Antioxidant and Antimicrobial Potential of the Bifurcaria bifurcata Epiphytic Bacteria
Abstract
:1. Introduction
2. Results and Discussion
| Epiphytic Bacteria | Genus | Occurrence (%) |
|---|---|---|
| 1 | Citricoccus sp. | 2.56 |
| 2; 3; 10; 11; 13; 14; 15; 18; 20; 21; 22; 23; 29; 31; 35; 39; 40; 50; 51 | Vibrio sp. | 48.72 |
| 4 | Cellulophaga sp. | 2.56 |
| 6; 12; 28; 30; 34 | Alteromonas sp. | 12.82 |
| 13B | Serratia sp. | 2.56 |
| 16; 17; 25; 26 | Shewanella sp. | 12.26 |
| 27 | Ruegeria sp. | 2.56 |
| 32 | Staphylococcus sp. | 2.56 |
| 8; 9; 24; 36; 38; 44 | <90% BLAST match | 15.38 |
| Epiphytic Bacteria | TPC (mg GAE/g Extract) | DPPH Radical Scavenging Activity IC50 (µg/mL) | ORAC (µmol TE/g Extract) | |
|---|---|---|---|---|
| 1 | 3.040 ± 0.32 | >1000 | 417.245 ± 9.44 | |
| 2 | 7.120 ± 0.05 | >1000 | 43.220 ± 6.41 | |
| 3 | 2.080 ± 0.19 | >1000 | 254.218 ± 6.47 | |
| 4 | 1.352 ± 0.09 | >1000 | 154.985 ± 3.04 | |
| 6 | 2.923 ± 0.05 | >1000 | 242.595 ± 6.68 | |
| 8 | 14.222 ± 4.25 | 164.40 (109.40–247.10) | 912.621 ± 23.79 | |
| 9 | 4.870 ± 0.02 | >1000 | 34.420 ± 1.63 | |
| 10 | 6.090 ± 0.07 | >1000 | 20.400 ± 4.99 | |
| 11 | 6.530 ± 0.05 | 183.80 (118.50–285.20) | 31.220 ± 2.06 | |
| 12 | 2.540 ± 0.30 | >1000 | 215.289 ± 7.46 | |
| 13 | 9.750 ± 0.04 | >1000 | 33.950 ± 4.40 | |
| 14 | 5.990 ± 0.06 | >1000 | 47.950 ± 5.77 | |
| 15 | 5.390 ± 0.07 | >1000 | 29.370 ± 2.62 | |
| 16 | 53.854 ± 3.02 | 20.21 (14.41–28.34) | 3603.659 ± 53.38 | |
| 17 | 6.470 ± 0.09 | >1000 | 44.250 ± 2.28 | |
| 18 | 1.633 ± 0.19 | >1000 | 202.810 ± 4.75 | |
| 20 | 7.140 ± 0.01 | >1000 | 52.110 ± 8.20 | |
| 21 | 1.668 ± 0.06 | >1000 | 131.556 ± 2.36 | |
| 22 | 0.878 ± 0.06 | >1000 | 119.415 ± 3.05 | |
| 23 | 2.035 ± 0.15 | >1000 | 561.990 ± 12.69 | |
| 24 | 3.220 ± 0.25 | >1000 | 407.490 ± 5.39 | |
| 25 | 9.380 ± 0.06 | >1000 | 42.720 ± 4.08 | |
| 26 | 5.656 ± 0.13 | 521.00 (366.90–739.70) | 520.917 ± 14.13 | |
| 27 | 2.640 ± 0.01 | >1000 | 12.210 ± 0.68 | |
| 28 | 1.848 ± 0.21 | >1000 | 149.545 ± 5.53 | |
| 29 | 1.117 ± 0.13 | >1000 | 67.211 ± 0.99 | |
| 30 | 3.502 ± 0.12 | >1000 | 68.979 ± 0.93 | |
| 31 | 8.556 ± 1.89 | >1000 | 692.260 ± 13.61 | |
| 32 | 1.963 ± 0.21 | >1000 | 201.190 ± 4.99 | |
| 34 | 2.220 ± 0.16 | >1000 | 204.646 ± 1.89 | |
| 35 | 2.770 ± 0.04 | >1000 | 358.226 ± 14.32 | |
| 36 | 10.376 ± 1.58 | 587.70 (442.00–781.40) | 129.665 ± 0.91 | |
| 38 | 5.340 ± 0.07 | >1,000 | 55.290 ± 3.11 | |
| 39 | 7.150 ± 0.03 | 23.62 (19.45–28.68) | 36.100 ± 4.69 | |
| 40 | 5.900 ± 0.02 | >1000 | 71.000 ± 15.49 | |
| 44 | 3.772 ± 0.44 | >1000 | 97.450 ± 2.11 | |
| 50 | 4.106 ± 0.02 | >1000 | 33.120 ± 0.42 | |
| 51 | 1.586 ± 0.89 | >1000 | 246.910 ± 9.97 | |
| 13B | 4.270 ± 0.02 | >1000 | 18.880 ± 0.24 | |
| BHT | - | 40.55 (27.39–60.05) | - |
| Epiphytic bacteria | B. subtilis | S. aureus |
|---|---|---|
| IC50 (µg/mL) | IC50 (µg/mL) | |
| 2 | 95.06 (82.41–109.7) | >1000 |
| 3 | 122.4 (87.66–170.9) | 95.54 (67.26–135.7) |
