PrimPol—Prime Time to Reprime
Abstract
:1. Introduction
2. Discovery and Evolutionary History of PrimPol
3. What Can PrimPol Do? The Domain Architecture and Catalytic Activities of PrimPol
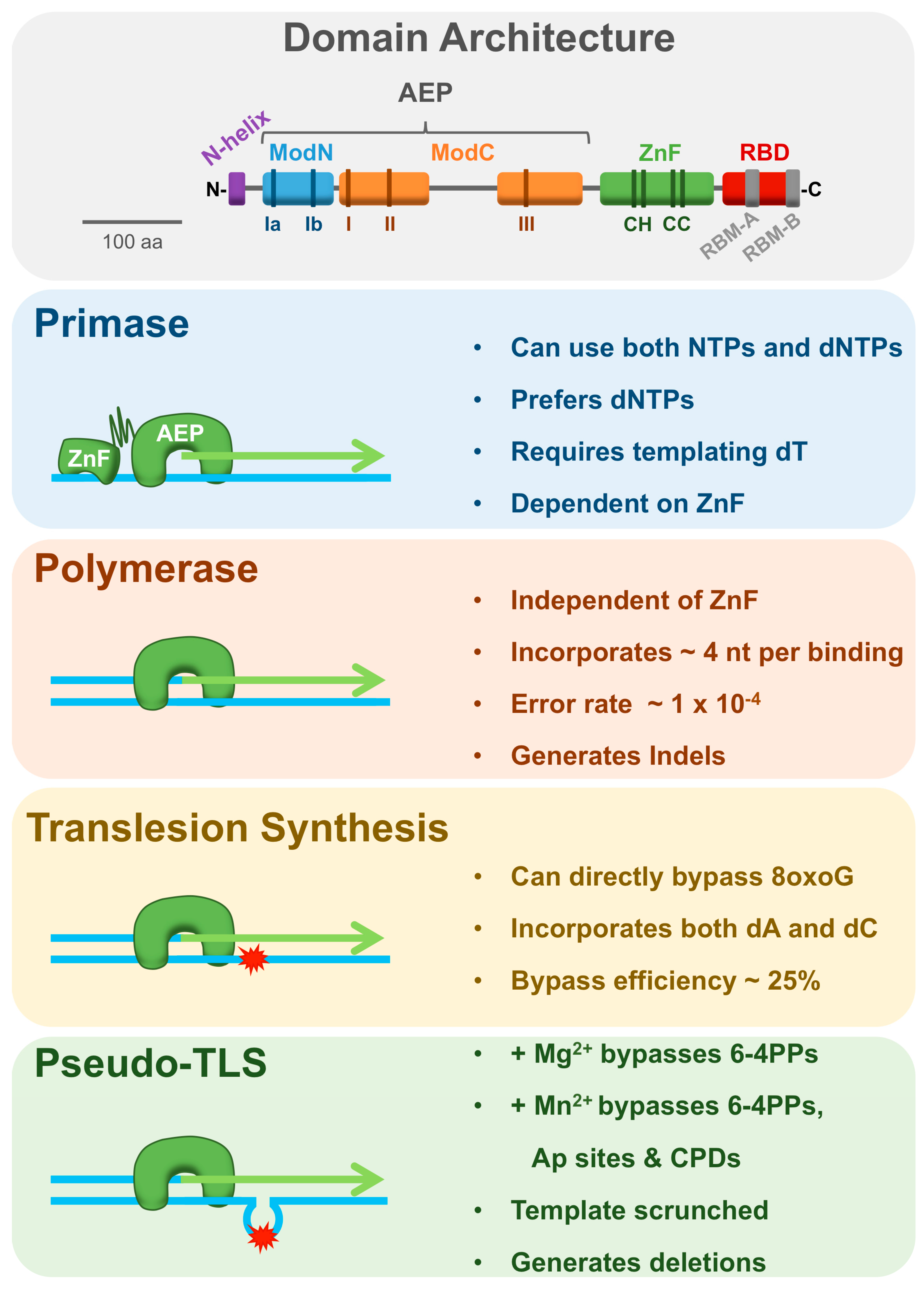
3.1. Domain Architecture and Structure
3.2. Primase Activity
3.3. Polymerase and Lesion Bypass Activities
3.4. Lesion Skipping and Template Independent Extension
3.5. Fidelity, Mutagenic Signature, and Processivity
4. What Does PrimPol Do? The Role of PrimPol in DNA Replication
4.1. PrimPol—A DNA Damage Tolerance Enzyme
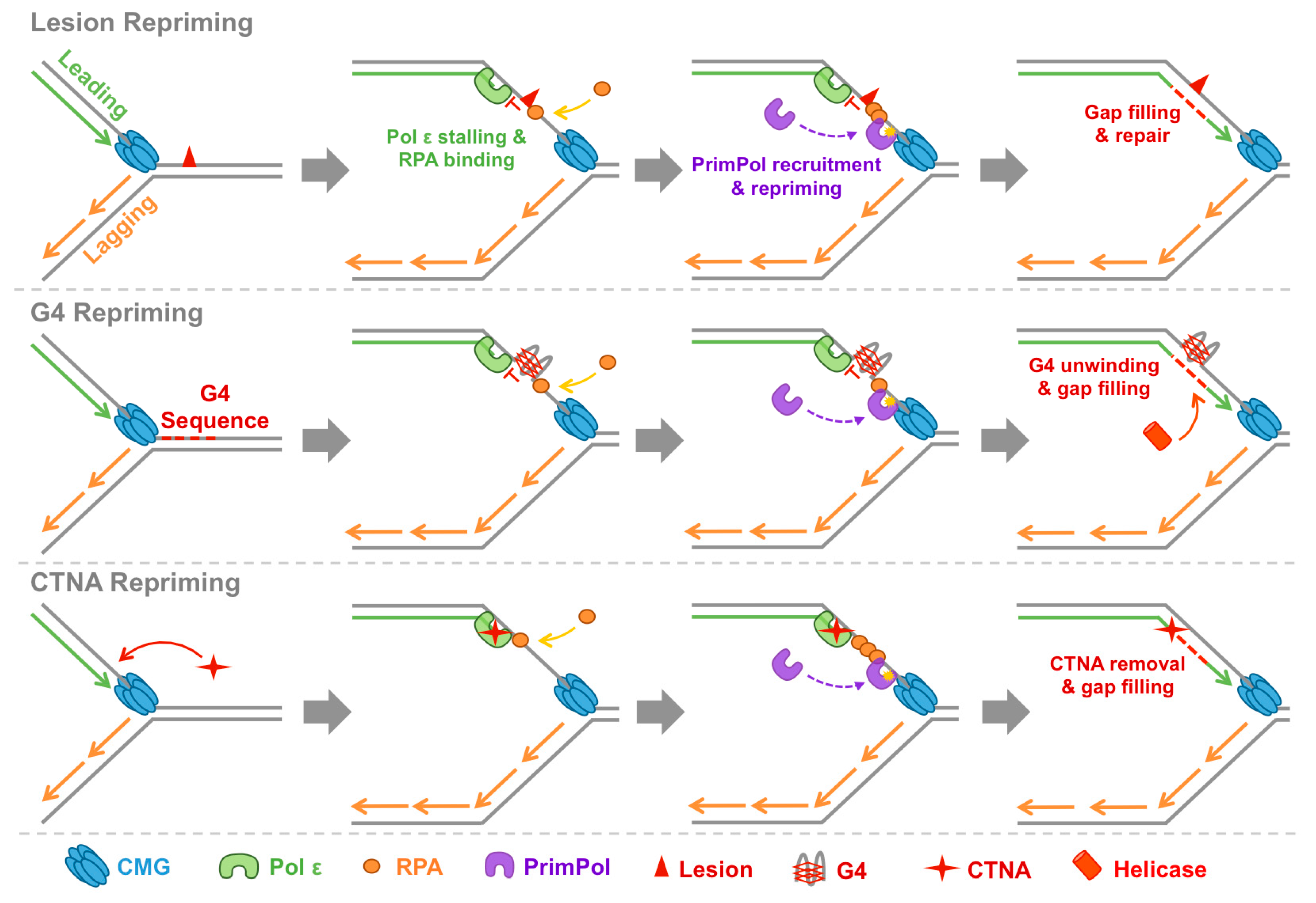
4.2. PrimPol Reprimes and Restarts Stalled Replication Forks
4.3. PrimPol Bypasses Non-Canonical Replication Impediments
4.4. A Role for PrimPol in Mitochondrial DNA Replication?
4.5. Is PrimPol Involved in Somatic Hypermutation?
4.6. Why Doesn’t the Pol α-Primase Complex Reprime Leading Strand Replication?
4.7. Why Is PrimPol Damage Tolerant In Vitro?
5. How Does PrimPol Get to Where It is Needed? The Recruitment of PrimPol to Stalled Replication Forks
5.1. PrimPol Interacts with Single-Strand Binding Proteins
5.2. RPA Recruits PrimPol to Stalled Replication Forks
6. Regulation of PrimPol during DNA Replication
6.1. Regulation of the Cellular Concentration of PrimPol
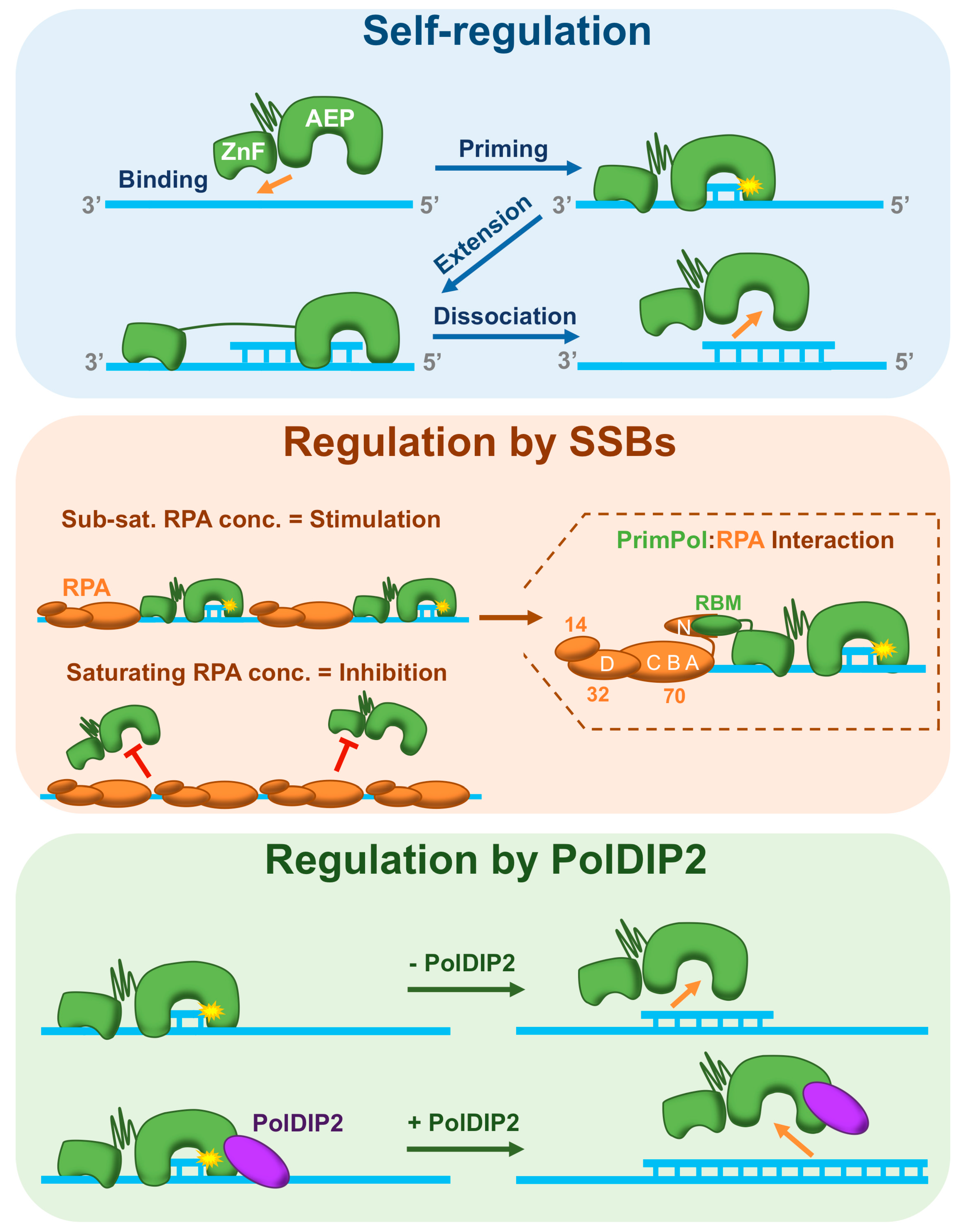
6.2. PrimPol Is Self-Regulating
6.3. Regulation by Single-Strand Binding Proteins
6.4. What Generates the ssDNA Interface Required for PrimPol Recruitment?
6.5. Regulation by PolDIP2
7. Conclusions and Perspectives
Acknowledgments
Data Statement
Conflicts of Interest
References
- Johansson, E.; Dixon, N. Replicative DNA Polymerases. Cold Spring Harb. Perspect. Biol. 2013, 5, a012799. [Google Scholar] [CrossRef] [PubMed]
- Zeman, M.K.; Cimprich, K.A. Causes and consequences of replication stress. Nat. Cell Biol. 2014, 16, 2–9. [Google Scholar] [CrossRef] [PubMed]
- Ciccia, A.; Elledge, S.J. The DNA damage response: Making it Safe to play with knives. Mol. Cell 2010, 40, 179–204. [Google Scholar] [CrossRef] [PubMed]
- Boyer, A.-S.; Grgurevic, S.; Cazaux, C.; Hoffmann, J.-S. The human specialized DNA polymerases and non-B DNA: Vital relationships to preserve genome integrity. J. Mol. Biol. 2013, 425, 4767–4781. [Google Scholar] [CrossRef] [PubMed]
- Brambati, A.; Colosio, A.; Zardoni, L.; Galanti, L.; Liberi, G. Replication and transcription on a collision course: Eukaryotic regulation mechanisms and implications for DNA stability. Front. Genet. 2015, 6, 166. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.C.; Mirkin, S.M. The balancing act of DNA repeat expansions. Curr. Opin. Genet. Dev. 2013, 23, 280–288. [Google Scholar] [CrossRef] [PubMed]
- Nick McElhinny, S.A.; Kumar, D.; Clark, A.B.; Watt, D.L.; Watts, B.E.; Lundström, E.-B.; Johansson, E.; Chabes, A.; Kunkel, T.A. Genome instability due to ribonucleotide incorporation into DNA. Nat. Chem. Biol. 2010, 6, 774–781. [Google Scholar] [CrossRef] [PubMed]
- Bester, A.C.; Roniger, M.; Oren, Y.S.; Im, M.M.; Sarni, D.; Chaoat, M.; Bensimon, A.; Zamir, G.; Shewach, D.S.; Kerem, B. Nucleotide deficiency promotes genomic instability in early stages of cancer development. Cell 2011, 145, 435–446. [Google Scholar] [CrossRef] [PubMed]
- Yeeles, J.T.P.; Poli, J.; Marians, K.J.; Pasero, P. Rescuing Stalled or Damaged Replication Forks. Cold Spring Harb. Perspect. Biol. 2013, 5, a012815. [Google Scholar] [CrossRef] [PubMed]
