Comparative Analysis of Glycoside Hydrolases Activities from Phylogenetically Diverse Marine Bacteria of the Genus Arenibacter
Abstract
:1. Introduction
2. Results and Discussion
2.1. Phenotypic and Phylogenetic Characterization of Arenibacter Isolates
| Strain number | Sources and allocation places of Pacific Ocean |
|---|---|
| Arenibacter certesii | |
| KMM 3941Т | Green alga Ulva fenestrata, Troitsa Bay, Gulf of Peter the Great, Sea of Japan, Russia. |
| Arenibacter echinorum | |
| KMM 6032 | Sea urchin Strongylocentrotus intermedius, Troitsa Bay, Gulf of Peter the Great, Sea of Japan, Russia. |
| KMM 6047 | |
| Arenibacter latericius | |
| KMM 426T | Sediments, depth of 20 m, Ku-Lao-Re Island, South China Sea, Vietnam. |
| KMM 3522 | Holothurian Apostichopus japonicus, Troitsa Bay, Gulf of Peter the Great, Sea of Japan, Russia. |
| KMM 3557 | Holothurian Apostichopus japonicus, Troitsa Bay, Gulf of Peter the Great, Sea of Japan, Russia. |
| KMM 3523 | Brown alga Chorda filum, Iturup Island, Sea of Okhotsk, Russia. |
| Arenibacter palladensis | |
| KMM 3961T | Green alga Ulva fenestrata, Troitsa Bay, Gulf of Peter the Great, Sea of Japan, Russia. |
| KMM 3980 | |
| Arenibacter troitsensis | |
| KMM 3674Т | Sediments, depth of 3 m, Troitsa Bay, Gulf of Peter the Great, Sea of Japan, Russia. |
| KMM 6037 | Green alga Acrosiphonia sonderi, Troitsa Bay, Gulf of Peter the Great, Sea of Japan, Russia. |
| KMM 6212 | |
| KMM 6195 | Brown alga Laminaria japonica, Troitsa Bay, Gulf of Peter the Great, Sea of Japan, Russia. |
| Arenibacter spp. | |
| KMM 6273 | Sea urchin Strongylocentrotus intermedius, Troitsa Bay, Gulf of Peter the Great, Sea of Japan, Russia. |
| KMM 6684 | Brown alga Chorda filum, Iturup Island, Sea of Okhotsk, Russia. |
| KMM 6685 | Green alga Ulva fenestrata, Troitsa Bay, Gulf of Peter the Great, Sea of Japan, Russia. |
| Characteristic | A. latericius | A. certesii | A. echinorum | A. palladensis | A. troitsensis | Arenibacter spp. | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| KMM 426T | KMM 3522 | KMM 3523 | KMM 3557 | KMM 3941T | KMM 6032T | KMM 6047 | KMM 3961T | KMM 3980 | KMM 3674T | KMM 6037 | KMM 6212 | KMM 6195 | KMM 6273 | KMM 6684 | KMM 6685 | |
