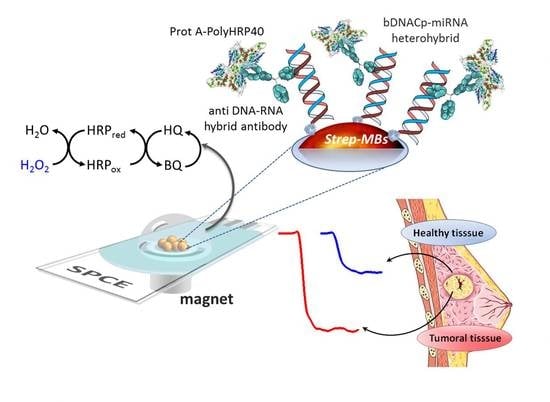
Magnetic Beads-Based Sensor with Tailored Sensitivity for Rapid and Single-Step Amperometric Determination of miRNAs
Abstract
:1. Introduction
2. Results and Discussion
2.1. Optimization of the Experimental Variables
2.2. Analytical Characteristics
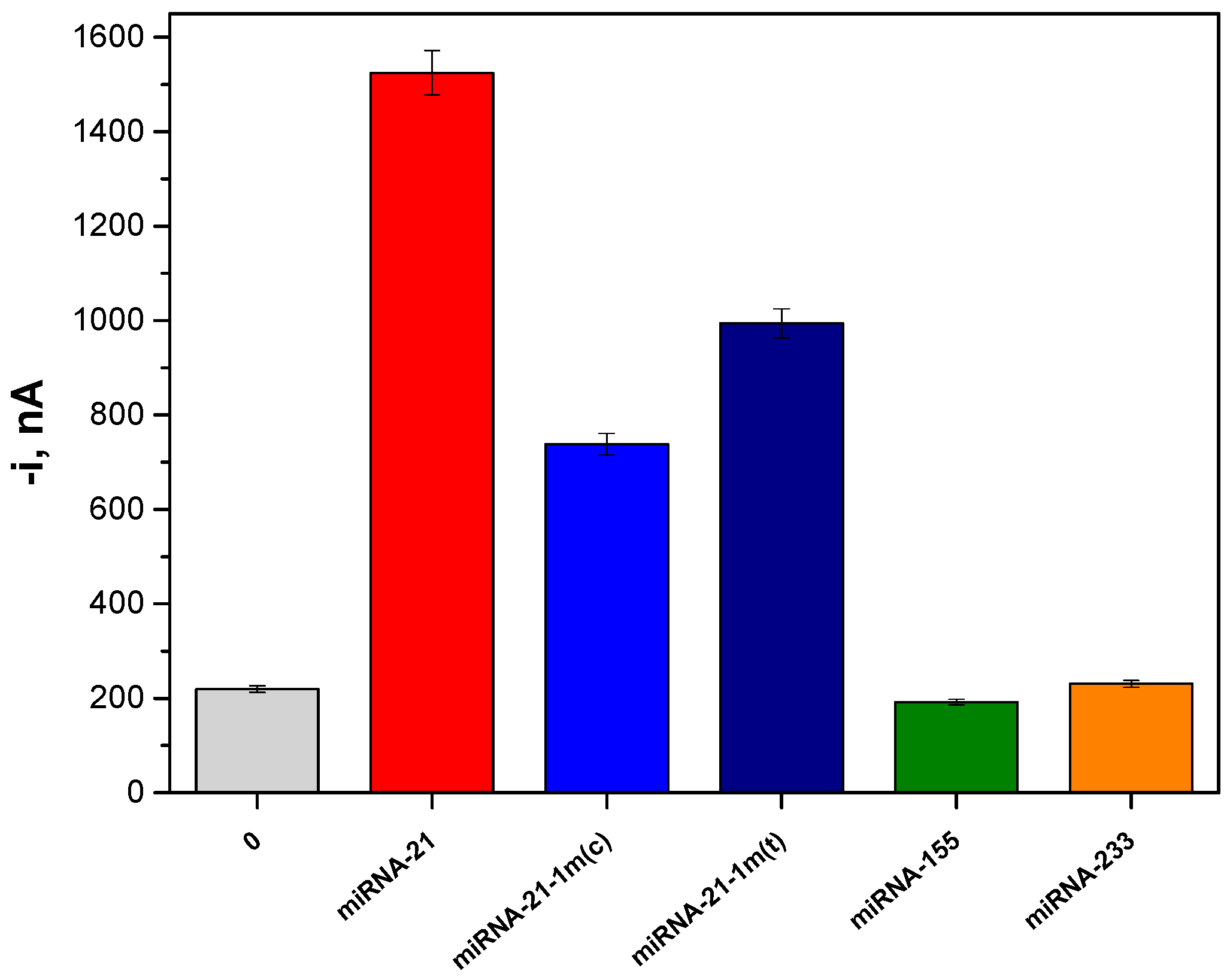
2.3. Selectivity
2.4. Determination of Mature miRNA-21 in RNAt Extracted from Cancer Cells and Tumor Tissues
3. Materials and Methods
3.1. Apparatus and Electrodes
3.2. Reagents and Solutions
3.3. MBs Modification
3.4. Electrochemical Measurements
3.5. Cell Culture, Human Tissues and RNAt Extraction
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Felekkis, K.; Touvana, E.; Stefanou, Ch.; Deltas, C. MicroRNAs: A newly described class of encoded molecules that play a role in health and disease. Hippokratia 2010, 14, 236–240. [Google Scholar] [PubMed]
- Hou, T.; Li, W.; Liu, X.; Li, F. Label-free and enzyme-free homogeneous electrochemical biosensing strategy based on hybridization chain reaction: A facile, sensitive, and highly specific microRNA assay. Anal. Chem. 2015, 87, 11368–11374. [Google Scholar] [CrossRef] [PubMed]
- Šípova, H.; Zhang, S.; Dudley, A.M.; Galas, D.; Wang, K.; Homola, J. Surface plasmon resonance biosensor for rapid label-free detection of microribonucleic acid at subfemtomole level. Anal. Chem. 2010, 82, 10110–10115. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Fu, Y.; Wu, Y.; Huang, R.; Zhou, X.; Xing, D. Ultrasensitive detection of microRNA in tumor cells and tissues via continuous assembly of DNA probe. Biomacromolecules 2015, 16, 3543–3551. [Google Scholar] [CrossRef] [PubMed]
