Quorum Sensing Activity of Enterobacter asburiae Isolated from Lettuce Leaves
Abstract
: Bacterial communication or quorum sensing (QS) is achieved via sensing of QS signaling molecules consisting of oligopeptides in Gram-positive bacteria and N-acyl homoserine lactones (AHL) in most Gram-negative bacteria. In this study, Enterobacteriaceae isolates from Batavia lettuce were screened for AHL production. Enterobacter asburiae, identified by matrix-assisted laser desorption ionization time of flight mass spectrometry (MALDI-TOF-MS) was found to produce short chain AHLs. High resolution triple quadrupole liquid chromatography mass spectrometry (LC/MS) analysis of the E. asburiae spent supernatant confirmed the production of N-butanoyl homoserine lactone (C4-HSL) and N–hexanoyl homoserine lactone (C6-HSL). To the best of our knowledge, this is the first report of AHL production by E. asburiae.1. Introduction
Quorum sensing (QS) refers to the capabilities of microorganisms to communicate via secreted signaling molecules called autoinducers which contribute to the regulation of gene expression in response to surrounding bacterial population density [1]. In general, Gram-negative bacteria use AHLs as autoinducers [2]. AHLs consist of 4- to 18-carbon side chains linked to a lactone ring [3]. The molecules are synthesized by the activity of LuxI synthase using S-adenosylmethionine (SAM) and acylated acyl carrier protein (Acyl-ACP) as substrates [4]. Gram-negative bacteria employ AHL as QS signals in their communication circuits to regulate a diverse array of physiological activities. These processes include symbiosis, virulence, competence, conjugation, antibiotic production, motility, sporulation, and biofilm formation [2].
The nutrient contents of vegetables and fruits are suitable for the growth of Gram-negative bacteria, particularly Enterobacteriaceae [5,6], resulting in accelerated spoilage as well as cases of food poisoning [7,8]. Studies on the microbial flora in vegetables and fruits indicate competitive inhibition within the microbial community which may affect food quality [9]. It is believed that this interaction among the microorganisms, and in particular, the availability of AHL-regulated systems may be responsible for the deterioration, toxicity and the ultimate safety of food products [10].
Enterobacter asburiae is a Gram-negative bacillus belonging to the Enterobacteriaceae family that has been isolated from soil, water and food products [11,12]. It is termed an epiphytic bacterium [13], which are generally described as microorganisms that live on plant surfaces and can either be favorable or harmful to the plant they reside on [14]. Although E. asburiae has been reported to possess autoinducer-2 (AI-2) receptors [15] there are no reports on the production of AHLs by this bacterial species. In this study, we aimed to identify the AHL(s) produced by the isolated Gram-negative bacteria E. asburiae from the Enterobacteriaceae family, obtained from Batavia lettuce.
2. Experimental Section
2.1. Sample Collection and Processing
Batavia lettuce samples were collected from a local market in Malaysia in a sterile container. The samples were processed within half an hour of sample collection. Briefly, 5 g of the leaf were placed in 50 mL of Brain Heart Infusion (BHI) broth and incubated overnight at 37 °C agitated at 200 rpm.
2.2. Isolation and Identification of Bacterial Strains
A 10 μL tenfold serial dilution (10−1, 10−2, 10−3, 10−4 and 10−5) of the overnight culture were plated on MacConkey agar. Bacterial isolates of interest were identified using a MALDI-TOF-MS (Bruker, Germany) [16] extraction method with UV laser wavelength of 337 nm and acceleration voltage of 20 kV. Each spot on the target plate was measured by the MBT-autoX.axe autoExecute method. The bacterial spectra were then analyzed in the Bruker MALDI Biotyper Real Time Classification (RTC) Version 3.1 (Build 65) software. The dendrogram was generated using the standard MALDI Biotyper MSP creation method.
