Proteomic Analysis of Rice Nonhost Resistance to Puccinia striiformis f. sp. tritici Using Two-Dimensional Electrophoresis
Abstract
:1. Introduction
2. Results
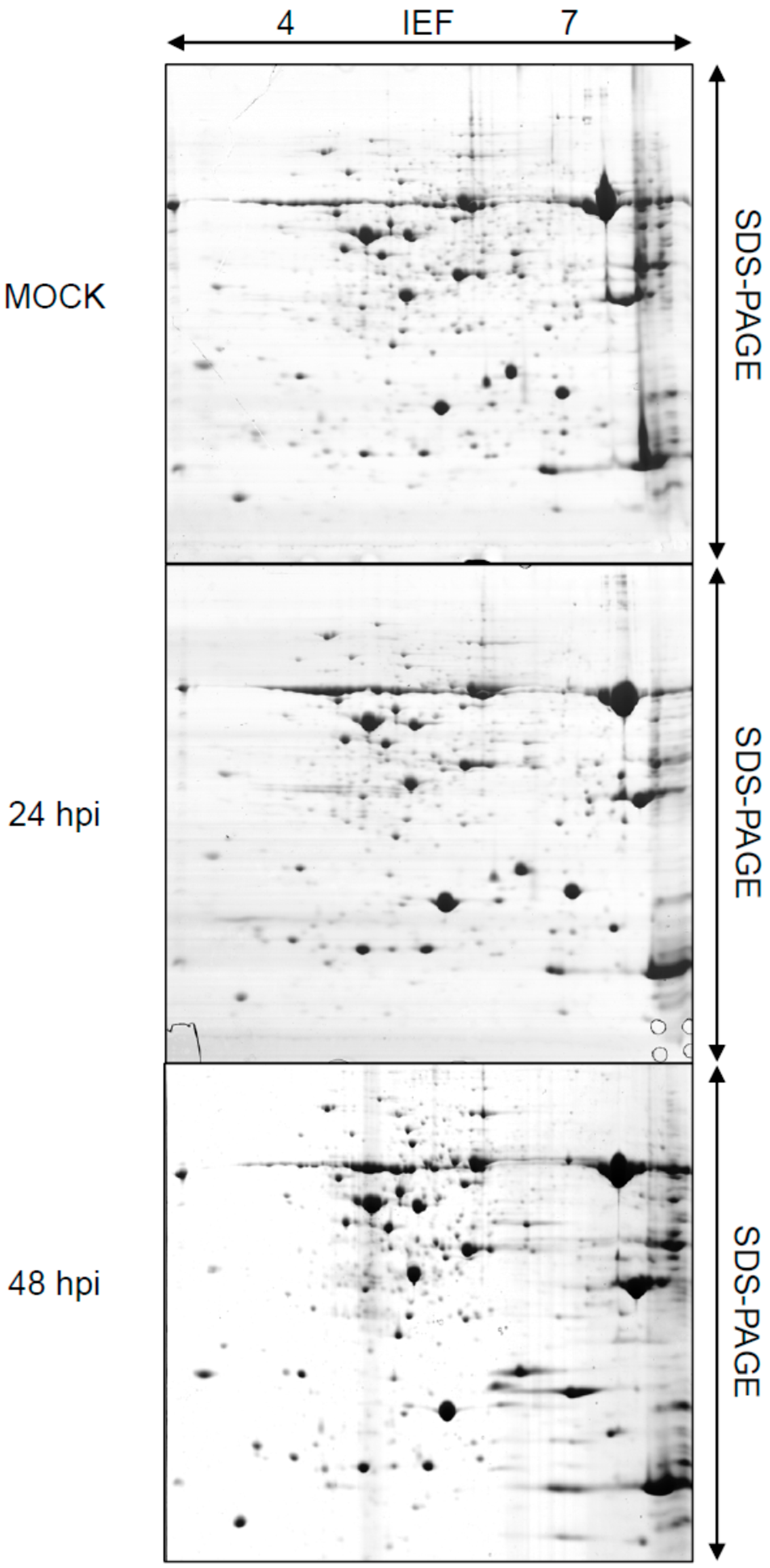
2.1. Rice Proteome Induced by Pst Infection
| Spot No. | Identification | GenBank Accession No. | pI/MW (kDa) a | Peptides Count b | Coverage (%) c | Protein Score/Ion Score | Mock Average Qty | Inoculated Average Qty | Fold Change |
|---|---|---|---|---|---|---|---|---|---|
| Up-Regulated Expression | |||||||||
| 124 | Ribulosebiphosphate carboxylase | gi|224612718 | 5.96/21.5 | 6 | 34.7 | 95/143 | 307.4 ± 3.1 | 1180.6 ± 19.3 | 3.84 ± 0.04 |
| 1125 | Ribosomal protein S6 | gi|125588601 | 5.97/23.1 | 5 | 29.8 | 138/171 | 444.5 ± 0.5 | 1118.3 ± 93.7 | 2.51 ± 0.21 |
| 1407 | Phosphoglycerate kinase | gi|125552851 | 6.86/30.5 | 9 | 43.3 | 568/634 | 55.5 ± 9.0 | 572.5 ± 25.2 | 10.59 ± 1.71 |
| 1507 | Chloroplast phosphoglycerate kinase | gi|46981258 | 9.93/32.5 | 10 | 38.4 | 229/300 | 0 | 238.9 ± 9.8 | − |
| 1709 | OsAPx8, Thylakoid-bound ascorbateperoxidase | gi|125582491 | 4.79/43.6 | 12 | 35.7 | 250/333 | 0 | 176.8 ± 3.9 | − |
| 1710 | OsAPx8, Thylakoid-bound ascorbateperoxidase | gi|115446663 | 5.36/51.4 | 21 | 44.8 | 348/538 | 78.4 ± 0.4 | 424.2 ± 0.4 | 5.41 ± 0.01 |
| 2521 | Glyceraldehyde-3-phosphate dehydrogenase b | gi|115450493 | 6.22/47.5 | 19 | 48.4 | 648/810 | 355.2 ± 7.4 | 1333.9 ± 4.5 | 3.76 ± 0.08 |
| 3816 | OsFtsH1, FtsH protease | gi|115470052 | 5.87/67.0 | 15 | 24.5 | 149/247 | 52.4 ± 3.7 | 122.9 ± 10.1 | 2.34 ± 0.01 |
| 4416 | Glyceraldehyde-3-phosphate dehydrogenase | gi|115458768 | 7.62/43.0 | 18 | 54.2 | 864/1030 | 660.8 ± 23.2 | 1636.4 ± 31.3 | 2.48 ± 0.09 |
| 7122 | Ribulose-bisphosphate carboxylase oxygenase large subunit | gi|297720471 | 6.35/29.1 | 14 | 39.7 | 408/621 | 1607.2 ± 32.7 | 8024.3 ± 34.2 | 5.22 ± 1.08 |
| 7413 | Glyceraldehyde-3-phosphate dehydrogenase | gi|115458768 | 7.62/43.0 | 21 | 59.2 | 716/923 | 342.25 ± 11.9 | 1922.6 ± 31.3 | 6.16 ± 1.83 |
| 0217 | ATP synthase F1, delta subunit family protein | gi|115448701 | 4.98/26.2 | 8 | 31.7 | 378/439 | 739.1 ± 3.1 | 30.3 ± 0.3 | 0.04 ± 0.04 |
| 5515 | Aminotransferase | gi|115477483 | 6.48/50.4 | 17 | 46.9 | 704/847 | 916.1 ± 31.5 | 282.5 ± 134.8 | 0.31 ± 0.15 |