- Válóczi, A.; Hornyik, C.; Varga, N.; Burgyán, J.; Kauppinen, S.; Havelda, Z. Sensitive and specific detection of microRNAs by northern blot analysis using LNA-modified oligonucleotide probes. Nucleic Acids Res. 2004, 32, e175. [Google Scholar] [CrossRef] [PubMed]
- Pall, G.S.; Codony-Servat, C.; Byrne, J.; Ritchie, L.; Hamilton, A. Carbodiimide-mediated cross-linking of RNA to nylon membranes improves the detection of siRNA, miRNA and piRNA by northern blot. Nucleic Acids Res. 2007, 35, e60. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Ridzon, D.A.; Broomer, A.J.; Zhou, Z.; Lee, D.H.; Nguyen, J.T.; Barbisin, M.; Xu, N.L.; Mahuvakar, V.R.; Andersen, M.R.; et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res. 2005, 33, e179. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Yao, B.; Huang, H.; Wang, Z.; Sun, C.; Fan, Y.; Chang, Q.; Li, S.; Wang, X.; Xi, J. Real-time polymerase chain reaction microRNA detection based on enzymatic stem-loop probes ligation. Anal. Chem. 2009, 81, 5446–5451. [Google Scholar] [CrossRef] [PubMed]
- Kroh, E.M.; Parkin, R.K.; Mitchell, P.S.; Tewari, M. Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods 2010, 50, 298–301. [Google Scholar] [CrossRef] [PubMed]
- Nelson, P.T.; Baldwin, D.A.; Scearce, L.M.; Oberholtzer, J.C.; Tobias, J.W.; Mourelatos, Z. Microarray-based, high-throughput gene expression profiling of microRNAs. Nat. Methods 2004, 1, 155–161. [Google Scholar] [CrossRef] [PubMed]
- Liang, R.-Q.; Li, W.; Li, Y.; Tan, C.; Li, J.X.; Jin, Y.X.; Ruan, K.C. An oligonucleotide microarray for microRNA expression analysis based on labeling RNA with quantum dot and nanogold probe. Nucleic Acids Res. 2005, 33, e17. [Google Scholar] [CrossRef] [PubMed]
