Differentiation and Distribution of Cordyline Viruses 1–4 in Hawaiian ti Plants (Cordyline fruticosa L.)
Abstract
:1. Introduction
2. Results and Discussion
2.1. Differentiation of CoVs
| Target | Primer | 5' to 3' Sequence (polarity) | Amplicon (bp) |
|---|---|---|---|
| CoV-1 | 1220 | CCTTCTTCGATAATAAGGTGGTG (+) | 1,075 |
| 1221 | AGGTGTAAGGAATTGGCATTGG (−) | ||
| CoV-2 | 1216 | GATTGTCAAATGGGAGGAATTGG (+) | 465 |
| 1217 | AGATACGGGAGAGATAAAGTTGG (−) | ||
| CoV-3 | 1214 | CATATGTCTGGTATTGGTACGG (+) | 375 |
| 1215 | CCAAGGAAAAGATCACCTTCTG (−) | ||
| CoV-4 | 1218 | TACGATCTAAAAAGATGGGTCGG (+) | 894 |
| 1219 | GGATCTTCTAGAAGAATGTGGAG (−) | ||
| C. fruticosa (rbcL) | 1212 | AGTATGGTCGTCCTTTTTTGGG (+) | 700 |
| 1213 | CATCTCCAAAGATTTCGGTTAGG (−) | ||
| Closterovirus HSP70h | 249 | GANVHRTCRAAMGTSCCTCCNCCRAARTC (−) | ~575 |
| 250 | GGARGTNGGNWWHGAMTTYGGNACNAC (+) | ||
| 1321 | GAYCTNAARMGNTGGGTNGGNG (+) | ~410 |
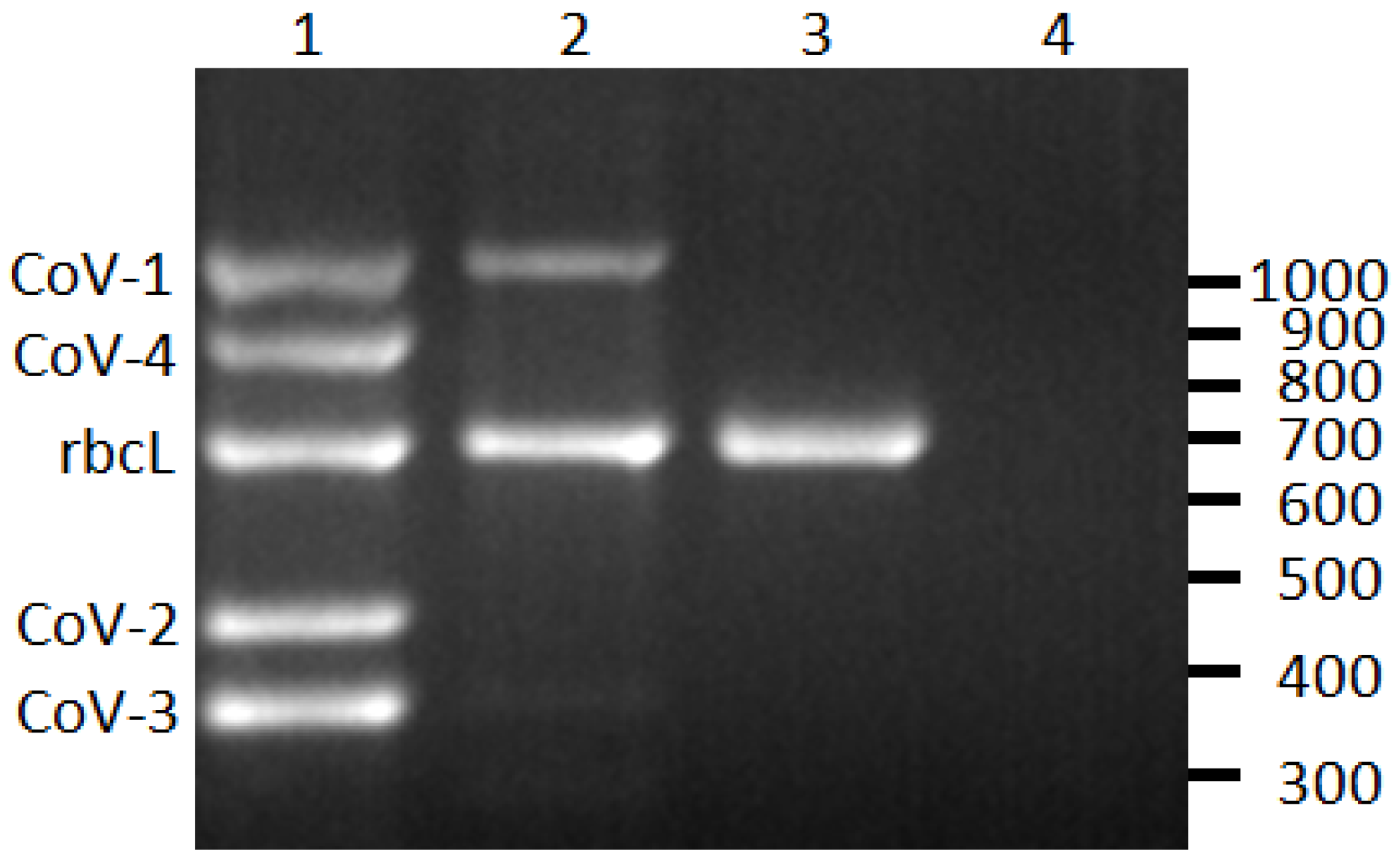
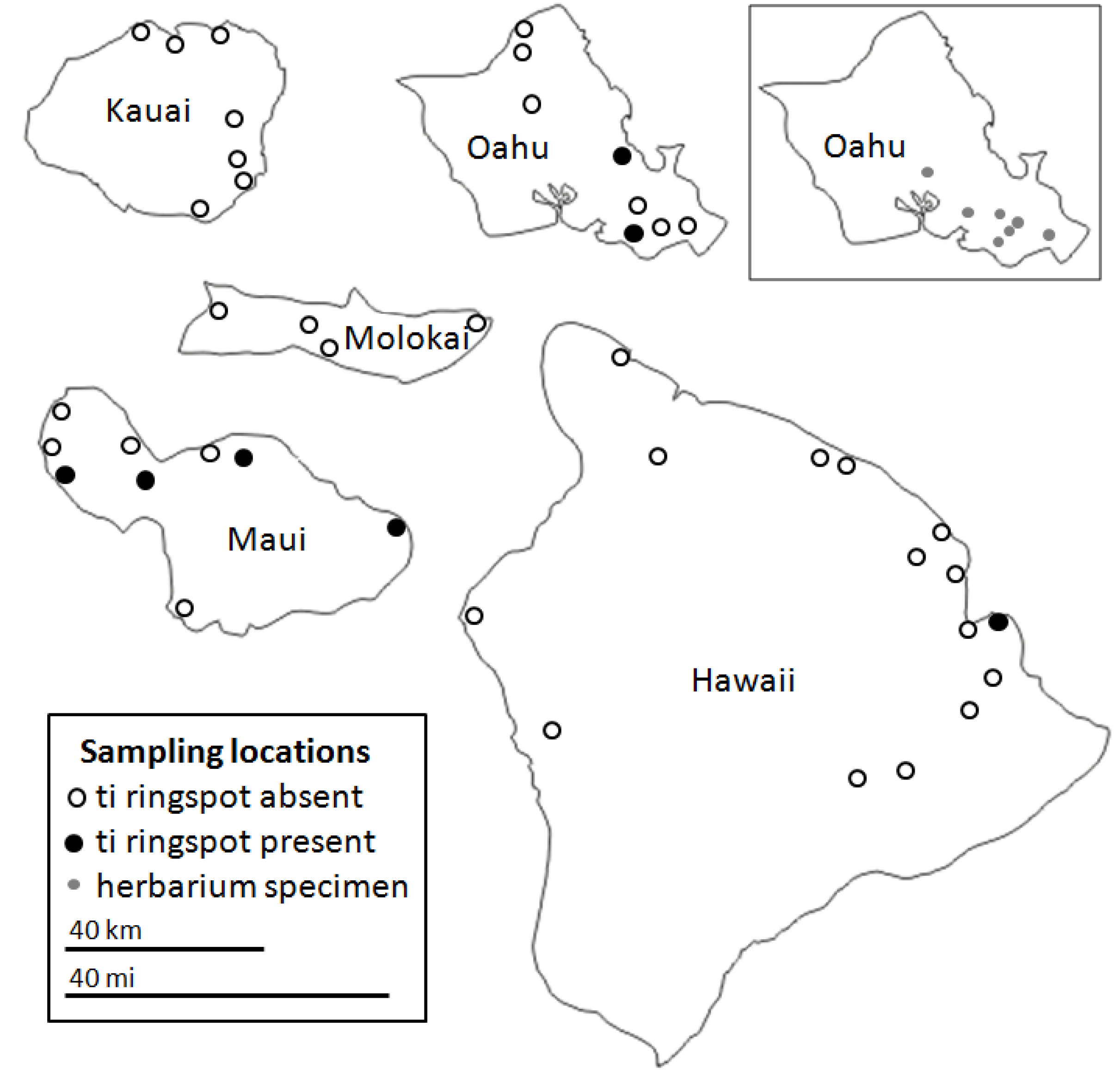
2.2. CoVs and ti Ringspot Disease
| Virus | Symptomatic | Asymptomatic | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Kauai | Oahu | Molokai | Maui | Hawaii | Total | Kauai | Oahu | Molokai | Maui | Hawaii | Total | |
| Negative | - | 1 | - | 3 | 1 | 5 | 4 | 6 | 8 | 8 | 20 | 46 |
| CoV-1 | - | - | - | 5 | - | 5 | - | 12 | 1 | - | 4 | 17 |
| CoV-2 | - | - | - | 1 | - | 1 | 3 | 1 | - | 5 | 1 | 10 |
| CoV-3 | - | - | - | - | - | - | 10 | - | - | 1 | 3 | 14 |
| CoV-4 | - | - | - | - | - | - | - | 1 | 1 | - | 1 | 3 |
| CoV-1 + CoV-2 | - | - | - | - | - | - | - | 1 | - | - | 3 | 4 |
| CoV-1 + CoV-3 | - | - | - | - | - | - | - | - | - | - | 1 | 1 |
| CoV-1 + CoV-4 | - | - | - | 3 | - | 3 | - | 1 | 1 | - | 1 | 3 |
| CoV-2 + CoV-3 | - | - | - | - | - | - | 3 | 1 | - | - | - | 4 |
| CoV-2 + CoV-4 | - | 1 | - | - | - | 1 | - | - | - | - | 3 | 3 |
| CoV-3 + CoV-4 | - | - | - | - | - | - | 1 | - | - | - | - | 1 |