| Down-Regulated Expression | |||||||||
| 6622 | NAD dependent epimerase/dehydratase family protein | gi|115482032 | 5.75/43.2 | 23 | 65.9 | 418/635 | 955.9 ± 47.3 | 353.9 ± 103.9 | 0.37 ± 0.11 |
| 7811 | Ferredoxin-nitrite reductase | gi|297599961 | 6.63/47.1 | 5 | 13.4 | 66/94 | 122.3 ± 14.9 | 52.2 ± 1.6 | 0.43 ± 0.01 |
| Spot No. | Identification | GenBank Accession No. | pI/MW (kDa) a | Peptides Count b | Coverage (%) c | Protein Score/Ion Score | Mock Average Qty | Inoculated Average Qty | Fold Change |
|---|---|---|---|---|---|---|---|---|---|
| Up-Regulated Expression | |||||||||
| 4532 | Glyceraldehyde-3-phosphate dehydrogenase b | gi|115450493 | 6.22/47.5 | 14 | 31.8 | 641/747 | 359.7 ± 13.7 | 1349.8 ± 37.4 | 3.76 ± 0.14 |
| 4827 | Transketolase, chloroplast precursor | gi|115466224 | 5.44/74.0 | 25 | 53.1 | 543/773 | 497.3 ± 23.6 | 1267.7 ± 35.4 | 2.55 ± 0.10 |
| 5318 | Chloroplast ATP synthase CF1 alpha chain | gi|20143564 | 5.27/29.4 | 3 | 23.0 | 150/166 | 324.5 ± 10.6 | 732.9 ± 21.7 | 2.26 ± 0.10 |
| 6124 | Ribulose-1,5-bisphophate carboxylase oxygenase | gi|354618517 | 5.80/19.7 | 8 | 38.2 | 268/340 | 729.0 ± 17.0 | 1702.1 ± 31.2 | 2.34 ± 0.08 |
| 6824 | DnaK family protein, chloroplast Hsp70 | gi|125578088 | 5.33/69.8 | 23 | 37.3 | 586/763 | 344.4 ± 24.8 | 1192.8 ± 84.6 | 3.48 ± 0.25 |
| 6826 | OsFtsH1, FtsH protease | gi|115470052 | 5.51/72.9 | 13 | 25.8 | 587/668 | 44.2 ± 2.7 | 335.6 ± 21.2 | 7.59 ± 0.68 |
| 7028 | Carbonic anhydrase | gi|5917783 | 8.41/29.6 | 7 | 41.8 | 203/253 | 451.3 ± 37.2 | 1196.4 ± 79.5 | 2.65 ± 0.25 |
| 7430 | Chloroplast 28 kDa ribonucleo protein | gi|149391365 | 5.03/21.0 | 12 | 75.3 | 234/351 | 316.9 ± 0.3 | 1552.1 ± 13.0 | 4.90 ± 0.06 |
| 7824 | Putative LR10 resistance protein | gi|305691143 | 6.08/105.4 | 8 | 14.7 | 13/37 | 284.7 ± 3.5 | 763.3 ± 17.7 | 2.68 ± 0.13 |
| 8428 | Phosphoribulo kinase | gi|115448091 | 5.68/45.2 | 12 | 25.6 | 153/230 | 210.4 ± 1.3 | 432.7 ± 57.8 | 2.05 ± 0.26 |
| 8722 | Rhodanese-like domain-containing protein chloroplastic-like | gi|115445387 | 5.05/48.4 | 6 | 15.5 | 215/238 | 108.0 ± 1.8 | 650.0 ± 15.9 | 6.03 ± 0.35 |
| 8805 | Chloroplast Hsp70 | gi|15233779 | 5.07/76.6 | 16 | 26.9 | 789/887 | 703.9 ± 13.5 | 2818.1 ± 105.0 | 4.00 ± 0.22 |
| Down-Regulated Expression | |||||||||