| 4 | >1000 | 722.9 (541.11–965.9) |
| 6 | 102.5 (68.34–153.9) | 63.48 (52.77–76.38) |
| 9 | 532.3 (452.2–626.6) | >1000 |
| 13 | 621.4 (508.2–759.9) | >1000 |
| 14 | 148.5 (115.9–190.4) | >1000 |
| 15 | 126.4 (91.68–174.2) | >1000 |
| 16 | 2.29 (1.79–2.94) | >1000 |
| 17 | 36.97 (29.97–45.60) | >1000 |
| 18 | 98.32 (84.47–114.4) | >1000 |
| 23 | 739 (525.5–1039) | 136.6 (102.8–181.6) |
| 24 | 47.39 (30.12–74.58) | >1000 |
| 25 | 31.01 (21.49–44.75) | >1000 |
| 30 | 71.35 (54.19–93.94) | >1000 |
| 32 | >1000 | 127.1 (85.79–188.4) |
| 34 | 24.17 (14.9–39.20) | 50.85 (40.72–63.50) |
| 35 | 303.3 (144.5–636.5) | >1000 |
| 38 | 58.87 (41.39–83.74) | >1000 |
| 39 | 58.86 (43.60–79.46) | >1000 |
| 40 | 411.5 (323.8–522.9) | >1000 |
| 50 | 62.54 (39.57–98.86) | >1000 |
| 13 B | 52.42 (37.00–74.27) | >1000 |
| Bacitracin | 4.09 (3.30–5.06) | 4.05 (3.35–4.90) |
| Chloramphenicol | 48.14 (33.73–68.71) | 26.01 (19.06–35.5) |
| Oxytetracycline | 0.16 (0.128–0.19) | 0.40 (0.265–0.610) |
| Ampicillin | 0.16 (0.124–0.21) | 0.04 (0.028–0.05) |
3. Experimental Section
3.1. Isolation and Purification of Epiphytic Marine Bacteria
3.2. Bacterial DNA Extraction and Identification
3.3. Preparation of Epiphytic Bacteria Extracts
3.4. Biological Activities
3.4.1. Antioxidant Activities
3.4.1.1. Quantification of Total Phenolic Content (TPC)
3.4.1.2. DPPH Radical Scavenging Activity
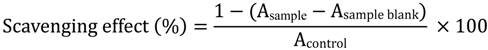
3.4.1.3. Oxygen Radical Absorbent Capacity (ORAC)
3.4.2. Antimicrobial Activity
3.5. Statistical Analysis
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Debbab, A.; Aly, A.H.; Lin, W.H.; Proksch, P. Bioactive compounds from marine bacteria and fungi. Microb. Biotechnol. 2010, 3, 544–563. [Google Scholar] [CrossRef]
- Zheng, L.-H.; Wang, Y.-J.; Sheng, J.; Wang, F.; Zheng, Y.; Lin, X.-K.; Sun, M. Antitumor peptides from marine organisms. Mar. Drugs 2011, 9, 1840–1859. [Google Scholar] [CrossRef]
- Kim, S.-K. Marine Cosmeceuticals: Trends and Prospects; CRS Press: New York, NY, USA, 2012. [Google Scholar]
- Aneiros, A.; Garateix, A. Bioactive peptides from marine sources: Pharmacological properties and isolation procedures. J. Chromatogr. B 2004, 803, 41–53. [Google Scholar] [CrossRef]
- Kim, S.-K. Marine Pharmacognosy: Trends and Applications; CRS Press: New York, NY, USA, 2013. [Google Scholar]
- Kijjoa, A.; Sawangwong, P. Drugs and cosmetics from the sea. Mar. Drugs 2004, 2, 73–82. [Google Scholar] [CrossRef]
- Graça, A.P.; Bondoso, J.; Gaspar, H.; Xavier, J.R.; Monteiro, M.C.; de la Cruz, M.; Oves-Costales, D.; Vicente, F.; Lage, O.M. Antimicrobial activity of heterotrophic bacterial communities from the marine sponge Erylus discophorus (astrophorida, geodiidae). PLoS One 2013, 8, e78992. [Google Scholar]
- Penesyan, A.; Marshall-Jones, Z.; Holmstrom, C.; Kjelleberg, S.; Egan, S. Antimicrobial activity observed among cultured marine epiphytic bacteria reflects their potential as a source of new drugs. FEMS Microbiol. Ecol. 2009, 69, 113–124. [Google Scholar] [CrossRef]