- Svoboda, D.L.; Vos, J.M. Differential replication of a single, UV-induced lesion in the leading or lagging strand by a human cell extract: Fork uncoupling or gap formation. Proc. Natl. Acad. Sci. USA 1995, 92, 11975–11979. [Google Scholar] [CrossRef] [PubMed]
- Lopes, M.; Foiani, M.; Sogo, J.M. Multiple mechanisms control chromosome integrity after replication fork uncoupling and restart at irreparable UV lesions. Mol. Cell 2006, 21, 15–27. [Google Scholar] [CrossRef] [PubMed]
- Elvers, I.; Johansson, F.; Groth, P.; Erixon, K.; Helleday, T. UV stalled replication forks restart by re-priming in human fibroblasts. Nucleic Acids Res. 2011, 39, 7049–7057. [Google Scholar] [CrossRef] [PubMed]
- Yeeles, J.T.P.; Marians, K.J. The Escherichia coli replisome is inherently DNA damage tolerant. Science 2011, 334, 235–238. [Google Scholar] [CrossRef] [PubMed]
- Yeeles, J.T.P.; Marians, K.J. Dynamics of leading-strand lesion skipping by the replisome. Mol. Cell 2013, 52, 855–865. [Google Scholar] [CrossRef] [PubMed]
- Goodman, M.F.; Woodgate, R. Translesion DNA Polymerases. Cold Spring Harb. Perspect. Biol. 2013, 5, a010363. [Google Scholar] [CrossRef] [PubMed]
- Sale, J.E.; Lehmann, A.R.; Woodgate, R. Y-family DNA polymerases and their role in tolerance of cellular DNA damage. Nat. Rev. Mol. Cell Biol. 2012, 13, 141–152. [Google Scholar] [CrossRef] [PubMed]
- Daigaku, Y.; Davies, A.A.; Ulrich, H.D. Ubiquitin-dependent DNA damage bypass is separable from genome replication. Nature 2010, 465, 951–955. [Google Scholar] [CrossRef] [PubMed]
- Karras, G.I.; Jentsch, S. The RAD6 DNA damage tolerance pathway operates uncoupled from the replication fork and is functional beyond S phase. Cell 2010, 141, 255–267. [Google Scholar] [CrossRef] [PubMed]
- Ulrich, H.D. Timing and spacing of ubiquitin-dependent DNA damage bypass. FEBS Lett. 2011, 585, 2861–2867. [Google Scholar] [CrossRef] [PubMed]
- Edmunds, C.E.; Simpson, L.J.; Sale, J.E. PCNA ubiquitination and REV1 define temporally distinct mechanisms for controlling translesion synthesis in the avian cell line DT40. Mol. Cell 2008, 30, 519–529. [Google Scholar] [CrossRef] [PubMed]
- Ge, X.Q.; Jackson, D.A.; Blow, J.J. Dormant origins licensed by excess Mcm2–7 are required for human cells to survive replicative stress. Genes Dev. 2007, 21, 3331–3341. [Google Scholar] [CrossRef] [PubMed]
- Branzei, D.; Szakal, B. DNA damage tolerance by recombination: Molecular pathways and DNA structures. DNA Repair 2016, 44, 68–75. [Google Scholar] [CrossRef] [PubMed]
- Neelsen, K.J.; Lopes, M. Replication fork reversal in eukaryotes: From dead end to dynamic response. Nat. Rev. Mol. Cell Biol. 2015, 16, 207–220. [Google Scholar] [CrossRef] [PubMed]
- Guilliam, T.A.; Keen, B.A.; Brissett, N.C.; Doherty, A.J. Primase-polymerases are a functionally diverse superfamily of replication and repair enzymes. Nucleic Acids Res. 2015, 43, 6651–6664. [Google Scholar] [CrossRef] [PubMed]
- Jozwiakowski, S.K.; Borazjani Gholami, F.; Doherty, A.J. Archaeal replicative primases can perform translesion DNA synthesis. Proc. Natl. Acad. Sci. USA 2015, 112, E633–E638. [Google Scholar] [CrossRef] [PubMed]
- Bianchi, J.; Rudd, S.G.; Jozwiakowski, S.K.; Bailey, L.J.; Soura, V.; Taylor, E.; Stevanovic, I.; Green, A.J.; Stracker, T.H.; Lindsay, H.D.; et al. PrimPol bypasses UV photoproducts during eukaryotic chromosomal DNA replication. Mol. Cell 2013, 52, 566–573. [Google Scholar] [CrossRef] [PubMed]
- García-Gómez, S.; Reyes, A.; Martínez-Jiménez, M.I.; Chocrón, E.S.; Mourón, S.; Terrados, G.; Powell, C.; Salido, E.; Méndez, J.; Holt, I.J.; et al. PrimPol, an archaic primase/polymerase operating in human cells. Mol. Cell 2013, 52, 541–553. [Google Scholar] [CrossRef] [PubMed]
