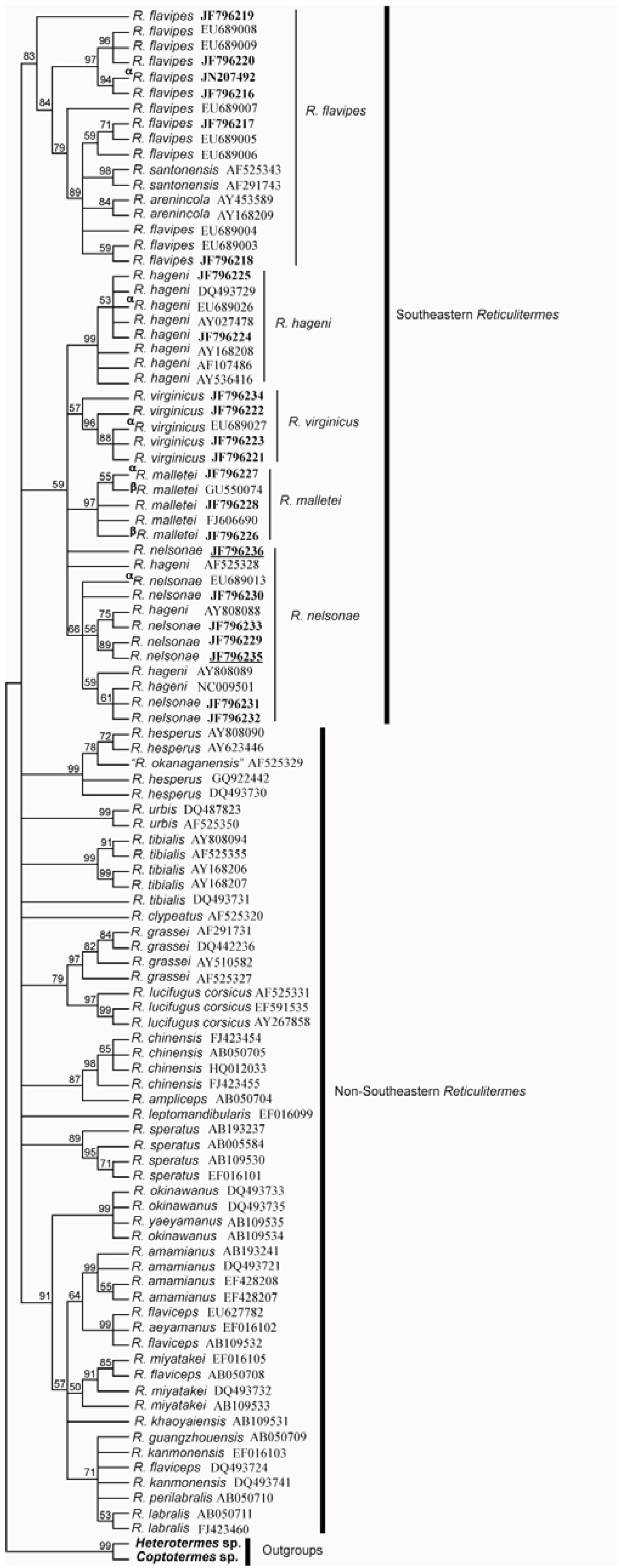
3.4. Molecular Data
| Species | R. flavipes | R. virginicus | R. hageni | R. malletei | R. nelsonae |
|---|---|---|---|---|---|
| R. flavipes | |||||
| R. virginicus | 0.052 | ||||
| R. hageni | 0.047 | 0.033 | |||
| R. malletei | 0.046 | 0.036 | 0.031 | ||
| R. nelsonae | 0.059 | 0.049 | 0.044 | 0.039 |
| Species | R. flavipes | R. virginicus | R. hageni | R. malletei | R. nelsonae |
|---|---|---|---|---|---|
| R. flavipes | |||||
| R. virginicus | 0.051 | ||||
| R. hageni | 0.048 | 0.040 | |||
| R. malletei | 0.042 | 0.037 | 0.027 | ||
| R. nelsonae | 0.050 | 0.044 | 0.036 | 0.029 |
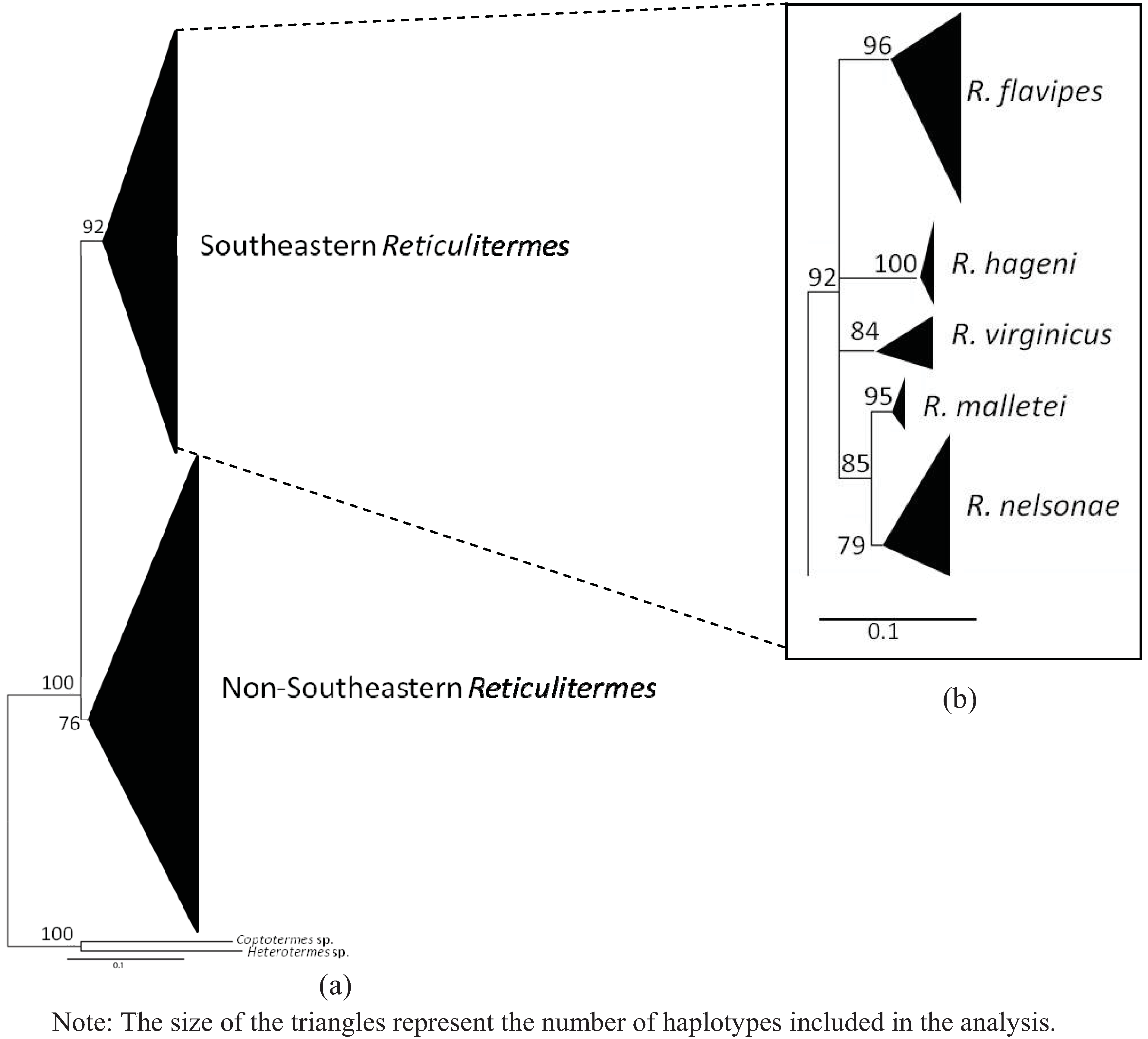
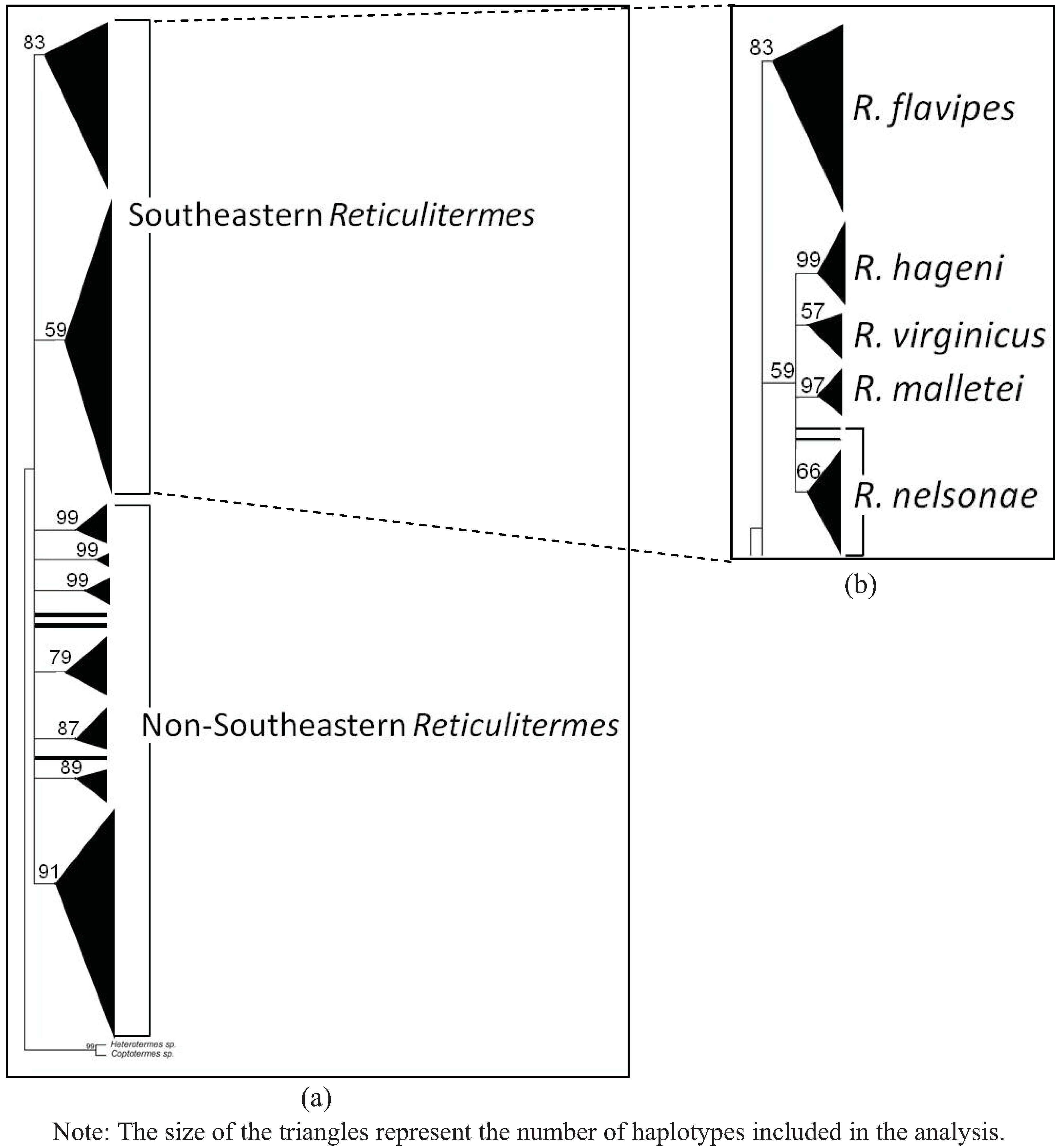
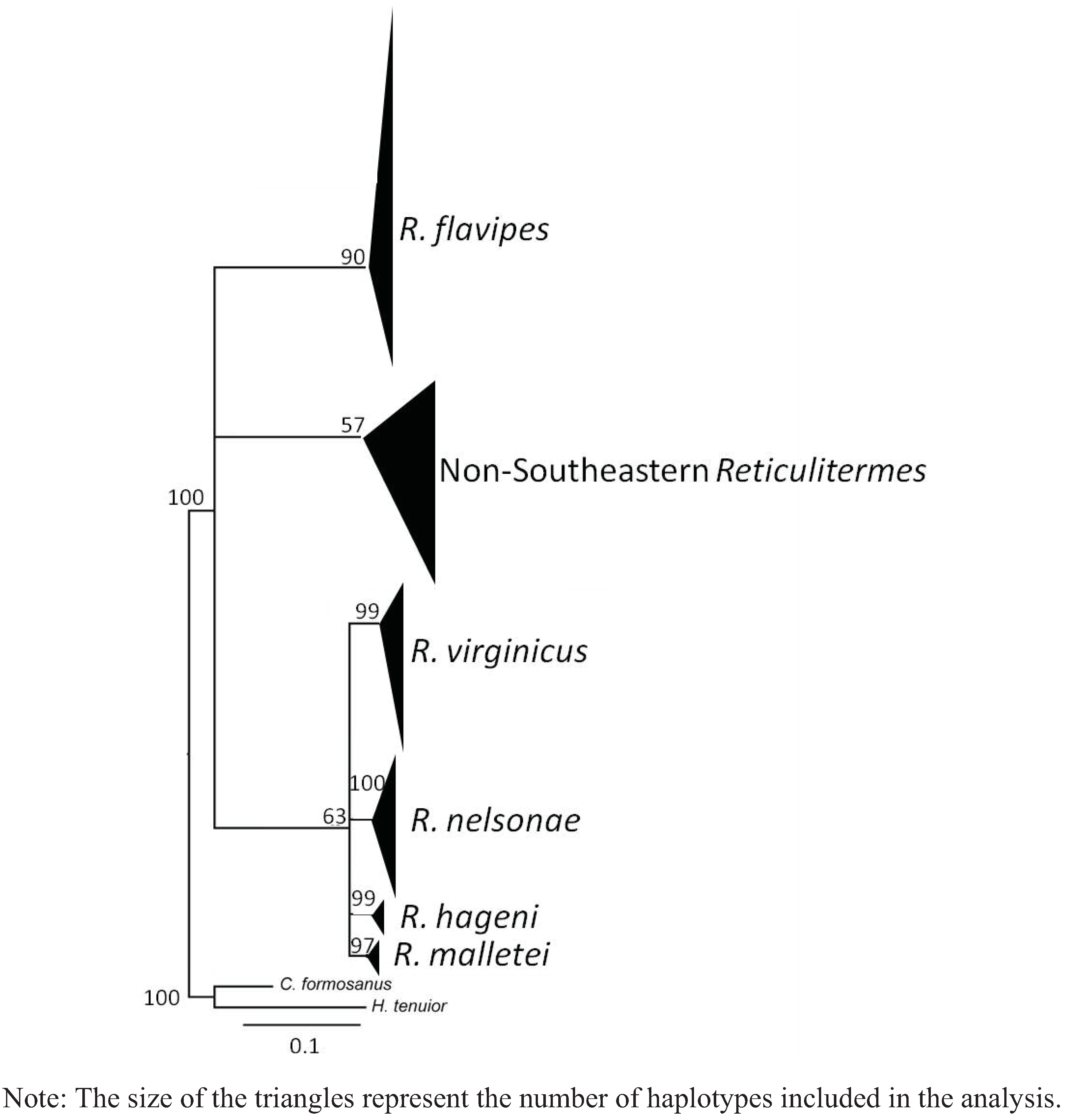

4. Systematics
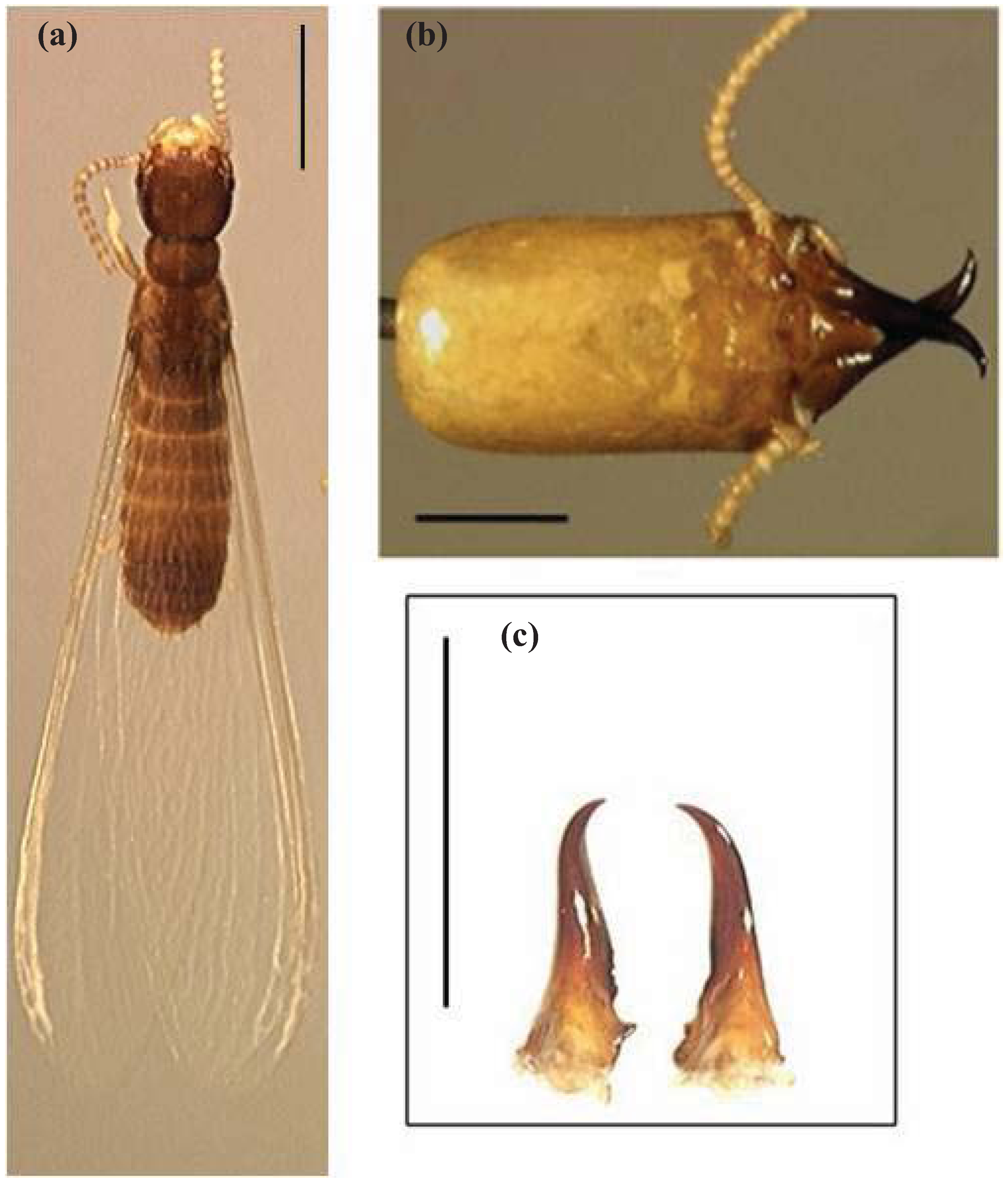

4.1. Diagnosis
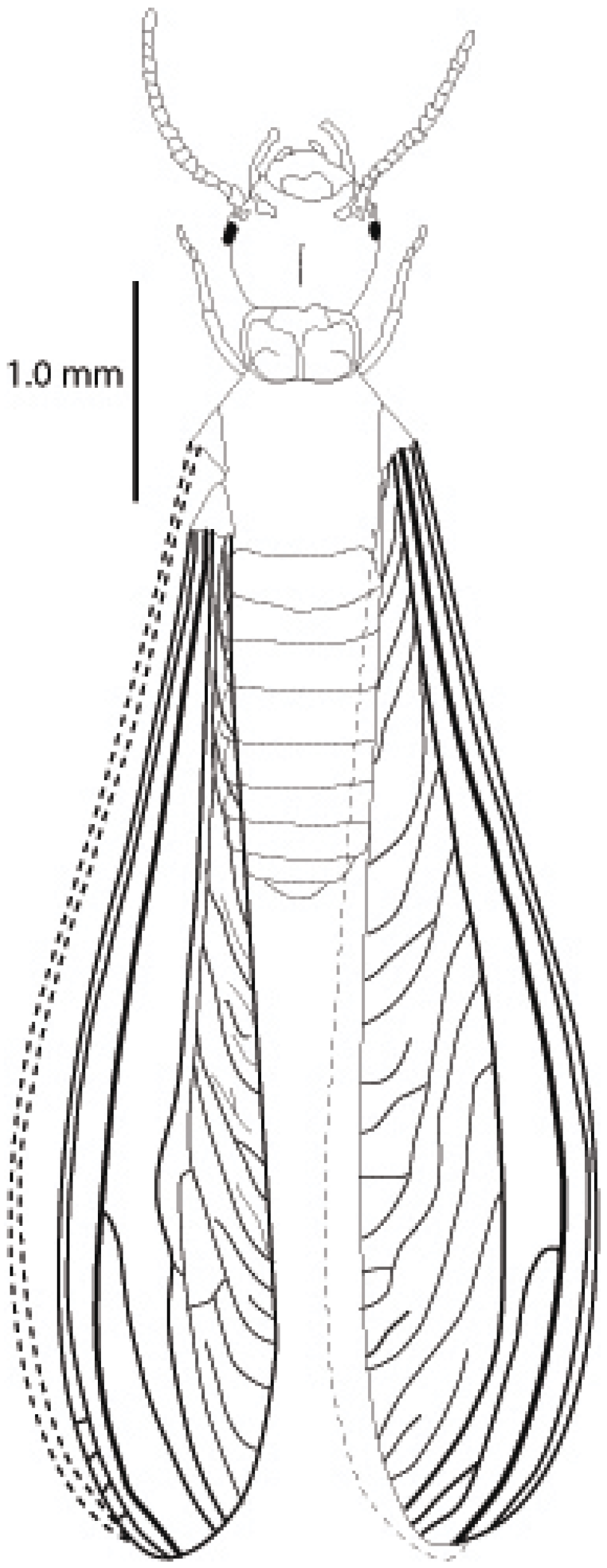
4.2. Description
4.3. Etymology

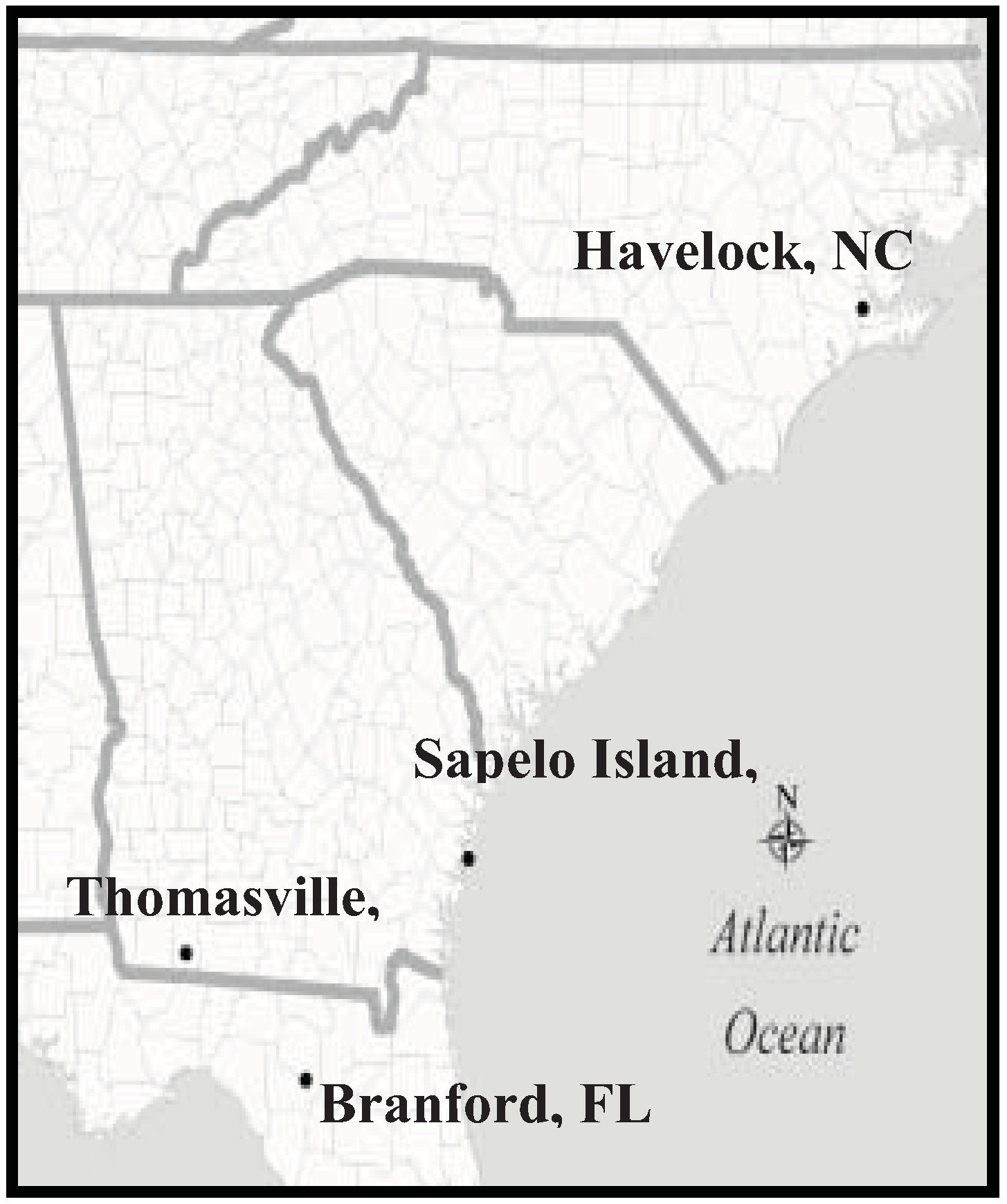
4.4. Distribution
4.5. Comments/Remarks
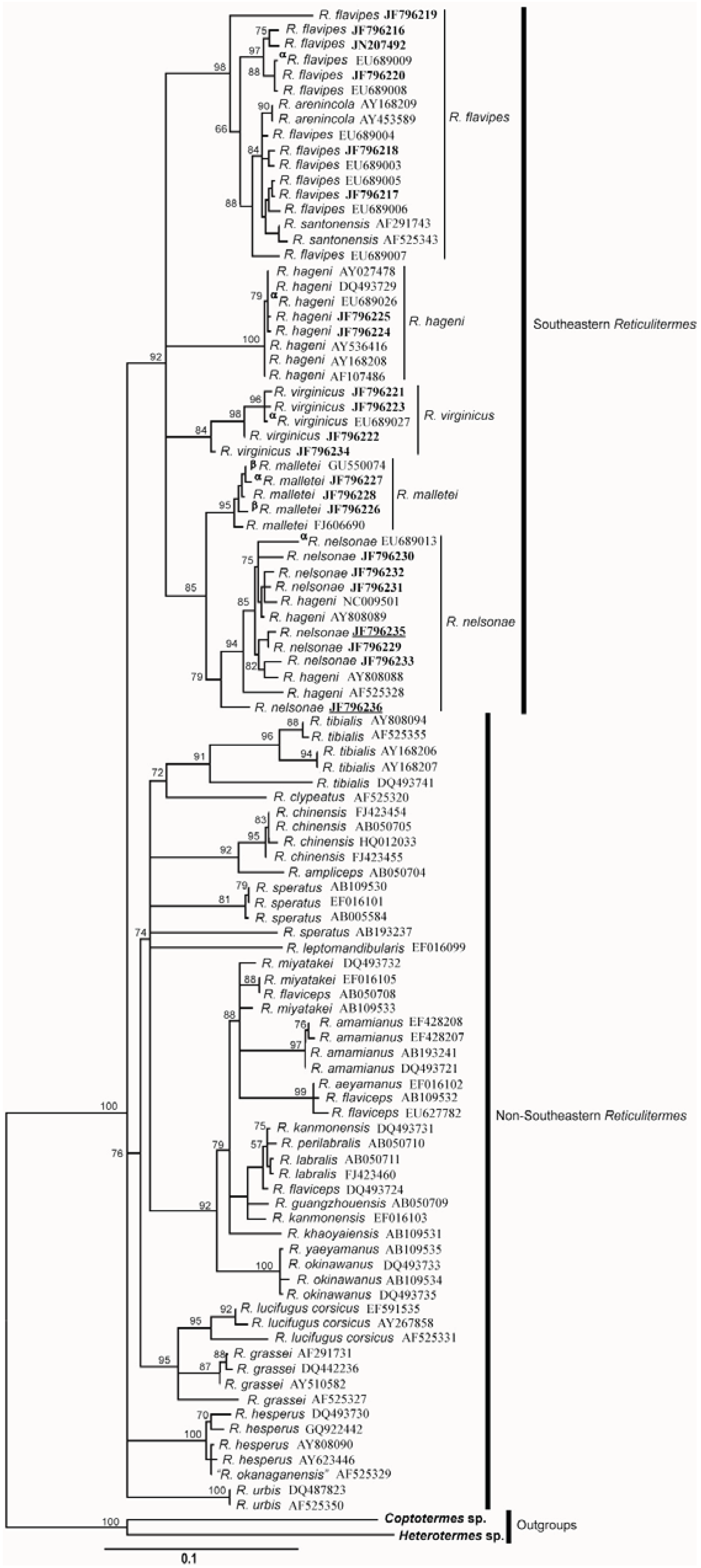
4.6. Genetics
5. Discussion
5.1. Cuticular Hydrocarbon
5.2. Morphology
5.3. Genetics
6. Conclusions
Acknowledgments
References