2.3. AHL Detection of Bacteria Isolates
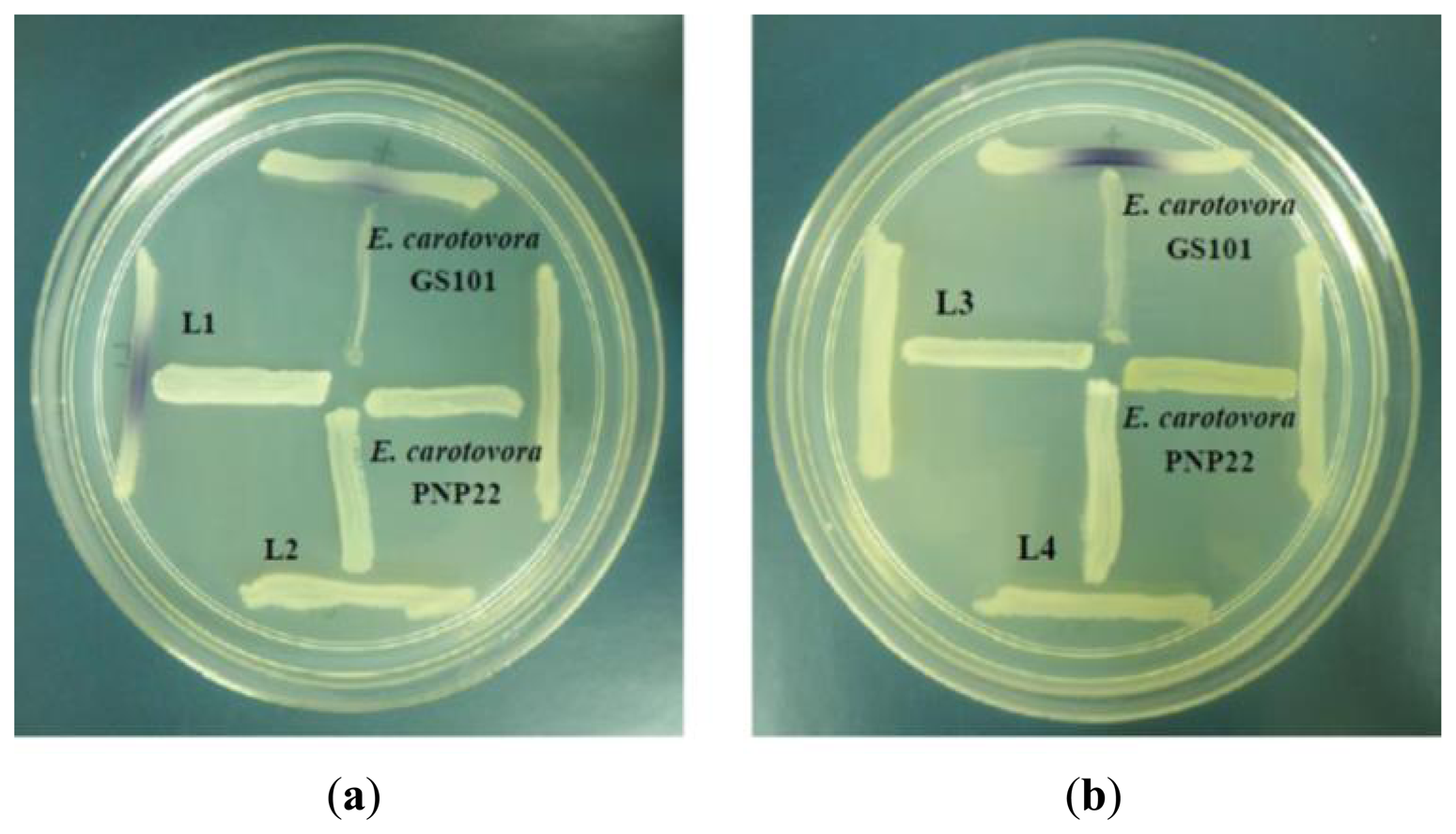
Chromobacterium violaceum CV026 which detects the presence of exogenous short chain AHLs ranging from four to eight carbons was used as an AHL biosensor. The bacterial isolates were screened using cross-streaking with CV026 [17]. Erwinia carotovora GS101 and E. carotovora PNP22 were used as positive control and negative controls, respectively [17].
2.4. AHL Extraction and Measurement of Bioluminescence
Bacterial colony that showed positive result for AHL detection via cross streaking with C. violaceum CV026 was incubated overnight in buffered Luria Bertani (LB) medium, pH 6.5, with 3-(N-morpholino)propanesulfonic acid (MOPS, 50 mM, pH 6.5) at 37 °C with shaking (220 rpm). The spent supernatant was then extracted thrice with an equal volume of acidified (0.1% v/v acetate acid) ethyl acetate. The extract was dried and stored at −20 °C prior to liquid chromatography-mass spectrometry (LC/MS) analysis.