- Wan, L.; Lou, J.; Xia, Y.; Su, B.; Liu, T.; Cui, J.; Sun, Y.; Lou, H.; Huang, J. hPrimpol1/CCDC111 is a human DNA primase-polymerase required for the maintenance of genome integrity. EMBO Rep. 2013, 14, 1104–1112. [Google Scholar] [CrossRef] [PubMed]
- Aravind, L.; Leipe, D.D.; Koonin, E.V. Toprim—A conserved catalytic domain in type IA and II topoisomerases, DnaG-type primases, OLD family nucleases and RecR proteins. Nucleic Acids Res. 1998, 26, 4205–4213. [Google Scholar] [CrossRef] [PubMed]
- Koonin, E.V.; Wolf, Y.I.; Kondrashov, A.S.; Aravind, L. Bacterial homologs of the small subunit of eukaryotic DNA primase. J. Mol. Microbiol. Biotechnol. 2000, 2, 509–512. [Google Scholar] [PubMed]
- Della, M.; Palmbos, P.L.; Tseng, H.-M.; Tonkin, L.M.; Daley, J.M.; Topper, L.M.; Pitcher, R.S.; Tomkinson, A.E.; Wilson, T.E.; Doherty, A.J. Mycobacterial Ku and ligase proteins constitute a two-component NHEJ repair machine. Science 2004, 306, 683–685. [Google Scholar] [CrossRef] [PubMed]
- Iyer, L.M.; Koonin, E.V.; Leipe, D.D.; Aravind, L. Origin and evolution of the archaeo-eukaryotic primase superfamily and related palm-domain proteins: Structural insights and new members. Nucleic Acids Res. 2005, 33, 3875–3896. [Google Scholar] [CrossRef] [PubMed]
- Rudd, S.G.; Glover, L.; Jozwiakowski, S.K.; Horn, D.; Doherty, A.J. PPL2 translesion polymerase is essential for the completion of chromosomal DNA replication in the African trypanosome. Mol. Cell 2013, 52, 554–565. [Google Scholar] [CrossRef] [PubMed]
- Keen, B.A.; Jozwiakowski, S.K.; Bailey, L.J.; Bianchi, J.; Doherty, A.J. Molecular dissection of the domain architecture and catalytic activities of human PrimPol. Nucleic Acids Res. 2014, 42, 5830–5845. [Google Scholar] [CrossRef] [PubMed]
- Rechkoblit, O.; Gupta, Y.K.; Malik, R.; Rajashankar, K.R.; Johnson, R.E.; Prakash, L.; Prakash, S.; Aggarwal, A.K. Structure and mechanism of human PrimPol, a DNA polymerase with primase activity. Sci. Adv. 2016, 2, e1601317. [Google Scholar] [CrossRef] [PubMed]
- Mourón, S.; Rodriguez-Acebes, S.; Martínez-Jiménez, M.I.; García-Gómez, S.; Chocrón, S.; Blanco, L.; Méndez, J. Repriming of DNA synthesis at stalled replication forks by human PrimPol. Nat. Struct. Mol. Biol. 2013, 20, 1383–1389. [Google Scholar] [CrossRef] [PubMed]
- Frick, D.N.; Richardson, C.C. DNA Primases. Annu. Rev. Biochem. 2001, 70, 39–80. [Google Scholar] [CrossRef] [PubMed]
- Biswas, N.; Weller, S.K. A Mutation in the C-terminal putative Zn2+ finger motif of UL52 severely affects the biochemical activities of the HSV-1 helicase-primase subcomplex. J. Biol. Chem. 1999, 274, 8068–8076. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Carrington-Lawrence, S.D.; Bai, P.; Weller, S.K. Mutations in the putative Zinc-binding motif of UL52 demonstrate a complex interdependence between the UL5 and UL52 subunits of the human herpes simplex virus type 1 helicase/primase complex. J. Virol. 2005, 79, 9088–9096. [Google Scholar] [CrossRef] [PubMed]
- Zafar, M.K.; Ketkar, A.; Lodeiro, M.F.; Cameron, C.E.; Eoff, R.L. Kinetic analysis of human PrimPol DNA polymerase activity reveals a generally error-prone enzyme capable of accurately bypassing 7,8-dihydro-8-oxo-2’-deoxyguanosine. Biochemistry (Mosc.) 2014, 53, 6584–6594. [Google Scholar] [CrossRef] [PubMed]
- Guilliam, T.A.; Bailey, L.J.; Brissett, N.C.; Doherty, A.J. PolDIP2 interacts with human PrimPol and enhances its DNA polymerase activities. Nucleic Acids Res. 2016, 44, 3317–3329. [Google Scholar] [CrossRef] [PubMed]
