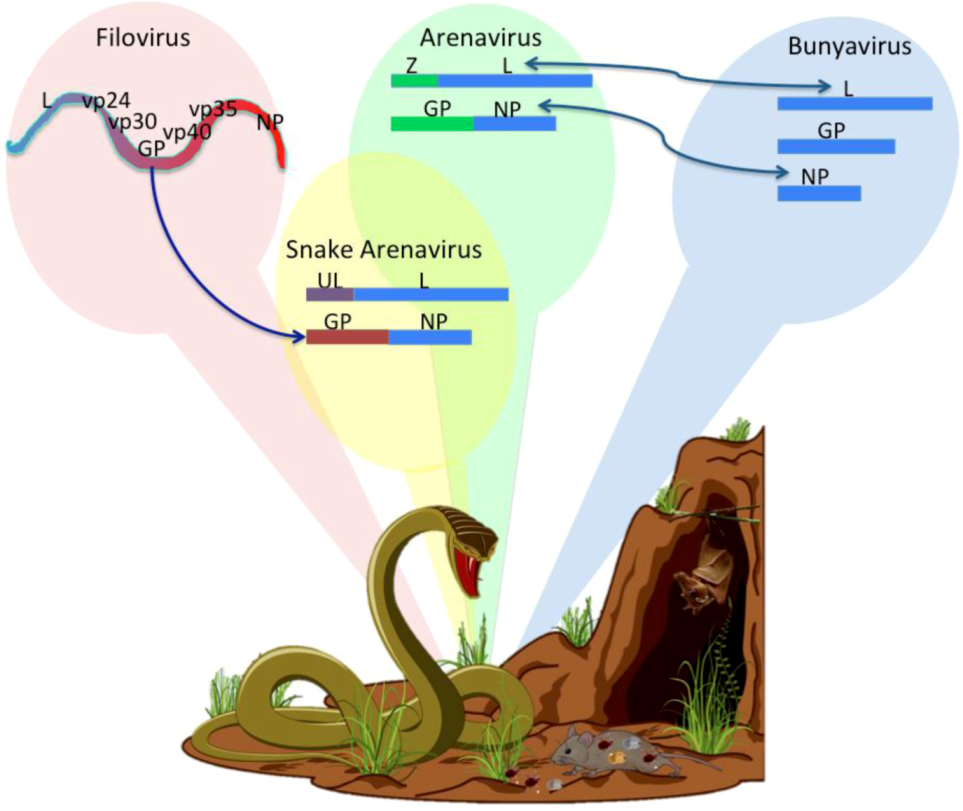
Arenavirus Variations Due to Host-Specific Adaptation
Abstract
:1. Background
1.3. Antigenic Characteristics
1.4. Virus-Host Coevolution
3. Conclusion
Conflict of Interest
| Virus Species | Origen of name | Year | Distribution | Host | Associated Disease |
|---|---|---|---|---|---|
| Old World or LASV-LCMV serocomplex | |||||
| Lymphocytic choriomeningitis virus (LCMV) | Disease | 1933 [162] | Worldwide | House mouse (Mus musculus) and (Mus domesticus) and Syrian hamster (Mesocricetus auratus) | Flu like symptoms Meningitis |
| Encephalitis | |||||
| Congenital abnormalities/Abortion | |||||
| Multisystem organ failure in transplanted patients | |||||
| Lassa (LASV) | Town, Nigeria | 1969 [163] | West Africa | Multimammate mouse (Mastomys genus) | Hemorrhagic fever |
| Mopeia (MOPV) | Town, Mozambique | 1977 [164] | Southern Africa | Multimammate mouse (Mastomis natalensis) | Not associated with human disease |
| Merino Walk | Farm, South Africa | 1985 [55] | Eastern Cape | Karoo rat (Myotomys unisulcatus) | Unknown pathogenicity for humans |
| (MWV) | South Africa | ||||
| Mopeia/Lassa | M/L reassortant clone 29 | 1992 [72] | Russia | Laboratory virus. Passaged in Vero E6 and BHK cells | Vaccine candidate against LHF |
| reassortant | |||||
| (ML29) | |||||
| Morogoro | City, Tanzania | 2009 [165] | East Africa | Multimammate mouse (Mastomys natalensis) | Unknown pathogenicity for humans |
| Tanzania | |||||
| Mobala (MOBV) | Region, DR of Congo | 1983 [166] | Central African Republic | Soft-furred rat (Praomys sp.) | Not associated with human disease |
| IPPY (IPPYV) | Town, Central African Republic | 1985 [167] | Central African | Nile grass rat (Arvicanthis sp.) | Not associated with human disease |
| Lujo (LUJV) | Lusaka, Zambia Johannesburg, South Africa | 2009 [54] | Southern Africa | Unknown | Hemorrhagic fever |
| Luna (LUNV) | Lusaka-Namwala, Zambia | 2009 [168] | Southern Africa Zambia | Multimammate mouse (Mastomys natalensis) | Unknown pathogenicity for humans |
| New World arenavirus or Tacaribe serocomplex South American group | |||||
| Group A | |||||
| Pichindé (PICV) | Valley, Colombia | 1965 [169] | South America | Tome’s rice rat (Oryzomys albigularis) | Not associated with human disease |
| Colombia | |||||
| Paraná (PARV) | River, Paraguay, Brazil, Argentina | 1965 [170] | South America | Paraguayan rice rat (Oryzomys buccinatus, Oryzomys angouya) | Not associated with human disease |
| Paraguay | |||||
| Flexal (FLEV) | Brazil | 1975 [171] | South America | Tome’s rice rat (Oryzomys albigularis, Nephelomys albigularis), Paraguayan rice rat (Oryzomys angouya, Oryzomys buccinatus), | Febril illness Associated with nonfatal laboratory-acquired infection |
| Brazil | |||||
| Pirital (PIRV) | Community, Venezuela | 1995 [172] | South America | Alston’s cotton rat (Sigmodon alstoni) | Not associated with human disease |
| Venezuela | |||||
| Allpaahuayo (ALLV) | National reserve, Peru | 1997 [173] | South America | Arboreal rice rats (Oecomys bicolor and Oecomys paricola) | Unknown pathogenicity for humans |
| Peru | |||||
| Group B | |||||
| Tacaribe (TCRV) | Beach, Trinidad | 1956 [174] | Caribbean Sea | fruit-eating bat (Artibeus sp.) | Associated only with single, nonfatal, laboratory-acquired infection. |
| Trinidad | |||||
| Junin (JUNV) | Town, Argentina | 1958 [175] | South America | Corn mouse, drylands vesper mouse (Calomys masculinus), grass field mouse (Akodon azarae), dark field mouse (Bolomys obscurus) | Hemorrhagic fever |
| Argentina | |||||
| Candid#1 | Argentina | 1985 [176] | South America | Passaged guinea pigs (GP2), mouse | Vaccine against Argentinian hemorrhagic fever |
| Argentina | (MB44), followed by clonal selection | ||||
| in fetal rhesus monkey lung cells (FRhL19). | |||||
| Machupo (MACV) | River, Bolivia | 1962 [28] | South America | large vesper mouse (Calomys callosus) | Hemorrhagic fever |
| Bolivia | |||||
| Amapari (AMAV) | Amapá region, Brazil | 1964 [177] | South America | rice rat (Oryzomys goeldii), bristly mouse (Neacomys guianae) | Not associated with human disease |
| Brazil | |||||
| Cupixi (CPXV) | Town, Brazil | 1970 [77] | South America | Large-headed Rice Rat (Oryzomys capito) | Not associated with human disease |
| Brazil | |||||
| Guanarito (GTOV) | Region, Venezuela | 1989 [178] | South America | Cane mouse (Zygodontomys brevicauda) | Hemorrhagic fever |
| Venezuela | |||||
| Sabiá (SABV) | Town, Brazil | 1990 [179] | South America | Unknown (Suspected rodent) | Hemorrhagic fever, Hemorrhagic fever associated with nonfatal laboratory-acquired infection |
| Brazil | |||||
| Chapare (CHPV) | Town, Bolivia | 2005 [180] | South America | Unknown | Hemorrhagic fever |
| Bolivia | |||||
| Group C | |||||
| Latino (LATV) | Bolivia | 1965 [31] | South America | Large vesper mouse (Calomys callosus) | Not associated with human disease |
| Bolivia, Brazil | |||||
| Oliveros (OLVV) | Town, Argentina | 1990 [181] | South America | Dark bolo mouse (Bolomys obscurus) | Not associated with human disease |
| Argentina | |||||
| Pampa virus (PAMV) | Region, Argentina | 1997 [182] | South America | Dark bolo mouse (Bolomys sp.) | Not associated with human disease |
| Argentina | |||||
| North American group | |||||
| Group A | |||||
| Tamiami (TAMV)* | Everglades, USA | 1964 [183] | North America Florida | Hispid cotton rat (Sigmodon hispidus) | Not associated with human disease |
| Whitewater Arroyo* (WWAV) | Whitewater Creek | 1993 [184] | North America New Mexico | White-throated wood rat (Neotoma albigula) | Febrile infection |
| Respiratory distress syndrome | |||||
| Catarina (CTNV) | Town, USA | 1999 [185] | North America Texas | Southern Plains Woodrat (Neotoma micropus) | Unknown Pathogenicity for humans |
| Skinner Tank virus | Reservoir, USA | 2002 [186] | North America Arizona | Mexican woodrat (Neotoma Mexicana) | Unknown pathogenicity for humans |
| (SKTV) | |||||
| Big Brushy Tank | USA | 2008 [187] | North America Arizona | White-throated woodrat (Neotoma albigula) | Unknown pathogenicity for humans |
| (BBTV) | |||||
| Tonto Creek | Creek, USA | 2008 [187] | North America Arizona | White-throated woodrat (Neotoma albigula) | Unknown pathogenicity for humans |
| (TTCV) | |||||
| Bear Canyon (BCNV)* | Trailhead, USA | 2002 [188] | North America California | California mouse (Peromyscus californico), Large-eared woodrat (Neotoma macrotis) | Unknown pathogenicity for humans |