| 7718 | T-complex protein | gi|115488160 | 5.12/61.1 | 23 | 38.6 | 683/882 | 2229.3 ± 60.5 | 859.0 ± 35.2 | 0.39 ± 0.01 |
| 8029 | Thioredoxin | gi|297728925 | 8.16/18.9 | 3 | 23.8 | 82/99 | 897.8 ± 22.2 | 275.5 ± 7.4 | 0.31 ± 0.01 |
| 9208 | Ribonucleoprotein chloroplastic-like | gi|149392545 | 4.45/22.2 | 6 | 35.1 | 420/462 | 1228.8 ± 35.4 | 403.5 ± 5.8 | 0.33 ± 0.01 |

2.2. Identification of Differentially Expressed Proteins by Mass Spectrometry
2.3. Validation of Up-Regulated Proteins by qRT-PCR

| Accession No. | Gene | 12 hpi | 24 hpi | 48 hpi | 72 hpi |
|---|---|---|---|---|---|
| gi|125588601 | Os03g62630 | 0.94 ± 0.32 | 2.65 ± 1.16 | 2.86 ± 0.84 | 1.25 ± 0.16 |
| gi|46981258 | Os05g41640 | 0.96 ± 0.19 | 2.06 ± 0.27 | 3.60 ± 1.18 | 0.85 ± 0.40 |
| gi|125582491 | Os02g34810 | 0.96 ± 0.14 | 2.31 ± 0.43 | 2.12 ± 0.75 | 0.87 ± 0.20 |
| gi|115450493 | Os03g03720 | 1.09 ± 0.05 | 3.77 ± 0.33 | 4.23 ± 0.41 | 1.91 ± 1.26 |
| gi|115470052 | Os06g51029 | 1.03 ± 0.16 | 0.68 ± 0.05 | 1.15 ± 0.29 | 0.63 ± 0.22 |
| gi|115458768 | Os04g38600 | 0.79 ± 0.50 | 3.41 ± 0.40 | 6.00 ± 2.13 | 1.11 ± 0.18 |
| gi|115466224 | Os06g04270 | 2.86 ± 0.74 | 2.86 ± 0.74 | 1.07 ± 0.31 | 0.51 ± 0.11 |
| gi|20143564 | OSJNBa0034L04.44 | 2.19 ± 0.70 | 0.89 ± 0.02 | 2.43 ± 0.68 | 2.30 ± 0.41 |
| gi|5917783 | Os01g45274 | 1.15 ± 0.23 | 0.88 ± 0.13 | 13.39 ± 0.54 | 0.84 ± 0.24 |
| gi|149391365 | Os09g39180 | 0.63 ± 0.19 | 0.83 ± 0.27 | 4.67 ± 0.64 | 1.84 ± 0.64 |
| gi|115448091 | Os02g47020 | 7.54 ± 0.85 | 2.91 ± 0.90 | 1.81 ± 0.48 | 1.13 ± 0.27 |
| gi|115445387 | Os02g15750 | 2.91 ± 1.70 | 0.70 ± 0.12 | 4.65 ± 0.74 | 0.99 ± 0.28 |
| gi|15233779 | Os12g14070 | 0.77 ± 0.09 | 1.35 ± 0.16 | 3.09 ± 0.19 | 1.33 ± 0.07 |
| gi|305691143 | Os11g14380 | 0.64 ± 0.09 | 1.84 ± 0.12 | 2.17 ± 0.19 | 2.39 ± 0.55 |
| gi|125578088 | Os11g47760 | 36.82 ± 9.84 | 12.18 ± 5.57 | 0.69 ± 0.21 | 1.20 ± 0.31 |
3. Discussion
4. Experimental Section
4.1. Plants and Pathogens
4.2. Protein Extraction
4.3. Two-Dimensional Electrophoresis
4.4. Image Analysis
4.5. Mass Spectrometric Analysis and Protein Identification
4.6. Gene Ontology Analysis
4.7. RNA Isolation and qRT-PCR Assays
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Dean, R.; van Kan, J.; Pretorius, Z.A.; Hammond-Kosack, K.E.; Di Pietro, A.; Spanu, P.D.; Rudd, J.J.; Dickman, M.; Kahmann, R.; Ellis, J.; et al. The top 10 fungal pathogens in molecular plant pathology. Mol. Plant Pathol. 2012, 13, 414–430. [Google Scholar]
