Transfer RNA Methyltransferases from Thermoplasma acidophilum, a Thermoacidophilic Archaeon
Abstract
:1. Introduction
2. Results and Discussion
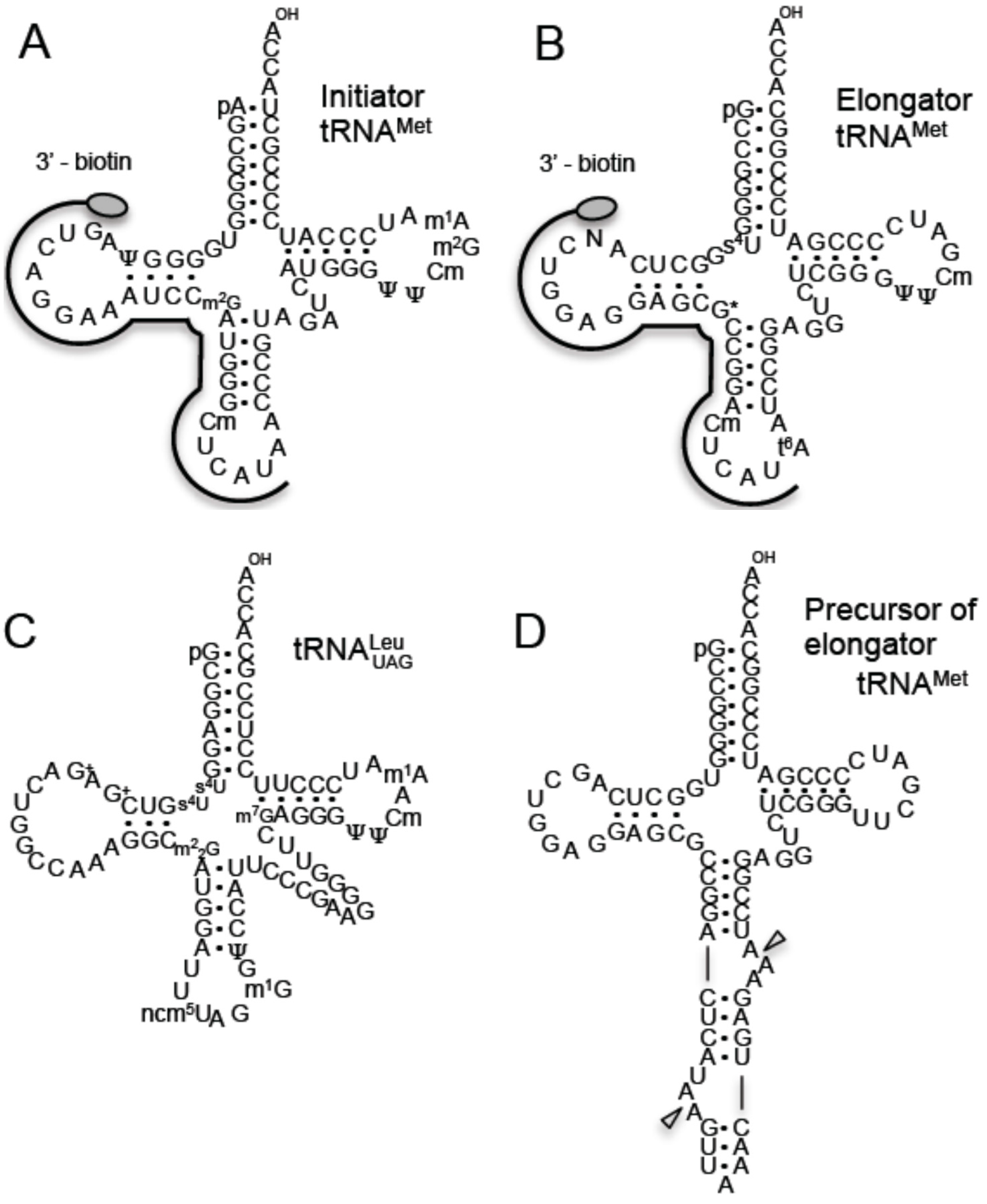
2.1. Methylated Nucleosides in T. acidophilum tRNAs
| Modification and Position | Candidate Enzyme and Gene | Results from This Study | |
|---|---|---|---|
| Intiator tRNAMet | |||
| m2G26 | Trm1 | Ta0997 | m22G26 and m2G26 formed by Trm1. |
| Cm32 | TrmJ? | Ta1010m? | 1 We did not analyze this gene product. |
| Cm56 | Trm56 | Ta0931 | Cm56 formed by Trm56. |
| m2G57 | ? | 2 This position in the tRNA gene is A57. | |
| m1A58 | TrmI | Ta0852? | 3 We could not detect m1A58 formation activity in the recombinant protein. |
| Elongator tRNAMet | |||
| G*26 | Trm1 | Ta0997 | m22G26 and m2G26 formed by Trm1. |
| Cm32 | TrmJ? | Ta1010m? | 1 We did not analyze this gene product. |
| Cm56 | Trm56 | Ta0931 | Cm56 formed by Trm56. |
| tRNALeuUAG | |||
| m22G26 | Trm1 | Ta0997 | m22G26 formed by Trm1. |
| m1G37 | Trm5 | Ta0836 | m1G37 formed by Trm5. |
| m7G49 | ? | Ta0679? +α? | We could not obtain soluble recombinant protein. |
| Cm56 | Trm56 | Ta0931 | Cm56 formed by Trm56. |
| m1A58 | TrmI | Ta0852? | 3 We could not detect m1A58 formation activity in the recombinant protein. |
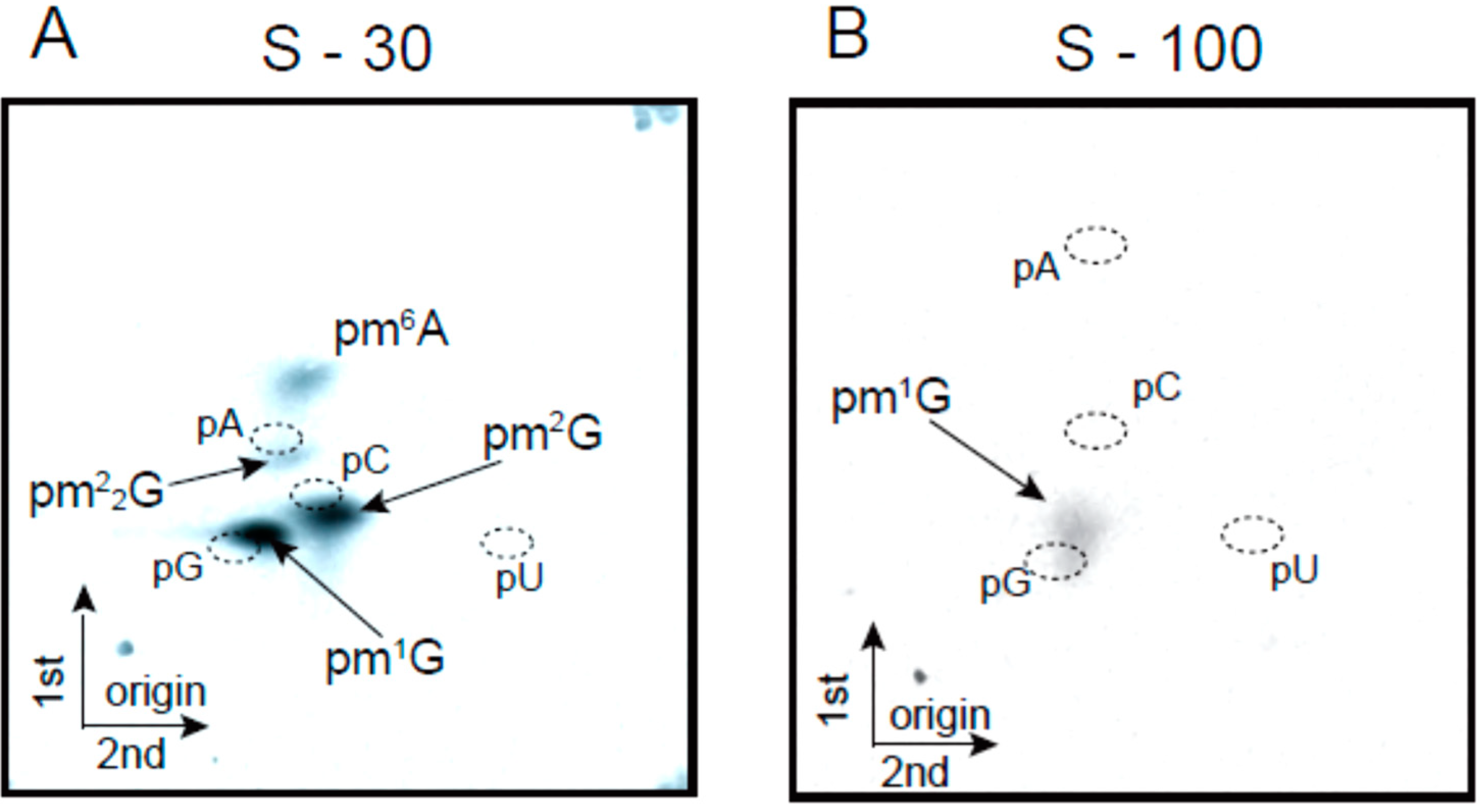
2.2. Transfer RNA Methyltransferase Activities in the Crude Cell Extract
2.3. Modified Nucleosides in Purified tRNAs

2.4. Formation of m1G37 in tRNALeuUAG Transcript by Ta0836 Gene Product

2.5. Ta0997 Gene Product Is a Single Site-Specific Trm1
2.6. T. acidophilum Trm1 Can Modify G26 in Initiator tRNAMet Transcript to m22G26 via m2G26

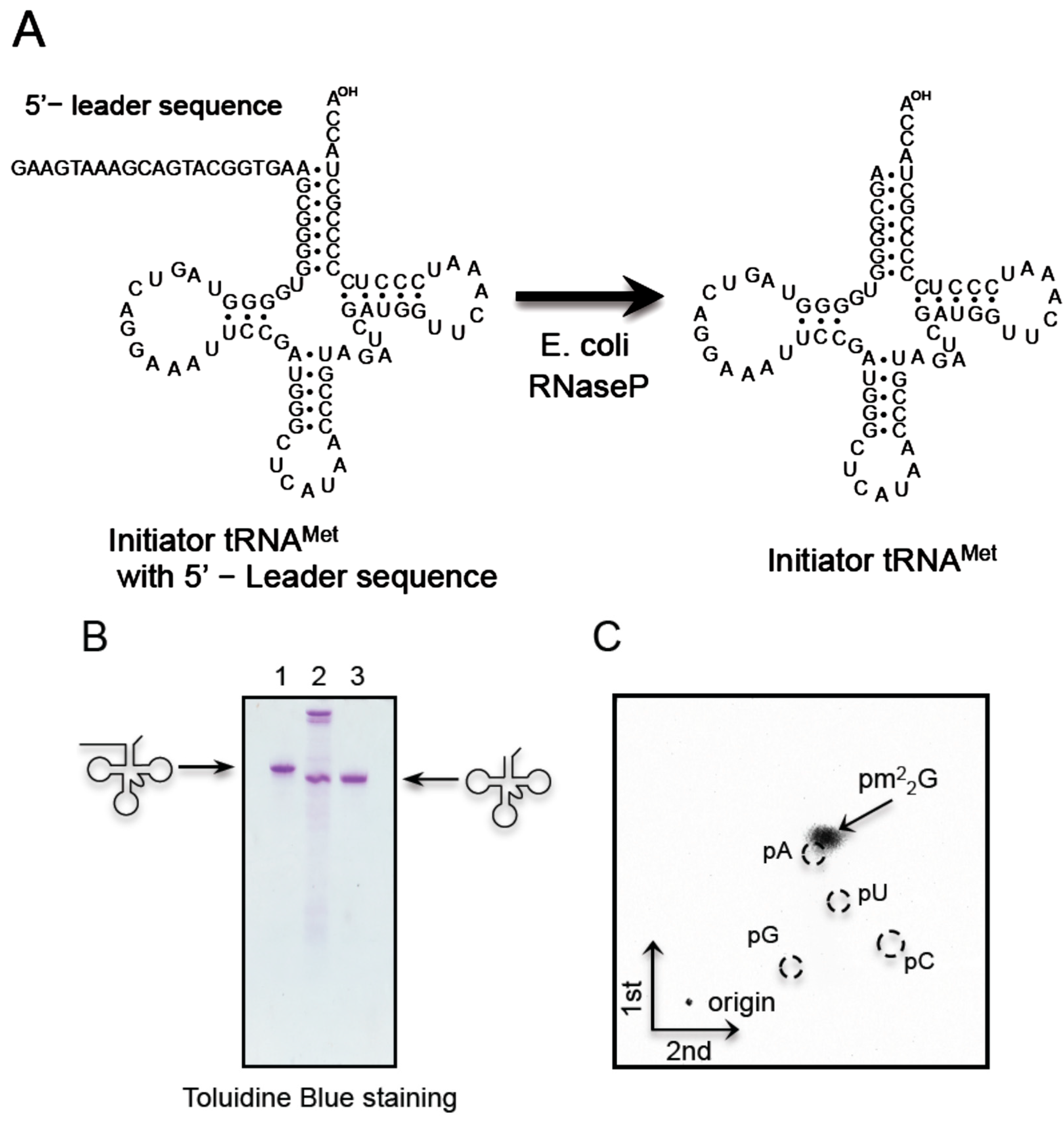
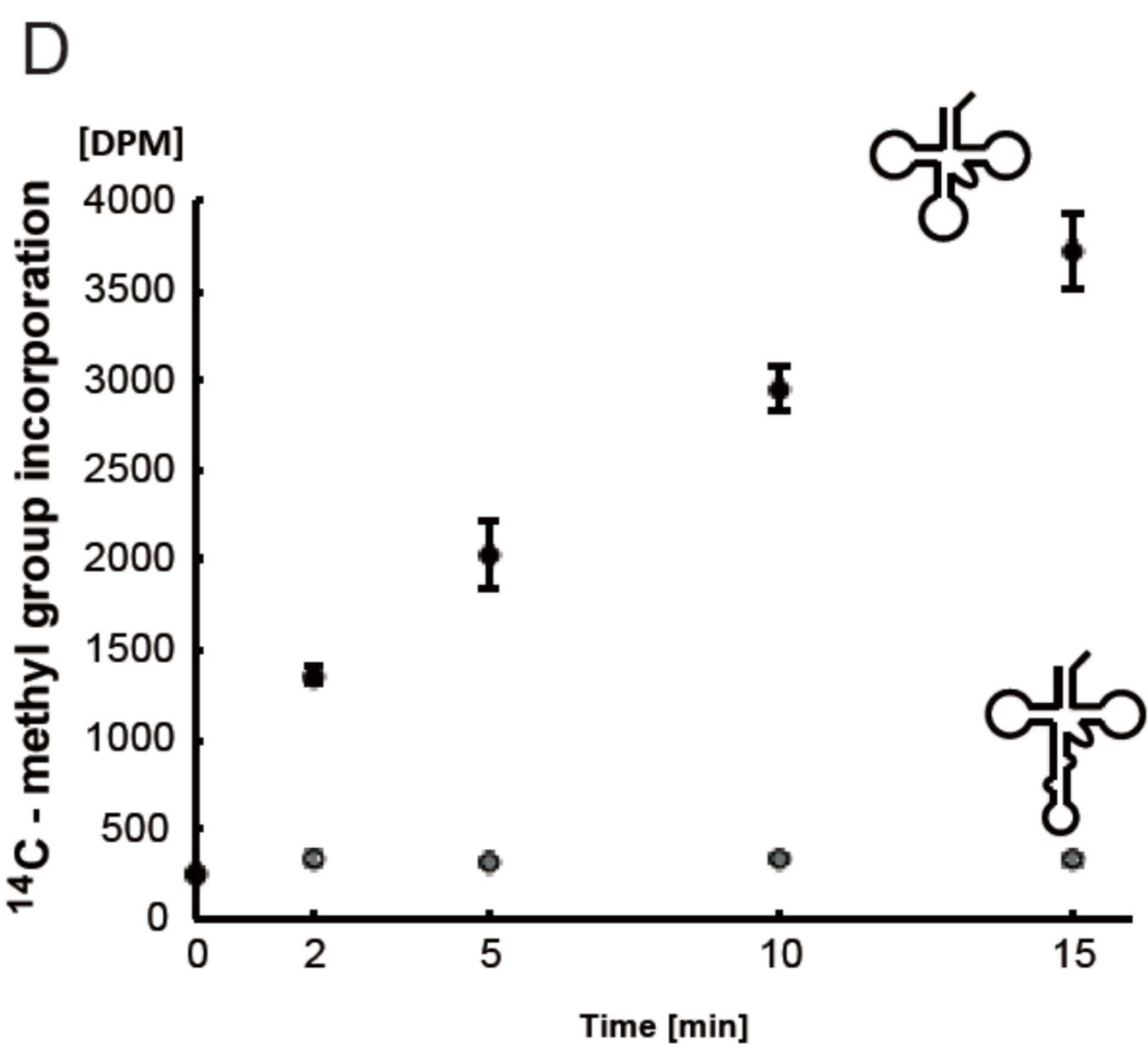
2.7. T. acidophilum Trm1 Can Methylate the Precursor of Elongator tRNAMet with an Intron

2.8. Ta0931 Gene Product Is Trm56

3. Discussion
4. Experimental Section
4.1. Materials
4.2. Strain, Medium, and Culture
4.3. Preparation of S-30 and S-100 Fractions, and Detection of tRNA Methyltransferase Activities
4.4. Purification of Initiator and Elongator tRNAMet by the Solid-Phase DNA Probe Method
4.5. Nucleoside Analysis
4.6. Selection of Candidate Genes
4.7. Cloning of the Candidate Genes, and Expression of Gene Products
4.7.1. Cloning of Ta0836 (trm5)
4.7.2. Cloning of Ta0997 (trm1)
4.7.3. Cloning of Ta0931 (trm56)
4.7.4. Expression of Gene Products
4.8. Purification of Recombinant Proteins
4.8.1. Purification of Trm5