- Miller, E.M. Biology of Termites; University of Miami Press: Coral Gables, FL, USA, 1964; p. 30. [Google Scholar]
- Thorne, B.L.; Russek-Cohen, E.; Forschler, B.T.; Breisch, N.L.; Traniello, J.F.A. Evaluation for mark-release-recapture methods for estimating forager population size of subterranean termite (Isoptera: Rhinotermitidae) colonies. Environ. Entomol. 1996, 25, 938–951. [Google Scholar]
- Su, N.-Y.; Scheffrahn, R.H. Economically important termites in the United States and their control. Sociobiology 1990, 17, 77–94. [Google Scholar]
- Mallis, A. Handbook of Pest Control, 10 ed.; The Mallis Handbook Company: Cleveland, Ohio, USA, 2011; p. 1599. [Google Scholar]
- Thorne, B.L.; Forschler, B.T. NPMA Research Report on Subterranean Termite: Biology of Subterranean Termites of the Genus Reticulitermes & Subterranean Termite Biology in Relation to Prevention and Removal of Structural Infestation; NPMA: Dunn Loring, VA, USA, 1999. [Google Scholar]
- Abe, T.; Bignell, D.E.; Higashi, M. Termites: Evolution, Sociality, Symbioses, Ecology; Kluwer Academic Publishers: Dordrecht, The Netherlands; Boston, MA, USA, 2000. [Google Scholar]