- Chen, W.; Wu, L.; Liu, T.; Xu, S. Race dynamics, diversity, and virulence evolution in Puccinia striiformis f. sp. tritici, the causal agent of wheat stripe rust in China from 2003 to 2007. Plant Dis. 2009, 93, 1093–1101. [Google Scholar]
- Fan, J.; Doerner, P. Genetic and molecular basis of nonhost disease resistance: Complex, yes; silver bullet, no. Curr. Opin. Plant Biol. 2012, 15, 400–406. [Google Scholar]
- Mysore, K.S. Nonhost resistance against bacterial pathogens: Retrospectives and prospects. Annu. Rev. Phytopathol. 2013, 51, 407–427. [Google Scholar]
- Mysore, K.S.; Ryu, C.M. Nonhost resistance: How much do we know? Trends Plant Sci. 2004, 9, 97–104. [Google Scholar]
- Schulze-Lefert, P.; Panstruga, R. A molecular evolutionary concept connecting nonhost resistance, pathogen host range, and pathogen speciation. Trends Plant Sci. 2011, 16, 117–125. [Google Scholar]
- Azinheira, H.G.; do Céu Silva, M.; Talhinhas, P.; Medeira, C.; Maia, I.; Petitot, A.S.; Fernandez, D. Nonhost resistance responses of Arabidopsis thaliana to the coffee leaf rust fungus (Hemileia vastatrix). Botany 2010, 88, 621–629. [Google Scholar]
- Loehrer, M.; Langenbach, C.; Goellner, K.; Conrath, U.; Schaffrath, U. Characterization of nonhost resistance of Arabidopsis to the Asian soybean rust. Mol. Plant-Microbe Interact. 2008, 21, 1421–1430. [Google Scholar]
- Shafiei, R.; Hang, C.; Kang, J.G.; Loake, G.J. Identification of loci controlling non-host disease resistance in Arabidopsis against the leaf rust pathogen Puccinia triticina. Mol. Plant Pathol. 2007, 8, 773–784. [Google Scholar]
- Mellersh, D.G.; Heath, M.C. An investigation into the involvement of defense signaling pathways in components of the nonhost resistance of Arabidopsis thaliana to rust fungi also reveals a model system for studying rust fungal compatibility. Mol. Plant-Microbe Interact. 2003, 16, 398–404. [Google Scholar]
- Jafary, H.; Albertazzi, G.; Marcel, T.C.; Niks, R.E. High diversity of genes for nonhost resistance of barley to heterologous rust fungi. Genetics 2008, 178, 2327–2339. [Google Scholar]