| CoV-1 + CoV-2 + CoV-3 | - | - | - | - | - | - | - | - | - | - | 2 | 2 |
| CoV-1 + CoV-2 + CoV-4 | - | 1 | - | 1 | - | 2 | - | 2 | 2 | - | - | 4 |
| CoV-1 + CoV-3 + CoV-4 | - | 3 | - | - | - | 3 | - | - | - | - | - | - |
| CoV-2 + CoV-3 + CoV-4 | - | - | - | - | - | - | - | - | - | - | 1 | 1 |
| CoV-1 + CoV-2 + CoV-3 + CoV-4 | - | 2 | - | - | - | 2 | - | - | - | 1 | 1 | 2 |
| Total | - | 8 | - | 13 | 1 | 22 | 21 | 25 | 13 | 15 | 41 | 115 |
2.3. Spatial and Varietal Distribution of CoVs
| Variety | CoV-1 | CoV-2 | CoV-3 | CoV-4 |
|---|---|---|---|---|
| Apple Juno | + | − | − | + |
| Charles Nii #3 | − | − | − | − |
| Haole Girl | − | − | − | + |
| Harold Yamamoto #12 | − | − | + | + |
| Heather Spray | − | − | − | − |
| Kahili | + | − | − | + |
| Kalani Akaka | − | − | − | + |
| Kauai Beauty (miniature) | − | + | − | + |
| Lilian Olivera | − | − | − | + |
| Mauna Kea (miniature) | − | − | − | − |
| Tachibana | − | + | − | + |
| Total | 2/11 (18%) | 2/11 (18%) | 1/11 (9%) | 8/11 (73%) |
2.4. Herbarium Specimens
| Specimen ID | Date Collected | Tissue (weight) | Location | rbcL | CoV-1 | CoV-2 | CoV-3 | CoV-4 |
|---|---|---|---|---|---|---|---|---|
| BISH 697966 | Jan 13, 2003 | leaf petiole (15 mg) | Lyon Arboretum | + | − | − | + | + |
| BISH 504184 | Jan 19, 1985 | flower, leaf (9 mg) | Manoa Falls | + | − | − | − | − |
| BISH 504185 | Jan 19, 1985 | flower (7 mg) | Manoa Falls | − | − | − | − | − |
| BISH 76555 | Dec 22, 1970 | flower (5 mg) | Lyon Arboretum | − | − | − | − | − |
| BISH 487514 | Apr 9, 1962 | flower, leaf (17 mg) | University of Hawaii Manoa Campus | − | − | − | − | − |
| BISH 06727 | Dec 10, 1945 | flower, pedicel (27 mg) | University of Hawaii Manoa Campus | − | − | − | − | − |
| BISH 446936 | Mar 22, 1936 | flower (11 mg) | Kipapa Gulch | − | − | − | − | − |
| BISH 121224 | Jan 12, 1920 | flower (5 mg) | Wailupe Valley | − | − | − | − | − |