- Hollants, J.; Leliaert, F.; de Clerck, O.; Willems, A. What we can learn from sushi: A review on seaweed-bacterial associations. FEMS Microbiol. Ecol. 2013, 83, 1–16. [Google Scholar]
- Rahman, A. Studies in Natural Products Chemistry, Bioactive Natural Products (Part F); Elsevier Science: Oxford, UK, 2001; Volume 25. [Google Scholar]
- Kim, S.-K. Marine Microbiology: Bioactive Compounds and Biotechnological Applications; Wiley-VCH: Weinheim, Germany, 2013. [Google Scholar]
- Zheng, L.; Han, X.; Chen, H.; Lin, W.; Yan, X. Marine bacteria associated with marine macroorganisms: The potential antimicrobial resources. Ann. Microbiol. 2005, 55, 119–124. [Google Scholar]
- Goecke, F.; Labes, A.; Wiese, J.; Imhoff, J. Dual effects of macroalgal extracts on growth of bacteria in western baltic sea. Rev. Biol. Mar. Oceanogr. 2012, 47, 75–86. [Google Scholar] [CrossRef]
- Soria-Mercado, I.E.; Villarreal-Gómez, L.J.; Rivas, G.G.; Sánchez, N.E.A. Bioactive Compounds from Bacteria Associated to Marine Algae; InTech: Rijeka, Croatia, 2012. [Google Scholar]
- Penesyan, A.; Tebben, J.; Lee, M.; Thomas, T.; Kjelleberg, S.; Harder, T.; Egan, S. Identification of the antibacterial compound produced by the marine epiphytic bacterium Pseudovibrio sp. D323 and related sponge-associated bacteria. Mar. Drugs 2011, 9, 1391–1402. [Google Scholar] [CrossRef]
- Weisburg, W.; Barns, S.; Pelletier, D.; Lane, D. 16S Ribosomal DNA Amplification for Phylogenetic Study. J. Bacteriol. 1991, 697–703. [Google Scholar]
- Murray, P.M.; Moane, S.; Collins, C.; Beletskaya, T.; Thomas, O.P.; Duarte, A.W.F.; Nobre, F.S.; Owoyemi, I.O.; Pagnocca, F.C.; Sette, L.D.; et al. Sustainable production of biologically active molecules of marine based origin. New Biotechnol. 2013, 30, 839–850. [Google Scholar] [CrossRef]
- Muñoz, J.; Culioli, G.; Köck, M. Linear diterpenes from the marine brown alga Bifurcaria bifurcata: A chemical perspective. Phytochem. Rev. 2013, 12, 407–424. [Google Scholar] [CrossRef]
- Spavieri, J.; Allmendinger, A.; Kaiser, M.; Casey, R.; Hingley-Wilson, S.; Lalvani, A.; Guiry, M.D.; Blunden, G.; Tasdemir, D. Antimycobacterial, antiprotozoal and cytotoxic potential of twenty-one brown algae (phaeophyceae) from british and irish waters. Phytother. Res. 2010, 24, 1724–1729. [Google Scholar] [CrossRef]
- Jiménez-Escrig, A.; Gómez-Ordóñez, E.; Rupérez, P. Brown and red seaweeds as potential sources of antioxidant nutraceuticals. J. Appl. Phycol. 2012, 24, 1123–1132. [Google Scholar] [CrossRef]
- Ducklow, H. Bacterial Production and Biomass in the Ocean. In Microbial Ecology of the Oceans, 1st ed.; Kirchman, D.L., Ed.; Wiley: New York, NY, USA, 2000; pp. 85–120. [Google Scholar]
- Hamid, R.; Usup, G.; Ahmad, A. Antimicrobial activity of bacteria associated with various marine sources. Adv. Environ. Biol. 2013, 7, 356–365. [Google Scholar]