References
- Auperin, D.D.; Romanowski, V.; Galinski, M.; Bishop, D.H. Sequencing studies of pichinde arenavirus S RNA indicate a novel coding strategy, an ambisense viral S RNA. J. Virol. 1984, 52, 897–904. [Google Scholar]
- Pinschewer, D.D.; Perez, M.; de la Torre, J.C. Role of the virus nucleoprotein in the regulation of lymphocytic choriomeningitis virus transcription and RNA replication. J. Virol. 2003, 77, 3882–3887. [Google Scholar] [CrossRef]
- Eichler, R.; Strecker, T.; Kolesnikova, L.; ter Meulen, J.; Weissenhorn, W.; Becker, S.; Klenk, H.D.; Garten, W.; Lenz, O. Characterization of the lassa virus matrix protein Z: Electron microscopic study of virus-like particles and interaction with the nucleoprotein (NP). Virus Res. 2004, 100, 249–255. [Google Scholar] [CrossRef]
- Casabona, J.C.; Levingston Macleod, J.M.; Loureiro, M.E.; Gomez, G.A.; Lopez, N. The ring domain and the l79 residue of Z protein are involved in both the rescue of nucleocapsids and the incorporation of glycoproteins into infectious chimeric arenavirus-like particles. J. Virol. 2009, 83, 7029–7039. [Google Scholar] [CrossRef]
- Shtanko, O.; Imai, M.; Goto, H.; Lukashevich, I.S.; Neumann, G.; Watanabe, T.; Kawaoka, Y. A role for the C terminus of mopeia virus nucleoprotein in its incorporation into Z protein-induced virus-like particles. J. Virol. 2010, 84, 5415–5422. [Google Scholar] [CrossRef]
- Martinez-Sobrido, L.; Zuniga, E.I.; Rosario, D.; Garcia-Sastre, A.; de la Torre, J.C. Inhibition of the type i interferon response by the nucleoprotein of the prototypic arenavirus lymphocytic choriomeningitis virus. J. Virol. 2006, 80, 9192–9199. [Google Scholar]
- Martinez-Sobrido, L.; Emonet, S.; Giannakas, P.; Cubitt, B.; Garcia-Sastre, A.; de la Torre, J.C. Identification of amino acid residues critical for the anti-interferon activity of the nucleoprotein of the prototypic arenavirus lymphocytic choriomeningitis virus. J. Virol. 2009, 83, 11330–11340. [Google Scholar]
- Hastie, K.M.; Kimberlin, C.R.; Zandonatti, M.A.; MacRae, I.J.; Saphire, E.O. Structure of the lassa virus nucleoprotein reveals a dsRNA-specific 3' to 5' exonuclease activity essential for immune suppression. P. Natl. Acad. Sci. U.S.A. 2011, 108, 2396–2401. [Google Scholar]
- Qi, X.; Lan, S.; Wang, W.; Schelde, L.M.; Dong, H.; Wallat, G.D.; Ly, H.; Liang, Y.; Dong, C. Cap binding and immune evasion revealed by lassa nucleoprotein structure. Nature 2010, 468, 779–783. [Google Scholar]
- Salvato, M.S.; Schweighofer, K.J.; Burns, J.; Shimomaye, E.M. Biochemical and immunological evidence that the 11 kda zinc-binding protein of lymphocytic choriomeningitis virus is a structural component of the virus. Virus Res. 1992, 22, 185–198. [Google Scholar] [CrossRef]
- Borden, K.L.; Campbelldwyer, E.J.; Carlile, G.W.; Djavani, M.; Salvato, M.S. Two ring finger proteins, the oncoprotein PML and the arenavirus Z protein, colocalize with the nuclear fraction of the ribosomal P proteins. J. Virol. 1998, 72, 3819–3826. [Google Scholar]
- Campbell Dwyer, E.J.; Lai, H.; MacDonald, R.C.; Salvato, M.S.; Borden, K.L. The lymphocytic choriomeningitis virus ring protein Z associates with eukaryotic initiation factor 4e and selectively represses translation in a ring-dependent manner. J. Virol. 2000, 74, 3293–3300. [Google Scholar] [CrossRef]
- Kentsis, A.; Dwyer, E.C.; Perez, J.M.; Sharma, M.; Chen, A.; Pan, Z.Q.; Borden, K.L. The ring domains of the promyelocytic leukemia protein PML and the arenaviral protein Z repress translation by directly inhibiting translation initiation factor eIf4e. J. Mol. Biol. 2001, 312, 609–623. [Google Scholar] [CrossRef]
- Lukashevich, I.S.; Salvato, M.S. Lassa virus genome. Curr. Genomics 2006, 7, 351–379. [Google Scholar] [CrossRef]
- Lee, K.J.; Novella, I.S.; Teng, M.N.; Oldstone, M.B.; de La Torre, J.C. Np and l proteins of lymphocytic choriomeningitis virus (LCMV) are sufficient for efficient transcription and replication of LCMV genomic rna analogs. J. Virol. 2000, 74, 3470–3477. [Google Scholar] [CrossRef]
- Lopez, N.; Jacamo, R.; Franze-Fernandez, M.T. Transcription and rna replication of tacaribe virus genome and antigenome analogs require N and L proteins: Z protein is an inhibitor of these processes. J. Virol. 2001, 75, 12241–12251. [Google Scholar] [CrossRef]
- Salvato, M.; Shimomaye, E.; Oldstone, M.B. The primary structure of the lymphocytic choriomeningitis virus L gene encodes a putative rna polymerase. Virology 1989, 169, 377–384. [Google Scholar] [CrossRef]
- Salvato, M.S.; Shimomaye, E.M. The completed sequence of lymphocytic choriomeningitis virus reveals a unique RNA structure and a gene for a zinc finger protein. Virology 1989, 173, 1–10. [Google Scholar] [CrossRef]
- Lenz, O.; ter Meulen, J.; Feldmann, H.; Klenk, H.D.; Garten, W. Identification of a novel consensus sequence at the cleavage site of the Lassa virus glycoprotein. J. Virol. 2000, 74, 11418–11421. [Google Scholar] [CrossRef]
- York, J.; Nunberg, J.H. Intersubunit interactions modulate ph-induced activation of membrane fusion by the junin virus envelope glycoprotein gpc. J. Virol. 2009, 83, 4121–4126. [Google Scholar] [CrossRef]
- Pfau, C.J. Biochemical and biophysical properties of the arenaviruses. Prog. Med. Virol. 1974, 18, 64–80. [Google Scholar]
- Salvato, M.S.; Clegg, J.C.S.; Buchmeier, M.J.; Charrel, R.N.; Gonzalez, J.P.; Lukashevich, I.S.; Peters, C.J.; Romanowski, V. Arenaviridae. In Virus Taxonomy, Ninth Report of the International Committee on Taxonomy of Viruses; King, A.M.Q., Adams, M.J., Carstens, E.B., Lefkowitz, E.J., Eds.; Elsevier: San Diego, CA, USA, 2012; pp. 715–723. [Google Scholar]
- Weissenbacher, M.C.; Coto, C.E.; Calello, M.A.; Rondinone, S.N.; Damonte, E.B.; Frigerio, M.J. Cross-protection in nonhuman primates against argentine hemorrhagic fever. Infect. Immun. 1982, 35, 425–430. [Google Scholar]
- Gonzalez, J.D.; Duplantier, J.M. The Arenaviruses and Rodent Co-Evolution Process : A Global View of a Theory. In Factors in the Emergence and Control of Rodent-Borne Viral Diseases (Hantaviral and Arenaviral Diseases); Saluzzo, J.F., Dodet, B., Eds.; Elsevier: Paris, 1999; pp. 39–42. [Google Scholar]
- Charrel, R.N.; Lemasson, J.J.; Garbutt, M.; Khelifa, R.; De Micco, P.; Feldmann, H.; de Lamballerie, X. New insights into the evolutionary relationships between arenaviruses provided by comparative analysis of small and large segment sequences. Virology 2003, 317, 191–196. [Google Scholar] [CrossRef]