| Gliding motility | - | - | - | - | - | + | + | + | + | - | - | - | - | - | - | + |
| Na+ requirement | + | + | + | + | + | - | - | - | - | + | + | - | + | - | + | - |
| Growth with: | ||||||||||||||||
| 8% NaCl | + | + | + | + | + | + | + | + | + | - | - | - | - | + | + | + |
| 10% NaCl | - | - | - | - | + | - | - | + | + | - | - | - | - | - | - | + |
| Maximum growth temperature (°C) | 42 | 42 | 42 | 42 | 38 | 35 | 32 | 38 | 38 | 42 | 37 | 38 | 42 | 40 | 42 | 38 |
| Nitrate reduction | + | + | + | + | + | - | - | - | - | + | + | + | - | + | + | - |
| H2S production | - | - | - | - | - | - | - | - | - | + | - | - | - | - | - | - |
| Hydrolysis of: | ||||||||||||||||
| Casein | - | - | - | - | - | - | - | - | - | - | + | - | - | - | - | - |
| Gelatin | - | - | - | - | - | - | - | - | - | + | + | - | - | + | - | - |
| Tween 20 | - | - | + | + | - | - | + | - | - | - | - | - | - | - | + | - |
| Tween 40 | - | - | + | - | - | + | + | - | + | + | + | - | + | - | - | + |
| Tween 80 | - | - | - | - | - | - | - | - | + | - | - | - | + | - | - | - |
| DNA | - | + | - | + | - | - | - | - | - | - | - | - | - | - | + | - |
| Urea | + | + | + | + | + | - | - | - | - | - | - | - | - | - | + | - |
| Acid from: | ||||||||||||||||
| Arabinose | - | + | + | + | - | - | - | + | - | - | - | - | - | - | + | - |
| Galactose | + | + | + | + | + | - | - | + | + | - | + | - | + | + | + | + |
| Glucose | + | + | + | + | + | + | + | + | + | - | + | + | + | + | + | + |
| Lactose | + | + | + | + | + | + | + | + | + | - | + | + | + | - | + | + |
| Melibiose | + | - | + | + | + | + | + | + | + | - | + | + | + | + | + | + |
| Raffinose | + | + | + | + | + | + | - | - | + | - | + | + | + | - | + | + |
| Rhamnose | + | - | + | - | - | + | + | + | + | - | + | + | - | + | - | + |
| Sucrose | + | + | + | + | + | + | + | + | + | + | + | + | + | - | + | + |
| Xylose | - | - | - | - | - | + | + | + | + | - | + | + | + | + | - | + |
| N-Acetylglucosamine | + | - | + | - | + | - | - | - | - | - | - | - | - | + | - | + |
| Glycerol | + | + | + | + | - | - | - | - | - | - | - | - | - | - | + | - |
| Utilization of: | ||||||||||||||||
| Arabinose | + | + | + | + | + | + | + | + | + | + | + | - | + | + | + | + |