- Git, A.; Dvinge, H.; Salmon-Divon, M.; Osborne, M.; Kutter, C.; Hadfield, J.; Bertone, P.; Caldas, C. Systematic comparison of microarray profiling, real-time PCR, and next-generation sequencing technologies for measuring differential microRNA expression. RNA 2010, 16, 991–1006. [Google Scholar] [CrossRef] [PubMed]
- Xia, N.; Zhang, L.; Wang, G.; Feng, Q.; Liu, L. Label-free and sensitive strategy for microRNAs detection based on the formation of boronate ester bonds and the dual-amplification of gold nanoparticles. Biosens. Bioelectron. 2013, 47, 461–466. [Google Scholar] [CrossRef] [PubMed]
- Yao, B.; Liu, Y.; Tabata, M.; Zhu, H.; Miyahara, Y. Sensitive detection of microRNA by chronocoulometry and rolling circle amplification on a gold electrode. Chem. Commun. 2014, 50, 9704–9706. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.; Jiang, J.; Yu, R. A sensitive electrochemical biosensor for microRNA detection based on streptavidin-gold nanoparticles and enzymatic amplification. Anal. Methods 2014, 6, 2889–2893. [Google Scholar] [CrossRef]
- Liu, L.; Xia, N.; Liu, H.; Kang, X.; Liu, X.; Xue, C.; He, X. Highly sensitive and label-free electrochemical detection of microRNAs based on triple signal amplification of multifunctional gold nanoparticles, enzymes and redox-cycling reaction. Biosens. Bioelectron. 2014, 53, 399–405. [Google Scholar] [CrossRef] [PubMed]
- Xia, N.; Zhang, Y.; Wei, X.; Huang, Y.; Liu, L. An electrochemical microRNAs biosensor with the signal amplification of alkaline phosphatase and electrochemical-chemical-chemical redox cycling. Anal. Chim. Acta 2015, 878, 95–101. [Google Scholar] [CrossRef] [PubMed]