- Jones, M.S., 2nd; Harrach, B.; Ganac, R.D.; Gozum, M.M.; Dela Cruz, W.P.; Riedel, B.; Pan, C.; Delwart, E.L.; Schnurr, D.P. New adenovirus species found in a patient presenting with gastroenteritis. J. Virol. 2007, 81, 5978–5984. [Google Scholar] [CrossRef]
- Mackenzie, R.B.; Webb, P.A.; Johnson, K.M. Detection of complement-fixing antibody after bolivian hemorrhagic fever, employing machupo, junin and tacaribe virus antigens. Am. J. Trop. Med. Hyge. 1965, 14, 1079–1084. [Google Scholar]
- Johnson, K.M.; Wiebenga, N.H.; Mackenzie, R.B.; Kuns, M.L.; Tauraso, N.M.; Shelokov, A.; Webb, P.A.; Justines, G.; Beye, H.K. Virus isolations from human cases of hemorrhagic fever in Bolivia. P. Soc. Exp. Biol. Med. 1965, 118, 113–118. [Google Scholar]
- Webb, P.A.; Johnson, K.M.; Mackenzie, R.B. The measurement of specific antibodies in bolivian hemorrhagic fever by neutralization of virus plaques. P. Soc. Exp. Biol. Med. 1969, 130, 1013–1019. [Google Scholar]
- Murphy, F.A.; Webb, P.A.; Johnson, K.M.; Whitfield, S.G. Morphological comparison of machupo with lymphocytic choriomeningitis virus: Basis for a new taxonomic group. J. Virol. 1969, 4, 535–541. [Google Scholar]
- Rowe, W.P.; Murphy, F.A.; Bergold, G.H.; Casals, J.; Hotchin, J.; Johnson, K.M.; Lehmann-Grube, F.; Mims, C.A.; Traub, E.; Webb, P.A. Arenoviruses: Proposed name for a newly defined virus group. J. Virol. 1970, 5, 651–652. [Google Scholar]
- Casals, J.; Buckley, S.M.; Cedeno, R. Antigenic properties of the arenaviruses. B. World Health Organ. 1975, 52, 421–427. [Google Scholar]
- Buchmeier, M.J.; Parekh, B.S. Protein structure and expression among arenaviruses. Curr. Top. Microbiol. Immunol. 1987, 133, 41–57. [Google Scholar] [CrossRef]
- Ruo, S.L.; Mitchell, S.W.; Kiley, M.P.; Roumillat, L.F.; Fisher-Hoch, S.P.; McCormick, J.B. Antigenic relatedness between arenaviruses defined at the epitope level by monoclonal antibodies. J. Gen. Virol. 1991, 72, 549–555. [Google Scholar] [CrossRef]
- Sanchez, A.; Pifat, D.Y.; Kenyon, R.H.; Peters, C.J.; McCormick, J.B.; Kiley, M.P. Junin virus monoclonal antibodies: Characterization and cross-reactivity with other arenaviruses. J. Gen. Virol. 1989, 70, 1125–1132. [Google Scholar] [CrossRef]
- Howard, C. Antigenic Diversity among the Arenaviruses. In The Arenaviridae; Salvato, M, Ed.; Plenum Press: New York, New York, USA, 1993; pp. 37–49. [Google Scholar]
- Weissenbacher, M.C.; de Guerrero, L.B.; Boxaca, M.C. Experimental biology and pathogenesis of junin virus infection in animals and man. B. World Health Organ. 1975, 52, 507–515. [Google Scholar]
- Jahrling, P.B.; Trotter, R.W.; Barrera Oro, J.G.; et al. Cross-Protection against Machupo Virus with Candid #1 Junin Virus Vaccine: Iii, Post-Challenge Clinical Findings. In Proceedings of the Second International Conference on the Impact of Viral Diseases on the Development of Latin American Countries and the Caribbean Region; Kurstak, E., Ed.; Mar del Plata, Argentina, 1988. [Google Scholar]
- Lukashevich, I.S.; Carrion, R., Jr.; Salvato, M.S.; Mansfield, K.; Brasky, K.; Zapata, J.; Cairo, C.; Goicochea, M.; Hoosien, G.E.; Ticer, A.; et al. Safety, immunogenicity, and efficacy of the ML29 reassortant vaccine for lassa fever in small non-human primates. Vaccine 2008, 26, 5246–5254. [Google Scholar]
- McCormick, J.B. Epidemiology and control of lassa fever. Curr. Top. Microbiol. Immunol. 1987, 134, 69–78. [Google Scholar] [CrossRef]
- Justines, G.; Johnson, K.M. Use of oral swabs for detection of machupo-virus infection in rodents. Am. J. Trop. Med. Hyg. 1968, 17, 788–790. [Google Scholar]
- Skinner, H.H.; Knight, E.H.; Grove, R. Murine lymphocytic choriomeningitis: The history of a natural cross-infection from wild to laboratory mice. Lab. Anim. 1977, 11, 219–222. [Google Scholar] [CrossRef]
- Peralta, L.A.; Laguens, R.P.; Cossio, P.M.; Sabattini, M.S.; Maiztegui, J.I.; Arana, R.M. Presence of viral particles in the salivary gland of calomys musculinus infected with junin virus by a natural route. Intervirology 1979, 11, 111–116. [Google Scholar] [CrossRef]
- Vitullo, A.D.; Hodara, V.L.; Merani, M.S. Effect of persistent infection with junin virus on growth and reproduction of its natural reservoir, calomys musculinus. Am. J. Trop. Med. Hyg. 1987, 37, 663–669. [Google Scholar]
- Vitullo, A.D.; Merani, M.S. Is vertical transmission sufficient to maintain junin virus in nature? J. Gen. Virol. 1988, 69, 1437–1440. [Google Scholar] [CrossRef]
- Borremans, B.; Leirs, H.; Gryseels, S.; Gunther, S.; Makundi, R.; de Bellocq, J.G. Presence of mopeia virus, an African arenavirus, related to biotope and individual rodent host characteristics: Implications for virus transmission. Vector Borne and Zoonotic Dis. 2011, 11, 1125–1131. [Google Scholar] [CrossRef]
- Webb, P.A.; Justines, G.; Johnson, K.M. Infection of wild and laboratory animals with machupo and latino viruses. B. World Health Organ. 1975, 52, 493–499. [Google Scholar]
- Vitullo, A.D.; Merani, M.S. Vertical transmission of junin virus in experimentally infected adult calomys musculinus. Intervirology 1990, 31, 339–344. [Google Scholar]
- Mills, J.N.; Ellis, B.A.; Childs, J.E.; McKee, K.T., Jr.; Maiztegui, J.I.; Peters, C.J.; Ksiazek, T.G.; Jahrling, P.B. Prevalence of infection with junin virus in rodent populations in the epidemic area of argentine hemorrhagic fever. Am. J. Trop. Med. Hyg. 1994, 51, 554–562. [Google Scholar]
- Calisher, C.H.; Nabity, S.; Root, J.J.; Fulhorst, C.F.; Beaty, B.J. Transmission of an arenavirus in white-throated woodrats (neotoma albigula), southeastern Colorado, 1995-1999. Emerg. Infect. Dis. 2001, 7, 397–402. [Google Scholar]
- Ostfeld, S.R.; Mills, J.N. Social Behavior, Demography, and Rodent-Borne Pathogens. In Rodent Societies: An Ecological and Evolutionary Perspective; Wolf, J.O., Sherman, P.W., Eds.; University of Chigago, Press: Chicago, Illinois, USA, 2007; pp. 478–486. [Google Scholar]
- Jay, M.T.; Glaser, C.; Fulhorst, C.F. The arenaviruses. J. Am. Vet. Med. Assoc. 2005, 227, 904–915. [Google Scholar] [CrossRef]
- Gonzalez, J.P.; Barbazan, P.; Baillon, F.; Capelle, J.; Chevallier, D.; Cornet, J.P.; Fournet, F.; Herbreteau, V.; Hugot, J.P.; Le Gouilh, M.; et al. Fundamentals, domains, and diffusion of disease emergence: Tools and strategies for a new paradigm. In encyclopedia of infectious diseases: Modern methodologies. Tibayrenc, M., Ed.; Wiley & Sons, Inc.: Hoboken, New Jersey, USA, 2007; pp. 525–568. [Google Scholar]