- Su, N.Y.; Scheffrahn, R.H.; Cabrera, B. Native Subterranean Termites: Reticulitermes Flavipes (Kollar),Reticultiermes Virginicus (Banks),Reticulitermes Hageni Banks (Insecta: Isoptera: Rhinotermitidae); University of Florida Cooperative Extension Service: Lackawanna, PA, USA, 2001; Publication #EENY-212. [Google Scholar]
- Logan, J.W.M.; Rajagopal, D.; Wightman, J.A.; Pearce, M.J. Control of termites and other soil pests of groundnuts with special reference to controlled release formulations of non-persistent insecticides in India and Sudan. Bull. Environ. Res. 1992, 82, 57–66. [Google Scholar] [CrossRef]
- Pearce, M.J. Termites: Biology and Pest Management; CAB International: Cambridge, MA, USA, 1997. [Google Scholar]
- Banks, N.; Snyder, T.E. A Revision of the Nearctic Termites; Smithsonian Institution, United States National Museum: Washington DC, USA, 1920; Volume 108, p. 228. [Google Scholar]
- Clément, J.L.; Howard, R.; Blum, M.; Lloyd, H. L'isolement spécifique des termites du genre Reticulitermes (Isoptera) du sud-est des États-Unis. Mise en evidence grace a la chimie et au portement d' une espéce jumelle de R. virginicus = R. malletei sp.nov. et d' une semi-species de R. flavipes. Comptesrendus de l'Académie des sciences.Série 3, Sciences de la vie 1986, 302, 67–70. [Google Scholar]