- Miao, P.; Wang, B.; Meng, F.; Yin, J.; Tang, Y. Ultrasensitive detection of microRNA through rolling circle amplification on a DNA tetrahedron decorated electrode. Bioconjug. Chem. 2015, 26, 602–607. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Chai, Y.; Zhang, P.; Yuan, R. An electrochemical biosensor for sensitive detection of microRNA-155: Combining target recycling with cascade catalysis for signal amplification. ACS Appl. Mater. Interfaces 2015, 7, 713–720. [Google Scholar] [CrossRef] [PubMed]
- Hu, T.; Zhang, L.; Wen, W.; Zhang, X.; Wang, S. Enzyme catalytic amplification of miRNA-155 detection with graphene quantum dot-based electrochemical biosensor. Biosens. Bioelectron. 2016, 77, 451–456. [Google Scholar] [CrossRef] [PubMed]
- Ren, Y.; Deng, H.; Shen, W.; Gao, Z. Highly sensitive and selective electrochemical biosensor for direct detection of microRNAs in serum. Anal. Chem. 2013, 85, 4784–4789. [Google Scholar] [CrossRef] [PubMed]
- Meng, X.; Zhou, Y.; Liang, Q.; Qu, X.; Yang, Q.; Yin, H.; Ai, S. Electrochemical determination of microRNA-21 based on bio bar code and hemin/G-quadruplet DNA enzyme. Analyst 2013, 138, 3409–3415. [Google Scholar] [CrossRef] [PubMed]
- Jolly, P.; Batistuti, M.R.; Miodek, A.; Zhurauski, P.; Mulato, M.; Lindsay, M.A.; Estrela, P. Highly sensitive dual mode electrochemical platform for microRNA detection. Sci. Rep. 2016, 6, 36719. [Google Scholar] [CrossRef] [PubMed]
- Campuzano, S.; Torrente-Rodríguez, R.M.; López-Hernández, E.; Conzuelo, F.; Granados, R.; Sánchez-Puelles, J.M.; Pingarrón, J.M. Magnetobiosensors based on viral protein p19 for microRNA determination in cancer cells and tissues. Angew. Chem. Int. Ed. 2014, 53, 6168–6171. [Google Scholar] [CrossRef] [PubMed]
- Torrente-Rodríguez, R.M.; Ruiz-Valdepeñas Montiel, V.; Campuzano, S.; Fachardo-Dinia, M.; Barderas, R.; San Segundo-Acosta, P.; Montoya, J.J.; Pingarrón, J.M. Fast electrochemical miRNAs determination in cancer cells and tumor tissues with antibody-functionalized magnetic microcarriers. ACS Sens. 2016, 1, 896–903. [Google Scholar] [CrossRef]