- Paweska, J.T.; Sewlall, N.H.; Ksiazek, T.G.; Blumberg, L.H.; Hale, M.J.; Lipkin, W.I.; Weyer, J.; Nichol, S.T.; Rollin, P.E.; McMullan, L.K.; et al. Nosocomial outbreak of novel arenavirus infection, southern Africa. Emerg. Infect. Dis. 2009, 15, 1598–1602. [Google Scholar]
- Palacios, G.; Savji, N.; Hui, J.; Travassos da Rosa, A.; Popov, V.; Briese, T.; Tesh, R.; Lipkin, W.I. Genomic and phylogenetic characterization of merino walk virus, a novel arenavirus isolated in south Africa. J. Gen. Virol. 2010, 91, 1315–1324. [Google Scholar] [CrossRef]
- Coulibaly-N'Golo, D.; Allali, B.; Kouassi, S.K.; Fichet-Calvet, E.; Becker-Ziaja, B.; Rieger, T.; Olschlager, S.; Dosso, H.; Denys, C.; Ter Meulen, J.; et al. Novel arenavirus sequences in hylomyscus sp. And mus (nannomys) setulosus from cote d'ivoire: Implications for evolution of arenaviruses in Africa. PLoS One 2011, 6, e20893. [Google Scholar]
- Bowen, M.D.; Peters, C.J.; Nichol, S.T. Phylogenetic analysis of the arenaviridae: Patterns of virus evolution and evidence for cospeciation between arenaviruses and their rodent hosts. Mol. Phylogenet. Evol. 1997, 8, 301–316. [Google Scholar] [CrossRef]
- Gonzalez, J.P.; Emonet, S.; de Lamballerie, X.; Charrel, R. Arenaviruses. Cur. Top. Microbiol. Iimmun. 2007, 315, 253–288. [Google Scholar]
- Irwin, N.R.; Bayerlova, M.; Missa, O.; Martinkova, N. Complex patterns of host switching in new world arenaviruses. Mol. Ecol. 2012, 21, 4137–4150. [Google Scholar] [CrossRef]
- Sabeti, P.C.; Varilly, P.; Fry, B.; Lohmueller, J.; Hostetter, E.; Cotsapas, C.; Xie, X.; Byrne, E.H.; McCarroll, S.A.; Gaudet, R.; et al. Genome-wide detection and characterization of positive selection in human populations. Nature 2007, 449, 913–918. [Google Scholar] [CrossRef]
- Andersen, K.G.; Shylakhter, I.; Tabrizi, S.; Grossman, S.R.; Happi, C.T.; Sabeti, P.C. Genome-wide scans provide evidence for positive selection of genes implicated in lassa fever. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 2012, 367, 868–877. [Google Scholar] [CrossRef]
- Cao, W.; Henry, M.D.; Borrow, P.; Yamada, H.; Elder, J.H.; Ravkov, E.V.; Nichol, S.T.; Compans, R.W.; Campbell, K.P.; Oldstone, M.B. Identification of alpha-dystroglycan as a receptor for lymphocytic choriomeningitis virus and lassa fever virus. Science 1998, 282, 2079–2081. [Google Scholar] [CrossRef]
- Radoshitzky, S.R.; Abraham, J.; Spiropoulou, C.F.; Kuhn, J.H.; Nguyen, D.; Li, W.; Nagel, J.; Schmidt, P.J.; Nunberg, J.H.; Andrews, N.C.; et al. Transferrin receptor 1 is a cellular receptor for new world haemorrhagic fever arenaviruses. Nature 2007, 446, 92–96. [Google Scholar]
- Spiropoulou, C.F.; Kunz, S.; Rollin, P.E.; Campbell, K.P.; Oldstone, M.B. New world arenavirus clade C, but not clade A and B viruses, utilizes alpha-dystroglycan as its major receptor. J. Virol. 2002, 76, 5140–5146. [Google Scholar] [CrossRef]
- Kunz, S. Receptor binding and cell entry of old world arenaviruses reveal novel aspects of virus-host interaction. Virology 2009, 387, 245–249. [Google Scholar] [CrossRef]
- Pasqual, G.; Rojek, J.M.; Masin, M.; Chatton, J.Y.; Kunz, S. Old world arenaviruses enter the host cell via the multivesicular body and depend on the endosomal sorting complex required for transport. PLoS pathogens 2011, 7, e1002232. [Google Scholar] [CrossRef]
- Botten, J.; Sidney, J.; Mothe, B.R.; Peters, B.; Sette, A.; Kotturi, M.F. Coverage of related pathogenic species by multivalent and cross-protective vaccine design: Arenaviruses as a model system. Microbiol. Mol. Biol. Rev. 2010, 74, 157–170. [Google Scholar] [CrossRef]
- Salvato, M.S.; Lukashevich, I.S. Vaccines Against Lassa Fever; Marcel Dekker, Inc: New York, New York, USA, 2009; p. 9. [Google Scholar]
- Albarino, C.G.; Posik, D.M.; Ghiringhelli, P.D.; Lozano, M.E.; Romanowski, V. Arenavirus phylogeny: A new insight. Virus Genes 1998, 16, 39–46. [Google Scholar] [CrossRef]
- Archer, A.M.; Rico-Hesse, R. High genetic divergence and recombination in arenaviruses from the Americas. Virology 2002, 304, 274–281. [Google Scholar] [CrossRef]
- Riviere, Y.; Ahmed, R.; Southern, P.; Oldstone, M.B. Perturbation of differentiated functions during viral infection in vivo. II. Viral reassortants map growth hormone defect to the S RNA of the lymphocytic choriomeningitis virus genome. Virology 1985, 142, 175–182. [Google Scholar] [CrossRef]
- Lukashevich, I.S. Generation of reassortants between African arenaviruses. Virology 1992, 188, 600–605. [Google Scholar] [CrossRef]
- Teng, M.N.; Borrow, P.; Oldstone, M.B.; de la Torre, J.C. A single amino acid change in the glycoprotein of lymphocytic choriomeningitis virus is associated with the ability to cause growth hormone deficiency syndrome. J. Virol. 1996, 70, 8438–8443. [Google Scholar]
- Chen, M.; Lan, S.; Ou, R.; Price, G.E.; Jiang, H.; de la Torre, J.C.; Moskophidis, D. Genomic and biological characterization of aggressive and docile strains of lymphocytic choriomeningitis virus rescued from a plasmid-based reverse-genetics system. J. Gen. Virol. 2008, 89, 1421–1433. [Google Scholar] [CrossRef]
- Ortiz-Riano, E.; Cheng, B.Y.; de la Torre, J.C.; Martinez-Sobrido, L. The C-terminal region of lymphocytic choriomeningitis virus nucleoprotein contains distinct and segregable functional domains involved in NP-Z interaction and counteraction of the type I interferon response. J. Virol. 2011, 85, 13038–13048. [Google Scholar] [CrossRef]
- Kerber, R.; Rieger, T.; Busch, C.; Flatz, L.; Pinschewer, D.D.; Kummerer, B.M.; Gunther, S. Cross-species analysis of the replication complex of old world arenaviruses reveals two nucleoprotein sites involved in l protein function. J. Virol. 2011, 85, 12518–12528. [Google Scholar] [CrossRef]
- Charrel, R.N.; Feldmann, H.; Fulhorst, C.F.; Khelifa, R.; de Chesse, R.; de Lamballerie, X. Phylogeny of new world arenaviruses based on the complete coding sequences of the small genomic segment identified an evolutionary lineage produced by intrasegmental recombination. Biochem. Bioph. Res. Co. 2002, 296, 1118–1124. [Google Scholar] [CrossRef]
- Emonet, S.; Lemasson, J.J.; Gonzalez, J.P.; de Lamballerie, X.; Charrel, R.N. Phylogeny and evolution of old world arenaviruses. Virology 2006, 350, 251–257. [Google Scholar] [CrossRef]