- Stojkovič, G.; Makarova, A.V.; Wanrooij, P.H.; Forslund, J.; Burgers, P.M.; Wanrooij, S. Oxidative DNA damage stalls the human mitochondrial replisome. Sci. Rep. 2016, 6, 28942. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Jiménez, M.I.; García-Gómez, S.; Bebenek, K.; Sastre-Moreno, G.; Calvo, P.A.; Díaz-Talavera, A.; Kunkel, T.A.; Blanco, L. Alternative solutions and new scenarios for translesion DNA synthesis by human PrimPol. DNA Repair 2015, 29, 127–138. [Google Scholar] [CrossRef] [PubMed]
- Pitcher, R.S.; Brissett, N.C.; Picher, A.J.; Andrade, P.; Juarez, R.; Thompson, D.; Fox, G.C.; Blanco, L.; Doherty, A.J. Structure and function of a mycobacterial NHEJ DNA repair polymerase. J. Mol. Biol. 2007, 366, 391–405. [Google Scholar] [CrossRef] [PubMed]
- Yakovleva, L.; Shuman, S. Nucleotide misincorporation, 3′-mismatch extension, and responses to abasic sites and DNA adducts by the polymerase component of cacterial DNA ligase D. J. Biol. Chem. 2006, 281, 25026–25040. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.S.-F.; Eichler, D.C.; Korn, D. Effect of manganese(2+) ions on the in vitro activity of human deoxyribonucleic acid polymerase .beta. Biochemistry (Mosc.) 1977, 16, 4927–4934. [Google Scholar] [CrossRef]
- Pelletier, H.; Sawaya, M.R.; Wolfle, W.; Wilson, S.H.; Kraut, J. A Structural basis for metal ion mutagenicity and nucleotide selectivity in human DNA polymerase β. Biochemistry (Mosc.) 1996, 35, 12762–12777. [Google Scholar] [CrossRef] [PubMed]
- Goodman, M.F.; Keener, S.; Guidotti, S.; Branscomb, E.W. On the enzymatic basis for mutagenesis by manganese. J. Biol. Chem. 1983, 258, 3469–3475. [Google Scholar] [PubMed]
- El-Deiry, W.S.; Downey, K.M.; So, A.G. Molecular mechanisms of manganese mutagenesis. Proc. Natl. Acad. Sci. USA 1984, 81, 7378–7382. [Google Scholar] [CrossRef] [PubMed]
- Vaisman, A.; Ling, H.; Woodgate, R.; Yang, W. Fidelity of Dpo4: Effect of metal ions, nucleotide selection and pyrophosphorolysis. EMBO J. 2005, 24, 2957–2967. [Google Scholar] [CrossRef] [PubMed]
- Chabes, A.; Georgieva, B.; Domkin, V.; Zhao, X.; Rothstein, R.; Thelander, L. Survival of DNA damage in yeast directly depends on increased dNTP levels allowed by relaxed feedback inhibition of ribonucleotide reductase. Cell 2003, 112, 391–401. [Google Scholar] [CrossRef]
- Shimizu, K.; Hashimoto, K.; Kirchner, J.M.; Nakai, W.; Nishikawa, H.; Resnick, M.A.; Sugino, A. Fidelity of DNA polymerase ε holoenzyme from budding yeast Saccharomyces cerevisiae. J. Biol. Chem. 2002, 277, 37422–37429. [Google Scholar] [CrossRef] [PubMed]
- Minko, I.G.; Washington, M.T.; Kanuri, M.; Prakash, L.; Prakash, S.; Lloyd, R.S. Translesion synthesis past acrolein-derived DNA adduct, γ-hydroxypropanodeoxyguanosine, by yeast and human DNA polymerase η. J. Biol. Chem. 2003, 278, 784–790. [Google Scholar] [CrossRef] [PubMed]
- Sabouri, N.; Viberg, J.; Goyal, D.K.; Johansson, E.; Chabes, A. Evidence for lesion bypass by yeast replicative DNA polymerases during DNA damage. Nucleic Acids Res. 2008, 36, 5660–5667. [Google Scholar] [CrossRef] [PubMed]
- Guilliam, T.A.; Jozwiakowski, S.K.; Ehlinger, A.; Barnes, R.P.; Rudd, S.G.; Bailey, L.J.; Skehel, J.M.; Eckert, K.A.; Chazin, W.J.; Doherty, A.J. Human PrimPol is a highly error-prone polymerase regulated by single-stranded DNA binding proteins. Nucleic Acids Res. 2015, 43, 1056–1068. [Google Scholar] [CrossRef] [PubMed]