| Mannitol | + | + | + | + | - | + | + | - | + | - | - | - | - | - | + | + |
| DNA G + C content (mol%) | 37.5 | 38.0 | 38.2 | 37.9 | 37.7 | 39.4 | 39.2 | 40.2 | 39.2 | 40.0 | 38.0 | 40.2 | 39.9 | 41.7 | 37.7 | 39.4 |
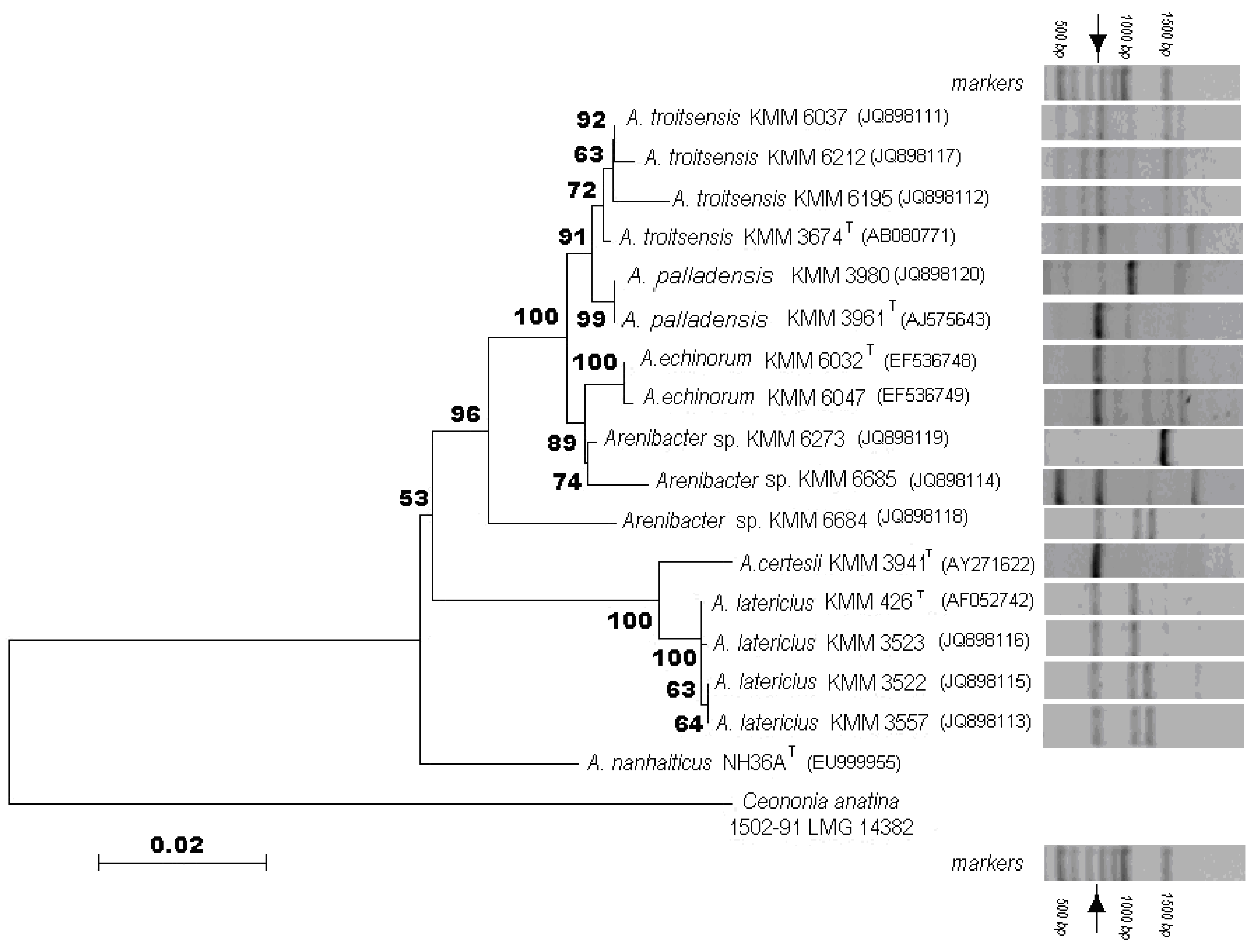
2.2. Analysis of Glycosidase Activities of the Arenibacter Isolates
| Number KMM | Specific activities (mU/mg protein) | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| β- N-aсetyl-glucosaminidase | α- N-aсetyl-galactosaminidase | α-fucosidase | α-galactosidase | α-glucosidase | α-mannosidase | α-xylosidase | β-galactosidase | β-glucosidase | |
| A. latericius | |||||||||
| 426T | 96.3 ± 7.1 * | 12.0 ± 0.9 | 3.4 ± 0.4 | 0.77 ± 0.04 | 1.11 ± 0.07 | 0.49 ± 0.04 | 0.66 ± 0.04 | 0.024 ± 0.005 | 0.3 ± 0.03 |
| 522 | 122.5 ± 4.8 | 27.3 ± 1.3 | 4.04 ± 0.02 | 1.61 ± 0.09 | 4.82 ± 0.19 | 2.54 ± 0.18 | 0.68 ± 0.03 | 1.78 ± 0.13 | 1.53 ± 0.03 |
| 3557 | 143.1 ± 4.3 | 25.6 ± 3.8 | 8.23 ± 0.4 | 0.23 ± 0.03 | 1.65 ± 0.04 | 2.06 ± 0.09 | 0.28 ± 0.01 | 0 | 1.56 ± 0.17 |
| 3523 | 113.29 ± 1.54 | 18.53 ± 0.24 | 7.64 ± 0.04 | 0.28 ± 0.004 | 3.86 ± 0.14 | 2.17 ± 0.17 | 1.23 ± 0.11 | 2.51 ± 0.26 | 1.27 ± 0.04 |
| A. certesii | |||||||||
| 3941Т | 204.81 ± 4.03 | 24.45 ± 1.42 | 1.7 ± 0.12 | 0 | 1.11 ± 0.10 | 1.43 ± 0.11 | 0 | 0 | 1.58 ± 0.13 |
| A. echinorum | |||||||||
| 6032T | 60.13 ± 2.43 | 11.93 ± 0.57 | 8.48 ± 0.48 | 0.62 ± 0.06 | 0.91 ± 0.19 | 0.82 ± 0.05 | 0.34 ± 0.09 | 1.5 ± 0.13 | 1.58 ± 0.19 |
| 6047 | 125.5 ± 1.98 | 25.86 ± 2.32 | 10.38 ± 0.23 | 0.7 ± 0.12 | 2.08 ± 0.05 | 1.88 ± 0.048 | 0.51 ± 0.07 | 0.79 ± 0.04 | 1.58 ± 0.09 |
| A. palladensis | |||||||||
| 3961T | 50.97 ± 0.89 | 14.06 ± 0.68 | 5.5 ± 0.6 | 0.91 ± 0.09 | 0.56 ± 0.05 | 1.21 ± 0.10 | 0 | 0 | 0.84 ± 0.025 |
| 3980 | 25.85 ± 1.60 | 2.26 ± 0.28 | 2.36 ± 0.38 | 1.81 ± 0.11 | 0.77 ± 0.14 | 0.48 ± 0.08 | 0 | 0 | 0 |
| A. troitsensis | |||||||||
| 3674Т | 26.91 ± 1.65 | 7.9 ± 0.36 | 2.18 ± 0.19 | 0 | 0.64 ± 0.04 | 3.34 ± 0.28 | 0 | 0 | 0 |
| 6037 | 24.4 ± 2.3 | 4.86 ± 0.71 | 3.62 ± 0.41 | 0 | 0.93 ± 0.14 | 0.25 ± 0.06 | 0 | 0 | 0 |
| 6195 | 23.81 ± 0.41 | 7.9 ± 0.3 | 2.71 ± 0.27 | 0.52 ± 0.09 | 0.35 ± 0.02 | 0.63 ± 0.05 | 0.36 ± 0.05 | 0 | 1.36 ± 0.18 |