- Holland, J.J.; De La Torre, J.C.; Steinhauer, D.A. RNA virus populations as quasispecies. Curr. Top. Microbiol. Immunol. 1992, 176, 1–20. [Google Scholar] [CrossRef]
- Drake, J.W.; Holland, J.J. Mutation rates among RNA viruses. P. Natl. Acad. Sci. U.S.A. 1999, 96, 13910–13913. [Google Scholar] [CrossRef]
- Grande-Perez, A.; Sierra, S.; Castro, M.G.; Domingo, E.; Lowenstein, P.R. Molecular indetermination in the transition to error catastrophe: Systematic elimination of lymphocytic choriomeningitis virus through mutagenesis does not correlate linearly with large increases in mutant spectrum complexity. P. Natl. Acad. Sci. U.S.A. 2002, 99, 12938–12943. [Google Scholar]
- Ruiz-Jarabo, C.M.; Ly, C.; Domingo, E.; de la Torre, J.C. Lethal mutagenesis of the prototypic arenavirus lymphocytic choriomeningitis virus (LCMV). Virology 2003, 308, 37–47. [Google Scholar] [CrossRef]
- Sevilla, N.; de la Torre, J.C. Arenavirus diversity and evolution: Quasispecies in vivo. Curr. Top. Microbiol. Immunol. 2006, 299, 315–335. [Google Scholar]
- Bui, H.H.; Botten, J.; Fusseder, N.; Pasquetto, V.; Mothe, B.; Buchmeier, M.J.; Sette, A. Protein sequence database for pathogenic arenaviruses. Immunome. Res. 2007, 3, 1. [Google Scholar] [CrossRef] [Green Version]
- Fulhorst, C.F.; Charrel, R.N.; Weaver, S.C.; Ksiazek, T.G.; Bradley, R.D.; Milazzo, M.L.; Tesh, R.B.; Bowen, M.D. Geographic distribution and genetic diversity of whitewater arroyo virus in the southwestern united states. Emerg. Infect. Dis. 2001, 7, 403–407. [Google Scholar]
- Blasdell, K.R.; Becker, S.D.; Hurst, J.; Begon, M.; Bennett, M. Host range and genetic diversity of arenaviruses in rodents, United Kingdom. Emerg. Infect. Dis. 2008, 14, 1455–1458. [Google Scholar] [CrossRef]
- Bowen, M.D.; Rollin, P.E.; Ksiazek, T.G.; Hustad, H.L.; Bausch, D.G.; Demby, A.H.; Bajani, M.D.; Peters, C.J.; Nichol, S.T. Genetic diversity among lassa virus strains. J. Virol. 2000, 74, 6992–7004. [Google Scholar]
- Weaver, S.C.; Salas, R.A.; de Manzione, N.; Fulhorst, C.F.; Travasos da Rosa, A.P.; Duno, G.; Utrera, A.; Mills, J.N.; Ksiazek, T.G.; Tovar, D.; et al. Extreme genetic diversity among pirital virus (arenaviridae) isolates from western Venezuela. Virology 2001, 285, 110–118. [Google Scholar] [CrossRef]
- Domingo, E.; Martin, V.; Perales, C.; Grande-Perez, A.; Garcia-Arriaza, J.; Arias, A. Viruses as quasispecies: Biological implications. Curr. Top. Microbiol. Immunol. 2006, 299, 51–82. [Google Scholar]
- Mas, A.; Lopez-Galindez, C.; Cacho, I.; Gomez, J.; Martinez, M.A. Unfinished stories on viral quasispecies and darwinian views of evolution. J. Mol. Biol. 2010, 397, 865–877. [Google Scholar] [CrossRef]
- de la Torre, J.C.; Holland, J.J. RNA virus quasispecies populations can suppress vastly superior mutant progeny. J. Virol. 1990, 64, 6278–6281. [Google Scholar]
- Buesa-Gomez, J.; Teng, M.N.; Oldstone, C.E.; Oldstone, M.B.; de la Torre, J.C. Variants able to cause growth hormone deficiency syndrome are present within the disease-nil WE strain of lymphocytic choriomeningitis virus. J. Virol. 1996, 70, 8988–8992. [Google Scholar]
- Stocker, C.; Martinez Peralta, L.; Kratzberg, T.; Lohmann, F.; Bruns, M. Characterization of a virus variant produced by l cells persistently infected with lymphocytic choriomeningitis virus. J. Gen. Virol. 1994, 75, 3431–3439. [Google Scholar] [CrossRef]
- Teng, M.N.; Oldstone, M.B.; de la Torre, J.C. Suppression of lymphocytic choriomeningitis virus--induced growth hormone deficiency syndrome by disease-negative virus variants. Virology 1996, 223, 113–119. [Google Scholar] [CrossRef]
- Perales, C.; Mateo, R.; Mateu, M.G.; Domingo, E. Insights into rna virus mutant spectrum and lethal mutagenesis events: Replicative interference and complementation by multiple point mutants. J. Mol. Biol. 2007, 369, 985–1000. [Google Scholar] [CrossRef]
- Romanowski, V.; Bishop, D.H. The formation of arenaviruses that are genetically diploid. Virology 1983, 126, 87–95. [Google Scholar] [CrossRef]
- Pulkkinen, A.J.; Pfau, C.J. Plaque size heterogeneity: A genetic trait of lymphocytic choriomeningitis virus. Appl. Microbiol. 1970, 20, 123–128. [Google Scholar]
- Hotchin, J.; Sikora, E. Low-pathogenicity variant of lymphocytic choriomeningitis virus. Infect. Immun. 1973, 7, 825–826. [Google Scholar]
- Loeb, L.A.; Essigmann, J.M.; Kazazi, F.; Zhang, J.; Rose, K.D.; Mullins, J.I. Lethal mutagenesis of hiv with mutagenic nucleoside analogs. P. Natl. Acad. Sci. U.S.A. 1999, 96, 1492–1497. [Google Scholar]
- Moreno, H.; Gallego, I.; Sevilla, N.; de la Torre, J.C.; Domingo, E.; Martin, V. Ribavirin can be mutagenic for arenaviruses. J. Virol. 2011, 85, 7246–7255. [Google Scholar] [CrossRef]
- Grande-Perez, A.; Gomez-Mariano, G.; Lowenstein, P.R.; Domingo, E. Mutagenesis-induced, large fitness variations with an invariant arenavirus consensus genomic nucleotide sequence. J. Virol. 2005, 79, 10451–10459. [Google Scholar] [CrossRef]
- Grande-Perez, A.; Lazaro, E.; Lowenstein, P.; Domingo, E.; Manrubia, S.C. Suppression of viral infectivity through lethal defection. P. Natl. Acad. Sci. U.S.A. 2005, 102, 4448–4452. [Google Scholar]
- Pfeiffer, J.K.; Kirkegaard, K. Increased fidelity reduces poliovirus fitness and virulence under selective pressure in mice. PLoS pathogens 2005, 1, e11. [Google Scholar] [CrossRef]
- Vignuzzi, M.; Stone, J.K.; Arnold, J.J.; Cameron, C.E.; Andino, R. Quasispecies diversity determines pathogenesis through cooperative interactions in a viral population. Nature 2006, 439, 344–348. [Google Scholar] [CrossRef]
- Lukashevich, I.S.; Patterson, J.; Carrion, R.; Moshkoff, D.; Ticer, A.; Zapata, J.; Brasky, K.; Geiger, R.; Hubbard, G.B.; Bryant, J.; et al. A live attenuated vaccine for lassa fever made by reassortment of lassa and mopeia viruses. J. Virol. 2005, 79, 13934–13942. [Google Scholar]
- Moshkoff, D.A.; Salvato, M.S.; Lukashevich, I.S. Molecular characterization of a reassortant virus derived from lassa and mopeia viruses. Virus Genes 2007, 34, 169–176. [Google Scholar] [CrossRef]
- Zapata, J.; Goicochea, M.; Bryan, J.; Davis, H.; Pauza, C.; Sadzewicz, L.; Tallon, L.; Mahurkar, A.; Myers, G.; Fraser-Liggett, C.; et al. Genetic Stability of a Lassa Vaccine Candidate (ml29) in Vaccinated Animals. XV International Congress of Virology, Sopporo, Japan, 11–16 September 2011. VI-PO5–61.