Cell density bioluminescence measurements were done using an Infinite M200 luminometer-spectrophotometer (Tecan, Männedorf, Switzerland). Aliquots of 200 μL of diluted (1:100) E. coli strain [pSB401] overnight culture in LB supplemented with tetracycline (20 μg/mL) was added with 1 μL of extracted AHL to every well of a 96-well optical bottom microtitre plate [18,19]. The E. coli strain used harbors lux from the pSB401 plasmid [20]. Acetonitrile and synthetic 3-oxo-C6-HSL (250 pg/μL) were used as the negative and positive standards, respectively. Results were indicated as Relative Light Units (RLU)/OD495 nm against incubation time.
2.5. AHL Identification by Triple Quadrupole LC/MS
Extracted AHL was reconstituted in acetonitrile followed by LC/MS analysis using an Agilent 1290 Infinity LC system (Agilent Technologies, Santa Clara, CA, USA) equipped with an Agilent ZORBAX Rapid Resolution High Definition SB-C18 Threaded Column (2.1 mm × 50 mm, 1.8 μm particle size). The flow rate was set at 0.3 mL/min and the temperature at 37 °C. Injection volume was 2 μL. Mobile phases A and B used included water and acetonitrile (both containing 0.1% v/v formic acid). The gradient profile was set at A:B 80:20 at 0 min, 50:50 at 7 min, 20:80 at 12 min, and 80:20 at 14 min. Subsequent MS detection of separated compounds was performed on the Agilent 6490 Triple Quadrupole LC/MS system. Precursor ion-scanning analysis were performed in positive ion mode with Q3 set to monitor for m/z 102 and Q1 set to scan a mass range of m/z 80 to m/z 400. Molecular mass of m/z 102 refers to the lactone ring thus indicating presence of AHLs. The MS parameters were: probe capillary voltage set at 3 kV, sheath gas at 11 mL/h, nebulizer pressure of 20 p.s.i. and desolvation temperature of 200 °C. The Agilent MassHunter software was used for the MS data analysis to confirm the presence of AHLs. Analysis was based on the retention index and the comparison of the EI mass spectra with AHL standards.
3. Results and Discussion
3.1. Identification of Bacterial Isolates from Batavia Lettuce
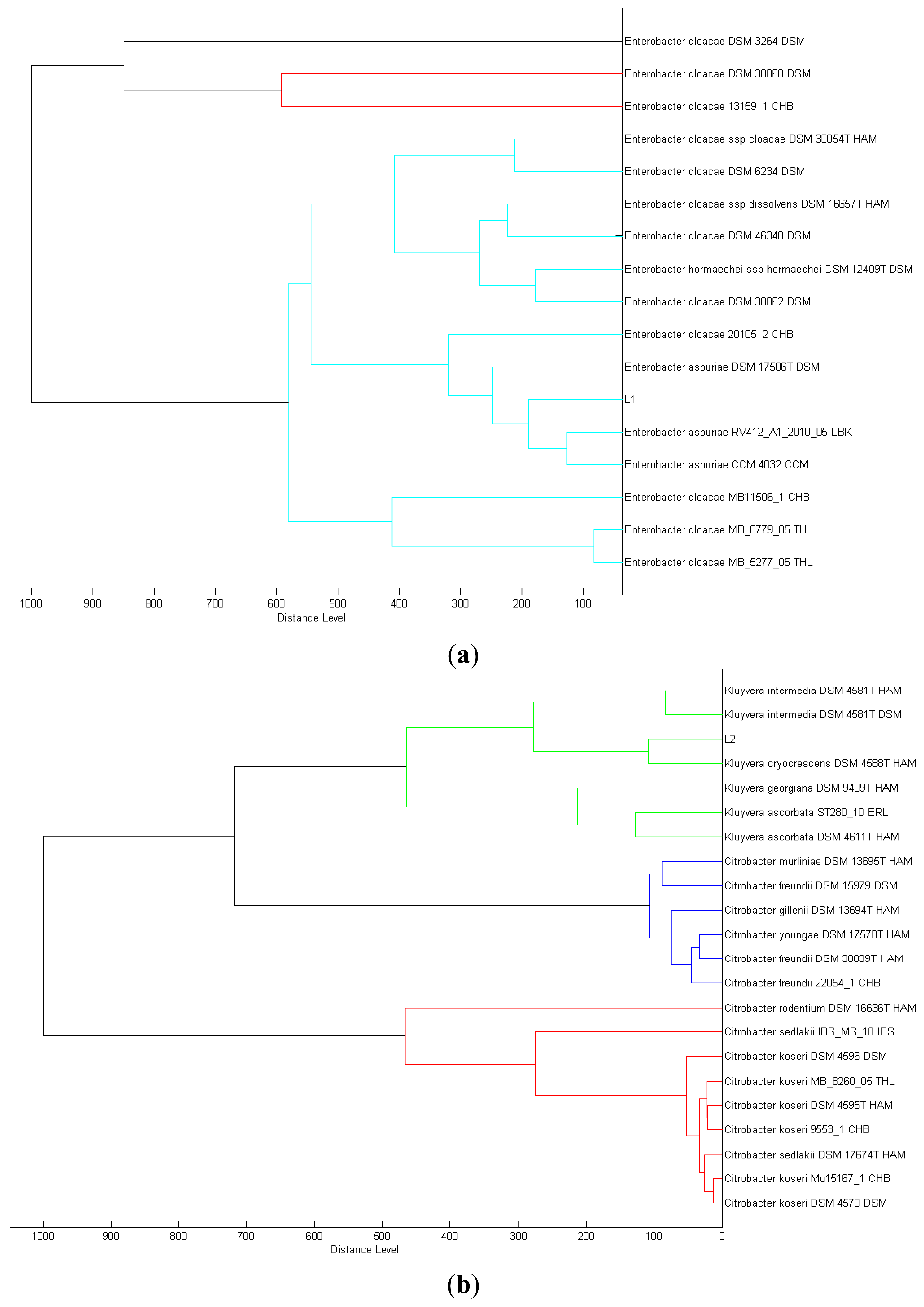