| 6212 | 24.05 ± 0.49 | 5.31 ± 0.14 | 3.19 ± 0.3 | 0.68 ± 0.06 | 0 | 1.51 ± 0.11 | 0 | 0 | 0 |
| Arenibacterspp. | |||||||||
| 6273 | 20.53 ± 1.08 | 5.73 ± 0.21 | 3.04 ± 0.34 | 2.71 ± 0.26 | 3.10 ± 0.38 | 2.93 ± 0.23 | 0.90 ± 0.04 | 2.43 ± 0.22 | 2.0 ± 0.12 |
| 6684 | 96.77 ± 1.58 | 36.42 ± 0.88 | 4.73 ± 0.40 | 3.93 ± 0.32 | 5.7 ± 0.1 | 5.3 ± 0.62 | 1.49 ± 0.04 | 3.13 ± 0.67 | 6.01 ± 0.96 |
| 6685 | 13.71 ± 1.24 | 2.24 ± 0.22 | 2.29 ± 0.21 | 4.11 ± 0.87 | 1.60 ± 0.12 | 0.18 ± 0.01 | 0.28 ± 0.06 | 0.10 ± 0.03 | 0.09 ± 0.02 |
Structural Characteristics of A. latericius α-N-acetylgalactosaminidase
2.3. Molecular Genetic Analysis of Glycosidases
2.3.1. Band Pattern Analysis
- A. latericius KMM 426T and KMM 3523;
- A. latericius KMM 3522 and KMM 3557, Arenibacter sp. KMM 6684;
- A. troitsensis KMM 6037, KMM 6212 and KMM 6195;
- A. troitsensis KMM 3674T;
- A. palladensis KMM 3980;
- A. palladensis KMM 3961T;
- A. echinorum KMM 6032T and KMM 6047;
- A. certesii KMM 3941Т;
- A. Arenibacter sp. KMM 6273;
- A. Arenibacter sp. KMM 6685.
2.3.2. GH20 β-N-acetylglucosaminidase of Arenibacter Isolates

2.3.3. GH109 α-N-acetylgalactosaminidase from Arenibacter latericius

3. Materials and Methods
3.1. Strains Isolation and Purification
3.2. Morphological, Biochemical, and Physiological Characterization
Antimicrobial Activity
3.3. Protein Assays
3.4. Enzymatic Assays
3.5. DNA Preparation and PCR Amplification
- (1)
- 5′-CC(T/G)CA(A/G)ATGGG(N)TGGAA-3′,
- (2)
- 5′-ACTCC(G/A/T)ATGGGNTG-3′,
- (3)
- 5′-CC(G/C/T)ATGGGNTT(T/C)AA(T/C)AA(T/C)TGG-3′,
- (4)
- 5′-(A/G)TNGA(T/C)GA(T/C)GGNTGGTT-3′;
- (5)
- 5′-AA(G/A)TCNGG(G/A)TC(G/A)TT(G/A)AA-3′,
- (6)
- 5′-CAT(G/A)TCNCC(G/A)TC(A/G)TTCCA-3′,
- (7)
- 5′-C(T/G)(G/A)TT(G/A/C)A(T/A)(G/A)TCCCA(C/T)TT-3′,
- (8)
- 5′-AT(A/C/T)CCCAT(G/A)TGNCC(G/A)AA-3′.
- Nend-acetyl_forward1—5′-GG(G/A/T)GC(A/T)AA(A/G)TA(T/C)ATGGGNGGNTT(T/C)TC-3′,
- Nend-acetyl_forward2—5′-AA(A/G)TA(T/C)ATGGGNGGNTT(T/C)-3′,
- N-acetyl_forward1—5′-CA(T/C)GCNTT(T/C)GTNGA(A/G)GTNCC-3′,
- N-acetyl_forward2—5′-ATGATGATGGA(A/G)AA(T/C)GTNAA(T/C)TA-3′,
- N-acetyl_reverse1—5′-CCNCTNGT(G/A)AA(G/A)TCNGG(G/A)AA-3′,
- N-acetyl_reverse2—5′-GG(A/G)TG(G/A)TC(G/A)TA(C/T)TTNTC-3′.