4.8.2. Purification of Trm1
4.8.3. Purification of Trm56
4.9. Measurement of tRNA Methyltransferase Activities
4.10. Preparation of E. coli RNase P and Removal of 5'-Leader Sequence
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Darland, G.; Brock, T.D.; Samsonoff, W.; Conti, S.F. A thermophilic, acidophilic mycoplasma isolated from a coal refuse pile. Science 1970, 170, 1416–1418. [Google Scholar] [CrossRef] [PubMed]
- Mayberry-Carson, K.J.; Roth, I.L.; Harris, J.L.; Smith, P.F. Scanning electron microscopy of Thermoplasma acidophilum. J. Bactriol. 1974, 120, 1472–1475. [Google Scholar]
- Yasuda, M.; Oyaizu, H.; Yamagishi, A.; Oshima, T. Morphological variation of new Thermoplasma acidophilum isolates from Japanese hot springs. Appl. Environ. Microbiol. 1995, 61, 3482–3485. [Google Scholar] [PubMed]
- Shimada, H.; Nemoto, N.; Shida, Y.; Oshima, T.; Yamagishi, A. Complete polar lipid composition of Thermoplasma acidophilum HO-62 determined by high-performance liquid chromatography with evaporative light-scattering detection. J. Bacteriol. 2002, 184, 556–563. [Google Scholar] [CrossRef] [PubMed]
- Posner, M.G.; Upadhyay, A.; Bagby, S.; Hough, D.W.; Danson, M.J. A unique lipoylation system in the Archaea lipoylation in Thermoplamsa acidophilum requires two proteins. FEBS J. 2009, 276, 4012–4022. [Google Scholar] [CrossRef] [PubMed]
- Xu, Q.; Eguchi, T.; Mathews, I.I.; Rife, C.L.; Chiu, H.J.; Farr, C.L.; Feuerhelm, J.; Jaroszewski, L.; Klock, H.E.; Knuth, M.W.; et al. Insights into substrate specificity of geranylgeranyl reductases revealed by the structure of digeranylgeranylglycerophospholipid reductase, an essential enzyme in the biosynthesis of archaeal membrane lipids. J. Mol. Biol. 2010, 404, 403–417. [Google Scholar] [CrossRef] [PubMed]
- Vinogradov, E.; Deschatelets, L.; Lamoureux, M.; Patel, G.B.; Tremblay, T.L.; Robotham, A.; Goneau, M.F.; Cummings-Lorbetskie, C.; Watson, D.C.; Brisson, J.R.; et al. Cell surface glycoproteins from Thermoplasma acidophilum are modified with an N-linked glycan containing 6-C-sulfofucose. Glycobiology 2012, 22, 1256–1267. [Google Scholar] [CrossRef] [PubMed]
- Kloda, A.; Martinac, B. Mechanosensitive channel of Thermoplasma, the cell wall-less archaea: Cloning and molecular characterization. Cell Biochem. Biophys. 2001, 34, 321–347. [Google Scholar] [CrossRef] [PubMed]
- Stein, D.B.; Sercy, D.G. Physiologically important stabilization of DNA by a prokaryotic histone-like protein. Science 1978, 202, 219–221. [Google Scholar] [CrossRef] [PubMed]