- Campuzano, S.; Yánez-Sedeño, P.; Pingarrón, J.M. Electrochemical biosensing of microribonucleic acids using antibodies and viral proteins with affinity for ribonucleic acid duplexes. Electrochim. Acta 2017, 230, 271–278. [Google Scholar] [CrossRef]
- Conzuelo, F.; Gamella, M.; Campuzano, S.; Reviejo, A.J.; Pingarrón, J.M. Disposable amperometric magneto-immunosensor for direct detection of tetracyclines antibiotics residues in milk. Anal. Chim. Acta 2012, 737, 29–36. [Google Scholar] [CrossRef] [PubMed]
- Qavi, A.J.; Kindt, J.T.; Gleeson, M.A.; Bailey, R.C. Anti-DNA: RNA antibodies and silicon photonic microring resonators: Increased sensitivity for multiplexed microRNA detection. Anal. Chem. 2011, 83, 5949–5956. [Google Scholar] [CrossRef] [PubMed]
- Novara, C.; Chiadò, A.; Paccotti, N.; Catuogno, S.; Esposito, C.L.; Condorelli, G.; de Franciscis, V.; Geobaldo, F.; Rivolo, P.; Giorgis, F. SERS-active metal-dielectric nanostructures integrated in microfluidic devices for label-free quantitative detection of miRNA. Faraday Discuss. 2017. [Google Scholar] [CrossRef] [PubMed]
- Si, M.-L.; Zhu, S.; Wu, H.; Lu, Z.; Wu, F.; Mo, Y.-Y. miR-21-mediated tumor growth. Oncogene 2007, 26, 2799–2803. [Google Scholar] [CrossRef] [PubMed]
- Elgamal, O.A.; Park, J.-K.; Gusev, Y.; Azevedo-Pouly, A.C.P.; Jiang, J.; Roopra, A.; Schmittgen, T.D. Tumor suppressive function of miR-205 in breast cancer is linked to HMGB3 regulation. PLoS ONE 2013, 8, e76402. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Feng, J.; Liu, H.; Li, Q.; Tong, L.; Tang, B. Two-color imaging of microRNA with enzyme-free signal amplification via hybridization chain reactions in living cells. Chem. Sci. 2016, 7, 1940–1945. [Google Scholar] [CrossRef]