- Austin, J.W.; Bagnères, A.-G.; Szalanski, A.L.; Scheffrahn, R.H.; Heintschel, B.P.; Messenger, M.T.; Clement, J.-L.; Gold, R.E. Reticulitermesmalletei (Isoptera: Rhinotermitidae): A valid Nearctic subterranean termite from Eastern North America. Zootaxa 2007, 1554, 1–26. [Google Scholar]
- Weesner, F.M. The Termites of the United States: A Handbook; The National Pest Control Association: Elizabeth, NJ, USA, 1965; p. 70. [Google Scholar]
- Nutting, W.L. Insecta: Isoptera. In Soil Biology Guide; Dindal, D.L., Ed.; John Wiley & Sons: New York, NY, USA, 1990; Volume 33, pp. 997–1032. [Google Scholar]
- Scheffrahn, R.H.; Su, N.-Y. Keys to soldier and winged adult termites (Isoptera) of Florida. Fla. Entomol. 1994, 77, 460–474. [Google Scholar]
- Hostettler, N.C.; Hall, D.W.; Scheffrahn, R.H. Intracolonymorphometric variation and labral shape in Florida Reticulitermes (Isoptera: Rhinotermitidae) soldiers: Significance for identification. Fla. Entomol. 1995, 78, 119–129. [Google Scholar]
