The Genetics of Diabetic Nephropathy
Abstract
:1. Background and Introduction
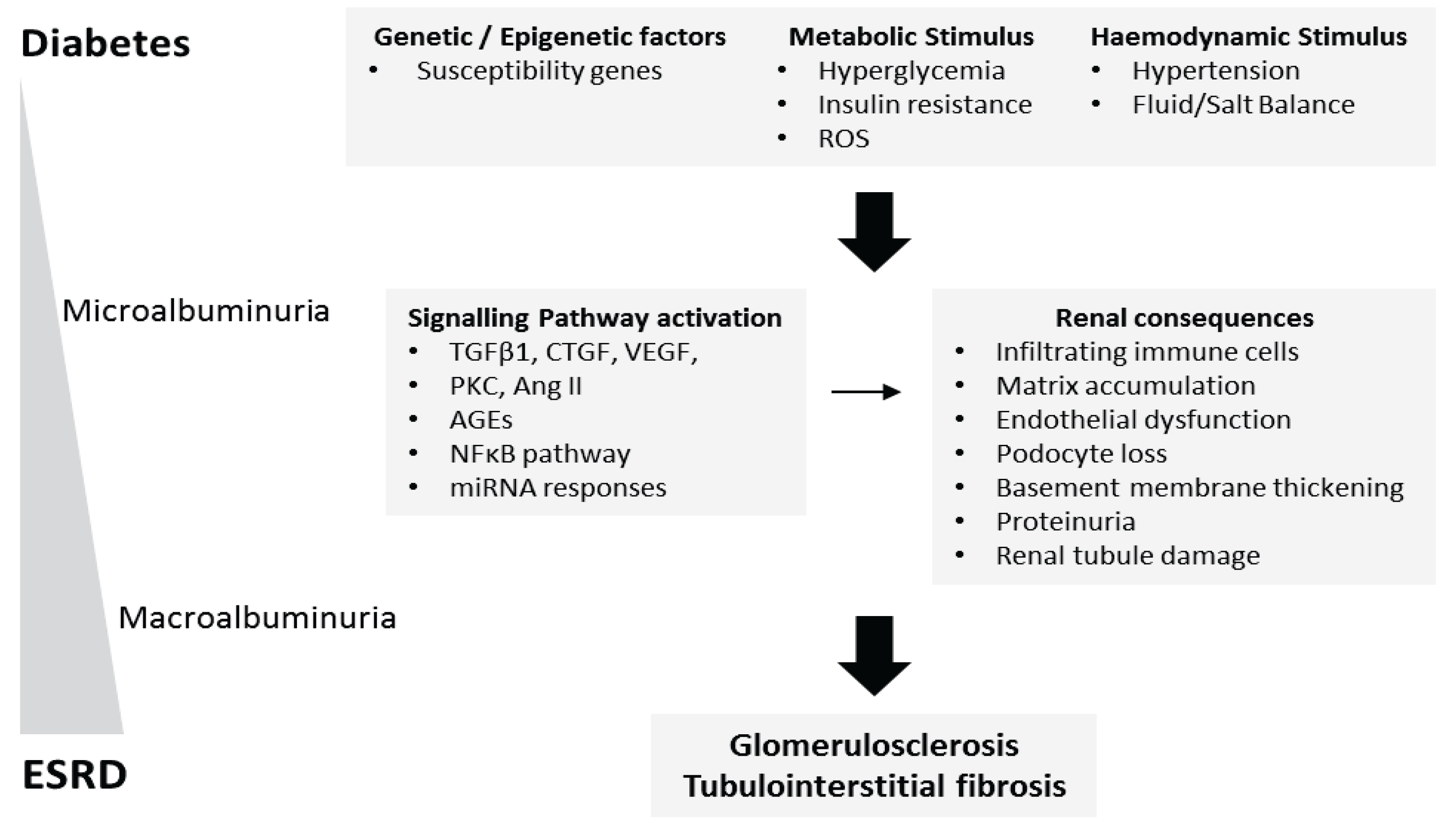
2. High Glucose in Diabetic Nephropathy
3. Profibrotic Growth Factors in Diabetic Nephropathy
4. Animal Models of Diabetic Nephropathy
5. Human Studies in Diabetic Nephropathy
5.1. Gene Expression Studies in DN
5.2. Epigenetic Studies in DN
6. Genetic Architecture of Diabetic Nephropathy
Gene Discovery in Diabetic Nephropathy
7. Integrating Multi-Layered Datasets in Diabetic Nephropathy
8. Discussion and Conclusions
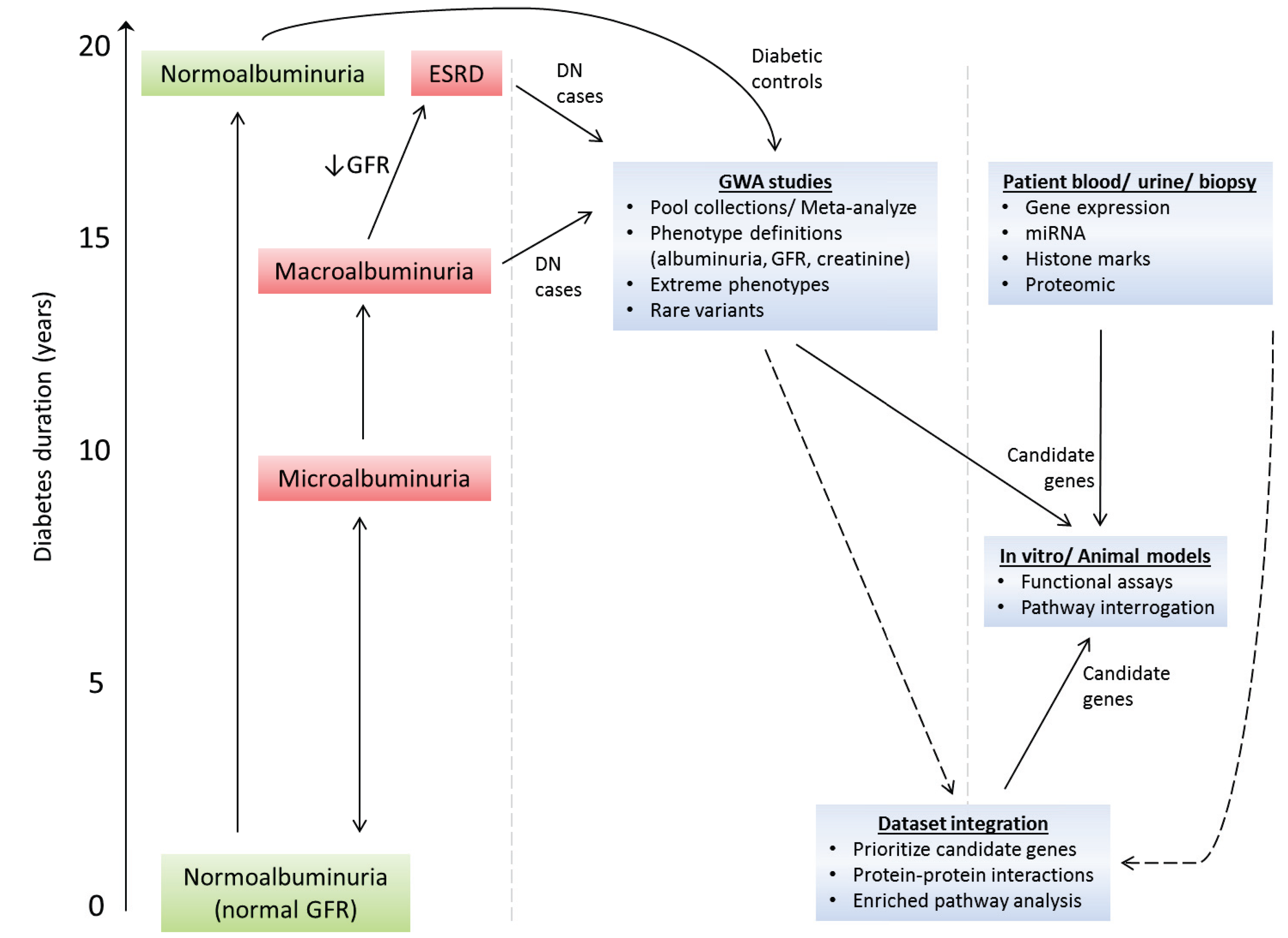
Acknowledgments
Conflicts of Interest
References
- Laing, S.P.; Swerdlow, A.J.; Slater, S.D.; Burden, A.C.; Morris, A.; Waugh, N.R.; Gatling, W.; Bingley, P.J.; Patterson, C.C. Mortality from heart disease in a cohort of 23,000 patients with insulin-treated diabetes. Diabetologia 2003, 46, 760–765. [Google Scholar] [CrossRef] [Green Version]
- Groop, P.H.; Thomas, M.C.; Moran, J.L.; Waden, J.; Thorn, L.M.; Makinen, V.P.; Rosengard-Barlund, M.; Saraheimo, M.; Hietala, K.; Heikkila, O.; et al. The presence and severity of chronic kidney disease predicts all-cause mortality in type 1 diabetes. Diabetes 2009, 58, 1651–1658. [Google Scholar] [CrossRef]
- Haffner, S.M.; Lehto, S.; Ronnemaa, T.; Pyorala, K.; Laakso, M. Mortality from coronary heart disease in subjects with type 2 diabetes and in nondiabetic subjects with and without prior myocardial infarction. N. Engl. J. Med. 1998, 339, 229–234. [Google Scholar] [CrossRef]
- Byrne, C.; Ford, D.; Gilg, J.; Ansell, D.; Feehally, J. UK Renal Registry 12th Annual Report (December 2009): Chapter 3: UK ESRD incident rates in 2008: National and centre-specific analyses. Nephron. Clin. Pract. 2010, 115, c9–c39. [Google Scholar] [CrossRef]
- Collins, A.J.; Foley, R.N.; Chavers, B.; Gilbertson, D.; Herzog, C.; Johansen, K.; Kasiske, B.; Kutner, N.; Liu, J.; St Peter, W.; et al. United States renal data system 2011 annual data report: Atlas of chronic kidney disease & end-stage renal disease in the United States. Am. J. Kidney Dis. 2012, 59, e1–e420. [Google Scholar] [CrossRef]
- Collins, A.J.; Foley, R.N.; Herzog, C.; Chavers, B.; Gilbertson, D.; Ishani, A.; Kasiske, B.; Liu, J.; Mau, L.W.; McBean, M.; et al. US renal data system 2010 annual data report. Am. J. Kidney Dis. 2011, 57, e1–e526. [Google Scholar] [CrossRef]
- Parving, H.H.; Oxenboll, B.; Svendsen, P.A.; Christiansen, J.S.; Andersen, A.R. Early detection of patients at risk of developing diabetic nephropathy. A longitudinal study of urinary albumin excretion. Acta Endocrinol. 1982, 100, 550–555. [Google Scholar]
- Viberti, G.C.; Hill, R.D.; Jarrett, R.J.; Argyropoulos, A.; Mahmud, U.; Keen, H. Microalbuminuria as a predictor of clinical nephropathy in insulin-dependent diabetes mellitus. Lancet 1982, 1, 1430–1432. [Google Scholar]
- Mogensen, C.E.; Christensen, C.K. Predicting diabetic nephropathy in insulin-dependent patients. N. Engl. J. Med. 1984, 311, 89–93. [Google Scholar] [CrossRef]
- Stephenson, J.; Fuller, J.H. Microvascular and acute complications in IDDM patients: The EURODIAB IDDM complications study. Diabetologia 1994, 37, 278–285. [Google Scholar] [CrossRef]
- Krolewski, A.S.; Warram, J.H.; Christlieb, A.R.; Busick, E.J.; Kahn, C.R. The changing natural history of nephropathy in type I diabetes. Am. J. Med. 1985, 78, 785–794. [Google Scholar] [CrossRef]
- Bojestig, M.; Arnqvist, H.J.; Hermansson, G.; Karlberg, B.E.; Ludvigsson, J. Declining incidence of nephropathy in insulin-dependent diabetes mellitus. N. Engl. J. Med. 1994, 330, 15–18. [Google Scholar] [CrossRef]
- Forsblom, C.M.; Groop, P.H.; Ekstrand, A.; Groop, L.C. Predictive value of microalbuminuria in patients with insulin-dependent diabetes of long duration. BMJ 1992, 305, 1051–1053. [Google Scholar] [CrossRef]
- Steinke, J.M.; Sinaiko, A.R.; Kramer, M.S.; Suissa, S.; Chavers, B.M.; Mauer, M. The early natural history of nephropathy in Type 1 Diabetes: III. Predictors of 5-year urinary albumin excretion rate patterns in initially normoalbuminuric patients. Diabetes 2005, 54, 2164–2171. [Google Scholar] [CrossRef]
- Perkins, B.A.; Ficociello, L.H.; Silva, K.H.; Finkelstein, D.M.; Warram, J.H.; Krolewski, A.S. Regression of microalbuminuria in type 1 diabetes. N. Engl. J. Med. 2003, 348, 2285–2293. [Google Scholar] [CrossRef]
- Hovind, P.; Tarnow, L.; Rossing, P.; Jensen, B.R.; Graae, M.; Torp, I.; Binder, C.; Parving, H.H. Predictors for the development of microalbuminuria and macroalbuminuria in patients with type 1 diabetes: Inception cohort study. BMJ 2004, 328, 1105. [Google Scholar]
- UK Prospective Diabetes Study Group. Tight blood pressure control and risk of macrovascular and microvascular complications in type 2 diabetes: UKPDS 38. BMJ 1998, 317, 703–713. [Google Scholar] [CrossRef]
- Spinas, G.A. Blood glucose monitoring and late complications of diabetes—Consequences of the Diabetes Control and Complications Trial (DCCT). Schweiz Med. Wochenschr. 1993, 123, 2387–2389. [Google Scholar]
- Chrisholm, D.J. The Diabetes Control and Complications Trial (DCCT). A milestone in diabetes management. Med. J. Aust. 1993, 159, 721–723. [Google Scholar]