- Niks, R.E. How specific is non-hypersensitive host and nonhost resistance of barley to rust and mildew fungi? J. Integr. Agric. 2014, 13, 244–254. [Google Scholar]
- Ayliffe, M.; Devilla, R.; Mago, R.; White, R.; Talbot, M.; Pryor, A.; Leung, H. Nonhost resistance of rice to rust pathogens. Mol. Plant-Microbe Interact. 2011, 24, 1143–1155. [Google Scholar]
- Rampitsch, C.; Bykova, N.V. Proteomics and plant disease: Advances in combating a major threat to the global food supply. Proteomics 2012, 12, 673–690. [Google Scholar]
- Shen, G.; Adam, Z.; Zhang, H. The E3 ligase AtCHIP ubiquitylates FtsH1, a component of the chloroplast FtsH protease, and affects protein degradation in chloroplasts. Plant J. 2007, 52, 309–321. [Google Scholar]
- Vogel, C.; Marcotte, E.M. Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nat. Rev. Genet. 2012, 13, 227–232. [Google Scholar]
- Yang, Y.; Zhao, J.; Xing, H.; Wang, J.; Zhou, K.; Zhan, G.; Zhang, H.; Kang, Z. Different non-host resistance responses of two rice subspecies, japonica and indica, to Puccinia striiformis f. sp. tritici. Plant Cell Rep. 2013, 33, 423–433. [Google Scholar]
- Liang, G.; Ji, H.; Zhang, Z.; Wei, H.; Kang, Z.; Peng, Y.; Li, Y. Proteome analysis of slow-rusting variety Chuanmai 107 inoculated by wheat stripe rust (Puccina striiformis). J. Triticeae Crops 2007, 27, 335–340. [Google Scholar]
- Ma, C.; Xu, S.; Xu, Q.; Zhang, Z.; Pan, Y. Proteomic analysis of stripe rust resistance wheat line Taichung29*6/Yr5 inoculated with stripe rust race CYR32. Sci. Agric. Sin. 2009, 42, 1616–1623. [Google Scholar]
- Li, H.; Goodwin, P.H.; Han, Q.; Huang, L.; Kang, Z. Microscopy and proteomic analysis of the non-host resistance of Oryza sativa to the wheat leaf rust fungus, Puccinia triticina f. sp. tritici. Plant Cell Rep. 2012, 31, 637–650. [Google Scholar]
- Tsunezuka, H.; Fujiwara, M.; Kawasaki, T.; Shimamoto, K. Proteome analysis of programmed cell death and defense signaling using the rice lesion mimic mutant cdr2. Mol. Plant-Microbe Interact. 2005, 18, 52–59. [Google Scholar]
- Kim, S.T.; Kim, S.G.; Kang, Y.H.; Wang, Y.; Kim, J.Y.; Yi, N.; Kim, J.K.; Rakwal, R.; Koh, H.J.; Kang, K.Y. Proteomics analysis of rice lesion mimic mutant (spl1) reveals tightly localized probenazole-induced protein (PBZ1) in cells undergoing programmed cell death. J. Proteome Res. 2008, 7, 1750–1760. [Google Scholar]
