Over-Production, Crystallization, and Preliminary X-ray Crystallographic Analysis of a Coiled-Coil Region in Human Pericentrin
Abstract
:1. Introduction
2. Materials and Methods
2.1. Macromolecule Production
2.2. Circular Dichroism (CD) Studies for Secondary Structure Estimation
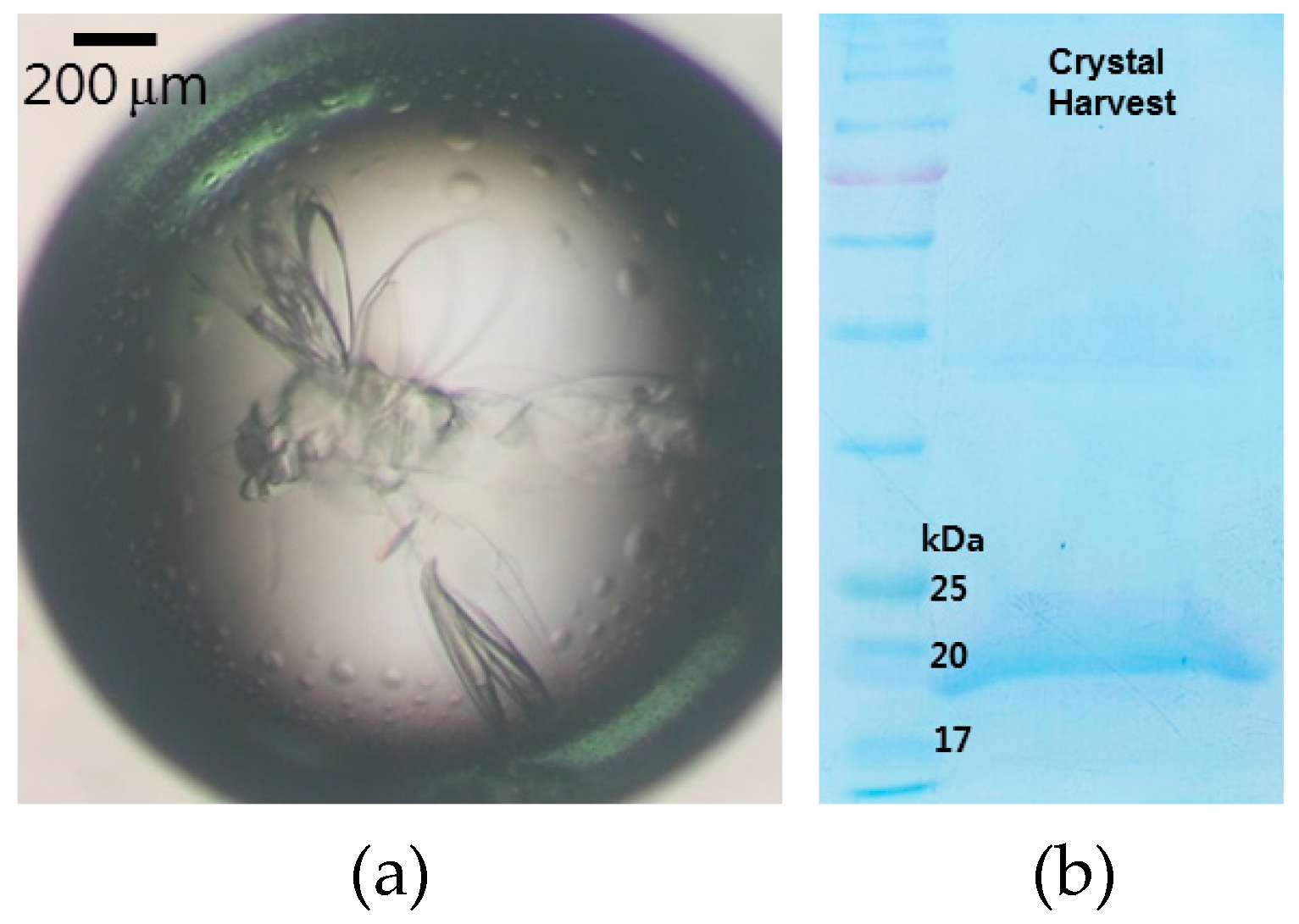
2.3. Crystallization
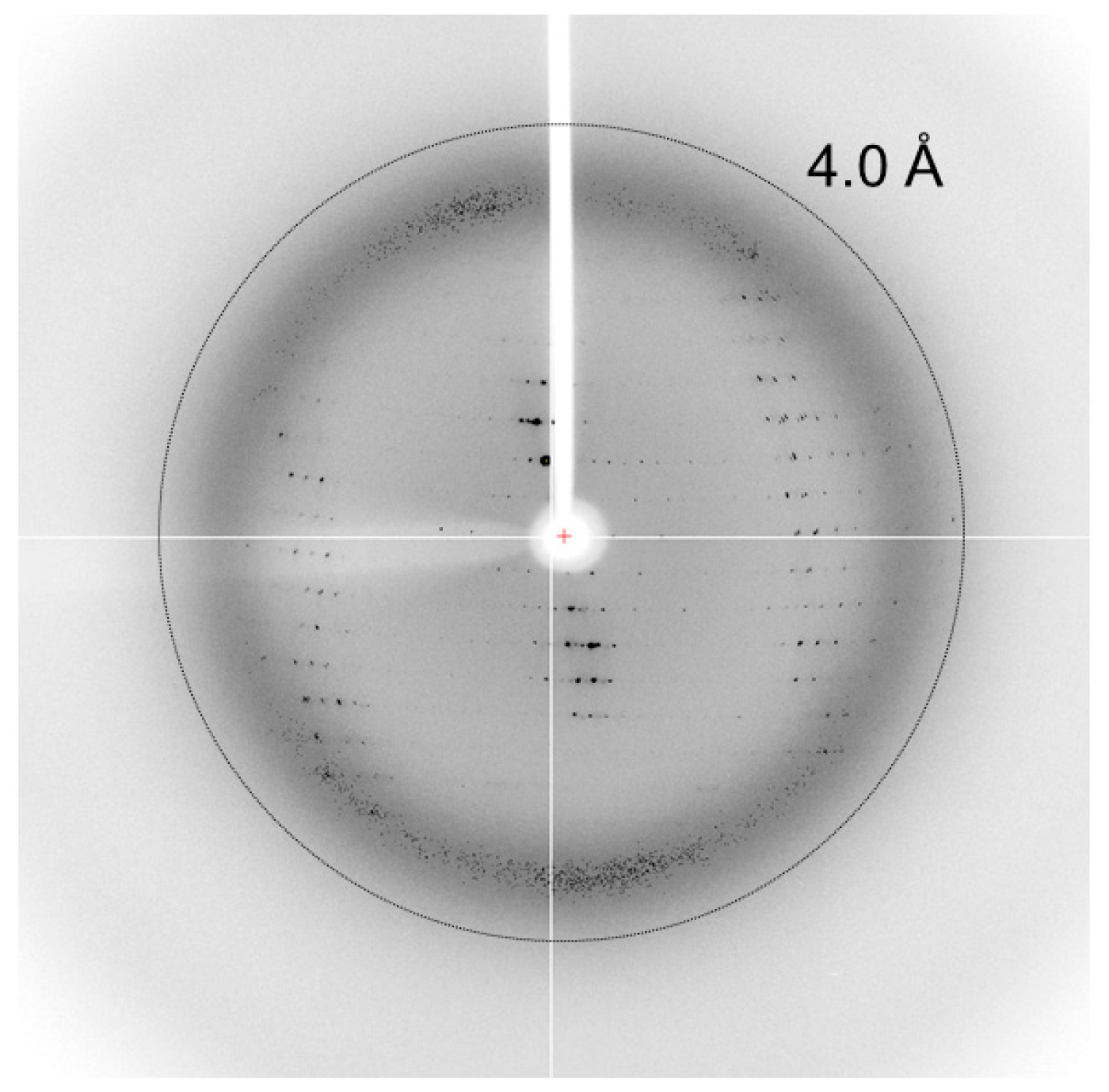
2.4. Data Collection and Processing
3. Results and Discussion
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Sluder, G. Two-way traffic: Centrosomes and the cell cycle. Nat. Rev. Mol. Cell Biol. 2005, 6, 743–748. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, M.; Yamagiwa, A.; Nishimura, T.; Mukai, H.; Ono, Y. Centrosomal proteins CG-NAP and kendrin provide microtubule nucleation sites by anchoring gamma-tubulin ring complex. Mol. Biol. Cell 2002, 13, 3235–3245. [Google Scholar] [CrossRef] [PubMed]
- Zimmerman, W.C.; Sillibourne, J.; Rosa, J.; Doxsey, S.J. Mitosis-specific anchoring of gamma tubulin complexes by pericentrin controls spindle organization and mitotic entry. Mol. Biol. Cell 2004, 15, 3642–3657. [Google Scholar] [CrossRef] [PubMed]