We isolated a total of four different Enterobacteriaceae bacterial colonies from the Batavia lettuce sample. In order to identify the isolates, MALDI-TOF-MS was performed [21]. Two of the isolates could be identified to the species level with score values above 2.3, namely E. asburiae (L1) and Morganella morganii (L3). However, we were only able to identify the other two isolates, Kluyvera (L2) and Rahnella (L4) at the genus level with score values of 2.1 and 2.0, respectively [22]. Phylogenetic trees generated on all of the isolates are shown in Figure 1.
3.2. Production of AHLs by E. Asburiae
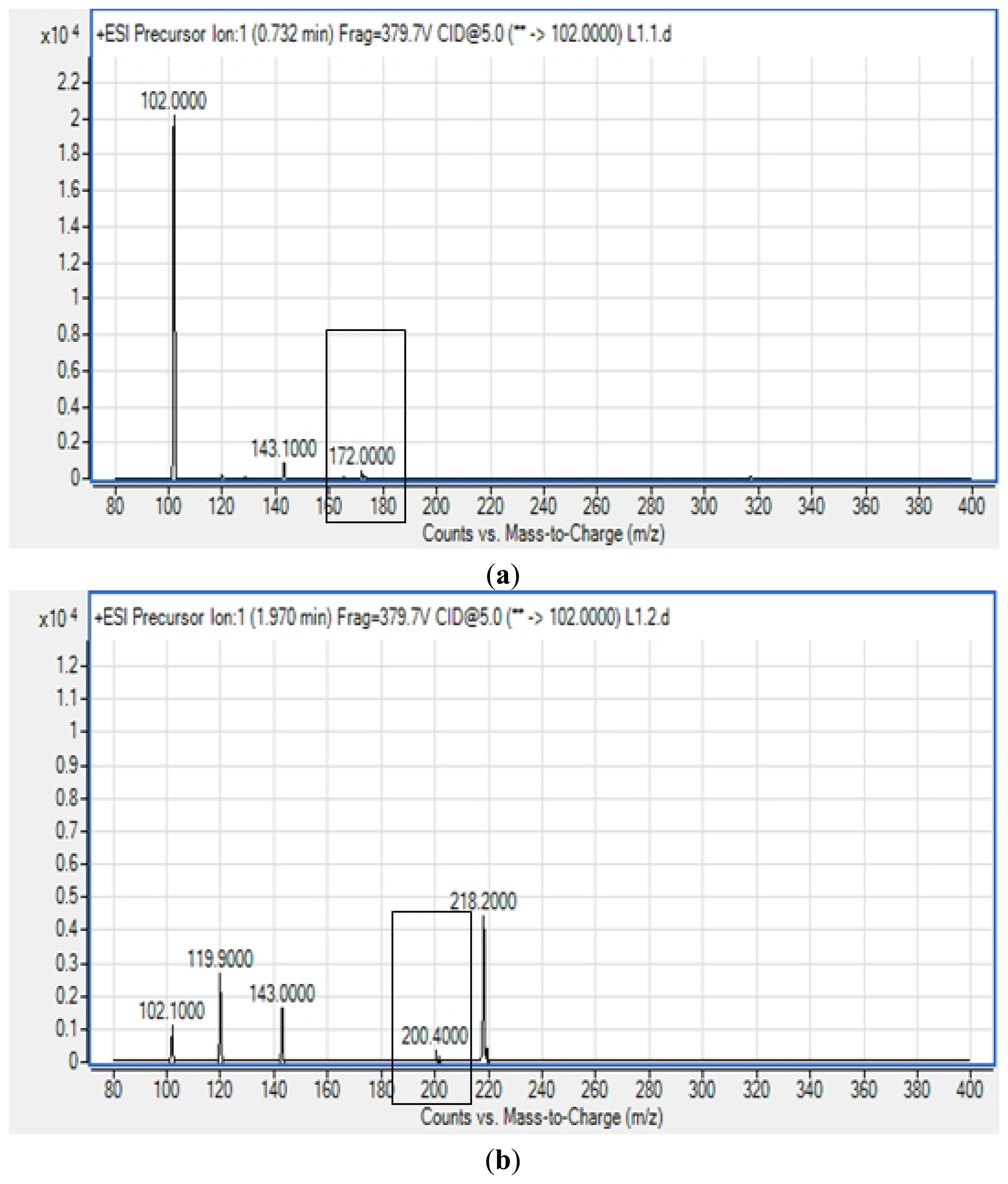
All four isolates of the Batavia lettuce sample were screened for the production of AHLs. Only E. asburiae (L1) was positive for the production of short chain AHLs (Figure 2), which triggered C. violaceum CV026 violacein production. The production of AHLs by the bacterium was further confirmed by using the luminometer-spectrophotometer, whereby the activation of bioluminescence of E. coli [pSB401] was observed (Figure 3). In order to identify the AHLs and to further confirm the production, high resolution triple quadrupole LC/MS system was used. The MS analysis results of the spent culture supernatant of E. asburiae, presented in Figure 4, provided evidence for the presence of C4-HSL (m/z 172.0000) and C6-HSL (m/z 200.4000).
Enterobacteriaceae colonization has been reported in numerous studies and is believed to cause the spoilage of food products [23,24]. Furthermore, members of the family Enterobacteriaceae have been indicated to cause gastrointestinal illnesses around the World and these outbreaks have been commonly connected to consumption of contaminated vegetable and fruit products [9,25]. This relation of both food spoilage and food-borne illnesses with enteric bacteria has caused an increase in multidisciplinary interest in researching the production of signaling molecules to better understand how these interactions may affect food safety. Hence, in this study, we focused on the isolation of the bacterial community from the Enterobacteriaceae family in fresh vegetable produce, specifically Batavia lettuce, which is commonly served as a garden salad.