3.6. PCR-Product and Sequence Analysis
4. Conclusions
Acknowledgements
References
- Ivanova, E.P.; Nedashkovskaya, O.I.; Chun, J.; Lysenko, A.M.; Frolova, G.M.; Svetashev, V.I.; Vysotskii, M.V.; Mikhailov, V.V.; Huq, A.; Colwell, R.R. Arenibacter gen. nov., a new genus of the family Flavobacteriaceae and description of new species, Arenibacter latericius sp. nov. Int. J. Syst. Evol. Microbiol. 2001, 51, 1987–1995. [Google Scholar] [CrossRef]
- Nedashkovskaya, O.I.; Suzuki, M.; Vysotskii, M.V.; Mikhailov, V.V. Arenibacter troitsensis sp. nov., isolated from marine bottom sediment. Int. J. Syst. Evol. Microbiol. 2003, 53, 1287–1290. [Google Scholar] [CrossRef]
- Nedashkovskaya, O.I.; Kim, S.B.; Han, S.K.; Lysenko, A.M.; Mikhailov, V.V.; Bae, K.S. Arenibacter certesii sp. nov., a novel marine bacterium isolated from the green alga Ulva fenestrata. Int. J. Syst. Evol. Microbiol. 2004, 54, 1173–1176. [Google Scholar] [CrossRef]
- Nedashkovskaya, O.I.; Vancanneyt, M.; Cleenwerck, I.; Snauwaert, C.; Kim, S.B.; Lysenko, A.M.; Shevchenko, L.S.; Lee, K.H.; Park, M.S.; Frolova, G.M.; et al. Arenibacter palladensis sp. nov., a novel marine bacterium isolated from the green alga Ulva fenestrata, and emended description of the genus Arenibacter. Int. J. Syst. Evol. Microbiol. 2006, 56, 155–160. [Google Scholar] [CrossRef]
- Nedashkovskaya, O.I.; Kim, S.B.; Shin, D.S.; Beleneva, I.A.; Vancanneyt, M.; Mikhailov, V.V. Echinicola vietnamencis sp. nov., a new bacterium of the phylum Bacteroidesetes isolated from sea water. Int. J. Syst. Evol. Microbiol. 2007, 57, 761–763. [Google Scholar] [CrossRef]
- Sun, F.; Wang, B.; Du, Y.; Liu, X.; Lai, Q.; Li, G.; Luo, J.; Shao, Z. Arenibacter nanhaiticus sp. nov., isolated from marine sediment of the South China Sea. Int. J. Syst. Evol. Microbiol. 2010, 60, 78–83. [Google Scholar] [CrossRef]
- Riemann, L.; Azam, F. Widespread N-acetyl-d-glucosamine uptake among pelagic marine bacteria and its ecological implications. Appl. Environ. Microbial. 2002, 68, 5554–5562. [Google Scholar] [CrossRef]
- Khoushab, F.; Yamabhai, M. Chitin research revisited. Mar. Drugs 2010, 8, 1988–2012. [Google Scholar] [CrossRef]
- Sharon, N. Carbohydrates as future anti-adhesion drugs for infectious diseases. Biochim. Biophys. Acta 2006, 1760, 527–537. [Google Scholar] [CrossRef]
- Shen, K.T.; Chen, M.H.; Chan, H.Y.; Jeng, J.H.; Wang, Y.J. Inhibitory effects of chitooligosaccharides on tumor growth and metastasis. Food Chem. Toxicol. 2009, 47, 1864–1871. [Google Scholar] [CrossRef]
- Zerkak, D.; Dougados, M. The use of glucosamine therapy in osteoarthritis. J. Bone Miner. Metab. 2002, 20, 298–302. [Google Scholar] [CrossRef]
- Ozkan, F.U.; Ozkan, K.; Ramadan, S.; Guven, Z. Chondroprotective effect of N-acetylglucosamine and hyaluronate in early stages of osteoarthritis—An experimental study in rabbits. Bull. NYU Hosp. Jt. Dis. 2009, 67, 352–357. [Google Scholar]
- Scigelova, M.; Crout, D.H.G. Microbial β-N-acetylglucosaminidases and their biotechnological applications. Enzyme Microb. Technol. 1999, 25, 3–14. [Google Scholar] [CrossRef]
- Ivanova, E.P.; Bakunina, I.Y.; Nedashkovskaya, O.I.; Gorshkova, N.M.; Mikhailov, V.V.; Elyakova, L.A. Incidence of marine microorganisms producing β-N-Acetylglucosaminidases, α-galactosidases and α-N-Acetylgalactosaminidases. Russ. J. Mar. Biol. 1998, 24, 365–372. [Google Scholar]