- Delaval, B.; Doxsey, S.J. Pericentrin in cellular function and disease. J. Cell Biol. 2010, 188, 181–190. [Google Scholar] [CrossRef] [PubMed]
- Doxsey, S.; Zimmerman, W.; Mikule, K. Centrosome control of the cell cycle. Trends Cell Biol. 2005, 15, 303–311. [Google Scholar] [CrossRef] [PubMed]
- Doxsey, S.J.; Stein, P.; Evans, L.; Calarco, P.D.; Kirschner, M. Pericentrin, a highly conserved centrosome protein involved in microtubule organization. Cell 1994, 76, 639–650. [Google Scholar] [CrossRef]
- Gillingham, A.K.; Munro, S. The PACT domain, a conserved centrosomal targeting motif in the coiled-coil proteins AKAP450 and pericentrin. EMBO Rep. 2000, 1, 524–529. [Google Scholar] [CrossRef] [PubMed]
- Rauch, A.; Thiel, C.T.; Schindler, D.; Wick, U.; Crow, Y.J.; Ekici, A.B.; van Essen, A.J.; Goecke, T.O.; Al-Gazali, L.; Chrzanowska, K.H.; et al. Mutations in the pericentrin (PCNT) gene cause primordial dwarfism. Science 2008, 319, 816–819. [Google Scholar] [CrossRef] [PubMed]