- Haverty, M.I.; Forschler, B.T.; Nelson, L.J. An assessment of the taxonomy of Reticulitermes (Isoptera: Rhinotermitidae) from the Southeastern United States based on cuticular hydrocarbons. Sociobiology 1996, 28, 287–318. [Google Scholar]
- Haverty, M.I.; Nelson, L.J.; Forschler, B.T. New cuticular hydrocarbon phenotypes of Reticulitermes (Isoptera: Rhinotermitidae) from the United States. Sociobiology 1999, 34, 1–21. [Google Scholar]
- Jenkins, T.M.; Haverty, M.I.; Basten, C.J.; Nelson, L.J.; Page, M.; Forschler, B.T. Correlation of mitochondrial haplotypes with cuticular hydrocarbon phenotypes of sympatric Reticulitermes species from the southeastern United States. J. Chem. Ecol. 2000, 26, 1525–1542. [Google Scholar] [CrossRef]
- Nelson, L.J.; Cool, L.G.; Forschler, B.T.; Haverty, M.I. Correspondence of soldier defense secretion mixtures with cuticular hydrocarbon phenotypes for chemotaxonomy of the termite genus Reticulitermes in North America. J. Chem. Ecol. 2001, 27, 1449–1479. [Google Scholar] [CrossRef]
- Nelson, L.J.; Cool, L.G.; Solek, C.W.; Haverty, M.I. Cuticular hydrocarbons and soldier defense secretions of Reticulitermes in Southern California: A critical analysis of taxonomy of the Genus in North America. J. Chem. Ecol. 2008, 34, 1452–1475. [Google Scholar] [CrossRef]