- DeLange, R.J. A histone-like protein (HTa) from Thermoplasma acidophilum. J. Biol. Chem. 1981, 256, 900–904. [Google Scholar] [PubMed]
- Haugland, G.T.; Shin, J.H.; Birkeland, N.K.; Kelman, Z. Stimulation of MCM helicase activity by a Cdc6 protein in the archaeon Thermoplasma acidophilum. Nucleic Acids Res. 2006, 34, 6337–6344. [Google Scholar] [CrossRef] [PubMed]
- Ogino, H.; Ishino, S.; Mayanagi, K.; Haugland, G.T.; Birkeland, N.K.; Yamagishi, A.; Ishino, Y. The GINS complex from the thermophilic archaeon, Thermoplasma acidophilum may function as a homotetramer in DNA replication. Extremophiles 2011, 15, 529–539. [Google Scholar] [CrossRef] [PubMed]
- Roth, H.M.; Tessmer, I.; van Houten, B.; Kisker, C. Bax1 is a novel endonuclease: Implications for archaeal nucleotide excision repair. J. Biol. Chem. 2009, 284, 32272–32278. [Google Scholar] [CrossRef] [PubMed]
- Moen, M.N.; Knævelsrud, I.; Haugland, G.T.; Grøsvik, K.; Birkeland, N.K.; Klungland, A.; Bjelland, S. Uracil-DNA glycosylase of Thermoplasma acidophilum directs long-patch base excision repair, which is promoted by deoxynucleoside triphosphates and ATP/ADP, into short-patch repair. J. Bacteriol. 2011, 193, 4495–4508. [Google Scholar] [CrossRef] [PubMed]
- Kuper, J.; Wolski, S.C.; Michels, G.; Kisker, C. Functional and structural studies of the nucleotide excision repair helicase XPD suggest a polarity for DNA translocation. EMBO J. 2012, 31, 494–502. [Google Scholar] [CrossRef] [PubMed]
- Buechner, C.N.; Heil, K.; Michels, G.; Carell, T.; Kisker, C.; Tessmer, I. Strand-specific recognition of DNA damages by XPD provides insights into nucleotide excision repair substrate versatility. J. Biol. Chem. 2014, 289, 3613–3624. [Google Scholar] [CrossRef] [PubMed]
- Sun, N.; Pan, C.; Nickell, S.; Mann, M.; Baumeister, W.; Nagy, I. Quantitative proteome and transcriptome analysis of the archaeon Thermoplasma acidophilum cultured under aerobic and anaerobic conditions. J. Proteome Res. 2010, 9, 4839–4850. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.M.; Paek, K.H.; Lee, S.B. Characterization of NADP+-specific L-rhamnose dehydrogenase from the thermoacidophilic Archaeon Thermoplasma acidophilum. Extremophiles 2012, 16, 447–454. [Google Scholar] [CrossRef] [PubMed]
- Ruepp, A.; Graml, W.; Santos-Martinez, M.L.; Koretke, K.K.; Volker, C.; Mewes, H.W.; Frishman, D.; Stocker, S.; Lupas, A.N.; Baumeister, W. The genome sequence of the thermoacidophilic scavenger Thermoplasma acidophilum. Nature 2000, 407, 508–813. [Google Scholar] [CrossRef] [PubMed]