- Yang, S.-H.; Kwon, K.K.; Lee, H.-S.; Kim, S.-J. Shewanella spongiae sp. nov., isolated from a marine sponge. Int. J. Syst. Evol. Microbiol. 2006, 56, 2879–2882. [Google Scholar] [CrossRef]
- Sawabe, T.; Ohtsuka, M.; Ezura, Y. Novel alginate lyases from marine bacterium Alteromonas sp. Strain h-4. Carbohydr. Res. 1997, 304, 69–76. [Google Scholar] [CrossRef]
- Jafarzade, M.; Mohamad, S.; Usup, G.; Ahmad, A. Heavy-metal tolerance and antibiotic susceptibility of red pigmented bacteria isolated from marine environment. Nat. Resour. 2012, 3, 171–174. [Google Scholar]
- Huo, Y.-Y.; Xu, X.-W.; Li, X.; Liu, C.; Cui, H.-L.; Wang, C.-S.; Wu, M. Ruegeria marina sp. nov., isolated from marine sediment. Int. J. Syst. Evol. Microbiol. 2011, 61, 347–350. [Google Scholar] [CrossRef]
- Gunn, B.; Colwell, R. Numerical taxonomy of Staphylococci isolated from the marine environment. Int. J. Syst. Bacteriol. 1983, 33, 751–759. [Google Scholar] [CrossRef]
- Kalinovskaya, N.I.; Romanenko, L.A.; Irisawa, T.; Ermakova, S.P.; Kalinovsky, A.I. Marine isolate Citricoccus sp. Kmm 3890 as a source of a cyclic siderophore nocardamine with antitumor activity. Microbiol. Res. 2011, 166, 654–661. [Google Scholar] [CrossRef]
- Karthik, L.; Kumar, G.; Rao, K.V.B. Antioxidant activity of newly discovered lineage of marine actinobacteria. Asian Pac. J. Trop. Med. 2013, 6, 325–332. [Google Scholar] [CrossRef]
- Silva, J.; Alves, C.; Pinteus, S.; Horta, A.; Pedrosa, R. High antioxidant activity of Sargassum muticum and Padina pavonica collected from peniche coast (portugal). Curr. Opin. Biotechnol. 2013, 24, S116. [Google Scholar]
- Kalirajan, A.; Ranjit Singh, A. Antioxidant potential and biochemical evaluation of metabolites from the marine bacteria Virgibacillus sp. associated with the sponge Callyspongia diffusa. Free Radic. Antioxid. 2013, 3, 47–51. [Google Scholar] [CrossRef]
- Kindleysides, S.; Quek, S.-Y.; Miller, M.R. Inhibition of fish oil oxidation and the radical scavenging activity of new zealand seaweed extracts. Food Chem. 2012, 133, 1624–1631. [Google Scholar] [CrossRef]
- Zulueta, A.; Esteve, M.J.; Frígola, A. Orac and teac assays comparison to measure the antioxidant capacity of food products. Food Chem. 2009, 114, 310–316. [Google Scholar] [CrossRef]
- Wang, T.; Jonsdottir, R.; Ólafsdóttir, G. Total phenolic compounds, radical scavenging and metal chelation of extracts from icelandic seaweeds. Food Chem. 2009, 116, 240–248. [Google Scholar] [CrossRef]
- Okuyama, H.; Orikasa, Y.; Nishida, T. Significance of antioxidative functions of eicosapentaenoic and docosahexaenoic acids in marine microorganisms. Appl. Environ. Microbiol. 2008, 74, 570–574. [Google Scholar] [CrossRef]
- Barja, J.L.; Lemos, M.L.; Toranzo, A.E. Purification and characterization of an antibacterial substance produced by a marine Alteromonas species. Antimicrob. Agents Chemother. 1989, 33, 1674–1679. [Google Scholar] [CrossRef]