- Gill, S.C.; von Hippel, P.H. Calculation of protein extinction coefficients from amino acid sequence data. Anal. Biochem. 1989, 182, 319–326. [Google Scholar] [CrossRef]
- Perez-Iratxeta, C.; Andrade-Navarro, M.A. K2D2: Estimation of protein secondary structure from circular dichroism spectra. BMC Struct. Biol. 2008, 8, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Louis-Jeune, C.; Andrade-Navarro, M.A.; Perez-Iratxeta, C. Prediction of protein secondary structure from circular dichroism using theoretically derived spectra. Proteins 2012, 80, 374–381. [Google Scholar] [CrossRef] [PubMed]
- Otwinowski, Z.; Minor, W. Processing of X-ray Diffraction Data Collected in Oscillation Mode. Methods Enzymol. 1997, 276, 307–326. [Google Scholar] [PubMed]
- Lupas, A.; Van Dyke, M.; Stock, J. Predicting Coiled Coils from Protein Sequences. Science 1991, 252, 1162–1164. [Google Scholar] [CrossRef] [PubMed]
- Matthews, B.W. Solvent content of protein crystals. J. Mol. Biol. 1968, 33, 491–497. [Google Scholar] [CrossRef]
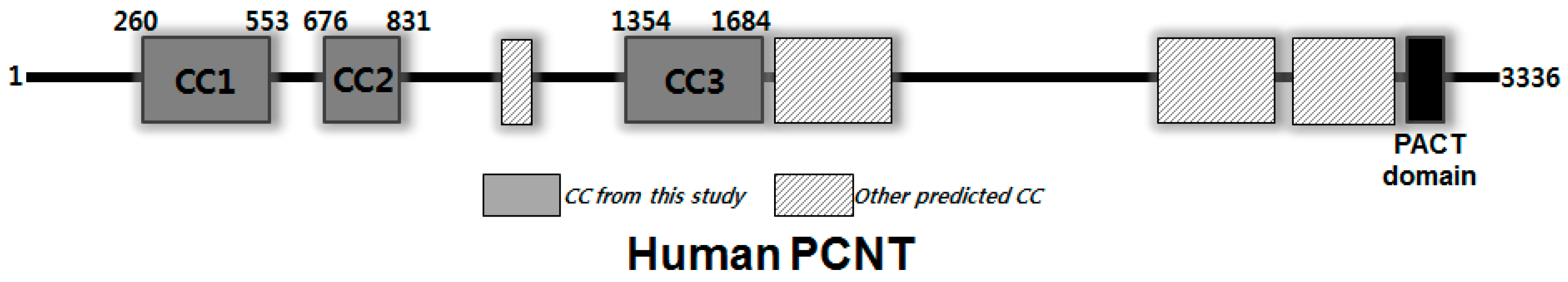
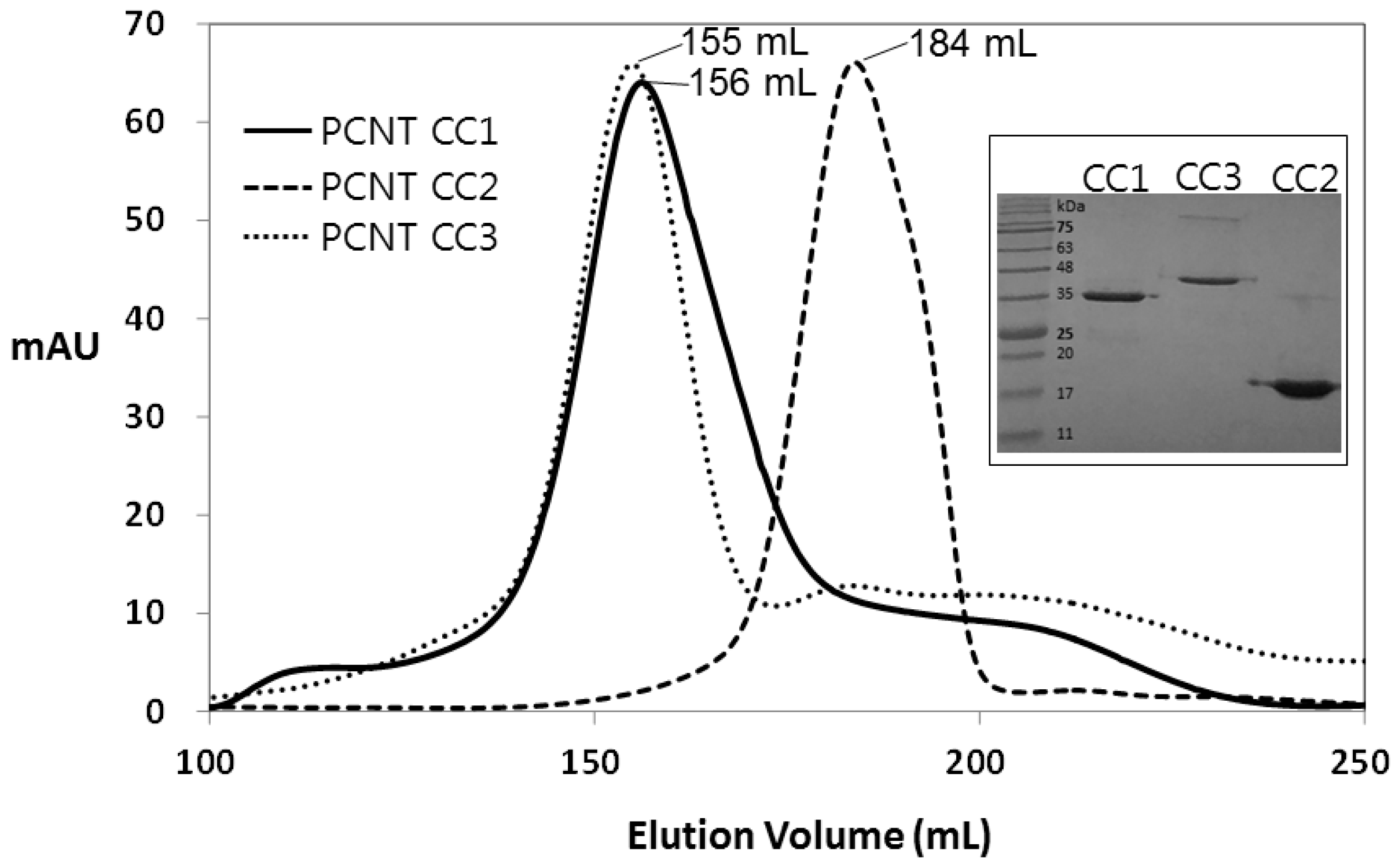