- Brandstetter, H.; Kim, J.-S.; Groll, M.; Huber, R. Crystal structure of the tricorn protease reveals a protein disassembly line. Nature 2001, 414, 466–470. [Google Scholar] [CrossRef] [PubMed]
- Steinbacher, S.; Ditzel, L. Review: Nucleotide binding to the Thermoplasma thermosome: Implications for the functional cycle of group II chaperonins. J. Struct. Biol. 2001, 135, 147–156. [Google Scholar] [CrossRef] [PubMed]
- Noi, K.; Hirai, H.; Hongo, K.; Mizobata, T.; Kawata, Y. Implications for the functional cycle of group II chaperonins from Thermoplasma acidophilum. Biochemistry 2009, 48, 9405–9415. [Google Scholar] [CrossRef] [PubMed]
- Ruschak, A.M.; Religa, T.L.; Breuer, S.; Witt, S.; Kay, L.E. The proteasome antechamber maintains substrates in an unfolded state. Nature 2010, 467, 868–873. [Google Scholar] [CrossRef] [PubMed]
- Kilpatrick, M.W.; Walker, R.T. The nucleotide sequence of the tRNAMMet from the archaebacterium Thermoplasma acidophilum. Nucleic Acids Res. 1981, 9, 4387–4390. [Google Scholar] [CrossRef] [PubMed]
- Kuchino, Y.; Ihara, M.; Yabusaki, Y.; Nishimura, S. Initiator tRNAs from archaebacteria show common unique sequence characteristics. Nature 1982, 298, 684–685. [Google Scholar] [CrossRef] [PubMed]
- Walker, R.T. Mycoplasma evolution: A review of the use of ribosomal and transfer RNA nucleotide sequences in the determination of phylogenetic relationships. Yale J. Biol. Med. 1983, 56, 367–372. [Google Scholar] [PubMed]
- Edmonds, C.G.; Crain, P.F.; Gupta, R.; Hashizume, T.; Hocart, C.H.; Kowalak, J.A.; Pomerantz, S.C.; Stetter, K.O.; McCloskey, J.A. Posttranscriptional modification of tRNA in thermophilic archaea (Archaebacteria). J. Bacteriol. 1991, 173, 3138–3148. [Google Scholar] [PubMed]
- Tomikawa, C.; Ohira, T.; Inoue, Y.; Kawamura, T.; Yamagishi, A.; Suzuki, T.; Hori, H. Distinct tRNA modifications in the thermo-acidophilic archaeon, Thermoplasma acidophilum. FEBS Lett. 2013, 587, 3575–3580. [Google Scholar] [CrossRef] [PubMed]
- Gregson, J.M.; Crain, P.F.; Edmonds, C.G.; Gupta, R.; Hashizume, T.; Phillipson, D.W.; McCloskey, J.A. Structure of the archaeal transfer RNA nucleoside G*-15 (2-amino-4,7-dihydro-4-oxo-7-beta-d-ribofuranosyl-1H-pyrrolo[2,3-d]pyrimidine-5-carboximidamide (archaeosine)). J. Biol. Chem. 1993, 268, 10076–10086. [Google Scholar] [PubMed]