- Yan, L.-X.; Huang, X.-F.; Shao, Q.; Huang, M.-Y.; Deng, L.; Wu, Q.-L.; Zeng, Y.-X.; Shao, J.-Y. MicroRNA miR-21 overexpression in human breast cancer is associated with advanced clinical stage, lymph node metastasis and patient poor prognosis. RNA 2008, 14, 2348–2360. [Google Scholar] [CrossRef] [PubMed]
- Vetrone, S.A.; Huarng, M.C.; Alocilja, E.C. Detection of non-PCR amplified S. enteritidis genomic DNA from food matrices using a gold-nanoparticle DNA biosensor: A proof-of-concept study. Sensors 2012, 12, 10487–10499. [Google Scholar] [CrossRef] [PubMed]
- Xu, F.; Yang, T.; Chen, Y. Quantification of microRNA by DNA−peptide probe and liquid chromatography-tandem mass spectrometry-based quasi-targeted proteomics. Anal. Chem. 2016, 88, 754–763. [Google Scholar] [CrossRef] [PubMed]
- Torrente-Rodríguez, R.M.; Campuzano, S.; López-Hernández, E.; Granados, R.; Sánchez-Puelles, J.M.; Pingarrón, J.M. Direct determination of miR-21 in total RNA extracted from breast cancer samples using magnetosensing platforms and the p19 viral protein as detector bioreceptor. Electroanalysis 2014, 26, 2080–2087. [Google Scholar] [CrossRef]


| Experimental Variable | Tested Range | Selected Value |
|---|---|---|
| [b-DNACp], μM | 0.0–1.0 | 0.1 |
| tb-DNACp incubation, min | 0–60 | 30 |
| Strep-MBs, μL | 2.5–10.0 | 5.0 |
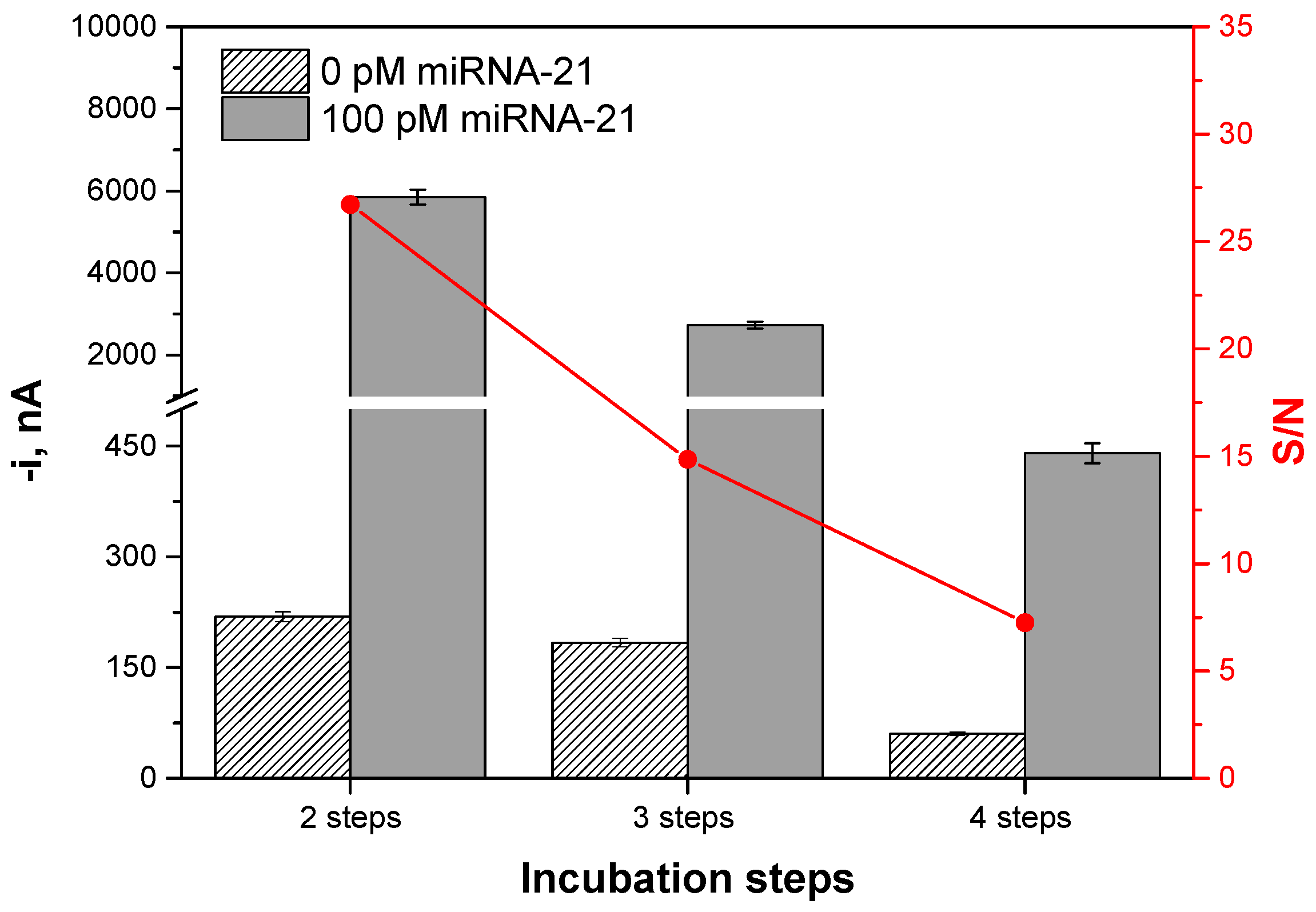
| Number of incubation steps | 2–4 | 2 |
| Anti-DNA-RNA hybrid antibody dilution | 1:100–1:10,000 | 1:1000 |
| ProtA–PolyHRP40 dilution | 1:5–1:250 | 1:25 |
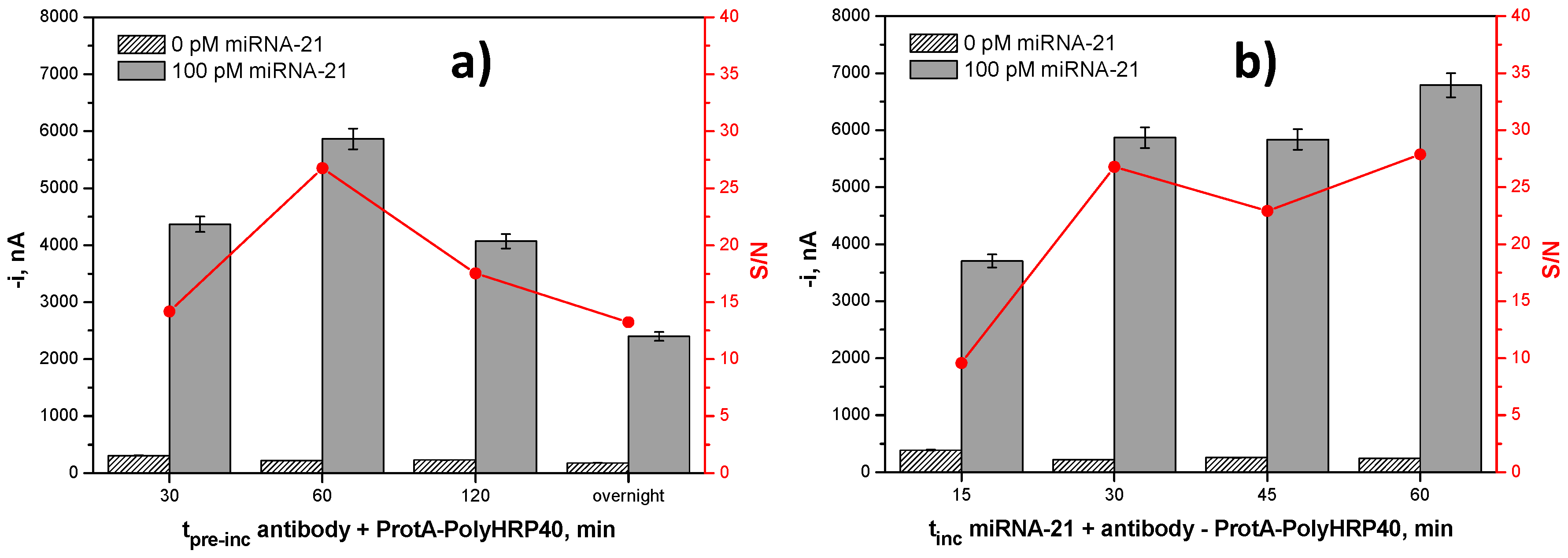
| tProtA–HRP40 + AbS9.6 pre-incubation, min | 30–overnight | 60 |
| tmixture solution incubation, min | 15–60 | 30 |
| Incubation Steps (30 min Each) | Protocol | Total Assay Time, min |