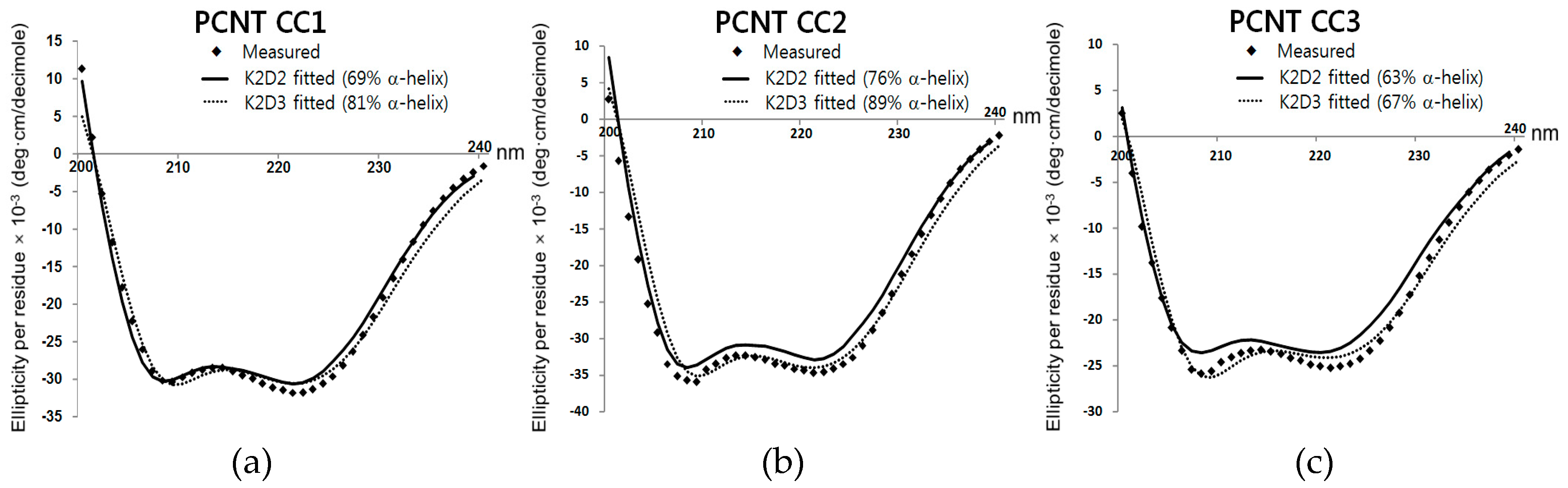


| Source Organism | Homo Sapiens |
|---|---|
| DNA source | Synthesized DNA |
| Cloning sites | NdeI and BamHI |
| Cloning vector | pET28a |
| Expression vector | pET28a |
| Expression host | E. coli BL21 (DE3) |
| Complete amino acid sequence of the construct produced 1 | MGSSHHHHHHSSGLVPRGSHM-676EHKVQ680 681LSLLQTELKEEIELLKIENRNLYGKLQHET710 711RLKDDLEKVKHNLIEDHQKELNNAKQKTEL740 741MKQEFQRKETDWKVMKEELQREAEEKLTLM770 771LLELREKAESEKQTIINKFELREAEMRQLQ800 801DQQAAQILDLERSLTEQQGRLQQLEQDLTSD831 |
| Peptide Sequence 1 | MH+ (Da) |
|---|---|
| GSHM-676EHKVQLSLLQTELKEEIELLK696 | 2932.59 |
| 679VQLSLLQTELKEEIELLKIENR700 | 2638.52 |
| 678KVQLSLLQTELKEEIELLK696 | 2254.33 |
| 683LLQTELKEEIELLKIENR700 | 2211.26 |
| 676EHKVQLSLLQTELKEEIELLK696 | 2520.44 |
| 682SLLQTELKEEIELLKIENR700 | 2298.30 |
| 681LSLLQTELKEEIELLKIENR700 | 2411.38 |
| 793EAEMRQLQDQQAAQILDLER812 | 2385.19 |
| 684LQTELKEEIELLKIENR700 | 2098.18 |
| 679VQLSLLQTELKEEIELLK696 | 2126.23 |
| 682SLLQTELKEEIELLK696 | 1786.02 |
| 685QTELKEEIELLKIENR700 | 1985.09 |
| 681LSLLQTELKEEIELLK696 | 1899.11 |