We have isolated four different enteric bacteria from the lettuce sample. Members of the Enterobacteriaceae have been described as having AHL-mediated gene regulation [10,26]. This could have an important role for the food-borne Enterobacteriaceae community in terms of regulating the expressions of phenotypes and cellular responses such as secondary metabolite production, toxin production, competence, plasmid transfer and biofilm production [27–29]. Although Morganella morganii was isolated in this study and is a bacterium known to cause food spoilage and toxicity [30], it was found that this isolate did not produce any detectable AHLs, and among all the isolates tested in this work, only E. asburiae produced AHLs.
E. asburiae has been reported to cause competition to the growth of human pathogens Salmonella enterica and Escherichia coli O157:H7 [13]. Additionally, as reported previously the presence of E. asburiae could inhibit the growth of S. enterica serovar Newport and E. coli O157:H7 by more than 10- to 100-fold [9,31]. The system(s) involved in this antagonistic communication is still unclear. However, we speculate that the signaling molecules produced by E. asburiae in this study, namely C4-HSL and C6-HSL may play a part in inhibiting the proliferation of the surrounding enteric bacteria. Perhaps the AHLs regulate the production of exoenzymes or secondary metabolites that could act as an antagonists against other microorganisms. However, further study is required to verify the function of these AHLs produced by E. asburiae and their role in interspecies microbial growth suppression.
Recently, there is much interest in exploring QS as a novel anti-infectious therapy [32,33] because since it does not involve the use of antibiotics, theoretically this will reduce the drug resistance problem [32]. The discovery of QS in our E. asburiae isolate will allow the investigation of QS-mediated gene expression in this bacterium and also the development of novel anti-QS molecules [34–41].