- Ash, D.E.; Schramm, V.L. Determination of free and bound manganese(II) in hepatocytes from fed and fasted rats. J. Biol. Chem. 1982, 257, 9261–9264. [Google Scholar] [PubMed]
- Markesbery, W.R.; Ehmann, W.D.; Alauddin, M.; Hossain, T.I.M. Brain trace element concentrations in aging. Neurobiol. Aging 1984, 5, 19–28. [Google Scholar] [CrossRef]
- Versieck, J.; McCall, J.T. Trace elements in human body fluids and tissues. CRC Crit. Rev. Clin. Lab. Sci. 1985, 22, 97–184. [Google Scholar] [CrossRef] [PubMed]
- Goldschmidt, V.; Didierjean, J.; Ehresmann, B.; Ehresmann, C.; Isel, C.; Marquet, R. Mg2+ dependency of HIV-1 reverse transcription, inhibition by nucleoside analogues and resistance. Nucleic Acids Res. 2006, 34, 42–52. [Google Scholar] [CrossRef] [PubMed]
- Gee, J.B., II; Corbett, R.J.T.; Perlman, J.M.; Laptook, A.R. Hypermagnesemia does not increase brain intracellular magnesium in newborn swine. Pediatr. Neurol. 2001, 25, 304–308. [Google Scholar] [CrossRef]
- Rampazzo, C.; Ferraro, P.; Pontarin, G.; Fabris, S.; Reichard, P.; Bianchi, V. Mitochondrial deoxyribonucleotides, pool sizes, synthesis, and regulation. J. Biol. Chem. 2004, 279, 17019–17026. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Grosse, F. Accuracy of DNA primase. J. Mol. Biol. 1990, 216, 475–479. [Google Scholar] [CrossRef]
- Sheaff, R.J.; Kuchta, R.D. Misincorporation of nucleotides by calf thymus DNA primase and elongation of primers containing multiple noncognate nucleotides by DNA polymerase alpha. J. Biol. Chem. 1994, 269, 19225–19231. [Google Scholar] [PubMed]
- Cotterill, S.; Chui, G.; Lehman, I.R. DNA polymerase-primase from embryos of Drosophila melanogaster. DNA primase subunits. J. Biol. Chem. 1987, 262, 16105–16108. [Google Scholar] [PubMed]
- Bailey, L.J.; Bianchi, J.; Hégarat, N.; Hochegger, H.; Doherty, A.J. PrimPol-deficient cells exhibit a pronounced G2 checkpoint response following UV damage. Cell Cycle 2016, 15, 908–918. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, K.; Guilliam, T.A.; Tsuda, M.; Yamamoto, J.; Bailey, L.J.; Iwai, S.; Takeda, S.; Doherty, A.J.; Hirota, K. Repriming by PrimPol is critical for DNA replication restart downstream of lesions and chain-terminating nucleosides. Cell Cycle 2016, 15, 1997–2008. [Google Scholar] [CrossRef] [PubMed]
- Schiavone, D.; Jozwiakowski, S.K.; Romanello, M.; Guilbaud, G.; Guilliam, T.A.; Bailey, L.J.; Sale, J.E.; Doherty, A.J. PrimPol is required for replicative tolerance of G quadruplexes in vertebrate cells. Mol. Cell 2016, 61, 161–169. [Google Scholar] [CrossRef] [PubMed]
- Pilzecker, B.; Buoninfante, O.A.; Pritchard, C.; Blomberg, O.S.; Huijbers, I.J.; van den Berk, P.C.M.; Jacobs, H. PrimPol prevents APOBEC/AID family mediated DNA mutagenesis. Nucleic Acids Res. 2016, 44, 4734–4744. [Google Scholar] [CrossRef] [PubMed]
- Haracska, L.; Yu, S.-L.; Johnson, R.E.; Prakash, L.; Prakash, S. Efficient and accurate replication in the presence of 7,8-dihydro-8-oxoguanine by DNA polymerase η. Nat. Genet. 2000, 25, 458–461. [Google Scholar] [PubMed]
- Maga, G.; Crespan, E.; Markkanen, E.; Imhof, R.; Furrer, A.; Villani, G.; Hübscher, U.; Loon, B. van DNA polymerase δ-interacting protein 2 is a processivity factor for DNA polymerase λ during 8-oxo-7,8-dihydroguanine bypass. Proc. Natl. Acad. Sci. USA 2013, 110, 18850–18855. [Google Scholar] [CrossRef] [PubMed]
- Zahn, K.E.; Wallace, S.S.; Doublié, S. DNA polymerases provide a canon of strategies for translesion synthesis past oxidatively generated lesions. Curr. Opin. Struct. Biol. 2011, 21, 358–369. [Google Scholar] [CrossRef] [PubMed]