| 683LLQTELKEEIELLK696 | 1698.99 |
| 721HNLIEDHQKELNNAK735 | 1802.92 |
| 686TELKEEIELLKIENR700 | 1857.03 |
| 685QTELKEEIELLK696 | 1472.82 |
| 719VKHNLIEDHQKELNNAK735 | 2030.08 |
| 684LQTELKEEIELLK696 | 1585.91 |
| 778AESEKQTIINKFELR792 | 1805.98 |
| 798QLQDQQAAQILDLER812 | 1768.92 |
| 687ELKEEIELLKIENR700 | 1755.99 |
| 767LTLMLLELREKAESEK782 | 1903.06 |
| 688LKEEIELLKIENR700 | 1626.95 |
| 798QLQDQQAAQILDLERSLTEQQGR820 | 2668.37 |
| 757EELQREAEEKLTLMLLELR775 | 2343.26 |
| 762EAEEKLTLMLLELR775 | 1687.93 |
| 762EAEEKLTLMLLELREK777 | 1945.07 |
| Diffraction Source | Pohang Light Source (PLS 5C) (Pohang, Korea) |
|---|---|
| Wavelength (Å) | 0.9795 |
| Temperature (K) | 100 |
| Detector | ADSC Quantum 315r |
| Crystal–detector distance (mm) | 450 |
| Rotation range per image (°) | 1 |
| Total rotation range (°) | 180 |
| Exposure time per image (s) | 1 |
| Space group | C2 |
| a, b, c (Å) | 324.9, 35.7, 79.5 |
| α, β, γ (°) | 90.0, 101.6, 90.0 |
| Mosaicity (°) | 1.1 |
| Resolution range (Å) | 50.0–3.80 (3.87–3.80)1 |
| Total No. of reflections | 16,221 |