4. Conclusions/Outlook
This study confirmed the production of AHLs in E. asburiae isolated from lettuce. The identified AHLs could be implicated in the regulation of physiological activities important in the survival of E. asburiae or other enteric microorganisms. Further studies are required to determine the importance and functions of these signaling molecules in the hope of enhancing produce safety and elucidating the mechanisms of QS-regulation in E. asburiae.
Acknowledgments
This work was supported by the University of Malaya for High Impact Research Grant (UM-MOHE HIR Grant No. A000001-50001) awarded to Kok-Gan Chan.
Conflict of Interest
The authors declare no conflict of interest.
References
- Smith, R.S.; Iglewski, B.H. P. aeruginosa quorum-sensing systems and virulence. Curr. Opin. Microbiol. 2003, 6, 56–60. [Google Scholar]
- Miller, M.B.; Bassler, B.L. Quorum sensing in bacteria. Ann. Rev. Microbiol. 2001, 55, 165–199. [Google Scholar]
- Pearson, J.P.; van Delden, C.; Iglewski, B.H. Active efflux and diffusion are involved in transport of Pseudomonas aeruginosa cell-to-cell signals. J. Bacteriol. 1999, 181, 1203–1210. [Google Scholar]
- Swift, S.; Karlyshev, A.V.; Fish, L.; Durant, E.L.; Winson, M.K.; Chhabra, S.R.; Williams, P.; Macintyre, S.; Stewart, G. Quorum sensing in Aeromonas hydrophila and Aeromonas salmonicida: Identification of the LuxRI homologs AhyRI and AsaRI and their cognate N-acylhomoserine lactone signal molecules. J. Bacteriol. 1997, 179, 5271–5281. [Google Scholar]
- Liao, C.-H. Analysis of pectate lyases produced by soft rot bacteria associated with spoilage of vegetables. Appl. Envir. Microbiol. 1989, 55, 1677–1683. [Google Scholar]
- Liao, C.H.; Sullivan, J.; Grady, J.; Wong, L.J. Biochemical characterization of pectate lyases produced by fluorescent pseudomonads associated with spoilage of fresh fruits and vegetables. J. Appl. Microbiol. 1997, 83, 10–16. [Google Scholar]
- Mead, P.S.; Slutsker, L.; Dietz, V.; McCaig, L.F.; Bresee, J.S.; Shapiro, C.; Griffin, P.M.; Tauxe, R.V. Food-related illness and death in the United States. Emerg. Infect. Dis. 1999, 5, 607–625. [Google Scholar]
- Friedemann, M. Enterobacter sakazakii in food and beverages (other than infant formula and milk powder). Int. J. Food Microbiol. 2007, 116, 1–10. [Google Scholar]
- Teplitski, M.; Warriner, K.; Bartz, J.; Schneider, K.R. Untangling metabolic and communication networks: Interactions of enterics with phytobacteria and their implications in produce safety. Trends Microbiol. 2011, 19, 121–127. [Google Scholar]
- Gram, L.; Christensen, A.B.; Ravn, L.; Molin, S.; Givskov, M. Production of acylated homoserine lactones by psychrotrophic members of the Enterobacteriaceae isolated from foods. Appl. Environ. Microbiol. 1999, 65, 3458–3463. [Google Scholar]
- Koth, K.; Boniface, J.; Chance, E.A.; Hanes, M.C. Enterobacter asburiae and Aeromonas hydrophila: Soft tissue infection requiring debridement. Orthopedics 2012, 35, 996–999. [Google Scholar]
- Asis, C.; Adachi, K. Isolation of endophytic diazotroph Pantoea agglomerans and nondiazotroph Enterobacter asburiae from sweetpotato stem in Japan. Lett. Appl. Microbiol. 2004, 38, 19–23. [Google Scholar]
- Cooley, M.B.; Chao, D.; Mandrell, R.E. Escherichia coli O157: H7 survival and growth on lettuce is altered by the presence of epiphytic bacteria. J. Food Protect. 2006, 69, 2329–2335. [Google Scholar]
- Gnanamanickam, S.S.; Immanuel, J.E. Epiphytic Bacteria, Their Ecology Functions. In Plant-Associated Bacteria; Springer: Houten, The Netherlands, 2006; pp. 131–153. [Google Scholar]