- Keen, B.A.; Bailey, L.J.; Jozwiakowski, S.K.; Doherty, A.J. Human PrimPol mutation associated with high myopia has a DNA replication defect. Nucleic Acids Res. 2014, 42, 12102–12111. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.-H.; Lindsey-Boltz, L.A.; Kemp, M.; Mason, A.C.; Wold, M.S.; Sancar, A. Reconstitution of RPA-covered single-stranded DNA-activated ATR-Chk1 signaling. Proc. Natl. Acad. Sci. USA 2010, 107, 13660–13665. [Google Scholar] [CrossRef] [PubMed]
- Watson, J.D.; Crick, F.H. Molecular structure of nucleic acids; a structure for deoxyribose nucleic acid. Nature 1953, 171, 737–738. [Google Scholar] [CrossRef] [PubMed]
- Bochman, M.L.; Paeschke, K.; Zakian, V.A. DNA secondary structures: Stability and function of G-quadruplex structures. Nat. Rev. Genet. 2012, 13, 770–780. [Google Scholar] [CrossRef] [PubMed]
- Murat, P.; Balasubramanian, S. Existence and consequences of G-quadruplex structures in DNA. Curr. Opin. Genet. Dev. 2014, 25, 22–29. [Google Scholar] [CrossRef] [PubMed]
- Maizels, N.; Gray, L.T. The G4 Genome. PLoS Genet. 2013, 9, e1003468. [Google Scholar] [CrossRef] [PubMed]
- Cheung, I.; Schertzer, M.; Rose, A.; Lansdorp, P.M. Disruption of dog-1 in Caenorhabditis elegans triggers deletions upstream of guanine-rich DNA. Nat. Genet. 2002, 31, 405–409. [Google Scholar] [CrossRef] [PubMed]
- Ribeyre, C.; Lopes, J.; Boulé, J.-B.; Piazza, A.; Guédin, A.; Zakian, V.A.; Mergny, J.-L.; Nicolas, A. The yeast Pif1 helicase prevents genomic instability caused by G-quadruplex-forming CEB1 sequences in vivo. PLoS Genet. 2009, 5, e1000475. [Google Scholar] [CrossRef] [PubMed]
- Sarkies, P.; Reams, C.; Simpson, L.J.; Sale, J.E. Epigenetic instability due to defective replication of structured DNA. Mol. Cell 2010, 40, 703–713. [Google Scholar] [CrossRef] [PubMed]
- León-Ortiz, A.M.; Svendsen, J.; Boulton, S.J. Metabolism of DNA secondary structures at the eukaryotic replication fork. DNA Repair 2014, 19, 152–162. [Google Scholar] [CrossRef] [PubMed]
- Sarkies, P.; Murat, P.; Phillips, L.G.; Patel, K.J.; Balasubramanian, S.; Sale, J.E. FANCJ coordinates two pathways that maintain epigenetic stability at G-quadruplex DNA. Nucleic Acids Res. 2012, 40, 1485–1498. [Google Scholar] [CrossRef] [PubMed]
- Schiavone, D.; Guilbaud, G.; Murat, P.; Papadopoulou, C.; Sarkies, P.; Prioleau, M.-N.; Balasubramanian, S.; Sale, J.E. Determinants of G quadruplex-induced epigenetic instability in REV1-deficient cells. EMBO J. 2014, 33, 2507–2520. [Google Scholar] [CrossRef] [PubMed]
- Berdis, A.J. DNA Polymerases as therapeutic targets. Biochemistry (Mosc.) 2008, 47, 8253–8260. [Google Scholar] [CrossRef] [PubMed]
- De Clercq, E.; Field, H.J. Antiviral prodrugs—The development of successful prodrug strategies for antiviral chemotherapy. Br. J. Pharmacol. 2006, 147, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Alexeyev, M.; Shokolenko, I.; Wilson, G.; LeDoux, S. The maintenance of mitochondrial DNA integrity—critical analysis and update. Cold Spring Harb. Perspect. Biol. 2013, 5, a012641. [Google Scholar] [CrossRef] [PubMed]
- Bianchi, J. Investigating the Role of a Novel Primase-Polymerase, PrimPol, in DNA Damage Tolerance in Vertebrate Cells. Ph.D. Thesis, University of Sussex, Brighton, UK, 2013. [Google Scholar]
- Falkenberg, M.; Larsson, N.-G.; Gustafsson, C.M. DNA replication and transcription in mammalian mitochondria. Annu. Rev. Biochem. 2007, 76, 679–699. [Google Scholar] [CrossRef] [PubMed]
- Loeb, L.A.; Monnat, R.J. DNA polymerases and human disease. Nat. Rev. Genet. 2008, 9, 594–604. [Google Scholar] [CrossRef] [PubMed]