- Graci, J.D.; Cameron, C.E. Therapeutically targeting RNA viruses via lethal mutagenesis. Future Virol. 2008, 3, 553–566. [Google Scholar]
- Ahmed, R.; Oldstone, M.B. Organ-specific selection of viral variants during chronic infection. J. Exp. Med. 1988, 167, 1719–1724. [Google Scholar] [CrossRef]
- Evans, C.F.; Borrow, P.; de la Torre, J.C.; Oldstone, M.B. Virus-induced immunosuppression: Kinetic analysis of the selection of a mutation associated with viral persistence. J. Virol. 1994, 68, 7367–7373. [Google Scholar]
- Cheng-Mayer, C.; Weiss, C.; Seto, D.; Levy, J.A. Isolates of human immunodeficiency virus type 1 from the brain may constitute a special group of the AIDS virus. P. Natl. Acad. Sci. U.S.A. 1989, 86, 8575–8579. [Google Scholar] [CrossRef]
- Trivedi, P.; Meyer, K.K.; Streblow, D.N.; Preuninger, B.L.; Schultz, K.T.; Pauza, C.D. Selective amplification of simian immunodeficiency virus genotypes after intrarectal inoculation of rhesus monkeys. J. Virol. 1994, 68, 7649–7653. [Google Scholar]
- Hall, J.S.; French, R.; Hein, G.L.; Morris, T.J.; Stenger, D.C. Three distinct mechanisms facilitate genetic isolation of sympatric wheat streak mosaic virus lineages. Virology 2001, 282, 230–236. [Google Scholar] [CrossRef]
- Cabot, B.; Martell, M.; Esteban, J.I.; Sauleda, S.; Otero, T.; Esteban, R.; Guardia, J.; Gomez, J. Nucleotide and amino acid complexity of hepatitis C virus quasispecies in serum and liver. J. Virol. 2000, 74, 805–811. [Google Scholar] [CrossRef]
- Sanjuan, R.; Codoner, F.M.; Moya, A.; Elena, S.F. Natural selection and the organ-specific differentiation of HIV-1 v3 hypervariable region. Evolution 2004, 58, 1185–1194. [Google Scholar]
- Deforges, S.; Evlashev, A.; Perret, M.; Sodoyer, M.; Pouzol, S.; Scoazec, J.Y.; Bonnaud, B.; Diaz, O.; Paranhos-Baccala, G.; Lotteau, V.; et al. Expression of hepatitis C virus proteins in epithelial intestinal cells in vivo. J. Gen. Virol. 2004, 85, 2515–2523. [Google Scholar] [CrossRef]
- Jelcic, I.; Hotz-Wagenblatt, A.; Hunziker, A.; Zur Hausen, H.; de Villiers, E.M. Isolation of multiple TT virus genotypes from spleen biopsy tissue from a hodgkin's disease patient: Genome reorganization and diversity in the hypervariable region. J. Virol. 2004, 78, 7498–7507. [Google Scholar] [CrossRef]
- Jridi, C.; Martin, J.F.; Marie-Jeanne, V.; Labonne, G.; Blanc, S. Distinct viral populations differentiate and evolve independently in a single perennial host plant. J. Virol. 2006, 80, 2349–2357. [Google Scholar] [CrossRef]
- Brown, R.J.; Peters, P.J.; Caron, C.; Gonzalez-Perez, M.P.; Stones, L.; Ankghuambom, C.; Pondei, K.; McClure, C.P.; Alemnji, G.; Taylor, S.; et al. Intercompartmental recombination of HIV-1 contributes to env intrahost diversity and modulates viral tropism and sensitivity to entry inhibitors. J. Virol. 2011, 85, 6024–6037. [Google Scholar]
- Wright, C.F.; Morelli, M.J.; Thebaud, G.; Knowles, N.J.; Herzyk, P.; Paton, D.J.; Haydon, D.T.; King, D.P. Beyond the consensus: Dissecting within-host viral population diversity of foot-and-mouth disease virus by using next-generation genome sequencing. J. Virol. 2011, 85, 2266–2275. [Google Scholar] [CrossRef]
- Rodas, J.D.; Lukashevich, I.S.; Zapata, J.C.; Cairo, C.; Tikhonov, I.; Djavani, M.; Pauza, C.D.; Salvato, M.S. Mucosal arenavirus infection of primates can protect them from lethal hemorrhagic fever. J. Med. Virol. 2004, 72, 424–435. [Google Scholar] [CrossRef]
- Carrion, R., Jr.; Brasky, K.; Mansfield, K.; Johnson, C.; Gonzales, M.; Ticer, A.; Lukashevich, I.; Tardif, S.; Patterson, J. Lassa virus infection in experimentally infected marmosets: Liver pathology and immunophenotypic alterations in target tissues. J. Virol. 2007, 81, 6482–6490. [Google Scholar] [CrossRef]
- Baize, S.; Marianneau, P.; Loth, P.; Reynard, S.; Journeaux, A.; Chevallier, M.; Tordo, N.; Deubel, V.; Contamin, H. Early and strong immune responses are associated with control of viral replication and recovery in lassa virus-infected cynomolgus monkeys. J. Virol. 2009, 83, 5890–5903. [Google Scholar] [CrossRef]
- Aebischer, T.; Moskophidis, D.; Rohrer, U.H.; Zinkernagel, R.M.; Hengartner, H. In vitro selection of lymphocytic choriomeningitis virus escape mutants by cytotoxic T lymphocytes. P. Natl. Acad. Sci. U.S.A. 1991, 88, 11047–11051. [Google Scholar]
- Ciurea, A.; Hunziker, L.; Martinic, M.M.; Oxenius, A.; Hengartner, H.; Zinkernagel, R.M. CD4+ T-cell-epitope escape mutant virus selected in vivo. Nature Medicine 2001, 7, 795–800. [Google Scholar] [CrossRef]
- Ciurea, A.; Hunziker, L.; Zinkernagel, R.M.; Hengartner, H. Viral escape from the neutralizing antibody response: The lymphocytic choriomeningitis virus model. Immunogenetics 2001, 53, 185–189. [Google Scholar] [CrossRef]
- Wu-Hsieh, B.; Howard, D.H.; Ahmed, R. Virus-induced immunosuppression: A murine model of susceptibility to opportunistic infection. J. Infect. Dis. 1988, 158, 232–235. [Google Scholar] [CrossRef]
- Ahmed, R.; Hahn, C.S.; Somasundaram, T.; Villarete, L.; Matloubian, M.; Strauss, J.H. Molecular basis of organ-specific selection of viral variants during chronic infection. J. Virol. 1991, 65, 4242–4247. [Google Scholar]
- Salvato, M.; Shimomaye, E.; Southern, P.; Oldstone, M.B. Virus-lymphocyte interactions. Iv. Molecular characterization of lcmv armstrong (CTL+) small genomic segment and that of its variant, clone 13 (CTL-). Virology 1988, 164, 517–522. [Google Scholar]
- Salvato, M.; Borrow, P.; Shimomaye, E.; Oldstone, M.B. Molecular basis of viral persistence: A single amino acid change in the glycoprotein of lymphocytic choriomeningitis virus is associated with suppression of the antiviral cytotoxic T-lymphocyte response and establishment of persistence. J. Virol. 1991, 65, 1863–1869. [Google Scholar]
- Bergthaler, A.; Flatz, L.; Hegazy, A.N.; Johnson, S.; Horvath, E.; Lohning, M.; Pinschewer, D.D. Viral replicative capacity is the primary determinant of lymphocytic choriomeningitis virus persistence and immunosuppression. P. Natl. Acad. Sci. U.S.A. 2010, 107, 21641–21646. [Google Scholar]
- Sevilla, N.; Kunz, S.; Holz, A.; Lewicki, H.; Homann, D.; Yamada, H.; Campbell, K.P.; de La Torre, J.C.; Oldstone, M.B. Immunosuppression and resultant viral persistence by specific viral targeting of dendritic cells. J. Exp. Med. 2000, 192, 1249–1260. [Google Scholar] [CrossRef]
- Sullivan, B.M.; Emonet, S.F.; Welch, M.J.; Lee, A.M.; Campbell, K.P.; de la Torre, J.C.; Oldstone, M.B. Point mutation in the glycoprotein of lymphocytic choriomeningitis virus is necessary for receptor binding, dendritic cell infection, and long-term persistence. P. Natl. Acad. Sci. U.S.A. 2011, 108, 2969–2974. [Google Scholar]
- Flatz, L.; Bergthaler, A.; de la Torre, J.C.; Pinschewer, D.D. Recovery of an arenavirus entirely from RNA polymerase i/ii-driven cDNA. P. Natl. Acad. Sci. U.S.A. 2006, 103, 4663–4668. [Google Scholar] [CrossRef]
- Zapata, J.; Salvato, M.S. Areanvirus variation due to host-specifc adaptation. Viruses 2013. Reported data on this manuscript. [Google Scholar]
- Kunz, S.; Sevilla, N.; McGavern, D.B.; Campbell, K.P.; Oldstone, M.B. Molecular analysis of the interaction of LCMV with its cellular receptor [alpha]-dystroglycan. J. Cell Biol. 2001, 155, 301–310. [Google Scholar] [CrossRef]
- Traub, E. An epidemic in a mouse colony due to the virus of acute lymphocytic choriomeningitis. J. Exp. Med. 1936, 63, 533–546. [Google Scholar] [CrossRef]