- Brenner, B.M.; Cooper, M.E.; de Zeeuw, D.; Keane, W.F.; Mitch, W.E.; Parving, H.H.; Remuzzi, G.; Snapinn, S.M.; Zhang, Z.; Shahinfar, S. Effects of losartan on renal and cardiovascular outcomes in patients with type 2 diabetes and nephropathy. N. Engl. J. Med. 2001, 345, 861–869. [Google Scholar] [CrossRef]
- Rosolowsky, E.T.; Skupien, J.; Smiles, A.M.; Niewczas, M.; Roshan, B.; Stanton, R.; Eckfeldt, J.H.; Warram, J.H.; Krolewski, A.S. Risk for ESRD in type 1 diabetes remains high despite renoprotection. J. Am. Soc. Nephrol. 2011, 22, 545–553. [Google Scholar] [CrossRef]
- Brownlee, M. The pathobiology of diabetic complications: A unifying mechanism. Diabetes 2005, 54, 1615–1625. [Google Scholar] [CrossRef]
- Marrero, M.B.; Fulton, D.; Stepp, D.; Stern, D.M. Angiotensin II-induced signaling pathways in diabetes. Curr. Diabetes Rev. 2005, 1, 197–202. [Google Scholar] [CrossRef]
- Sharma, K.; Ziyadeh, F.N. Hyperglycemia and diabetic kidney disease. The case for transforming growth factor-beta as a key mediator. Diabetes 1995, 44, 1139–1146. [Google Scholar] [CrossRef]
- Natarajan, R.; Cai, Q. Monocyte retention in the pathology of atherosclerosis. Future Cardiol. 2005, 1, 331–340. [Google Scholar] [CrossRef]
- Yan, S.F.; Ramasamy, R.; Schmidt, A.M. Mechanisms of disease: Advanced glycation end-products and their receptor in inflammation and diabetes complications. Nat. Clin. Pract. Endocrinol. Metab. 2008, 4, 285–293. [Google Scholar] [CrossRef]
- McMahon, B.; Mitchell, D.; Shattock, R.; Martin, F.; Brady, H.R.; Godson, C. Lipoxin, leukotriene, and PDGF receptors cross-talk to regulate mesangial cell proliferation. FASEB J. 2002, 16, 1817–1819. [Google Scholar]
- Borgeson, E.; Godson, C. Molecular circuits of resolution in renal disease. Sci. World J. 2010, 10, 1370–1385. [Google Scholar] [CrossRef]
- King, G.L. The role of inflammatory cytokines in diabetes and its complications. J. Periodontol. 2008, 79, 1527–1534. [Google Scholar] [CrossRef]
- Geraldes, P.; King, G.L. Activation of protein kinase C isoforms and its impact on diabetic complications. Circ. Res. 2010, 106, 1319–1331. [Google Scholar] [CrossRef]
- Schmidt, A.M.; Yan, S.D.; Wautier, J.L.; Stern, D. Activation of receptor for advanced glycation end products: A mechanism for chronic vascular dysfunction in diabetic vasculopathy and atherosclerosis. Circ. Res. 1999, 84, 489–497. [Google Scholar] [CrossRef]
- Schmid, H.; Boucherot, A.; Yasuda, Y.; Henger, A.; Brunner, B.; Eichinger, F.; Nitsche, A.; Kiss, E.; Bleich, M.; Grone, H.J.; et al. Modular activation of nuclear factor-kappaB transcriptional programs in human diabetic nephropathy. Diabetes 2006, 55, 2993–3003. [Google Scholar] [CrossRef]
- Villeneuve, L.M.; Natarajan, R. The role of epigenetics in the pathology of diabetic complications. Am. J. Physiol. Ren. Physiol. 2010, 299, F14–F25. [Google Scholar] [CrossRef]
- Pezzolesi, M.G.; Poznik, G.D.; Mychaleckyj, J.C.; Paterson, A.D.; Barati, M.T.; Klein, J.B.; Ng, D.P.; Placha, G.; Canani, L.H.; Bochenski, J.; et al. Genome-wide association scan for diabetic nephropathy susceptibility genes in type 1 diabetes. Diabetes 2009, 58, 1403–1410. [Google Scholar] [CrossRef]
- Sandholm, N.; Salem, R.M.; McKnight, A.J.; Brennan, E.P.; Forsblom, C.; Isakova, T.; McKay, G.J.; Williams, W.W.; Sadlier, D.M.; Makinen, V.P.; et al. New susceptibility loci associated with kidney disease in type 1 diabetes. PLoS Genet. 2012, 8, e1002921. [Google Scholar] [CrossRef] [Green Version]
- The Diabetes Control and Complications Trial Research Group. The effect of intensive treatment of diabetes on the development and progression of long-term complications in insulin-dependent diabetes mellitus. N. Engl. J. Med. 1993, 329, 977–986. [Google Scholar] [CrossRef]
- UK Prospective Diabetes Study Group. Intensive blood-glucose control with sulphonylureas or insulin compared with conventional treatment and risk of complications in patients with type 2 diabetes (UKPDS 33). Lancet 1998, 352, 837–853. [Google Scholar] [CrossRef]
- Clarkson, M.R.; Murphy, M.; Gupta, S.; Lambe, T.; Mackenzie, H.S.; Godson, C.; Martin, F.; Brady, H.R. High glucose-altered gene expression in mesangial cells. Actin-regulatory protein gene expression is triggered by oxidative stress and cytoskeletal disassembly. J. Biol. Chem. 2002, 277, 9707–9712. [Google Scholar] [CrossRef]
- Murphy, M.; Godson, C.; Cannon, S.; Kato, S.; Mackenzie, H.S.; Martin, F.; Brady, H.R. Suppression subtractive hybridization identifies high glucose levels as a stimulus for expression of connective tissue growth factor and other genes in human mesangial cells. J. Biol. Chem. 1999, 274, 5830–5834. [Google Scholar]
- McKnight, A.J.; Patterson, C.C.; Pettigrew, K.A.; Savage, D.A.; Kilner, J.; Murphy, M.; Sadlier, D.; Maxwell, A.P. A GREM1 gene variant associates with diabetic nephropathy. J. Am. Soc. Nephrol. 2010, 21, 773–781. [Google Scholar] [CrossRef]
- McMahon, R.; Murphy, M.; Clarkson, M.; Taal, M.; Mackenzie, H.S.; Godson, C.; Martin, F.; Brady, H.R. IHG-2, a mesangial cell gene induced by high glucose, is human gremlin. Regulation by extracellular glucose concentration, cyclic mechanical strain, and transforming growth factor-beta1. J. Biol. Chem. 2000, 275, 9901–9904. [Google Scholar] [CrossRef]