- Shiozawa, H.; Kagasaki, T.; Kinoshita, T.; Haruyama, H.; Domon, H.; Utsui, Y.; Kodama, K.; Takahashi, S. Thiomarinol, a new hybrid antimicrobial antibiotic produced by a marine bacterium: Fermentation, isolation, structure, and antimicrobial activity. J. Antibiot. 1993, 46, 1834–1842. [Google Scholar] [CrossRef]
- Abraham, T. Antibacterial marine bacterium deter luminous vibriosis in shrimp larvae. NAGA WorldFish Cent. Quart. 2004, 27, 28–31. [Google Scholar]
- Wilson, G.S.; Raftos, D.A.; Nair, S.V. Antimicrobial activity of surface attached marine bacteria in biofilms. Microbiol. Res. 2011, 166, 437–448. [Google Scholar] [CrossRef]
- Sofidiya, M.O.; Odukoya, O.A.; Afolayan, A.J.; Familoni, O.B. Phenolic contents, antioxidant and antibacterial activities of Hymenocardia acida. Nat. Prod. Res. 2009, 23, 168–177. [Google Scholar] [CrossRef]
- Kanagasabhapathy, M.; Sasaki, H.; Haldar, S.; Yamasaki, S.; Nagata, S. Antibacterial activities of marine epibiotic bacteria isolated from brown algae of japan. Ann. Microbiol. 2006, 56, 167–173. [Google Scholar] [CrossRef]
- Sørensen, E.; Rasmussen, E.M.; Eriksen, B.; Larsen, H.J.; Morling, N. Evaluation and validation of quality assessment of mitochondrial control region sequence data by phred. Int. Congr. Ser. 2004, 1261, 94–96. [Google Scholar]
- Singleton, V.L.; Rossi, J.A. Colorimetry of total phenolics with phosphomolybdic-phosphotungstic acid reagents. Am. J. Enol. Vitic. 1965, 16, 144–158. [Google Scholar]
- Brand-Williams, W.; Cuvelier, M.E.; Berset, C. Use of a free radical method to evaluate antioxidant activity. LWT Food Sci. Technol. 1995, 28, 25–30. [Google Scholar] [CrossRef]
- Dávalos, A.; Gómez-Cordovés, C.; Bartolomé, B. Extending applicability of the oxygen radical absorbance capacity (ORAC-fluorescein) assay. J. Agric. Food Chem. 2003, 52, 48–54. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Horta, A.; Pinteus, S.; Alves, C.; Fino, N.; Silva, J.; Fernandez, S.; Rodrigues, A.; Pedrosa, R. Antioxidant and Antimicrobial Potential of the Bifurcaria bifurcata Epiphytic Bacteria. Mar. Drugs 2014, 12, 1676-1689. https://doi.org/10.3390/md12031676
Horta A, Pinteus S, Alves C, Fino N, Silva J, Fernandez S, Rodrigues A, Pedrosa R. Antioxidant and Antimicrobial Potential of the Bifurcaria bifurcata Epiphytic Bacteria. Marine Drugs. 2014; 12(3):1676-1689. https://doi.org/10.3390/md12031676
Chicago/Turabian StyleHorta, André, Susete Pinteus, Celso Alves, Nádia Fino, Joana Silva, Sara Fernandez, Américo Rodrigues, and Rui Pedrosa. 2014. "Antioxidant and Antimicrobial Potential of the Bifurcaria bifurcata Epiphytic Bacteria" Marine Drugs 12, no. 3: 1676-1689. https://doi.org/10.3390/md12031676
APA StyleHorta, A., Pinteus, S., Alves, C., Fino, N., Silva, J., Fernandez, S., Rodrigues, A., & Pedrosa, R. (2014). Antioxidant and Antimicrobial Potential of the Bifurcaria bifurcata Epiphytic Bacteria. Marine Drugs, 12(3), 1676-1689. https://doi.org/10.3390/md12031676