- Dalldorf, G. The simultaneous occurrence of the viruses of canine distemper and lymphocytic choriomeningitis : A correction of “canine distemper in the rhesus monkey”. J. Exp. Med. 1939, 70, 19–27. [Google Scholar] [CrossRef]
- Smadel, J.E.; Wall, M.J. A soluble antigen of lymphocytic choriomeningitis: II. Independence of anti-soluble substance antibodies and neutralizing antibodies, and the role of soluble antigen and inactive virus in immunity to infection. J. Exp. Med. 1940, 72, 389–405. [Google Scholar]
- Bowen, G.S.; Calisher, C.H.; Winkler, W.G.; Kraus, A.L.; Fowler, E.H.; Garman, R.H.; Fraser, D.W.; Hinman, A.R. Laboratory studies of a lymphocytic choriomeningitis virus outbreak in man and laboratory animals. Am. J. Epidemiol. 1975, 102, 233–240. [Google Scholar]
- Biggar, R.J.; Schmidt, T.J.; Woodall, J.P. Lymphocytic choriomeningitis in laboratory personnel exposed to hamsters inadvertently infected with LCM virus. J. Am. Vet. Med. Assoc. 1977, 171, 829–832. [Google Scholar]
- Montali, R.J.; Scanga, C.A.; Pernikoff, D.; Wessner, D.R.; Ward, R.; Holmes, K.V. A common-source outbreak of callitrichid hepatitis in captive tamarins and marmosets. J. Infect. Dis. 1993, 167, 946–950. [Google Scholar] [CrossRef]
- Greenwood, A.G.; Sanchez, S. Serological evidence of murine pathogens in wild grey squirrels (sciurus carolinensis) in north wales. Vet. Rec. 2002, 150, 543–546. [Google Scholar] [CrossRef]
- Zapata, J.C.; Pauza, C.D.; Djavani, M.M.; Rodas, J.D.; Moshkoff, D.; Bryant, J.; Ateh, E.; Garcia, C.; Lukashevich, I.S.; Salvato, M.S. Lymphocytic choriomeningitis virus (LCMV) infection of macaques: A model for lassa fever. Antivir. Res. 2011, 92, 125–138. [Google Scholar] [CrossRef]
- Oldstone, M.B.; Ahmed, R.; Buchmeier, M.J.; Blount, P.; Tishon, A. Perturbation of differentiated functions during viral infection in vivo. I. Relationship of lymphocytic choriomeningitis virus and host strains to growth hormone deficiency. Virology 1985, 142, 158–174. [Google Scholar] [CrossRef]
- Weaver, S.C.; Salas, R.A.; de Manzione, N.; Fulhorst, C.F.; Duno, G.; Utrera, A.; Mills, J.N.; Ksiazek, T.G.; Tovar, D.; Tesh, R.B. Guanarito virus (arenaviridae) isolates from endemic and outlying localities in venezuela: Sequence comparisons among and within strains isolated from venezuelan hemorrhagic fever patients and rodents. Virology 2000, 266, 189–195. [Google Scholar] [CrossRef]
- Matloubian, M.; Kolhekar, S.R.; Somasundaram, T.; Ahmed, R. Molecular determinants of macrophage tropism and viral persistence: Importance of single amino acid changes in the polymerase and glycoprotein of lymphocytic choriomeningitis virus. J. Virol. 1993, 67, 7340–7349. [Google Scholar]
- Barresi, R.; Campbell, K.P. Dystroglycan: From biosynthesis to pathogenesis of human disease. J. Cell Sci. 2006, 119, 199–207. [Google Scholar] [CrossRef]
- Kunz, S.; Sevilla, N.; Rojek, J.M.; Oldstone, M.B. Use of alternative receptors different than alpha-dystroglycan by selected isolates of lymphocytic choriomeningitis virus. Virology 2004, 325, 432–445. [Google Scholar] [CrossRef]
- Pasquato, A.; Rochat, C.; Burri, D.J.; Pasqual, G.; de la Torre, J.C.; Kunz, S. Evaluation of the anti-arenaviral activity of the subtilisin kexin isozyme-1/site-1 protease inhibitor pf-429242. Virology 2012, 423, 14–22. [Google Scholar] [CrossRef]
- Plotkin, J.B.; Robins, H.; Levine, A.J. Tissue-specific codon usage and the expression of human genes. P. Natl. Acad. Sci. U.S.A. 2004, 101, 12588–12591. [Google Scholar] [CrossRef]
- Pandit, A.; Sinha, S. Differential trends in the codon usage patterns in HIV-1 genes. PLoS One 2011, 6, e28889. [Google Scholar] [CrossRef]
- Gallaher, W.R.; DiSimone, C.; Buchmeier, M.J. The viral transmembrane superfamily: Possible divergence of arenavirus and filovirus glycoproteins from a common RNA virus ancestor. B.M.C. Microbiol. 2001, 1, 1. [Google Scholar]
- Carter, S.D.; Surtees, R.; Walter, C.T.; Ariza, A.; Bergeron, E.; Nichol, S.T.; Hiscox, J.A.; Edwards, T.A.; Barr, J.N. Structure, function, and evolution of the crimean-congo hemorrhagic fever virus nucleocapsid protein. J. Virol. 2012, 86, 10914–10923. [Google Scholar]
- Urquidi, V.; Bishop, D.H. Non-random reassortment between the tripartite RNA genomes of la crosse and snowshoe hare viruses. J. Gen. Virol. 1992, 73, 2255–2265. [Google Scholar] [CrossRef]
- Rodriguez, L.L.; Owens, J.H.; Peters, C.J.; Nichol, S.T. Genetic reassortment among viruses causing hantavirus pulmonary syndrome. Virology 1998, 242, 99–106. [Google Scholar] [CrossRef]
- Borucki, M.K.; Chandler, L.J.; Parker, B.M.; Blair, C.D.; Beaty, B.J. Bunyavirus superinfection and segment reassortment in transovarially infected mosquitoes. J. Gen. Virol. 1999, 80, 3173–3179. [Google Scholar]
- Stenglein, M.D.; Sanders, C.; Kistler, A.L.; Ruby, J.G.; Franco, J.Y.; Reavill, D.R.; Dunker, F.; Derisi, J.L. Identification, characterization, and in vitro culture of highly divergent arenaviruses from boa constrictors and annulated tree boas: Candidate etiological agents for snake inclusion body disease. mBio 2012, 3, e00180–00112. [Google Scholar]
- Foto, C.s. Available online: http://ec.Comps.Canstockphoto.Com/can-stock-photo_csp7080858.jpg (accessed on 12 November 2012).
- Klenerman, P.; Hengartner, H.; Zinkernagel, R.M. A non-retroviral RNA virus persists in DNA form. Nature 1997, 390, 298–301. [Google Scholar]
- Geuking, M.B.; Weber, J.; Dewannieux, M.; Gorelik, E.; Heidmann, T.; Hengartner, H.; Zinkernagel, R.M.; Hangartner, L. Recombination of retrotransposon and exogenous RNA virus results in nonretroviral cdna integration. Science 2009, 323, 393–396. [Google Scholar] [CrossRef]
- Armstrong, C.; Lillie, R.D. Experimental lymphocytic choriomeningitis of monkeys and mice produced by a virus encountered in studies of the 1933 St. Louis encephalitis epidemic. Pub. Health Rep. 1934, 49, 1019–1027. [Google Scholar] [CrossRef]
- Frame, J.D.; Baldwin, J.M., Jr.; Gocke, D.J.; Troup, J.M. Lassa fever, a new virus disease of man from west Africa. I. Clinical description and pathological findings. Am. J. Trop. Med. Hyg. 1970, 19, 670–676. [Google Scholar]
- Wulff, H.; McIntosh, B.M.; Hamner, D.B.; Johnson, K.M. Isolation of an arenavirus closely related to lassa virus from mastomys natalensis in south-east Africa. B. World Health Organ. 1977, 55, 441–444. [Google Scholar]
- Gunther, S.; Hoofd, G.; Charrel, R.; Roser, C.; Becker-Ziaja, B.; Lloyd, G.; Sabuni, C.; Verhagen, R.; van der Groen, G.; Kennis, J.; et al. Mopeia virus-related arenavirus in natal multimammate mice, Morogoro, Tanzania. Emerg. Infect. Dis. 2009, 15, 2008–2012. [Google Scholar] [CrossRef]
- Gonzalez, J.P.; McCormick, J.B.; Saluzzo, J.F.; Herve, J.P.; Georges, A.J.; Johnson, K.M. An arenavirus isolated from wild-caught rodents (pramys species) in the central African Republic. Intervirology 1983, 19, 105–112. [Google Scholar] [CrossRef]
- Swanepoel, R.; Leman, P.A.; Shepherd, A.J.; Shepherd, S.P.; Kiley, M.P.; McCormick, J.B. Identification of ippy as a lassa-fever-related virus. Lancet 1985, 1, 639. [Google Scholar]
- Ishii, A.; Thomas, Y.; Moonga, L.; Nakamura, I.; Ohnuma, A.; Hang'ombe, B.; Takada, A.; Mweene, A.; Sawa, H. Novel arenavirus, Zambia. Emerg. Infect. Dis. 2011, 17, 1921–1924. [Google Scholar] [CrossRef]