- Murphy, M.; Docherty, N.G.; Griffin, B.; Howlin, J.; McArdle, E.; McMahon, R.; Schmid, H.; Kretzler, M.; Droguett, A.; Mezzano, S.; et al. IHG-1 amplifies TGF-beta1 signaling and is increased in renal fibrosis. J. Am. Soc. Nephrol. 2008, 19, 1672–1680. [Google Scholar] [CrossRef]
- Krupa, A.; Jenkins, R.; Luo, D.D.; Lewis, A.; Phillips, A.; Fraser, D. Loss of MicroRNA-192 promotes fibrogenesis in diabetic nephropathy. J. Am. Soc. Nephrol. 2010, 21, 438–447. [Google Scholar] [CrossRef]
- Dey, N.; Das, F.; Mariappan, M.M.; Mandal, C.C.; Ghosh-Choudhury, N.; Kasinath, B.S.; Choudhury, G.G. MicroRNA-21 orchestrates high glucose-induced signals to TOR complex 1, resulting in renal cell pathology in diabetes. J. Biol. Chem. 2011, 286, 25586–25603. [Google Scholar] [CrossRef]
- Wang, Q.; Wang, Y.; Minto, A.W.; Wang, J.; Shi, Q.; Li, X.; Quigg, R.J. MicroRNA-377 is up-regulated and can lead to increased fibronectin production in diabetic nephropathy. FASEB J. 2008, 22, 4126–4135. [Google Scholar] [CrossRef]
- Sun, G.; Reddy, M.A.; Yuan, H.; Lanting, L.; Kato, M.; Natarajan, R. Epigenetic histone methylation modulates fibrotic gene expression. J. Am. Soc. Nephrol. 2010, 21, 2069–2080. [Google Scholar] [CrossRef]
- Gao, C.; Chen, G.; Liu, L.; Li, X.; He, J.; Jiang, L.; Zhu, J.; Xu, Y. Impact of high glucose and proteasome inhibitor MG132 on Histone H2A and H2B ubiquitination in rat glomerular mesangial cells. J. Diabetes Res. 2013, 2013, 589474. [Google Scholar]
- Shi, Y.; Massague, J. Mechanisms of TGF-beta signaling from cell membrane to the nucleus. Cell 2003, 113, 685–700. [Google Scholar] [CrossRef]
- Schmierer, B.; Hill, C.S. TGFbeta-SMAD signal transduction: Molecular specificity and functional flexibility. Nat. Rev. Mol. Cell Biol. 2007, 8, 970–982. [Google Scholar] [CrossRef]
- Chaudhury, A.; Howe, P.H. The tale of transforming growth factor-beta (TGFbeta) signaling: A soigne enigma. IUBMB Life 2009, 61, 929–939. [Google Scholar] [CrossRef]
- Reeves, W.B.; Andreoli, T.E. Transforming growth factor beta contributes to progressive diabetic nephropathy. Proc. Natl. Acad. Sci. USA 2000, 97, 7667–7669. [Google Scholar] [CrossRef]
- Russo, L.M.; del Re, E.; Brown, D.; Lin, H.Y. Evidence for a role of transforming growth factor (TGF)-beta1 in the induction of postglomerular albuminuria in diabetic nephropathy: Amelioration by soluble TGF-beta type II receptor. Diabetes 2007, 56, 380–388. [Google Scholar] [CrossRef]
- Sharma, K.; Jin, Y.; Guo, J.; Ziyadeh, F.N. Neutralization of TGF-beta by anti-TGF-beta antibody attenuates kidney hypertrophy and the enhanced extracellular matrix gene expression in STZ-induced diabetic mice. Diabetes 1996, 45, 522–530. [Google Scholar] [CrossRef]
- Ziyadeh, F.N.; Hoffman, B.B.; Han, D.C.; Iglesias-De La Cruz, M.C.; Hong, S.W.; Isono, M.; Chen, S.; McGowan, T.A.; Sharma, K. Long-term prevention of renal insufficiency, excess matrix gene expression, and glomerular mesangial matrix expansion by treatment with monoclonal antitransforming growth factor-beta antibody in db/db diabetic mice. Proc. Natl. Acad. Sci. USA 2000, 97, 8015–8020. [Google Scholar] [CrossRef]
- Houlihan, C.A.; Akdeniz, A.; Tsalamandris, C.; Cooper, M.E.; Jerums, G.; Gilbert, R.E. Urinary transforming growth factor-beta excretion in patients with hypertension, type 2 diabetes, and elevated albumin excretion rate: Effects of angiotensin receptor blockade and sodium restriction. Diabetes Care 2002, 25, 1072–1077. [Google Scholar] [CrossRef]
- Goumenos, D.S.; Tsakas, S.; El Nahas, A.M.; Alexandri, S.; Oldroyd, S.; Kalliakmani, P.; Vlachojannis, J.G. Transforming growth factor-beta(1) in the kidney and urine of patients with glomerular disease and proteinuria. Nephrol. Dial. Transplant. 2002, 17, 2145–2152. [Google Scholar] [CrossRef]
- Brennan, E.P.; Morine, M.J.; Walsh, D.W.; Roxburgh, S.A.; Lindenmeyer, M.T.; Brazil, D.P.; Gaora, P.O.; Roche, H.M.; Sadlier, D.M.; Cohen, C.D.; et al. Next-generation sequencing identifies TGF-beta1-associated gene expression profiles in renal epithelial cells reiterated in human diabetic nephropathy. Biochim. Biophys. Acta 2012, 1822, 589–599. [Google Scholar]
- Kalluri, R.; Weinberg, R.A. The basics of epithelial-mesenchymal transition. J. Clin. Invest. 2009, 119, 1420–1428. [Google Scholar] [CrossRef]
- LeBleu, V.S.; Taduri, G.; O’Connell, J.; Teng, Y.; Cooke, V.G.; Woda, C.; Sugimoto, H.; Kalluri, R. Origin and function of myofibroblasts in kidney fibrosis. Nat. Med. 2013, 19, 1047–1053. [Google Scholar] [CrossRef]
- Borges, F.T.; Melo, S.A.; Ozdemir, B.C.; Kato, N.; Revuelta, I.; Miller, C.A.; Gattone, V.H., 2nd; LeBleu, V.S.; Kalluri, R. TGF-beta1-containing exosomes from injured epithelial cells activate fibroblasts to initiate tissue regenerative responses and fibrosis. J. Am. Soc. Nephrol. 2013, 24, 385–392. [Google Scholar] [CrossRef]
- Yoshikawa, M.; Hishikawa, K.; Marumo, T.; Fujita, T. Inhibition of histone deacetylase activity suppresses epithelial-to-mesenchymal transition induced by TGF-beta1 in human renal epithelial cells. J. Am. Soc. Nephrol. 2007, 18, 58–65. [Google Scholar] [CrossRef]