- Sillam-Dussès, D.; Forschler, B.T. A dominant and undescribed species of Reticulitermes in Sapelo Island (Georgia, USA). Sociobiology 2010, 46, 137–147. [Google Scholar]
- Kollar, V. Naturgeschichte der Schadlichen Insekten. Verh. Landwirthschaft Gesellshaft in Wien 1837, 5, 411. [Google Scholar]
- Banks, N. A New Species of Termes. In Entomological News and Proceedings of the Entomological Section Academy of Natural Sciences of Philadephia; Academy of Natural Sciences of Philadelphia: Philadelphia, PA, USA, 1907; 18, pp. 392–393. [Google Scholar]
- Scheffrahn, R.H.; Su, N.-Y.; Chase, J.A.; Forschler, B.T. New termite (Isoptera: Kalotermitidae, rhinotermitidae) records from Georgia. J. Entomol. Sci. 2001, 36, 109–113. [Google Scholar]
- Feytaud, J. Le termite de saintonge. C.R. Acad. Sc. Paris 1924, 178, 241–244. [Google Scholar]
- Austin, J.W.; Szalanski, A.L.; Scheffrahn, R.H.; Messenger, M.T.; Dronnet, S.; Bagneres, A.-G. Genetic evidence for the synonymy of two reticulitermes species: Reticulitermes flavipes and Reticulitermes santonensis. Ann. Entomol. Soc. Am. 2005, 98, 395–401. [Google Scholar] [CrossRef]
- Jenkins, T.M.; Dean, R.E.; Verkerk, R.; Forschler, B.T. Phylogenetic analyses of two mitochondrial genes and one nuclear intron region illuminate European subterranean termite (Isoptera: Rhinotermitidae) gene flow, taxonomy, and introduction dynamics. Mol. Phylogenet. Evol. 2001, 20, 286–293. [Google Scholar] [CrossRef]
- Lim, S.Y. Taxonomic Review and Biogeography of Reticulitermes (Rhinotermitidae) Termites of Georgia; University of Georgia: Athens, GA, USA, 2011. [Google Scholar]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular Evolutionary Genetics Analysis using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Mol. Biol. Evol. 2011. [Google Scholar] [CrossRef]
- Dereeper, A.; Guignon, V.; Blanc, G.; Audic, S.; Buffet, S.; Chevenet, F.; Dufayar, J.-F.; Guindon, S.; Lefort, V.; Lescot, M.; et al. Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res. 2008, 1, W465–W469. [Google Scholar]
- Molecular evolution, phylogenetics and epidemiology. FigTree. Available online: http://tree.bio.ed.ac.uk/software/figtree/ (accessed on 2 December 2011).