- Wisnovsky, S.; Jean, S.R.; Kelley, S.O. Mitochondrial DNA repair and replication proteins revealed by targeted chemical probes. Nat. Chem. Biol. 2016, 12, 567–573. [Google Scholar] [CrossRef] [PubMed]
- Singh, B.; Li, X.; Owens, K.M.; Vanniarajan, A.; Liang, P.; Singh, K.K. Human REV3 DNA Polymerase Zeta Localizes to Mitochondria and Protects the Mitochondrial Genome. PLoS ONE 2015, 10, e0140409. [Google Scholar] [CrossRef] [PubMed]
- Bharti, S.K.; Sommers, J.A.; Zhou, J.; Kaplan, D.L.; Spelbrink, J.N.; Mergny, J.-L.; Brosh, R.M. DNA Sequences Proximal to Human Mitochondrial DNA Deletion Breakpoints Prevalent in Human Disease Form G-quadruplexes, a Class of DNA Structures Inefficiently Unwound by the Mitochondrial Replicative Twinkle Helicase. J. Biol. Chem. 2014, 289, 29975–29993. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bailey, L.J.; Doherty, A.J. Mitochondrial DNA replication—A PrimPol perspective. Biochem. Soc. Trans. (Under Revison).
- Noia, J.M.D.; Neuberger, M.S. Molecular mechanisms of antibody somatic hypermutation. Annu. Rev. Biochem. 2007, 76, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Sale, J.E.; Batters, C.; Edmunds, C.E.; Phillips, L.G.; Simpson, L.J.; Szüts, D. Timing matters: Error-prone gap filling and translesion synthesis in immunoglobulin gene hypermutation. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2009, 364, 595–603. [Google Scholar] [CrossRef] [PubMed]
- Williams, J.S.; Lujan, S.A.; Kunkel, T.A. Processing ribonucleotides incorporated during eukaryotic DNA replication. Nat. Rev. Mol. Cell Biol. 2016, 17, 350–363. [Google Scholar] [CrossRef] [PubMed]
- Williams, J.S.; Kunkel, T.A. Ribonucleotides in DNA: Origins, repair and consequences. DNA Repair 2014, 19, 27–37. [Google Scholar] [CrossRef] [PubMed]
- Williams, J.S.; Clausen, A.R.; Lujan, S.A.; Marjavaara, L.; Clark, A.B.; Burgers, P.M.; Chabes, A.; Kunkel, T.A. Evidence that processing of ribonucleotides in DNA by topoisomerase 1 is leading-strand specific. Nat. Struct. Mol. Biol. 2015, 22, 291–297. [Google Scholar] [CrossRef] [PubMed]
- Reijns, M.A.M.; Rabe, B.; Rigby, R.E.; Mill, P.; Astell, K.R.; Lettice, L.A.; Boyle, S.; Leitch, A.; Keighren, M.; Kilanowski, F.; et al. Enzymatic removal of ribonucleotides from DNA is essential for mammalian genome integrity and development. Cell 2012, 149, 1008–1022. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Baranovskiy, A.G.; Babayeva, N.D.; Zhang, Y.; Gu, J.; Suwa, Y.; Pavlov, Y.I.; Tahirov, T.H. Mechanism of concerted RNA-DNA primer synthesis by the human primosome. J. Biol. Chem. 2016, 291, 10006–10020. [Google Scholar] [PubMed]
- Baranovskiy, A.G.; Zhang, Y.; Suwa, Y.; Gu, J.; Babayeva, N.D.; Pavlov, Y.I.; Tahirov, T.H. Insight into the Human DNA Primase Interaction with Template-Primer. J. Biol. Chem. 2016, 291, 4793–4802. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Huang, M. Essential role of the iron-sulfur cluster binding domain of the primase regulatory subunit Pri2 in DNA replication initiation. Protein Cell 2015, 6, 194–210. [Google Scholar] [CrossRef] [PubMed]
- Zou, L.; Elledge, S.J. Sensing DNA Damage through ATRIP recognition of RPA-ssDNA complexes. Science 2003, 300, 1542–1548. [Google Scholar] [CrossRef] [PubMed]
- Guilliam, T.A.; Brissett, N.C.; Ehlinger, A.; Keen, B.A.; Kolesar, P.; Taylor, E.; Bailey, L.J.; Lindsay, H.D.; Chazin, W.J.; Doherty, A.J. Molecular basis for PrimPol recruitment to replication forks by RPA. Nat. Commun. (Under Review).