- Kato, M.; Putta, S.; Wang, M.; Yuan, H.; Lanting, L.; Nair, I.; Gunn, A.; Nakagawa, Y.; Shimano, H.; Todorov, I.; et al. TGF-beta activates Akt kinase through a microRNA-dependent amplifying circuit targeting PTEN. Nat. Cell Biol. 2009, 11, 881–889. [Google Scholar] [CrossRef]
- Kato, M.; Zhang, J.; Wang, M.; Lanting, L.; Yuan, H.; Rossi, J.J.; Natarajan, R. MicroRNA-192 in diabetic kidney glomeruli and its function in TGF-beta-induced collagen expression via inhibition of E-box repressors. Proc. Natl. Acad. Sci. USA 2007, 104, 3432–3437. [Google Scholar]
- Wang, B.; Herman-Edelstein, M.; Koh, P.; Burns, W.; Jandeleit-Dahm, K.; Watson, A.; Saleem, M.; Goodall, G.J.; Twigg, S.M.; Cooper, M.E.; et al. E-cadherin expression is regulated by miR-192/215 by a mechanism that is independent of the profibrotic effects of transforming growth factor-beta. Diabetes 2010, 59, 1794–1802. [Google Scholar] [CrossRef]
- Brennan, E.P.; Nolan, K.A.; Borgeson, E.; Gough, O.S.; McEvoy, C.M.; Docherty, N.G.; Higgins, D.F.; Murphy, M.; Sadlier, D.M.; Ali-Shah, S.T.; et al. Lipoxins attenuate renal fibrosis by inducing let-7c and suppressing TGFbetaR1. J. Am. Soc. Nephrol. 2013, 24, 627–637. [Google Scholar] [CrossRef]
- Gruden, G.; Thomas, S.; Burt, D.; Lane, S.; Chusney, G.; Sacks, S.; Viberti, G. Mechanical stretch induces vascular permeability factor in human mesangial cells: Mechanisms of signal transduction. Proc. Natl. Acad. Sci. USA 1997, 94, 12112–12116. [Google Scholar] [CrossRef]
- Crean, J.K.; Furlong, F.; Finlay, D.; Mitchell, D.; Murphy, M.; Conway, B.; Brady, H.R.; Godson, C.; Martin, F. Connective tissue growth factor [CTGF]/CCN2 stimulates mesangial cell migration through integrated dissolution of focal adhesion complexes and activation of cell polarization. FASEB J. 2004, 18, 1541–1543. [Google Scholar]
- Faherty, N.; Curran, S.P.; O’Donovan, H.; Martin, F.; Godson, C.; Brazil, D.P.; Crean, J.K. CCN2/CTGF increases expression of miR-302 microRNAs, which target the TGFbeta type II receptor with implications for nephropathic cell phenotypes. J. Cell Sci. 2012, 125, 5621–5629. [Google Scholar] [CrossRef]
- Martinez-Estrada, O.M.; Lettice, L.A.; Essafi, A.; Guadix, J.A.; Slight, J.; Velecela, V.; Hall, E.; Reichmann, J.; Devenney, P.S.; Hohenstein, P.; et al. Et1 is required for cardiovascular progenitor cell formation through transcriptional control of Snail and E-cadherin. Nat. Genet. 2010, 42, 89–93. [Google Scholar] [CrossRef]
- Breyer, M.D.; Bottinger, E.; Brosius, F.C., 3rd; Coffman, T.M.; Harris, R.C.; Heilig, C.W.; Sharma, K. Mouse models of diabetic nephropathy. J. Am. Soc. Nephrol. 2005, 16, 27–45. [Google Scholar] [CrossRef]
- Alpers, C.E.; Hudkins, K.L. Mouse models of diabetic nephropathy. Curr. Opin. Nephrol. Hypertens. 2011, 20, 278–284. [Google Scholar] [CrossRef]
- Brosius, F.C., 3rd; Alpers, C.E.; Bottinger, E.P.; Breyer, M.D.; Coffman, T.M.; Gurley, S.B.; Harris, R.C.; Kakoki, M.; Kretzler, M.; Leiter, E.H.; et al. Mouse models of diabetic nephropathy. J. Am. Soc. Nephrol. 2009, 20, 2503–2512. [Google Scholar] [CrossRef]
- Tesch, G.H.; Lim, A.K. Recent insights into diabetic renal injury from the db/db mouse model of type 2 diabetic nephropathy. Am. J. Physiol. Ren. Physiol. 2011, 300, F301–F310. [Google Scholar] [CrossRef]
- Higgins, D.F.; Kimura, K.; Bernhardt, W.M.; Shrimanker, N.; Akai, Y.; Hohenstein, B.; Saito, Y.; Johnson, R.S.; Kretzler, M.; Cohen, C.D.; et al. Hypoxia promotes fibrogenesis in vivo via HIF-1 stimulation of epithelial-to-mesenchymal transition. J. Clin. Invest. 2007, 117, 3810–3820. [Google Scholar]
- Chevalier, R.L. Specific molecular targeting of renal injury in obstructive nephropathy. Kidney Int. 2006, 70, 1200–1201. [Google Scholar] [CrossRef]
- Chevalier, R.L. Obstructive nephropathy: Towards biomarker discovery and gene therapy. Nat. Clin. Pract. Nephrol. 2006, 2, 157–168. [Google Scholar] [CrossRef]
- Chevalier, R.L. Pathogenesis of renal injury in obstructive uropathy. Curr. Opin. Pediatr. 2006, 18, 153–160. [Google Scholar] [CrossRef]
- Hong, S.W.; Isono, M.; Chen, S.; Iglesias-De La Cruz, M.C.; Han, D.C.; Ziyadeh, F.N. Increased glomerular and tubular expression of transforming growth factor-beta1, its type II receptor, and activation of the Smad signaling pathway in the db/db mouse. Am. J. Pathol. 2001, 158, 1653–1663. [Google Scholar] [CrossRef]
- Usui, H.; Shikata, K.; Matsuda, M.; Okada, S.; Ogawa, D.; Yamashita, T.; Hida, K.; Satoh, M.; Wada, J.; Makino, H. HMG-CoA reductase inhibitor ameliorates diabetic nephropathy by its pleiotropic effects in rats. Nephrol. Dial. Transplant. 2003, 18, 265–272. [Google Scholar] [CrossRef]
- Zhang, Q.; Xiao, X.; Li, M.; Li, W.; Yu, M.; Zhang, H.; Sun, X.; Mao, L.; Xiang, H. Gene expression profiling in glomeruli of diabetic nephropathy rat. Exp. Biol. Med. 2012, 237, 903–911. [Google Scholar] [CrossRef]
- Wada, J.; Zhang, H.; Tsuchiyama, Y.; Hiragushi, K.; Hida, K.; Shikata, K.; Kanwar, Y.S.; Makino, H. Gene expression profile in streptozotocin-induced diabetic mice kidneys undergoing glomerulosclerosis. Kidney Int. 2001, 59, 1363–1373. [Google Scholar] [CrossRef]