| BISH 92012 | Dec 29, 1909 | flower (8 mg) | Hillebrands Glen (Nuuanu Valley) | − | − | − | − | − |
| BISH 121219 | Dec 13, 1903 | flower (6 mg) | Moanalua Valley | − | − | − | − | − |
2.5. Degenerate Primer RT-PCR
3. Experimental
3.1. Differentiation of CoVs
3.2. Geographical and Varietal Distribution
3.3. Herbarium Specimens
3.4. Degenerate Primer RT-PCR
4. Conclusions
Acknowledgments
Conflict of Interest
References and Notes
- Karasev, A.V. Genetic diversity and evolution of closteroviruses. Annu. Rev. Phytopathol. 2000, 38, 293–324. [Google Scholar] [CrossRef]
- Martelli, G.P.; Agranovsky, A.A.; Bar-Joseph, M.; Boscia, D.; Candresse, T; Coutts, R.H.A.; Dolja, V.V.; Falk, B.W.; Gonsalves, D.; Jelkmann, W.; et al. The family Closteroviridae revised. Virol.2002 2002, 147, 2039–2044. [Google Scholar]
- Martelli, G.P.; Abou Ghanem-Sabanadzovic, N.; Agranovsky, A.A.; Al Rwahnih, M.; Dolja, V.V.; Dovas, C.I.; Fuchs, M.; Gugerli, P.; Hu, J.S.; Jelkmann, W.; et al. Taxonomic revision of the family Closteroviridae with special reference to the grapevine leafroll-associated members of the genus Ampelovirus and the putative species unassigned to the family. J. Plant Pathol. 2012, 94, 7–19. [Google Scholar]
- Al Rwahnih, M.; Dolja, V.V.; Daubert, S.; Koonin, E.V.; Rowhani, A. Genomic and biological analysis of Grapevine leafroll-associated virus 7 reveals a possible new genus within the family Closteroviridae. Virus Res. 2012, 163, 302–309. [Google Scholar] [CrossRef]
- Melzer, M.J.; Sugano, J.S.; Uchida, J.Y.; Borth, W.B.; Kawate, M.K.; Hu, J.S. Molecular characterization of closteroviruses infecting Cordyline fruticosa L. in Hawaii. Front. Microbiol. 2013, 4, e39. [Google Scholar]
- Hinkle, A.E. Population structure of Pacific Cordyline fruticosa (Laxmanniaceae) with implications for human settlement of Polynesia. Am. J. Bot. 2007, 94, 828–839. [Google Scholar] [CrossRef]