- Renalier, M.H.; Joseph, N.; Gaspin, C.; Thebault, P.; Mougin, A. The Cm56 tRNA modification in archaea is catalyzed either by a specific 2'-O-methylase, or a C/D sRNP. RNA 2005, 11, 1051–1063. [Google Scholar] [CrossRef] [PubMed]
- Jühling, F.; Mörl, M.; Hartmann, R.K.; Sprinzl, M.; Stadler, P.F.; Pütz, J. tRNAdb 2009: Compilation of tRNA sequences and tRNA genes. Nucleic Acids Res. 2009, 37, D159–D162. [Google Scholar] [CrossRef] [PubMed]
- Machnicka, M.A.; Milanowska, K.; Osman Oglou, O.; Purta, E.; Kurkowska, M.; Olchowik, A.; Januszewski, W.; Kalinowski, S.; Dunin-Horkawicz, S.; Rother, K.M.; et al. MODOMICS: A database of RNA modification pathways—2013 update. Nucleic Acids Res. 2013, 41, D262–D267. [Google Scholar] [CrossRef] [PubMed]
- De Bie, L.G.; Roovers, M.; Oudjama, Y.; Wattiez, R.; Tricot, C.; Stalon, V.; Droogmans, L.; Bujnicki, J.M. The yggH gene of Escherichia coli encodes a tRNA (m7G46) methyltransferase. J. Bacteriol. 2003, 185, 3238–3243. [Google Scholar] [CrossRef] [PubMed]
- Okamoto, H.; Watanabe, K.; Ikeuchi, Y.; Suzuki, T.; Endo, Y.; Hori, H. Substrate tRNA recognition mechanism of tRNA (m7G46) methyltransferase from Aquifex aeolicus. J. Biol. Chem. 2004, 279, 49151–49159. [Google Scholar] [CrossRef] [PubMed]
- Alexandrov, A.; Martzen, M.R.; Phizicky, E.M. Two proteins that form a complex are required for 7-methylguanosine modification of yeast tRNA. RNA 2002, 8, 1253–1266. [Google Scholar] [CrossRef] [PubMed]
- Kambampati, R.; Lauhon, C.T. Evidence for the transfer of sulfane sulfur from IscS to ThiI during the in vitro biosynthesis of 4-thiouridine in Escherichia coli tRNA. J. Biol. Chem. 2000, 275, 10727–10730. [Google Scholar] [CrossRef] [PubMed]
- Lauhon, C.T.; Kambampati, R. The iscS gene in Escherichia coli is required for the biosynthesis of 4-thiouridine, thiamin, and NAD. J. Biol. Chem. 2000, 275, 20096–20103. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zhu, X.; Nakamura, A.; Orlando, R.; Söll, D.; Whitman, W.B. Biosynthesis of 4-thiouridine in tRNA in the methanogenic archaeon Methanococcus maripaludis. J. Biol. Chem. 2012, 287, 36683–36692. [Google Scholar] [CrossRef] [PubMed]
- Miranda, H.V.; Nembhard, N.; Su, D.; Hepowit, N.; Krause, D.J.; Pritz, J.R.; Phillips, C.; Söll, D.; Maupin-Furlow, J.A. E1- and ubiquitin-like proteins provide a direct link between protein conjugation and sulfur transfer in archaea. Proc. Natl. Acad. Sci. USA 2011, 108, 4417–4422. [Google Scholar] [CrossRef] [PubMed]