- Woroniecka, K.I.; Park, A.S.; Mohtat, D.; Thomas, D.B.; Pullman, J.M.; Susztak, K. Transcriptome analysis of human diabetic kidney disease. Diabetes 2011, 60, 2354–2369. [Google Scholar] [CrossRef]
- Reich, H.N.; Tritchler, D.; Cattran, D.C.; Herzenberg, A.M.; Eichinger, F.; Boucherot, A.; Henger, A.; Berthier, C.C.; Nair, V.; Cohen, C.D.; et al. A molecular signature of proteinuria in glomerulonephritis. PLoS One 2010, 5, e13451. [Google Scholar] [CrossRef] [Green Version]
- Tonna, S.; El-Osta, A.; Cooper, M.E.; Tikellis, C. Metabolic memory and diabetic nephropathy: Potential role for epigenetic mechanisms. Nat. Rev. Nephrol. 2010, 6, 332–341. [Google Scholar] [CrossRef]
- El-Osta, A.; Brasacchio, D.; Yao, D.; Pocai, A.; Jones, P.L.; Roeder, R.G.; Cooper, M.E.; Brownlee, M. Transient high glucose causes persistent epigenetic changes and altered gene expression during subsequent normoglycemia. J. Exp. Med. 2008, 205, 2409–2417. [Google Scholar] [CrossRef]
- Miao, F.; Wu, X.; Zhang, L.; Yuan, Y.C.; Riggs, A.D.; Natarajan, R. Genome-wide analysis of histone lysine methylation variations caused by diabetic conditions in human monocytes. J. Biol. Chem. 2007, 282, 13854–13863. [Google Scholar]
- Bell, C.G.; Teschendorff, A.E.; Rakyan, V.K.; Maxwell, A.P.; Beck, S.; Savage, D.A. Genome-wide DNA methylation analysis for diabetic nephropathy in type 1 diabetes mellitus. BMC Med. Genomics 2010, 3, 33. [Google Scholar] [CrossRef] [Green Version]
- Brennan, E.P.; Ehrich, M.; Brazil, D.P.; Crean, J.K.; Murphy, M.; Sadlier, D.M.; Martin, F.; Godson, C.; McKnight, A.J.; van den Boom, D.; et al. Comparative analysis of DNA methylation profiles in peripheral blood leukocytes versus lymphoblastoid cell lines. Epigenetics 2009, 4, 159–164. [Google Scholar] [CrossRef]
- Bechtel, W.; McGoohan, S.; Zeisberg, E.M.; Muller, G.A.; Kalbacher, H.; Salant, D.J.; Muller, C.A.; Kalluri, R.; Zeisberg, M. Methylation determines fibroblast activation and fibrogenesis in the kidney. Nat. Med. 2010, 16, 544–550. [Google Scholar] [CrossRef]
- Brennan, E.P. Lipoxins attenuate renal fibrosis by induction of let-7c and suppression of TGFβR1. J. Am. Soc. Nephrol. 2013, in press. [Google Scholar]
- Zhong, X.; Chung, A.C.; Chen, H.Y.; Meng, X.M.; Lan, H.Y. Smad3-mediated upregulation of miR-21 promotes renal fibrosis. J. Am. Soc. Nephrol. 2011, 22, 1668–1681. [Google Scholar] [CrossRef]
- Wang, B.; Komers, R.; Carew, R.; Winbanks, C.E.; Xu, B.; Herman-Edelstein, M.; Koh, P.; Thomas, M.; Jandeleit-Dahm, K.; Gregorevic, P.; et al. Suppression of microRNA-29 expression by TGF-beta1 promotes collagen expression and renal fibrosis. J. Am. Soc. Nephrol. 2012, 23, 252–265. [Google Scholar] [CrossRef]
- Chung, A.C.; Huang, X.R.; Meng, X.; Lan, H.Y. miR-192 mediates TGF-beta/Smad3-driven renal fibrosis. J. Am. Soc. Nephrol. 2010, 21, 1317–1325. [Google Scholar] [CrossRef]
- Argyropoulos, C.; Wang, K.; McClarty, S.; Huang, D.; Bernardo, J.; Ellis, D.; Orchard, T.; Galas, D.; Johnson, J. Urinary microRNA profiling in the nephropathy of type 1 diabetes. PLoS One 2013, 8, e54662. [Google Scholar] [CrossRef] [Green Version]
- Beltrami, C.; Clayton, A.; Phillips, A.O.; Fraser, D.J.; Bowen, T. Analysis of urinary microRNAs in chronic kidney disease. Biochem. Soc. Trans. 2012, 40, 875–879. [Google Scholar]
- Bouchie, A. First microRNA mimic enters clinic. Nat. Biotechnol. 2013, 31, 577. [Google Scholar] [CrossRef]
- Borch-Johnsen, K.; Norgaard, K.; Hommel, E.; Mathiesen, E.R.; Jensen, J.S.; Deckert, T.; Parving, H.H. Is diabetic nephropathy an inherited complication? Kidney Int. 1992, 41, 719–722. [Google Scholar] [CrossRef]
- Quinn, M.; Angelico, M.C.; Warram, J.H.; Krolewski, A.S. Familial factors determine the development of diabetic nephropathy in patients with IDDM. Diabetologia 1996, 39, 940–945. [Google Scholar] [CrossRef]
- Seaquist, E.R.; Goetz, F.C.; Rich, S.; Barbosa, J. Familial clustering of diabetic kidney disease. Evidence for genetic susceptibility to diabetic nephropathy. N. Engl. J. Med. 1989, 320, 1161–1165. [Google Scholar] [CrossRef]
- Barzilay, J.; Warram, J.H.; Bak, M.; Laffel, L.M.; Canessa, M.; Krolewski, A.S. Predisposition to hypertension: Risk factor for nephropathy and hypertension in IDDM. Kidney Int. 1992, 41, 723–730. [Google Scholar] [CrossRef]
- Earle, K.; Walker, J.; Hill, C.; Viberti, G. Familial clustering of cardiovascular disease in patients with insulin-dependent diabetes and nephropathy. N. Engl. J. Med. 1992, 326, 673–677. [Google Scholar] [CrossRef]
- Moczulski, D.K.; Rogus, J.J.; Antonellis, A.; Warram, J.H.; Krolewski, A.S. Major susceptibility locus for nephropathy in type 1 diabetes on chromosome 3q: Results of novel discordant sib-pair analysis. Diabetes 1998, 47, 1164–1169. [Google Scholar] [CrossRef]
- Chistiakov, D.A.; Savost’anov, K.V.; Shestakova, M.V.; Chugunova, L.A.; Samkhalova, M.; Dedov, I.I.; Nosikov, V.V. Confirmation of a susceptibility locus for diabetic nephropathy on chromosome 3q23-q24 by association study in Russian type 1 diabetic patients. Diabetes Res. Clin. Pract. 2004, 66, 79–86. [Google Scholar] [CrossRef]