- Bakunina, I.Y.; Nedashkovskaya, O.I.; Kim, S.B.; Zvyagintseva, T.N.; Mikhailov, V.V. Diversity of glycosidase activities in the bacteria of the phylum Bacteroidetes isolated from marine algae. Microbiology 2012, 81, 688–695. [Google Scholar] [CrossRef]
- Intra, J.; Pavesi, G.; Horner, D.S. Phylogenetic analyses suggest multiple changes of substrate specificity within the Glycosyl hydrolase 20 family. BMC Evol. Biol. 2008, 8, 214–231. [Google Scholar] [CrossRef]
- Dowd, P.F.; Johnson, E.T.; Pinkerton, T.S. Oral toxicity of β-N-acetyl hexosaminidase to insects. J. Agric. Food Chem. 2007, 55, 3421–3428. [Google Scholar] [CrossRef]
- Clausen, H.; Hakomori, S.-I. ABH and related histo-blood group antigens; immunochemical differences in carrier isotypes and their distribution. Vox Sang. 1989, 56, 1–20. [Google Scholar] [CrossRef]
- Yamamoto, F.; Clausen, H.; White, T.; Marken, J.; Hakomori, S. Molecular genetic basis of the histo-blood group ABO system. Nature 1990, 345, 229–233. [Google Scholar] [CrossRef]
- Wu, A.M.; Wu, J.H.; Chen, Y.Y.; Tsai, M.S.; Herp, A. Forssman pentasaccharide and polyvalent Galβ1→4GlcNAc as major ligands with affinity for Caragana arborescens agglutinin. FEBS Lett. 1999, 463, 223–230. [Google Scholar]
- Kenne, L.; Lindberg, B. Bacterial Polysaccharides. In The Polysaccharides; Aspinoll, G.O., Ed.; Academic Press: New York, NY, USA, 1983; Volume 2, pp. 287–363. [Google Scholar]
- Tomshich, S.V.; Isakov, V.V.; Komandrova, N.A.; Shevchenko, L.S. Structure of the O-specific polysaccharide of the marine bacterium Arenibacter palladensis KMM 3961T containing 2-acetamido-2-deoxy-l-galacturonic acid. Biochemistry (Mosc.) 2012, 77, 87–91. [Google Scholar] [CrossRef]
- Urvantseva, A.M.; Bakunina, I.Y.; Nedashkovskaya, O.I.; Kim, S.B.; Zvyagintseva, T.N. Distribution of intracellular fucoidan hydrolases among marine bacteria of the family Flavobacteriaceae. Appl. Biochem. Microbiol. (Mosc.) 2006, 42, 484–491. [Google Scholar] [CrossRef]
- Bakunina, I.Y.; Nedashkovskaya, O.I.; Kim, S.B.; Zvyagintseva, T.N.; Mikhailov, V.V. Distribution of α-N-acetylgalactosaminidases among marine bacteria of the phylum Bacteroidetes, epiphytes of marine algae of the Seas of Okhotsk and Japan. Microbiology (Mosc.) 2012, 81, 373–378. [Google Scholar]
- Bakunina, I.Y.; Kuhlmann, R.A.; Likhosherstov, L.M.; Martynova, M.D.; Nedashkovskaya, O.I.; Mikhailov, V.V.; Elyakova, L.A. α-N-Acetylgalactosaminidase from marine bacterium Arenibacter latericius KMM 426T removing blood type specificity of A erythrocytes. Biochemistry (Mosc.) 2002, 67, 689–695. [Google Scholar] [CrossRef]
- Tsujibo, H.; Kondo, N.; Tanaka, K.; Miyamoto, K.; Baba, N.; Inamori, Y. Molecular analysis of the gene encoding a novel transglycosylative enzyme from Alteromonas sp. strain O-7 and its physiological role in the chitinolytic system. J. Bacteriol. 1999, 181, 5461–5466. [Google Scholar]
- Keyhani, N.O.; Roseman, S. The chitin catabolic cascade in the marine bacterium Vibrio furnissii. Molecular cloning, isolation, and characterization of a periplasmic β-N-acetylglucosaminidase. J. Biol. Chem. 1996, 271, 33425–33432. [Google Scholar] [CrossRef]
- Jung, B.O.; Roseman, S.; Park, J.K. The central concept for chitin catabolic cascade in marine bacterium, Vibrios. Macromol. Res. 2008, 16, 1–5. [Google Scholar] [CrossRef]
- Suginta, W.; Chuenark, D.; Mizuhara, M.; Fukamizo, T. Novel β-N-acetylglucosaminidases from Vibrio harveyi 650: Cloning, expression, enzymatic properties, and subsite identification. BMC Biochem. 2010, 11, 40–52. [Google Scholar] [CrossRef]