- Rezzonico, F.; Smits, T.H.; Duffy, B. Detection of AI-2 receptors in genomes of Enterobacteriaceae suggests a role of type-2 quorum sensing in closed ecosystems. Sensors 2012, 12, 6645–6665. [Google Scholar]
- Seng, P.; Drancourt, M.; Gouriet, F.; La Scola, B.; Fournier, P.-E.; Rolain, J.M.; Raoult, D. Ongoing revolution in bacteriology: Routine identification of bacteria by matrix-assisted laser desorption ionization time-of-flight mass spectrometry. Clin. Infect. Dis. 2009, 49, 543–551. [Google Scholar]
- McClean, K.H.; Winson, M.K.; Fish, L.; Taylor, A.; Chhabra, S.R.; Camara, M.; Daykin, M.; Lamb, J.H.; Swift, S.; Bycroft, B.W.; et al. Quorum sensing and Chromobacterium violaceum: Exploitation of violacein production and inhibition for the detection of N-acylhomoserine lactones. Microbiology 1997, 143, 3703–3711. [Google Scholar]
- Wong, C.-S.; Yin, W.-F.; Choo, Y.-M.; Sam, C.-K.; Koh, C.-L.; Chan, K.-G. Coexistence of quorum-quenching and quorum-sensing in tropical marine Pseudomonas aeruginosa strain MW3A. World J. Microbiol. Biotechnol. 2012, 28, 453–461. [Google Scholar]
- Yin, W.-F.; Purmal, K.; Chin, S.; Chan, X.-Y.; Koh, C.-L.; Sam, C.-K.; Chan, K.-G. N-acyl homoserine lactone production by Klebsiella pneumoniae isolated from human tongue surface. Sensors 2012, 12, 3472–3483. [Google Scholar]
- Winson, M.K.; Swift, S.; Fish, L.; Throup, J.P.; Jørgensen, F.; Chhabra, S.R.; Bycroft, B.W.; Williams, P.; Stewart, G.S. Construction and analysis of luxCDABE-based plasmid sensors for investigating N‐acyl homoserine lactone‐mediated quorum sensing. FEMS Microbiol. Lett. 1998, 163, 185–192. [Google Scholar]
- Chen, J.W.; Koh, C.L.; Sam, C.K.; Yin, W.F.; Chan, K.G. Short chain n-acyl homoserine lactone production by soil isolate Burkholderia sp. strain A9. Sensors 2013, 13, 13217–13227. [Google Scholar]
- Reich, M.; Bosshard, P.P.; Stark, M.; Beyser, K.; Borgmann, S. Species identification of bacteria and fungi from solid and liquid culture media by MALDI-TOF mass spectrometry. J. Bacteriol. Parasitol. 2013, 10, 2155–9597. [Google Scholar]
- Borch, E.; Kant-Muermans, M.-L.; Blixt, Y. Bacterial spoilage of meat and cured meat products. Int. J. Food Microbiol. 1996, 33, 103–120. [Google Scholar]
- Bennik, M.; Vorstman, W.; Smid, E.; Gorris, L. The influence of oxygen and carbon dioxide on the growth of prevalent Enterobacteriaceae and Pseudomonas species isolated from fresh and controlled-atmosphere-stored vegetables. Food Microbiol. 1998, 15, 459–469. [Google Scholar]
- Mandrell, R.E. Enteric Human Pathogens Associated with Fresh Produce: Sources, Transport and Ecology. In Microbial Safety of Fresh Produce; IFT Press/Wiley-Blackwell Publishing: Ames, IA, USA, 2009; pp. 5–41. [Google Scholar]
- Bruhn, J.B.; Christensen, A.B.; Flodgaard, L.R.; Nielsen, K.F.; Larsen, T.O.; Givskov, M.; Gram, L. Presence of acylated homoserine lactones (AHLs) and AHL-producing bacteria in meat and potential role of AHL in spoilage of meat. Appl. Environ. Microbiol. 2004, 70, 4293–4302. [Google Scholar]
- Chen, X.; Schauder, S.; Potier, N.; van Dorsselaer, A.; Pelczer, I.; Bassler, B.L.; Hughson, F.M. Structural identification of a bacterial quorum-sensing signal containing boron. Nature 2002, 415, 545–549. [Google Scholar]
- Hentzer, M.; Wu, H.; Andersen, J.B.; Riedel, K.; Rasmussen, T.B.; Bagge, N.; Kumar, N.; Schembri, M.A.; Song, Z.; Kristoffersen, P. Attenuation of Pseudomonas aeruginosa virulence by quorum sensing inhibitors. EMBO J 2003, 22, 3803–3815. [Google Scholar]