|---|---|---|
| 2 | (1) b-DNACp immobilization onto Strep-MBs→b-DNACp-MBs; (2) Simultaneous hybridization of the target miRNA with the b-DNACp-MBs and labeling of the b-DNACp–miRNA heteroduplexes, immobilized onto the MBs with an anti-DNA–RNA hybrid antibody and ProtA–PolyHRP40 mixture solution pre-incubated for 1 h. | 60 |
| 3 | (1) b-DNACp immobilization onto Strep-MBs→b-DNACp-MBs; (2) Hybridization of the target miRNA with the b-DNACp-MBs; (3) Recognition of the b-DNACp–miRNA heteroduplexes immobilized onto the MBs by the anti DNA–RNA hybrid antibody and ProtA–PolyHRP40 mixture solution. | 90 |
| 4 | (1) b-DNACp immobilization onto Strep-MBs→b-DNACp-MBs; (2) Hybridization of the target miRNA with the b-DNACp-MBs; (3) Recognition of the b-DNACp–miRNA heteroduplexes immobilized onto the MBs by the anti DNA-RNA hybrid antibody; (4) Labeling of the anti DNA–RNA hybrid antibody with ProtA–PolyHRP40. | 120 |
| Parameter | Value |
|---|---|
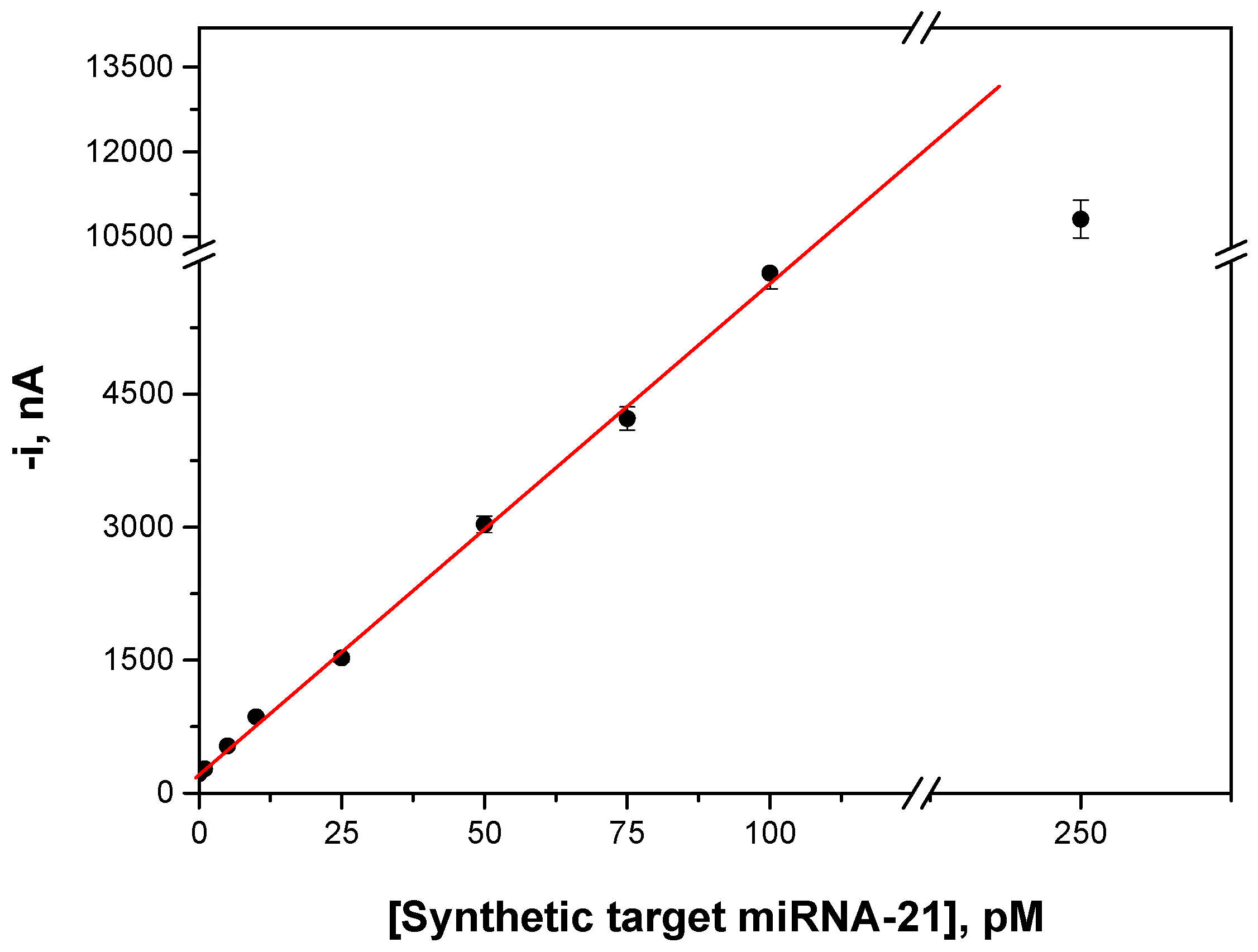
| R * | 0.9995 |
| Slope, nA·pM−1 | 55.3 ± 0.9 |
| Intercept, nA | 228 ± 45 |
| Linear range, pM | 1.0–100 |
| Limit of detection (LOD) **, pM | 0.4 |
| Limit of quantification (LQ) ***, pM | 1.0 |
| b-DNACp-MBs-Mixture Solution Incubation Time, min | [Anti DNA–RNA Hybrid Antibody], µg·mL−1 | [ProtA–HRP40], μg·mL−1 | Slope, nA·nM−1 | Sensitivity, % |
|---|---|---|---|---|
| 30 | 2.0 | 2.0 | 55,214 ± 921 | 100 |
| 15 | 2.0 | 2.0 | 34,843 ± 2542 | 63.1 |
| 4.0 | 2.0 | 30,844 ± 4494 | 55.9 | |
| 2.0 | 4.0 | 53,899 ± 659 | 97.6 | |
| 4.0 | 4.0 | 48,862 ± 4269 | 88.5 |
| Sample | miRNA-21 (n = 3) | T/NT * Ratio | Contents Found by Other Authors | |
|---|---|---|---|---|
| Cells | MCF-10A | 0.79 ± 0.14 | 2.95 | 0.93 [35] |
| 1.02 [25] | ||||