- Naumoff, D.G. Hierarchical classification of glycoside hydrolases. Biochemistry (Mosc.) 2011, 76, 622–635. [Google Scholar] [CrossRef]
- Lovatt, A.; Roberts, I.S. Cloning and expression in Escherichia coli of the nahA gene from Porphyromonas gingivalis indicates that β-N-acetylhexosaminidase is an outer-membrane-associated lipoprotein. Microbiology 1994, 140, 3399–3406. [Google Scholar] [CrossRef]
- Mayer, C.; Vocadlo, D.J.; Mah, M.; Rupitz, K.; Stoll, D.; Warren, R.A.J.; Withers, S.G. Characterization of a β-N-acetylhexosaminidase and a β-N-acetylglucosaminidase/β-glucosidase from Cellulomonas fimi. FEBS J. 2006, 273, 2929–2941. [Google Scholar] [CrossRef]
- Cantarel, B.L.; Coutinho, P.M.; Rancurel, C.; Bernard, T.; Lombard, V.; Henrissat, B. The Carbohydrate-active EnZymes database (CAZy): An expert resource for Glycogenomics. Nucl. Acids. Res. 2009, 37, D233–D238. [Google Scholar] [CrossRef]
- Henrissat, B. A classification of glycosyl hydrolases based on amino-acid sequence similarities. Biochem. J. 1991, 280, 309–316. [Google Scholar]
- Langley, D.B.; Harty, D.W.S.; Jacques, N.A.; Hunter, N.; Guss, J.M.; Collyer, C.A. Structure of N-acetyl-β-d-glucosaminidase (GcnA) from the endocarditis pathogen Streptococcus gordonii and its complex with the mechanism-based inhibitor NAG-thiazoline. J. Mol. Biol. 2008, 377, 104–116. [Google Scholar] [CrossRef]
- Liu, Q.P.; Sulzenbacher, G.; Yuan, H.; Bennett, E.P.; Pietz, G.; Saunders, K.; Spence, J.; Nudelman, E.; Levery, S.B.; White, T.; et al. Bacterial glycosidases for the production of universal red blood cells. Nat. Biotechnol. 2007, 25, 454–464. [Google Scholar] [CrossRef]
- PHYLIP (Phylogeny Inference Package), version 3.6; Department of Genome Sciences, University of Washington: Seattle, WA, USA, 2004.
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucl. Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef]
- Koshland, D.E.J. Stereochemistry and the mechanism of enzymatic reactions. Biol. Rev. Camb. Philos. Soc. 1953, 28, 416–436. [Google Scholar] [CrossRef]
- Van Passel, M.W.J.; Kant, R.; Zoetendal, E.G.; Plugge, C.M.; Derrien, M.; Malfatti, S.A.; Chain, P.S.G.; Woyke, T.; Palva, A.; de Vos, W.M.; et al. The Genome of Akkermansia muciniphila, a dedicated intestinal mucin degrader, and its use in exploring intestinal metagenomes. PLoS One 2011, 6, e16876. [Google Scholar] [CrossRef]
- Williams, T.J.; Wilkins, D.; Long, E.; Evans, F.; DeMaere, M.Z.; Raftery, M.J.; Cavicchioli, R. The role of planktonic Flavobacteria in processing algal organic matter in coastal East Antarctica revealed using metagenomics and metaproteomics. Environ. Microbiol. 2012, 15, 1302–1317. [Google Scholar]
- Smibert, R.M.; Krieg, N.R. Phenotypic Characterization. In Methods for General and Molecular Bacteriology; Gerhardt, P., Murray, R.G.E., Wood, W.A., Krieg, N.R., Eds.; American Society for Microbiology: Washington, DC, USA, 1994; pp. 607–654. [Google Scholar]
- Lemos, M.L.; Toranzo, A.E.; Barja, J.L. Modified medium for oxidation N-fermentation test in the identification of marine bacteria. Appl. Environ. Microbiol. 1985, 40, 1541–1543. [Google Scholar]