- Trapido, H.; Sanmartin, C. Pichinde virus, a new virus of the tacaribe group from Colombia. Am. J. Trop. Med. Hyg. 1971, 20, 631–641. [Google Scholar]
- Webb, P.A.; Johnson, K.M.; Hibbs, J.B.; Kuns, M.L. Parana, a new tacaribe complex virus from Paraguay. Arch. Gesamte Virusforsch 1970, 32, 379–388. [Google Scholar] [CrossRef]
- Carlton, M.L.; Gillespie, R.A.; Garver, J.; Draguljic, D.; Vela, E.M. The syrian golden hamster as a model to study flexal virus pathogenesis. Arch. Clin. Microbiol. 2012, 3, 1–9. [Google Scholar]
- Fulhorst, C.E.; Bowen, M.D.; Salas, R.A.; de Manzione, N.M.; Duno, G.; Utrera, A.; Ksiazek, T.G.; Peters, C.J.; Nichol, S.T.; De Miller, E.; et al. Isolation and characterization of pirital virus, a newly discovered south American arenavirus. Am. J. Trop. Med. Hyg. 1997, 56, 548–553. [Google Scholar]
- Moncayo, A.C.; Hice, C.L.; Watts, D.M.; Travassos de Rosa, A.P.; Guzman, H.; Russell, K.L.; Calampa, C.; Gozalo, A.; Popov, V.L.; Weaver, S.C.; et al. Allpahuayo virus: A newly recognized arenavirus (arenaviridae) from arboreal rice rats (oecomys bicolor and oecomys paricola) in northeastern Peru. Virology 2001, 284, 277–286. [Google Scholar] [CrossRef]
- Downs, W.G.; Anderson, C.R.; Spence, L.; Aitken, T.H.; Greenhall, A.H. Tacaribe virus, a new agent isolated from artibeus bats and mosquitoes in Trinidad, west Indies. Am. J. Trop. Med. Hyg. 1963, 12, 640–646. [Google Scholar]
- Parodi, A.S.; Rugiero, H.R.; Greenway, D.J.; Mettler, N.; Martinez, A.; Boxaca, M.; De La Barrera, J.M. Isolation of the junin virus (epidemic hemorrhagic fever) from the mites of the epidemic area (echinolaelaps echidninus, barlese). Prensa Med. Argent 1959, 46, 2242–2244. [Google Scholar]
- Barrera Oro, J.G.; McKee, K.T., Jr. Toward a vaccine against argentine hemorrhagic fever. Bull. Pan. Am. Health Organ. 1991, 25, 118–126. [Google Scholar]
- Pinheiro, F.; Shope, R.; Paes de Andrade, A.; Bensabath, H.; Cacios, G.; Casals, J. Amapari, a new virus of the tacaribe group from rodents and mites of amapa territory, Brazil. Proc. Soc. Exp. Biol. Med. 1966, 122, 531–535. [Google Scholar]
- Salas, R.; de Manzione, N.; Tesh, R.B.; Rico-Hesse, R.; Shope, R.E.; Betancourt, A.; Godoy, O.; Bruzual, R.; Pacheco, M.E.; Ramos, B.; et al. Venezuelan haemorrhagic fever. Lancet 1991, 338, 1033–1036. [Google Scholar]
- Lisieux, T.; Coimbra, M.; Nassar, E.S.; Burattini, M.N.; de Souza, L.T.; Ferreira, I.; Rocco, I.M.; da Rosa, A.P.; Vasconcelos, P.F.; Pinheiro, F.P.; et al. New arenavirus isolated in Brazil. Lancet 1994, 343, 391–392. [Google Scholar]
- Delgado, S.; Erickson, B.R.; Agudo, R.; Blair, P.J.; Vallejo, E.; Albarino, C.G.; Vargas, J.; Comer, J.A.; Rollin, P.E.; Ksiazek, T.G.; et al. Chapare virus, a newly discovered arenavirus isolated from a fatal hemorrhagic fever case in Bolivia. PLoS pathogens 2008, 4, e1000047. [Google Scholar] [CrossRef]
- Bowen, M.D.; Peters, C.J.; Mills, J.N.; Nichol, S.T. Oliveros virus: A novel arenavirus from Argentina. Virology 1996, 217, 362–366. [Google Scholar] [CrossRef]
- Lozano, M.E.; Posik, D.M.; Albarino, C.G.; Schujman, G.; Ghiringhelli, P.D.; Calderon, G.; Sabattini, M.; Romanowski, V. Characterization of arenaviruses using a family-specific primer set for RT-PCR amplification and RFLP analysis. Its potential use for detection of uncharacterized arenaviruses. Virus Res. 1997, 49, 79–89. [Google Scholar] [CrossRef]
- Calisher, C.H.; Tzianabos, T.; Lord, R.D.; Coleman, P.H. Tamiami virus, a new member of the tacaribe group. Am. J. Trop. Med. Hyg. 1970, 19, 520–526. [Google Scholar]
- Fulhorst, C.F.; Bowen, M.D.; Ksiazek, T.G.; Rollin, P.E.; Nichol, S.T.; Kosoy, M.Y.; Peters, C.J. Isolation and characterization of whitewater arroyo virus, a novel north American arenavirus. Virology 1996, 224, 114–120. [Google Scholar] [CrossRef]
- Cajimat, M.N.; Milazzo, M.L.; Bradley, R.D.; Fulhorst, C.F. Catarina virus, an arenaviral species principally associated with neotoma micropus (southern plains woodrat) in Texas. Am. J. Trop. Med. Hyg. 2007, 77, 732–736. [Google Scholar]
- Cajimat, M.N.; Milazzo, M.L.; Borchert, J.N.; Abbott, K.D.; Bradley, R.D.; Fulhorst, C.F. Diversity among tacaribe serocomplex viruses (family arenaviridae) naturally associated with the Mexican woodrat (neotoma mexicana). Virus Res. 2008, 133, 211–217. [Google Scholar] [CrossRef]
- Milazzo, M.L.; Cajimat, M.N.; Haynie, M.L.; Abbott, K.D.; Bradley, R.D.; Fulhorst, C.F. Diversity among tacaribe serocomplex viruses (family arenaviridae) naturally associated with the white-throated woodrat (neotoma albigula) in the southwestern United States. Vector Borne and Zoonotic Dis. 2008, 8, 523–540. [Google Scholar] [CrossRef]
- Fulhorst, C.F.; Bennett, S.G.; Milazzo, M.L.; Murray, H.L., Jr.; Webb, J.P., Jr.; Cajimat, M.N.; Bradley, R.D. Bear canyon virus: An arenavirus naturally associated with the california mouse (peromyscus californicus). Emerg. Infect. Dis. 2002, 8, 717–721. [Google Scholar] [CrossRef]
- Lecompte, E.; ter Meulen, J.; Emonet, S.; Daffis, S.; Charrel, R.N. Genetic identification of kodoko virus, a novel arenavirus of the African pigmy mouse (mus nannomys minutoides) in west africa. Virology 2007, 364, 178–183. [Google Scholar] [CrossRef]
- Palacios, G.; Druce, J.; Du, L.; Tran, T.; Birch, C.; Briese, T.; Conlan, S.; Quan, P.L.; Hui, J.; Marshall, J.; et al. A new arenavirus in a cluster of fatal transplant-associated diseases. The New England J. Med. 2008, 358, 991–998. [Google Scholar] [CrossRef]
- Ayers, S.; Bradley, R. Genetic diversity in the transferrin receptor 1 (tfr1) among natural hosts of the north american arenaviruses. Mus. Texas Tech. 2011, 304, 1–15. [Google Scholar]
- Virus Pathogens and Database and Analysis Resource (ViPR)-Arenaviridae. Available online: http://www.viprbrc.org/brc/viprDetails.do?ncbiAccession=EU938669&context=1332876702165 (accessed on 22 May 2012).
- Salvato, M.S.; Clegg, J.C.S.; Buchmeier, M.J.; Charrel, R.N.; Gonzalez, J.P.; Lukashevich, I.S.; Peters, C.J.; Rico-Hesse, R.; Romanowski, V. Index of viruses - arenaviridae (2006). In Ictvdb— the Universal Virus Database, Version 4; Büchen-Osmond, C., Ed.; Columbia University: New York, New York, USA, 2006. [Google Scholar]
- Inizan, C.C.; Cajimat, M.N.; Milazzo, M.L.; Barragan-Gomez, A.; Bradley, R.D.; Fulhorst, C.F. Genetic evidence for a tacaribe serocomplex virus, Mexico. Emerg. Infect. Dis. 2010, 16, 1007–1010. [Google Scholar] [CrossRef]
- Charrel, R.N.; de Lamballerie, X. Zoonotic aspects of arenavirus infections. Vet. Microbiol. 2010, 140, 213–220. [Google Scholar] [CrossRef]
- Cajimat, M.N.; Milazzo, M.L.; Bradley, R.D.; Fulhorst, C.F. Ocozocoautla de espinosa virus and hemorrhagic fever, Mexico. Emerg. Infect. Dis. 2012, 18, 401–405. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Zapata, J.C.; Salvato, M.S. Arenavirus Variations Due to Host-Specific Adaptation. Viruses 2013, 5, 241-278. https://doi.org/10.3390/v5010241
Zapata JC, Salvato MS. Arenavirus Variations Due to Host-Specific Adaptation. Viruses. 2013; 5(1):241-278. https://doi.org/10.3390/v5010241
Chicago/Turabian StyleZapata, Juan C., and Maria S. Salvato. 2013. "Arenavirus Variations Due to Host-Specific Adaptation" Viruses 5, no. 1: 241-278. https://doi.org/10.3390/v5010241
APA StyleZapata, J. C., & Salvato, M. S. (2013). Arenavirus Variations Due to Host-Specific Adaptation. Viruses, 5(1), 241-278. https://doi.org/10.3390/v5010241