- Phillips, G.; Swairjo, M.A.; Gaston, K.W.; Bailly, M.; Limbach, P.A.; Iwata-Reuyl, D.; de Crécy-Lagard, V. Diversity of archaeosine synthesis in Crenarchaeota. ACS Chem. Biol. 2012, 7, 300–305. [Google Scholar] [CrossRef] [PubMed]
- Keith, G.; Desgrès, J.; Pochart, P.; Heyman, T.; Kuo, K.C.; Gehrke, C.W. Eukaryotic tRNAs(Pro): Primary structure of the anticodon loop; presence of 5-carbamoylmethyluridine or inosine as the first nucleoside of the anticodon. Biochim. Biophys. Acta 1990, 1049, 255–260. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Huang, B.; Anderson, J.T.; Byström, A.S. Unexpected accumulation of ncm5U and ncm5s2U in a trm9 mutant suggests an additional step in the synthesis of mcm5U and mcm5s2U. PLoS One 2011, 6, e20783. [Google Scholar] [CrossRef] [PubMed]
- Grosjean, H.; Constantinesco, F.; Foiret, D.; Benachenhou, N. A novel enzymatic pathway leading to 1-methylinosine modification in Haloferax volcanii tRNA. Nucleic Acids Res. 1995, 23, 4312–4319. [Google Scholar] [CrossRef] [PubMed]
- Roovers, M.; Wouters, J.; Bujnicki, J.M.; Tricot, C.; Stalon, V.; Grosjean, H.; Droogmans, L. A primordial RNA modification enzyme: the case of tRNA (m1A) methyltransferase. Nucleic Acids Res. 2004, 32, 465–476. [Google Scholar] [CrossRef] [PubMed]
- Hamdane, D.; Guelorget, A.; Guérineau, V.; Golinelli-Pimpaneau, B. Dynamics of RNA modification by a multi-site-specific tRNA methyltransferase. Nucleic Acids Res. 2015, 42, 11697–11706. [Google Scholar] [CrossRef] [PubMed]
- Kuchino, Y.; Kato, M.; Sugisaki, H.; Nishimura, S. Nucleotide sequence of starfish initiator tRNA. Nucleic Acids Res. 1979, 6, 3459–3469. [Google Scholar] [CrossRef] [PubMed]
- Constantinesco, F.; Motorin, Y.; Grosjean, H. Transfer RNA modification enzymes from Pyrococcus furiosus: detection of the enzymatic activities in vitro. Nucleic Acids Res. 1999, 27, 1308–1315. [Google Scholar] [CrossRef] [PubMed]
- Sun, N.; Tamura, N.; Tamura, T.; Knispel, R.W.; Hrabe, T.; Kofler, C.; Nickell, S.; Nagy, I. Size distribution of native cytosolic proteins of Thermoplasma acidophilum. Proteomics 2009, 9, 3783–3786. [Google Scholar] [CrossRef] [PubMed]
- Droogmans, L.; Roovers, M.; Bujnicki, J.M.; Tricot, C.; Hartsch, T.; Stalon, V.; Grosjean, H. Cloning and characterization of tRNA (m1A58) methyltransferase (TrmI) from Thermus thermophilus HB27, a protein required for cell growth at extreme temperatures. Nucleic Acids Res. 2003, 31, 2148–2156. [Google Scholar] [CrossRef] [PubMed]