- Chen, X.; Fu, S.; Zhang, P.; Gu, Z.; Liu, J.; Qian, Q.; Ma, B. Proteomic analysis of a disease-resistance-enhanced lesion mimic mutant spotted leaf 5 in rice. Rice 2013. [Google Scholar] [CrossRef]
- Bolton, M.D. Primary metabolism and plant defense-fuel for the fire. Mol. Plant-Microbe Interact. 2009, 22, 487–497. [Google Scholar]
- Chanda, B.; Xia, Y.; Mandal, M.; Yu, K.; Sekine, K.; Gao, Q.; Selote, D.; Hu, Y.; Stromberg, A.; Navarre, D.; et al. Glycerol-3-phosphate is a critical mobile inducer of systemic immunity in plants. Nat. Genet. 2011, 43, 421–427. [Google Scholar]
- Yang, Y.; Zhao, J.; Liu, P.; Xing, H.; Li, C.; Wei, G.; Kang, Z. Glycerol-3-phosphate metabolism in wheat contributes to systemic acquired resistance against Puccinia striiformis f. sp. tritici. PLoS One 2013, 8, e81756. [Google Scholar]
- Guo, L.; Devaiah, S.P.; Narasimhan, R.; Pan, X.; Zhang, Y.; Zhang, W.; Wang, X. Cytosolic glyceraldehyde-3-phosphate dehydrogenases interact with phospholipase Dδ to transduce hydrogen peroxide signals in the Arabidopsis response to stress. Plant Cell 2012, 24, 2200–2212. [Google Scholar]
- Shingaki-Wells, R.N.; Huang, S.; Taylor, N.L.; Carroll, A.J.; Zhou, W.; Millar, A.H. Differential molecular responses of rice and wheat coleoptiles to anoxia reveal novel metabolic adaptations in amino acid metabolism for tissue tolerance. Plant Physiol. 2011, 156, 1706–1724. [Google Scholar]
- Thao, N.P.; Chen, L.; Nakashima, A.; Hara, S.; Umemura, K.; Takahashi, A.; Shirasu, K.; Kawasaki, T.; Shimamoto, K. RAR1 and HSP90 form a complex with Rac/Rop GTPase and function in innate-immune responses in rice. Plant Cell 2007, 19, 4035–4045. [Google Scholar]
- Qi, Y.; Wang, H.; Zou, Y.; Liu, C.; Liu, Y.; Wang, Y.; Zhang, W. Over-expression of mitochondrial heat shock protein 70 suppresses programmed cell death in rice. FEBS Lett. 2011, 585, 231–239. [Google Scholar]
- Feuillet, C.; Travella, S.; Stein, N.; Albar, L.; Nublat, A.; Keller, B. Map-based isolation of the leaf rust disease resistance gene Lr10 from the hexaploid wheat (Triticum aestivum L.) genome. Proc. Natl. Acad. Sci. USA 2003, 100, 15253–15258. [Google Scholar]
- Zhao, B.; Li0n, X.; Poland, J.; Trick, H.; Leach, J.; Hulbert, S. A maize resistance gene functions against bacterial streak disease in rice. Proc. Natl. Acad. Sci. USA 2005, 102, 15383–15388. [Google Scholar]