- Krishna, K.; Weesner, F.M. Biology of Termites; Academic Press Inc.: New York, NY, USA, 1970. [Google Scholar]
- Wang, C.; Zhou, X.; Li, S.; Schwinghammer, M.; Scharf, M.; Buczkowski, G.; Bennett, G. Survey and Identification of Termites (Isoptera: Rhinotermitidae) in Indiana. Ann. Entomol. Soc. Am. 2009, 102, 1029–1036. [Google Scholar] [CrossRef]
- Austin, J.W.; Szalanski, A.L.; Cabrera, B.J. Phylogenetic analysis of the subterranean termite family Rhinotermitidae (Isoptera) by using the mitochondrial cytochromeoxidase II gene. Ann. Entomol. Soc. Am. 2004, 97, 548–555. [Google Scholar] [CrossRef]
- Austin, J.W.; Szalanski, A.L.; Scheffrahn, R.H.; Messenger, M.T. Genetic variation of Reticulitermes flavipes (Isoptera: Rhinotermitidae) in north america applying the mitochondrial rRNA 16S gene. Ann. Entomol. Soc. Am. 2005, 98, 980–988. [Google Scholar] [CrossRef]
- Vargo, E.L.; Husseneder, C. Biology of subterranean termites: Insights from molecular studies of Reticulitermes and Coptotermes. Annu.Rev. Entomol. 2009, 54, 379–403. [Google Scholar] [CrossRef]
- Copren, K.A.; Nelson, L.J.; Vargo, E.L.; Haverty, M.I. Phylogenetic analyses of mtDNA sequences corroborate taxonomic designations based on cuticular hydrocarbons in subterranean termites. Mol. Phylogenet. Evol. 2005, 35, 689–700. [Google Scholar] [CrossRef]
- Cameron, S.L.; Whiting, M.F. Mitochondrial genomic comparisons of the subterranean termites from the Genus Reticulitermes (Insects: Isoptera: Rhinotermitidae). Genome 2007, 50, 188–202. [Google Scholar] [CrossRef]
- Ye, W.; Lee, C.-Y.; Scheffrahn, R.H.; Aleong, J.M.; Su, N.-Y.; Bennett, G.W.; Scharf, M.E. Phylogenetic relationships of neartic Reticulitermes species (Isoptera: Rhinotermitidae) with particular reference to Reticulitermes arenincola Goellner. Mol. Biol. Evol. 2004, 30, 815–822. [Google Scholar]
- Forschler, B.T.; Jenkins, T.M. Evaluation of subterranean termite biology using genetic, chemotaxonomic, and morphometric markers and ecological data: A testimonial for multi-disciplinary efforts. Trends Entomol. 1999, 2, 71–80. [Google Scholar]
- Lee, M.S.Y. The molecularisation of taxonomy. Invertebr.Syst. 2004, 18, 1–6. [Google Scholar] [CrossRef]
- Cognato, A.I. Standard percent DNA sequence difference for insects does not predict species boundaries. Entomol.Soc. Am. 2006, 1037–1045. [Google Scholar]
- Winston, J.E. Describing Species: Practical Taxonomic Procedure for Biologists; Columbia University Press: New York, NY, USA, 1999; p. 513. [Google Scholar]
- Cracraft, J. Speciation and Its Ontology: The Empirical Consequences of Alternative Species Concepts for Understanding Patterns and Processes of Differentiation. In Speciation and Its Consequences; Otte, D., Endleer, J.A., Eds.; Sinauer Associates: Sunderland, MA, USA, 1989; pp. 28–59. [Google Scholar]
- Baker, R.J.; Bradley, R.D. Speciation in mammals and the genetic species concept. J. Mammal. 2006, 87, 643–662. [Google Scholar] [CrossRef]
- Wilkins, S.J. Defining Species: A Sourcebook from Antiquity to Today; Peter Lang Publishing: New York, NY, USA, 2009; p. 224. [Google Scholar]
Appendix 1: Key 1
Appendix 2: Key 2


© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Lim, S.Y.; Forschler, B.T. Reticulitermes nelsonae, a New Species of Subterranean Termite (Rhinotermitidae) from the Southeastern United States. Insects 2012, 3, 62-90. https://doi.org/10.3390/insects3010062
Lim SY, Forschler BT. Reticulitermes nelsonae, a New Species of Subterranean Termite (Rhinotermitidae) from the Southeastern United States. Insects. 2012; 3(1):62-90. https://doi.org/10.3390/insects3010062
Chicago/Turabian StyleLim, Su Yee, and Brian T. Forschler. 2012. "Reticulitermes nelsonae, a New Species of Subterranean Termite (Rhinotermitidae) from the Southeastern United States" Insects 3, no. 1: 62-90. https://doi.org/10.3390/insects3010062
APA StyleLim, S. Y., & Forschler, B. T. (2012). Reticulitermes nelsonae, a New Species of Subterranean Termite (Rhinotermitidae) from the Southeastern United States. Insects, 3(1), 62-90. https://doi.org/10.3390/insects3010062