| MCF-7 | 2.33 ± 0.54 | 3.3 [35] | ||
| 3.1 [25] | ||||
| Breast tissues | NT1 | 0.94 ± 0.12 | 2.06–2.99 | 0.1−1.5 [35] |
| NT2 | 0.80 ± 0.20 | |||
| T1 | 2.81 ± 0.43 | 0.4−3.0 [35] | ||
| T2 | 1.65 ± 0.20 | |||
| Oligonucleotide | Sequence (5′→3′) |
|---|---|
| b-antiDNA-21 Cp | 5′-TCAACATCAGTCTGATAAGCTA-Biotin-3′ |
| Target miRNA-21 | 5′-UAGCUUAUCAGACUGAUGUUGA-3′ |
| 1-central base mismatched miRNA-21 (miRNA-21 1-m(c)) | 5′-UAGCUUAUCAAACUGAUGUUGA-3′ |
| 1-terminal base mismatched miRNA-21 (miRNA-21 1-m(t)) | 5′-UAGCUUAUCAGACUGAUGUUGG-3′ |
| miRNA-155 | 5′-UUAAUGCUAAUCGUGAUAGGGGU-3′ |
| miRNA-223 | 5′-CGUGUAUUUGACAAGCUGAGUU-3′ |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vargas, E.; Torrente-Rodríguez, R.M.; Ruiz-Valdepeñas Montiel, V.; Povedano, E.; Pedrero, M.; Montoya, J.J.; Campuzano, S.; Pingarrón, J.M. Magnetic Beads-Based Sensor with Tailored Sensitivity for Rapid and Single-Step Amperometric Determination of miRNAs. Int. J. Mol. Sci. 2017, 18, 2151. https://doi.org/10.3390/ijms18112151
Vargas E, Torrente-Rodríguez RM, Ruiz-Valdepeñas Montiel V, Povedano E, Pedrero M, Montoya JJ, Campuzano S, Pingarrón JM. Magnetic Beads-Based Sensor with Tailored Sensitivity for Rapid and Single-Step Amperometric Determination of miRNAs. International Journal of Molecular Sciences. 2017; 18(11):2151. https://doi.org/10.3390/ijms18112151
Chicago/Turabian StyleVargas, Eva, Rebeca M. Torrente-Rodríguez, Víctor Ruiz-Valdepeñas Montiel, Eloy Povedano, María Pedrero, Juan J. Montoya, Susana Campuzano, and José M. Pingarrón. 2017. "Magnetic Beads-Based Sensor with Tailored Sensitivity for Rapid and Single-Step Amperometric Determination of miRNAs" International Journal of Molecular Sciences 18, no. 11: 2151. https://doi.org/10.3390/ijms18112151