| No. of unique reflections | 9074 |
| Completeness (%) | 97.3 (89.7) |
| Redundancy | 1.7 (1.6) |
| 〈I/σ(I)〉 | 18.1 (3.9) |
| Rmerge | 0.463 (0.120) |
| Rp.i.m. | 0.081 (0.303) |
| CC1/2 | (0.918) |
| Overall B factor from Wilson plot (Å2) | 78.7 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, M.Y.; Park, J.K.; Sim, Y.; Kim, D.; Sim, J.Y.; Park, S. Over-Production, Crystallization, and Preliminary X-ray Crystallographic Analysis of a Coiled-Coil Region in Human Pericentrin. Crystals 2017, 7, 296. https://doi.org/10.3390/cryst7100296
Kim MY, Park JK, Sim Y, Kim D, Sim JY, Park S. Over-Production, Crystallization, and Preliminary X-ray Crystallographic Analysis of a Coiled-Coil Region in Human Pericentrin. Crystals. 2017; 7(10):296. https://doi.org/10.3390/cryst7100296
Chicago/Turabian StyleKim, Min Ye, Jeong Kuk Park, Yeowon Sim, Doheum Kim, Jeong Yeon Sim, and SangYoun Park. 2017. "Over-Production, Crystallization, and Preliminary X-ray Crystallographic Analysis of a Coiled-Coil Region in Human Pericentrin" Crystals 7, no. 10: 296. https://doi.org/10.3390/cryst7100296