- Osterholm, A.M.; He, B.; Pitkaniemi, J.; Albinsson, L.; Berg, T.; Sarti, C.; Tuomilehto, J.; Tryggvason, K. Genome-wide scan for type 1 diabetic nephropathy in the Finnish population reveals suggestive linkage to a single locus on chromosome 3q. Kidney Int. 2007, 71, 140–145. [Google Scholar] [CrossRef]
- Imperatore, G.; Hanson, R.L.; Pettitt, D.J.; Kobes, S.; Bennett, P.H.; Knowler, W.C. Sib-pair linkage analysis for susceptibility genes for microvascular complications among Pima Indians with type 2 diabetes. Pima Diabetes Genes Group. Diabetes 1998, 47, 821–830. [Google Scholar]
- Bowden, D.W.; Colicigno, C.J.; Langefeld, C.D.; Sale, M.M.; Williams, A.; Anderson, P.J.; Rich, S.S.; Freedman, B.I. A genome scan for diabetic nephropathy in African Americans. Kidney Int. 2004, 66, 1517–1526. [Google Scholar] [CrossRef]
- Krolewski, A.S.; Poznik, G.D.; Placha, G.; Canani, L.; Dunn, J.; Walker, W.; Smiles, A.; Krolewski, B.; Fogarty, D.G.; Moczulski, D.; et al. A genome-wide linkage scan for genes controlling variation in urinary albumin excretion in type II diabetes. Kidney Int. 2006, 69, 129–136. [Google Scholar] [CrossRef]
- Iyengar, S.K.; Abboud, H.E.; Goddard, K.A.; Saad, M.F.; Adler, S.G.; Arar, N.H.; Bowden, D.W.; Duggirala, R.; Elston, R.C.; Hanson, R.L.; et al. Genome-wide scans for diabetic nephropathy and albuminuria in multiethnic populations: The family investigation of nephropathy and diabetes (FIND). Diabetes 2007, 56, 1577–1585. [Google Scholar] [CrossRef]
- Vardarli, I.; Baier, L.J.; Hanson, R.L.; Akkoyun, I.; Fischer, C.; Rohmeiss, P.; Basci, A.; Bartram, C.R.; van der Woude, F.J.; Janssen, B. Gene for susceptibility to diabetic nephropathy in type 2 diabetes maps to 18q22.3–23. Kidney Int. 2002, 62, 2176–2183. [Google Scholar] [CrossRef]
- Schelling, J.R.; Abboud, H.E.; Nicholas, S.B.; Pahl, M.V.; Sedor, J.R.; Adler, S.G.; Arar, N.H.; Bowden, D.W.; Elston, R.C.; Freedman, B.I.; et al. Genome-wide scan for estimated glomerular filtration rate in multi-ethnic diabetic populations: The Family Investigation of Nephropathy and Diabetes (FIND). Diabetes 2008, 57, 235–243. [Google Scholar]
- McKnight, A.J.; Savage, D.A.; Patterson, C.C.; Brady, H.R.; Maxwell, A.P. Resequencing of the characterised CTGF gene to identify novel or known variants, and analysis of their association with diabetic nephropathy. J. Hum. Genet. 2006, 51, 383–386. [Google Scholar] [CrossRef]
- Pociot, F.; Hansen, P.M.; Karlsen, A.E.; Langdahl, B.L.; Johannesen, J.; Nerup, J. TGF-beta1 gene mutations in insulin-dependent diabetes mellitus and diabetic nephropathy. J. Am. Soc. Nephrol. 1998, 9, 2302–2307. [Google Scholar]
- Patel, A.; Scott, W.R.; Lympany, P.A.; Rippin, J.D.; Gill, G.V.; Barnett, A.H.; Bain, S.C. The TGF-beta 1 gene codon 10 polymorphism contributes to the genetic predisposition to nephropathy in Type 1 diabetes. Diabet. Med. 2005, 22, 69–73. [Google Scholar] [CrossRef]
- Ewens, K.G.; George, R.A.; Sharma, K.; Ziyadeh, F.N.; Spielman, R.S. Assessment of 115 candidate genes for diabetic nephropathy by transmission/disequilibrium test. Diabetes 2005, 54, 3305–3318. [Google Scholar] [CrossRef]
- McKnight, A.J.; Maxwell, A.P.; Patterson, C.C.; Brady, H.R.; Savage, D.A. Association of VEGF-1499C-->T polymorphism with diabetic nephropathy in type 1 diabetes mellitus. J. Diabetes Complicat. 2007, 21, 242–245. [Google Scholar]
- McKnight, A.J.; Pettigrew, K.A.; Patterson, C.C.; Kilner, J.; Sadlier, D.M.; Maxwell, A.P. Investigation of the association of BMP gene variants with nephropathy in Type 1 diabetes mellitus. Diabet. Med. 2010, 27, 624–630. [Google Scholar]
- Ma, J.; Mollsten, A.; Prazny, M.; Falhammar, H.; Brismar, K.; Dahlquist, G.; Efendic, S.; Gu, H.F. Genetic influences of the intercellular adhesion molecule 1 (ICAM-1) gene polymorphisms in development of Type 1 diabetes and diabetic nephropathy. Diabet. Med. 2006, 23, 1093–1099. [Google Scholar] [CrossRef]
- Yang, B.; Houlberg, K.; Millward, A.; Demaine, A. Polymorphisms of chemokine and chemokine receptor genes in Type 1 diabetes mellitus and its complications. Cytokine 2004, 26, 114–121. [Google Scholar] [CrossRef]
- Poirier, O.; Nicaud, V.; Vionnet, N.; Raoux, S.; Tarnow, L.; Vlassara, H.; Parving, H.H.; Cambien, F. Polymorphism screening of four genes encoding advanced glycation end-product putative receptors. Association study with nephropathy in type 1 diabetic patients. Diabetes 2001, 50, 1214–1218. [Google Scholar] [CrossRef]
- McKay, G.J.; Savage, D.A.; Patterson, C.C.; Lewis, G.; McKnight, A.J.; Maxwell, A.P. Association analysis of dyslipidemia-related genes in diabetic nephropathy. PLoS One 2013, 8, e58472. [Google Scholar]
- Doria, A.; Warram, J.H.; Krolewski, A.S. Genetic predisposition to diabetic nephropathy. Evidence for a role of the angiotensin I-converting enzyme gene. Diabetes 1994, 43, 690–695. [Google Scholar] [CrossRef]
- Marre, M.; Bernadet, P.; Gallois, Y.; Savagner, F.; Guyene, T.T.; Hallab, M.; Cambien, F.; Passa, P.; Alhenc-Gelas, F. Relationships between angiotensin I converting enzyme gene polymorphism, plasma levels, and diabetic retinal and renal complications. Diabetes 1994, 43, 384–388. [Google Scholar] [CrossRef]