- Tomikawa, C.; Yokogawa, T.; Kanai, T.; Hori, H. N7-Methylguanine at position 46 (m7G46) in tRNA from Thermus thermophilus is required for cell viability through a tRNA modification network. Nucleic Acids Res. 2010, 38, 942–957. [Google Scholar] [CrossRef] [PubMed]
- Yokogawa, T.; Kitamura, Y.; Nakamura, D.; Ohno, S.; Nishikawa, K. Optimization of the hybridization-based method for purification of thermostable tRNAs in the presence of tetraalkylammonium salts. Nucleic Acids Res. 2010, 38. [Google Scholar] [CrossRef] [PubMed]
- Awai, T.; Kimura, S.; Tomikawa, C.; Ochi, A.; Ihsanawati; Bessho, Y.; Yokoyama, S.; Ohno, S.; Nishikawa, K.; Yokogawa, T.; Suzuki, T.; Hori, H. Aquifex aeolicus tRNA (N2,N2-guanine)-dimethyltransferase (Trm1) catalyzes transfer of methyl groups not only to guanine 26 but also to guanine 27 in tRNA. J. Biol. Chem. 2009, 284, 20467–20478. [Google Scholar] [CrossRef] [PubMed]
- Ishida, K.; Kunibayashi, T.; Tomikawa, C.; Ochi, A.; Kanai, T.; Hirata, A.; Iwashita, C.; Hori, H. Pseudouridine at position 55 in tRNA controls the contents of other modified nucleotides for low-temperature adaptation in the extreme-thermophilic eubacterium Thermus thermophilus. Nucleic Acids Res. 2011, 39, 2304–2318. [Google Scholar] [CrossRef] [PubMed]
- Hirata, A.; Kitajima, T.; Hori, H. Cleavage of intron from the standard or non-standard position of the precursor tRNA by the splicing endonuclease of Aeropyrum pernix, a hyper-thermophilic Crenarchaeon, involves a novel RNA recognition site in the Crenarchaea specific loop. Nucleic Acids Res. 2011, 39, 9376–9389. [Google Scholar] [CrossRef] [PubMed]
- Christian, T.; Evilia, C.; Williams, S.; Hou, Y.M. Distinct origins of tRNA(m1G37) methyltransferase. J. Mol. Biol. 2004, 339, 707–719. [Google Scholar] [CrossRef] [PubMed]
- Goto-Ito, S.; Ito, T.; Ishii, R.; Muto, Y.; Bessho, Y.; Yokoyama, S. Crystal structure of archaeal tRNA(m(1)G37)methyltransferase aTrm5. Proteins 2008, 72, 1274–1289. [Google Scholar]
- Reinhart, M.P.; Lewis, J.M.; Leboy, P.S. A single tRNA (guanine)-methyltransferase from Tetrahymena with both mono- and di-methylating activity. Nucleic Acids Res. 1986, 14, 1131–1148. [Google Scholar] [CrossRef] [PubMed]
- Constantinesco, F.; Motorin, Y.; Grosjean, H. Characterisation and enzymatic properties of tRNA(guanine 26, N2,N2)-dimethyltransferase (Trm1p) from Pyrococcus furiosus. J. Mol. Biol. 1999, 291, 375–392. [Google Scholar] [CrossRef] [PubMed]