- Niu, Z.; Puri, K.; Chao, S.; Jin, Y.; Sun, Y.; Steffenson, B.; Maan, S.; Xu, S.; Zhong, S. Genetic analysis and molecular mapping of crown rust resistance in common wheat. Theor. Appl. Genet. 2014, 127, 609–619. [Google Scholar]
- Zhang, H.; Wang, C.; Cheng, Y.; Wang, X.; Li, F.; Han, Q.; Xu, J.; Chen, X.; Huang, L.; Wei, G.; et al. Histological and molecular studies of the non-host interaction between wheat and Uromyces fabae. Planta 2011, 234, 979–991. [Google Scholar]
- Xiang, X.; Ning, S.; Jiang, X.; Gong, X.; Zhu, R.; Zhu, L.; Wei, D. Protein extraction from rice (Oryza sativa L.) root for two-dimensional electrophresis. Front. Agric. China 2010, 1, 416–421. [Google Scholar]
- Kruger, N.J. The Bradford method for protein quantitation. In The Protein Protocols Handbook, 3rd ed.; Walker, J.M., Ed.; Humana Press: Totowa, NJ, USA, 2009; pp. 17–24. [Google Scholar]
- Shevchenko, A.; Wilm, M.; Vorm, O.; Mann, M. Mass spectrometric sequencing of proteins from silver-stained polyacrylamide gels. Anal. Chem. 1996, 68, 850–858. [Google Scholar]
- Gotz, S.; Garcia-Gomez, J.M.; Terol, J.; Williams, T.D.; Nagaraj, S.H.; Nueda, M.J.; Robles, M.; Talon, M.; Dopazo, J; Conesa, A. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 2008, 36, 3420–3435. [Google Scholar]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt Method. Methods 2001, 25, 402–408. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, J.; Yang, Y.; Kang, Z. Proteomic Analysis of Rice Nonhost Resistance to Puccinia striiformis f. sp. tritici Using Two-Dimensional Electrophoresis. Int. J. Mol. Sci. 2014, 15, 21644-21659. https://doi.org/10.3390/ijms151221644
Zhao J, Yang Y, Kang Z. Proteomic Analysis of Rice Nonhost Resistance to Puccinia striiformis f. sp. tritici Using Two-Dimensional Electrophoresis. International Journal of Molecular Sciences. 2014; 15(12):21644-21659. https://doi.org/10.3390/ijms151221644
Chicago/Turabian StyleZhao, Jing, Yuheng Yang, and Zhensheng Kang. 2014. "Proteomic Analysis of Rice Nonhost Resistance to Puccinia striiformis f. sp. tritici Using Two-Dimensional Electrophoresis" International Journal of Molecular Sciences 15, no. 12: 21644-21659. https://doi.org/10.3390/ijms151221644