- Tarnow, L.; Cambien, F.; Rossing, P.; Nielsen, F.S.; Hansen, B.V.; Lecerf, L.; Poirier, O.; Danilov, S.; Parving, H.H. Lack of relationship between an insertion/deletion polymorphism in the angiotensin I-converting enzyme gene and diabetic nephropathy and proliferative retinopathy in IDDM patients. Diabetes 1995, 44, 489–494. [Google Scholar] [CrossRef]
- Chowdhury, T.A.; Dronsfield, M.J.; Kumar, S.; Gough, S.L.; Gibson, S.P.; Khatoon, A.; MacDonald, F.; Rowe, B.R.; Dunger, D.B.; Dean, J.D.; et al. Examination of two genetic polymorphisms within the renin-angiotensin system: No evidence for an association with nephropathy in IDDM. Diabetologia 1996, 39, 1108–1114. [Google Scholar] [CrossRef]
- Boright, A.P.; Paterson, A.D.; Mirea, L.; Bull, S.B.; Mowjoodi, A.; Scherer, S.W.; Zinman, B. Genetic variation at the ACE gene is associated with persistent microalbuminuria and severe nephropathy in type 1 diabetes: The DCCT/EDIC Genetics Study. Diabetes 2005, 54, 1238–1244. [Google Scholar] [CrossRef]
- Hadjadj, S.; Tarnow, L.; Forsblom, C.; Kazeem, G.; Marre, M.; Groop, P.H.; Parving, H.H.; Cambien, F.; Tregouet, D.A.; Gut, I.G.; et al. Association between angiotensin-converting enzyme gene polymorphisms and diabetic nephropathy: Case-control, haplotype, and family-based study in three European populations. J. Am. Soc. Nephrol. 2007, 18, 1284–1291. [Google Scholar] [CrossRef]
- Ng, D.P.; Tai, B.C.; Koh, D.; Tan, K.W.; Chia, K.S. Angiotensin-I converting enzyme insertion/deletion polymorphism and its association with diabetic nephropathy: A meta-analysis of studies reported between 1994 and 2004 and comprising 14,727 subjects. Diabetologia 2005, 48, 1008–1016. [Google Scholar] [CrossRef]
- Osawa, N.; Koya, D.; Araki, S.; Uzu, T.; Tsunoda, T.; Kashiwagi, A.; Nakamura, Y.; Maeda, S. Combinational effect of genes for the renin-angiotensin system in conferring susceptibility to diabetic nephropathy. J. Hum. Genet. 2007, 52, 143–151. [Google Scholar] [CrossRef]
- Barrett, J.C.; Clayton, D.G.; Concannon, P.; Akolkar, B.; Cooper, J.D.; Erlich, H.A.; Julier, C.; Morahan, G.; Nerup, J.; Nierras, C.; et al. Genome-wide association study and meta-analysis find that over 40 loci affect risk of type 1 diabetes. Nat. Genet. 2009, 41, 703–707. [Google Scholar] [CrossRef] [Green Version]
- Zeggini, E.; Scott, L.J.; Saxena, R.; Voight, B.F.; Marchini, J.L.; Hu, T.; de Bakker, P.I.; Abecasis, G.R.; Almgren, P.; Andersen, G.; et al. Meta-analysis of genome-wide association data and large-scale replication identifies additional susceptibility loci for type 2 diabetes. Nat. Genet. 2008, 40, 638–645. [Google Scholar] [CrossRef]
- Voight, B.F.; Scott, L.J.; Steinthorsdottir, V.; Morris, A.P.; Dina, C.; Welch, R.P.; Zeggini, E.; Huth, C.; Aulchenko, Y.S.; Thorleifsson, G.; et al. Twelve type 2 diabetes susceptibility loci identified through large-scale association analysis. Nat. Genet. 2010, 42, 579–589. [Google Scholar] [CrossRef]
- Dupuis, J.; Langenberg, C.; Prokopenko, I.; Saxena, R.; Soranzo, N.; Jackson, A.U.; Wheeler, E.; Glazer, N.L.; Bouatia-Naji, N.; Gloyn, A.L.; et al. New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat. Genet. 2010, 42, 105–116. [Google Scholar] [CrossRef] [Green Version]
- Sladek, R.; Rocheleau, G.; Rung, J.; Dina, C.; Shen, L.; Serre, D.; Boutin, P.; Vincent, D.; Belisle, A.; Hadjadj, S.; et al. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 2007, 445, 881–885. [Google Scholar] [CrossRef]
- Bell, J.T.; Timpson, N.J.; Rayner, N.W.; Zeggini, E.; Frayling, T.M.; Hattersley, A.T.; Morris, A.P.; McCarthy, M.I. Genome-wide association scan allowing for epistasis in type 2 diabetes. Ann. Hum. Genet. 2011, 75, 10–19. [Google Scholar] [CrossRef]
- Kao, W.H.; Klag, M.J.; Meoni, L.A.; Reich, D.; Berthier-Schaad, Y.; Li, M.; Coresh, J.; Patterson, N.; Tandon, A.; Powe, N.R.; et al. MYH9 is associated with nondiabetic end-stage renal disease in African Americans. Nat. Genet. 2008, 40, 1185–1192. [Google Scholar] [CrossRef]
- Kopp, J.B.; Smith, M.W.; Nelson, G.W.; Johnson, R.C.; Freedman, B.I.; Bowden, D.W.; Oleksyk, T.; McKenzie, L.M.; Kajiyama, H.; Ahuja, T.S.; et al. MYH9 is a major-effect risk gene for focal segmental glomerulosclerosis. Nat. Genet. 2008, 40, 1175–1184. [Google Scholar] [CrossRef]
- Stanescu, H.C.; Arcos-Burgos, M.; Medlar, A.; Bockenhauer, D.; Kottgen, A.; Dragomirescu, L.; Voinescu, C.; Patel, N.; Pearce, K.; Hubank, M.; et al. Risk HLA-DQA1 and PLA(2)R1 alleles in idiopathic membranous nephropathy. N. Engl. J. Med. 2011, 364, 616–626. [Google Scholar] [CrossRef]
- Gharavi, A.G.; Yan, Y.; Scolari, F.; Schena, F.P.; Frasca, G.M.; Ghiggeri, G.M.; Cooper, K.; Amoroso, A.; Viola, B.F.; Battini, G.; et al. IgA nephropathy, the most common cause of glomerulonephritis, is linked to 6q22–23. Nat. Genet. 2000, 26, 354–357. [Google Scholar] [CrossRef]
- Lyons, P.A.; Rayner, T.F.; Trivedi, S.; Holle, J.U.; Watts, R.A.; Jayne, D.R.; Baslund, B.; Brenchley, P.; Bruchfeld, A.; Chaudhry, A.N.; et al. Genetically distinct subsets within ANCA-associated vasculitis. N. Engl. J. Med. 2012, 367, 214–223. [Google Scholar] [CrossRef]
- Palmer, N.D.; Freedman, B.I. Diabetic nephropathy: FRMD3 in diabetic nephropathy-guilt by association. Nat. Rev. Nephrol. 2013, 9, 313–314. [Google Scholar] [CrossRef]