- Gambello, M.J.; Iglewski, B.H. Cloning and characterization of the Pseudomonas aeruginosa lasR gene, a transcriptional activator of elastase expression. J. Bacteriol. 1991, 173, 3000–3009. [Google Scholar]
- Klausen, N.K.; Huss, H.H. Growth and histamine production by Morganella morganii under various temperature conditions. Int. J. Food Microbiol. 1987, 5, 147–156. [Google Scholar]
- Cooley, M.B.; Miller, W.G.; Mandrell, R.E. Colonization of Arabidopsis thaliana with Salmonella enterica and enterohemorrhagic Escherichia coli O157: H7 and competition by Enterobacter asburiae. Appl. Environ. Microbiol. 2003, 69, 4915–4926. [Google Scholar]
- Hong, K.W.; Koh, C.L.; Sam, C.K.; Yin, W.F.; Chan, K.G. Quorum quenching revisited-from signal decays to signalling confusion. Sensors 2012, 12, 4661–4696. [Google Scholar]
- Chan, K.G.; Atkinson, S.; Mathee, K.; Sam, C.K.; Chhabra, S.R.; Cámara, M.; Koh, C.L.; Williams, P. Characterization of N-acylhomoserine lactone-degrading bacteria associated with the Zingiber. officinale(ginger) rhizosphere: Co-existence of quorum quenching and quorum sensing in Acinetobacter. and Burkholderia. BMC Microbiol. 2011, 11. [Google Scholar] [CrossRef]
- Chong, Y.M.; Yin, W.F.; Ho, C.Y.; Mustafa, M.R.; Hadi, A.H.A.; Awang, K.; Narrima, P.; Koh, C.L.; Appleton, D.R.; Chan, K.G. Malabaricone C from Myristica cinnamomea exhibits anti-quorum sensing activity. J. Nat. Prod. 2011, 74, 2261–2264. [Google Scholar]
- Krishnan, T.; Yin, W.F.; Chan, K.G. Inhibition of quorum sensing-controlled virulence factor production in Pseudomonas aeruginosa PAO1 by Ayurveda spice clove (Syzygium. aromaticum) bud extract. Sensors 2012, 12, 4016–4030. [Google Scholar]
- Tan, L.Y.; Yin, W.F.; Chan, K.G. Silencing quorum sensing through extracts of Melicope lunu-ankenda. Sensors 2012, 12, 4339–4351. [Google Scholar]
- Koh, C.L.; Sam, C.K.; Yin, W.F.; Tan, L.Y.; Krishnan, T.; Chong, Y.M.; Chan, K.G. Plant-derived natural products as sources of anti-quorum sensing compounds. Sensors 2013, 13, 6217–6228. [Google Scholar]
- Norizan, S.N.M.; Yin, W.F.; Chan, K.G. Caffeine as a potential quorum sensing inhibitor. Sensors 2013, 13, 5117–5129. [Google Scholar]
- Tan, L.Y.; Yin, W.F.; Chan, K.G. Piper nigrum, Piper betle and Gnetum. gnemon natural food sources with anti-quorum sensing properties. Sensors 2013, 13, 3975–3985. [Google Scholar]
- Wong, C.S.; Yin, W.F.; Choo, Y.M.; Sam, C.K.; Koh, C.L.; Chan, K.G. Coexistence of quorum quenching and quorum sensing in tropical marine Pseudomonas aeruginosa strain MW3A. World J. Microbiol. Biotechnol. 2011, 28, 453–461. [Google Scholar]
- Yin, W.F.; Tung, H.J.; Sam, C.K.; Koh, C.L.; Chan, K.G. Quorum quenching Bacillus sonorensis isolated from soya sauce fermentation brine. Sensors 2012, 12, 4065–4073. [Google Scholar]



© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Lau, Y.Y.; Sulaiman, J.; Chen, J.W.; Yin, W.-F.; Chan, K.-G. Quorum Sensing Activity of Enterobacter asburiae Isolated from Lettuce Leaves. Sensors 2013, 13, 14189-14199. https://doi.org/10.3390/s131014189
Lau YY, Sulaiman J, Chen JW, Yin W-F, Chan K-G. Quorum Sensing Activity of Enterobacter asburiae Isolated from Lettuce Leaves. Sensors. 2013; 13(10):14189-14199. https://doi.org/10.3390/s131014189
Chicago/Turabian StyleLau, Yin Yin, Joanita Sulaiman, Jian Woon Chen, Wai-Fong Yin, and Kok-Gan Chan. 2013. "Quorum Sensing Activity of Enterobacter asburiae Isolated from Lettuce Leaves" Sensors 13, no. 10: 14189-14199. https://doi.org/10.3390/s131014189