- Phillips, J.H.; Kjellin-Straby, K. Studies on microbial ribonucleic acid. IV. Two mutants of Saccharomyces cerevisiae lacking N-2-dimethylguanine in soluble ribonucleic acid. J. Mol. Biol. 1967, 26, 509–518. [Google Scholar] [CrossRef] [PubMed]
- Ellis, S.R.; Morales, M.J.; Li, J.M.; Hopper, A.K.; Martin, N.C. Isolation and characterization of the TRM1 locus, a gene essential for the N2,N2-dimethylguanosine modification of both mitochondrial and cytoplasmic tRNA in Saccharomyces cerevisiae. J. Biol. Chem. 1986, 261, 9703–9709. [Google Scholar] [PubMed]
- Fukunaga, J.; Gouda, M.; Umeda, K.; Ohno, S.; Yokogawa, T.; Nishikawa, K. Use of RNase P for efficient preparation of yeast tRNATyr transcript and its mutants. J. Biochem. 2006, 139, 123–127. [Google Scholar]
- Goto-Ito, S.; Ito, T.; Kuratani, M.; Bessho, Y.; Yokoyama, S. Tertiary structure checkpoint at anticodon loop modification in tRNA functional maturation. Nat. Struct. Mol. Biol. 2009, 16, 1109–1115. [Google Scholar] [CrossRef] [PubMed]
- Stanley, J.; Vassilenko, S. A different approach to RNA sequencing. Nature 1978, 274, 87–89. [Google Scholar] [CrossRef]
- Grosjean, H.; Droogmans, L.; Roovers, M.; Keith, G. Detection of enzymatic activity of transfer RNA modification enzymes using radiolabeled tRNA substrates. Methods Enzymol. 2007, 425, 55–101. [Google Scholar] [PubMed]
- Somme, J.; van Laer, B.; Roovers, M.; Steyaert, J.; Versées, W.; Droogmans, L. Characterization of two homologous 2'-O-methyltransferases showing different specificities for their tRNA substrates. RNA 2014, 20, 1257–1271. [Google Scholar] [CrossRef] [PubMed]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kawamura, T.; Anraku, R.; Hasegawa, T.; Tomikawa, C.; Hori, H. Transfer RNA Methyltransferases from Thermoplasma acidophilum, a Thermoacidophilic Archaeon. Int. J. Mol. Sci. 2015, 16, 91-113. https://doi.org/10.3390/ijms16010091
Kawamura T, Anraku R, Hasegawa T, Tomikawa C, Hori H. Transfer RNA Methyltransferases from Thermoplasma acidophilum, a Thermoacidophilic Archaeon. International Journal of Molecular Sciences. 2015; 16(1):91-113. https://doi.org/10.3390/ijms16010091
Chicago/Turabian StyleKawamura, Takuya, Ryou Anraku, Takahiro Hasegawa, Chie Tomikawa, and Hiroyuki Hori. 2015. "Transfer RNA Methyltransferases from Thermoplasma acidophilum, a Thermoacidophilic Archaeon" International Journal of Molecular Sciences 16, no. 1: 91-113. https://doi.org/10.3390/ijms16010091