- Martini, S.; Nair, V.; Patel, S.R.; Eichinger, F.; Nelson, R.G.; Weil, E.J.; Pezzolesi, M.G.; Krolewski, A.S.; Randolph, A.; Keller, B.J.; et al. From single nucleotide polymorphism to transcriptional mechanism: A model for FRMD3 in diabetic nephropathy. Diabetes 2013, 62, 2605–2612. [Google Scholar] [CrossRef]
- Mooyaart, A.L.; Valk, E.J.; van Es, L.A.; Bruijn, J.A.; de Heer, E.; Freedman, B.I.; Dekkers, O.M.; Baelde, H.J. Genetic associations in diabetic nephropathy: A meta-analysis. Diabetologia 2011, 54, 544–553. [Google Scholar] [CrossRef]
- Wallace, C.; Rotival, M.; Cooper, J.D.; Rice, C.M.; Yang, J.H.; McNeill, M.; Smyth, D.J.; Niblett, D.; Cambien, F.; Tiret, L.; et al. Statistical colocalization of monocyte gene expression and genetic risk variants for type 1 diabetes. Hum. Mol. Genet. 2012, 21, 2815–2824. [Google Scholar] [CrossRef]
- Zeng, F.; Zhang, M.Z.; Singh, A.B.; Zent, R.; Harris, R.C. ErbB4 isoforms selectively regulate growth factor induced Madin-Darby canine kidney cell tubulogenesis. Mol. Biol. Cell 2007, 18, 4446–4456. [Google Scholar] [CrossRef]
- Veikkolainen, V.; Naillat, F.; Railo, A.; Chi, L.; Manninen, A.; Hohenstein, P.; Hastie, N.; Vainio, S.; Elenius, K. ErbB4 modulates tubular cell polarity and lumen diameter during kidney development. J. Am. Soc. Nephrol. 2012, 23, 112–122. [Google Scholar] [CrossRef]
- Sandholm, N.; McKnight, A.J.; Salem, R.M.; Brennan, E.P.; Forsblom, C.; Harjutsalo, V.; Makinen, V.P.; McKay, G.J.; Sadlier, D.M.; Williams, W.W.; et al. Chromosome 2q31.1 Associates with ESRD in Women with Type 1 Diabetes. J. Am. Soc. Nephrol. 2013, 24, 1537–1543. [Google Scholar] [CrossRef]
- Williams, W.W.; Salem, R.M.; McKnight, A.J.; Sandholm, N.; Forsblom, C.; Taylor, A.; Guiducci, C.; McAteer, J.B.; McKay, G.J.; Isakova, T.; et al. Association testing of previously reported variants in a large case-control meta-analysis of diabetic nephropathy. Diabetes 2012, 61, 2187–2194. [Google Scholar] [CrossRef]
- SUrroate markers for Micro- and Macro-vascular hard endpoints for Innovative diabetes Tools. Available online: www.imi-summit.eu/ (accessed on 31 October 2013).
- Nephromine. Available online: www.nephromine.org/ (accessed on 31 October 2013).
- The Kidney & Urinary Pathway Knowledge Base. Available online: www.kupkb.org/ (accessed on 31 October 2013).
- Jupp, S.; Klein, J.; Schanstra, J.; Stevens, R. Developing a kidney and urinary pathway knowledge base. J. Biomed. Semant. 2011, 2, S7. [Google Scholar]
- Human Kidney and Urine Proteome Project. Available online: www.hkupp.org/ (accessed on 31 October 2013).
- European Renal Genome Project. Available online: www.euregene.org/ (accessed on 31 October 2013).
- Fechete, R.; Heinzel, A.; Perco, P.; Monks, K.; Sollner, J.; Stelzer, G.; Eder, S.; Lancet, D.; Oberbauer, R.; Mayer, G.; et al. Mapping of molecular pathways, biomarkers and drug targets for diabetic nephropathy. Proteomics Clin. Appl. 2011, 5, 354–366. [Google Scholar] [CrossRef]
- Tranchevent, L.C.; Barriot, R.; Yu, S.; van Vooren, S.; van Loo, P.; Coessens, B.; de Moor, B.; Aerts, S.; Moreau, Y. ENDEAVOUR update: A web resource for gene prioritization in multiple species. Nucleic Acids Res. 2008, 36, W377–W384. [Google Scholar] [CrossRef]
- Su, A.I.; Wiltshire, T.; Batalov, S.; Lapp, H.; Ching, K.A.; Block, D.; Zhang, J.; Soden, R.; Hayakawa, M.; Kreiman, G.; et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl. Acad. Sci. USA 2004, 101, 6062–6067. [Google Scholar] [CrossRef]
- Pers, T.H.; Hansen, N.T.; Lage, K.; Koefoed, P.; Dworzynski, P.; Miller, M.L.; Flint, T.J.; Mellerup, E.; Dam, H.; Andreassen, O.A.; et al. Meta-analysis of heterogeneous data sources for genome-scale identification of risk genes in complex phenotypes. Genet. Epidemiol. 2011, 35, 318–332. [Google Scholar] [CrossRef]
- Pers, T.H.; Dworzynski, P.; Thomas, C.E.; Lage, K.; Brunak, S. MetaRanker 2.0: A web server for prioritization of genetic variation data. Nucleic Acids Res. 2013, 41, W104–W108. [Google Scholar] [CrossRef]
- Perkins, B.A.; Ficociello, L.H.; Roshan, B.; Warram, J.H.; Krolewski, A.S. In patients with type 1 diabetes and new-onset microalbuminuria the development of advanced chronic kidney disease may not require progression to proteinuria. Kidney Int. 2010, 77, 57–64. [Google Scholar] [CrossRef]
- Krolewski, A.S.; Niewczas, M.A.; Skupien, J.; Gohda, T.; Smiles, A.; Eckfeldt, J.H.; Doria, A.; Warram, J.H. Early progressive renal decline precedes the onset of microalbuminuria and its progression to macroalbuminuria. Diabetes Care 2013. [Epub ahead of print]. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Brennan, E.; McEvoy, C.; Sadlier, D.; Godson, C.; Martin, F. The Genetics of Diabetic Nephropathy. Genes 2013, 4, 596-619. https://doi.org/10.3390/genes4040596
Brennan E, McEvoy C, Sadlier D, Godson C, Martin F. The Genetics of Diabetic Nephropathy. Genes. 2013; 4(4):596-619. https://doi.org/10.3390/genes4040596
Chicago/Turabian StyleBrennan, Eoin, Caitríona McEvoy, Denise Sadlier, Catherine Godson, and Finian Martin. 2013. "The Genetics of Diabetic Nephropathy" Genes 4, no. 4: 596-619. https://doi.org/10.3390/genes4040596