- Xu, X.; Vaithiyalingam, S.; Glick, G.G.; Mordes, D.A.; Chazin, W.J.; Cortez, D. The basic cleft of RPA70N binds multiple checkpoint proteins, including RAD9, To Regulate ATR Signaling. Mol. Cell. Biol. 2008, 28, 7345–7353. [Google Scholar] [CrossRef] [PubMed]
- Vallerga, M.B.; Mansilla, S.F.; Federico, M.B.; Bertolin, A.P.; Gottifredi, V. Rad51 recombinase prevents Mre11 nuclease-dependent degradation and excessive PrimPol-mediated elongation of nascent DNA after UV irradiation. Proc. Natl. Acad. Sci. USA 2015, 112, E6624–6633. [Google Scholar] [CrossRef] [PubMed]
- Robison, J.G.; Elliott, J.; Dixon, K.; Oakley, G.G. Replication protein A and the Mre11·Rad50·Nbs1 complex co-localize and interact at sites of stalled replication forks. J. Biol. Chem. 2004, 279, 34802–34810. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Shell, S.M.; Zou, Y. Interaction and colocalization of Rad9/Rad1/Hus1 checkpoint complex with replication protein A in human cells. Oncogene 2005, 24, 4728–4735. [Google Scholar] [CrossRef] [PubMed]
- Oakley, G.G.; Patrick, S.M. Replication protein A: Directing traffic at the intersection of replication and repair. Front. Biosci. J. Virtual Libr. 2010, 15, 883–900. [Google Scholar] [CrossRef]
- Michel, B. After 30 Years of Study, the Bacterial SOS Response Still Surprises Us. PLoS Biol 2005, 3, e255. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beck, M.; Schmidt, A.; Malmstroem, J.; Claassen, M.; Ori, A.; Szymborska, A.; Herzog, F.; Rinner, O.; Ellenberg, J.; Aebersold, R. The quantitative proteome of a human cell line. Mol. Syst. Biol. 2011, 7, 549. [Google Scholar] [CrossRef] [PubMed]
- Kuchta, R.D.; Stengel, G. Mechanism and evolution of DNA primases. Biochim. Biophys. Acta BBA Proteins Proteom. 2010, 1804, 1180–1189. [Google Scholar] [CrossRef] [PubMed]
- Tsurimoto, T.; Stillman, B. Multiple replication factors augment DNA synthesis by the two eukaryotic DNA polymerases, alpha and delta. EMBO J. 1989, 8, 3883–3889. [Google Scholar] [PubMed]
- Oliveira, M.T.; Kaguni, L.S. Functional roles of the N- and C-terminal regions of the human mitochondrial single-stranded DNA-binding protein. PLoS ONE 2010, 5, e15379. [Google Scholar] [CrossRef] [PubMed]
- Collins, K.L.; Kelly, T.J. Effects of T antigen and replication protein A on the initiation of DNA synthesis by DNA polymerase alpha-primase. Mol. Cell. Biol. 1991, 11, 2108–2115. [Google Scholar] [CrossRef] [PubMed]
- Kolpashchikov, D.M.; Khodyreva, S.N.; Khlimankov, D.Y.; Wold, M.S.; Favre, A.; Lavrik, O.I. Polarity of human replication protein A binding to DNA. Nucleic Acids Res. 2001, 29, 373–379. [Google Scholar] [CrossRef] [PubMed]
- De Laat, W.L.; Appeldoorn, E.; Sugasawa, K.; Weterings, E.; Jaspers, N.G.J.; Hoeijmakers, J.H.J. DNA-binding polarity of human replication protein A positions nucleases in nucleotide excision repair. Genes Dev. 1998, 12, 2598–2609. [Google Scholar] [CrossRef] [PubMed]
- Iftode, C.; Borowiec, J.A. 5’ → 3’ Molecular Polarity of Human Replication Protein A (hRPA) Binding to Pseudo-Origin DNA Substrates. Biochemistry (Mosc.) 2000, 39, 11970–11981. [Google Scholar] [CrossRef]
- Fan, J.; Pavletich, N.P. Structure and conformational change of a replication protein A heterotrimer bound to ssDNA. Genes Dev. 2012, 26, 2337–2347. [Google Scholar] [CrossRef] [PubMed]
- Brosey, C.A.; Yan, C.; Tsutakawa, S.E.; Heller, W.T.; Rambo, R.P.; Tainer, J.A.; Ivanov, I.; Chazin, W.J. A new structural framework for integrating replication protein A into DNA processing machinery. Nucleic Acids Res. 2013, 41, 2313–2327. [Google Scholar] [CrossRef] [PubMed]
- Iftode, C.; Daniely, Y.; Borowiec, J.A. Replication Protein A (RPA): The Eukaryotic SSB. Crit. Rev. Biochem. Mol. Biol. 1999, 34, 141–180. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Rodriguez-Belmonte, E.M.; Mazloum, N.; Xie, B.; Lee, M.Y. Identification of a novel protein, PDIP38, that interacts with the p50 subunit of DNA polymerase δ and proliferating cell nuclear antigen. J. Biol. Chem. 2003, 278, 10041–10047. [Google Scholar] [CrossRef] [PubMed]
- Tissier, A.; Janel-Bintz, R.; Coulon, S.; Klaile, E.; Kannouche, P.; Fuchs, R.P.; Cordonnier, A.M. Crosstalk between replicative and translesional DNA polymerases: PDIP38 interacts directly with Polη. DNA Repair 2010, 9, 922–928. [Google Scholar] [CrossRef] [PubMed]
- Daigaku, Y.; Keszthelyi, A.; Müller, C.A.; Miyabe, I.; Brooks, T.; Retkute, R.; Hubank, M.; Nieduszynski, C.A.; Carr, A.M. A global profile of replicative polymerase usage. Nat. Struct. Mol. Biol. 2015, 22, 192–198. [Google Scholar] [CrossRef] [PubMed]
- Rupp, W.D.; Howard-Flanders, P. Discontinuities in the DNA synthesized in an excision-defective strain of Escherichia coli following ultraviolet irradiation. J. Mol. Biol. 1968, 31, 291–304. [Google Scholar] [CrossRef]
- Heller, R.C.; Marians, K.J. Replication fork reactivation downstream of a blocked nascent leading strand. Nature 2006, 439, 557–562. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Ma, L.; Yi, D.; Yoon, J.; Diercks, A.; Foltz, G.; Price, N.D.; Hood, L.E.; Tian, Q. A CD133-related gene expression signature identifies an aggressive glioblastoma subtype with excessive mutations. Proc. Natl. Acad. Sci. USA 2011, 108, 1591–1596. [Google Scholar] [CrossRef] [PubMed]

© 2017 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guilliam, T.A.; Doherty, A.J. PrimPol—Prime Time to Reprime. Genes 2017, 8, 20. https://doi.org/10.3390/genes8010020
Guilliam TA, Doherty AJ. PrimPol—Prime Time to Reprime. Genes. 2017; 8(1):20. https://doi.org/10.3390/genes8010020
Chicago/Turabian StyleGuilliam, Thomas A., and Aidan J. Doherty. 2017. "PrimPol—Prime Time to Reprime" Genes 8, no. 1: 20. https://doi.org/10.3390/genes8010020