- Melzer, M.J.; Sether, D.M.; Borth, W.B.; Mersino, E.F.; Hu, J.S. An assemblage of closteroviruses infects Hawaiian ti (Cordyline fruticosa L.). Virus Genes 2011, 42, 254–260. [Google Scholar] [CrossRef]
- Gugerli, F.; Parducci, L.; Petit, R.J. Ancient plant DNA: Review and prospects. New Phytol. 2005, 166, 409–418. [Google Scholar] [CrossRef]
- Taylor, J.W.; Swann, E.C. DNA from herbarium specimens. In Ancient DNA; Hermann, B., Hummel, S., Eds.; Springer: New York, NY, USA, 1994; pp. 166–181. [Google Scholar]
- Malmstrom, C.M.; Shu, R.; Linton, E.W.; Newton, L.A.; Cook, M.A. Barley yellow dwarf viruses (BYDVs) preserved in herbarium specimens illuminate historical disease ecology of invasive and native grasses. J. Ecol. 2007, 95, 1153–1166. [Google Scholar] [CrossRef]
- Guy, P.L. Ancient RNA? RT-PCR of 50-year-old RNA identifies peach latent mosaic viroid. Arch. Virol. 2013, 158, 691–694. [Google Scholar] [CrossRef]
- Sether, D.M.; Melzer, M.J.; Busto, J.; Hu, J.S. Diversity and mealybug transmissibility of ampeloviruses in pineapple. Plant Dis. 2005, 89, 450–456. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Melzer, M.; Ayin, C.; Sugano, J.; Uchida, J.; Kawate, M.; Borth, W.; Hu, J. Differentiation and Distribution of Cordyline Viruses 1–4 in Hawaiian ti Plants (Cordyline fruticosa L.). Viruses 2013, 5, 1655-1663. https://doi.org/10.3390/v5071655
Melzer M, Ayin C, Sugano J, Uchida J, Kawate M, Borth W, Hu J. Differentiation and Distribution of Cordyline Viruses 1–4 in Hawaiian ti Plants (Cordyline fruticosa L.). Viruses. 2013; 5(7):1655-1663. https://doi.org/10.3390/v5071655
Chicago/Turabian StyleMelzer, Michael, Caleb Ayin, Jari Sugano, Janice Uchida, Michael Kawate, Wayne Borth, and John Hu. 2013. "Differentiation and Distribution of Cordyline Viruses 1–4 in Hawaiian ti Plants (Cordyline fruticosa L.)" Viruses 5, no. 7: 1655-1663. https://doi.org/10.3390/v5071655