- Suzuki, M.; Nakagawa, Y.; Harayama, S.; Yamamoto, S. Phylogenetic analysis and taxonomic study of marine Cytophaga-like bacteria: Proposal for Tenacibaculum gen. nov. with Tenacibaculum maritimum comb. nov., and Tenacibaculum ovolyticum comb. nov., and description of Tenacibaculum mesophilum sp. nov. and Tenacibaculum amylolyticum sp. nov. Int. J. Syst. Evol. Microbiol. 2001, 51, 1639–1652. [Google Scholar] [CrossRef]
- Marmur, J. A procedure for the isolation of deoxyribonucleic acid from microorganisms. J. Mol. Biol. 1961, 3, 208–218. [Google Scholar] [CrossRef]
- Marmur, J.; Doty, P. Determination of the base composition of deoxyribonucleic acid from its thermal denaturation temperature. J. Mol. Biol. 1962, 5, 109–118. [Google Scholar] [CrossRef]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Nelson, T.E. A photometric adaptation of the Somogyi method for the determination of glucose. J. Biol. Chem. 1944, 153, 375–381. [Google Scholar]
- Zvyagintseva, T.N.; Shevchenko, N.M.; Chizhov, A.O.; Krupnova, T.N.; Sundukova, E.V.; Isakov, V.V. Water-soluble polysaccharides of some far-eastern brown seaweeds. Distribution, structure, and their dependence on the developmental conditions. J. Exp. Mar. Biol. Ecol. 2003, 294, 1–13. [Google Scholar] [CrossRef]
- Kusaykin, M.I.; Bakunina, I.Y.; Sova, V.V.; Ermakova, S.P.; Kuznetsova, T.S.; Besednova, N.N.; Zaporozhets, T.S.; Zvyagintseva, T.N. Structure and biological action of the polysaccharides and products of their transformation. J. Biotechnol. 2008, 3, 904–915. [Google Scholar] [CrossRef]
- Zvyagintseva, T.N.; Shevchenko, N.M.; Nazarenko, E.L.; Gorbach, V.I.; Urvantseva, A.M.; Kiseleva, M.I.; Isakov, V.V. Water-soluble polysaccharides of some brown algae of the Russian Far-East. Structure and biological action of low-molecular mass polyuronans. J. Exp. Mar. Biol. Ecol. 2005, 320, 123–131. [Google Scholar] [CrossRef]
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 2007, 24, 1596–1599. [Google Scholar] [CrossRef]
- Kim, O.S.; Cho, Y.J.; Lee, K.; Yoon, S.H.; Kim, M.; Na, H.; Park, S.C.; Jeon, Y.S.; Lee, J.H.; Yi, H.; et al. Introducing EzTaxon-e: A prokaryotic 16S rRNA Gene sequence database with phylotypes that represent uncultured species. Int. J. Syst. Evol. Microbiol. 2012, 62, 716–721. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Bakunina, I.; Nedashkovskaya, O.; Balabanova, L.; Zvyagintseva, T.; Rasskasov, V.; Mikhailov, V. Comparative Analysis of Glycoside Hydrolases Activities from Phylogenetically Diverse Marine Bacteria of the Genus Arenibacter. Mar. Drugs 2013, 11, 1977-1998. https://doi.org/10.3390/md11061977
Bakunina I, Nedashkovskaya O, Balabanova L, Zvyagintseva T, Rasskasov V, Mikhailov V. Comparative Analysis of Glycoside Hydrolases Activities from Phylogenetically Diverse Marine Bacteria of the Genus Arenibacter. Marine Drugs. 2013; 11(6):1977-1998. https://doi.org/10.3390/md11061977
Chicago/Turabian StyleBakunina, Irina, Olga Nedashkovskaya, Larissa Balabanova, Tatyana Zvyagintseva, Valery Rasskasov, and Valery Mikhailov. 2013. "Comparative Analysis of Glycoside Hydrolases Activities from Phylogenetically Diverse Marine Bacteria of the Genus Arenibacter" Marine Drugs 11, no. 6: 1977-1998. https://doi.org/10.3390/md11061977
APA StyleBakunina, I., Nedashkovskaya, O., Balabanova, L., Zvyagintseva, T., Rasskasov, V., & Mikhailov, V. (2013). Comparative Analysis of Glycoside Hydrolases Activities from Phylogenetically Diverse Marine Bacteria of the Genus Arenibacter. Marine Drugs, 11(6), 1977-1998. https://doi.org/10.3390/md11061977




