Molecular and Functional Analyses of the Fast Skeletal Myosin Light Chain2 Gene of the Korean Oily Bitterling, Acheilognathus koreensis
Abstract
:1. Introduction
2. Results and Discussion
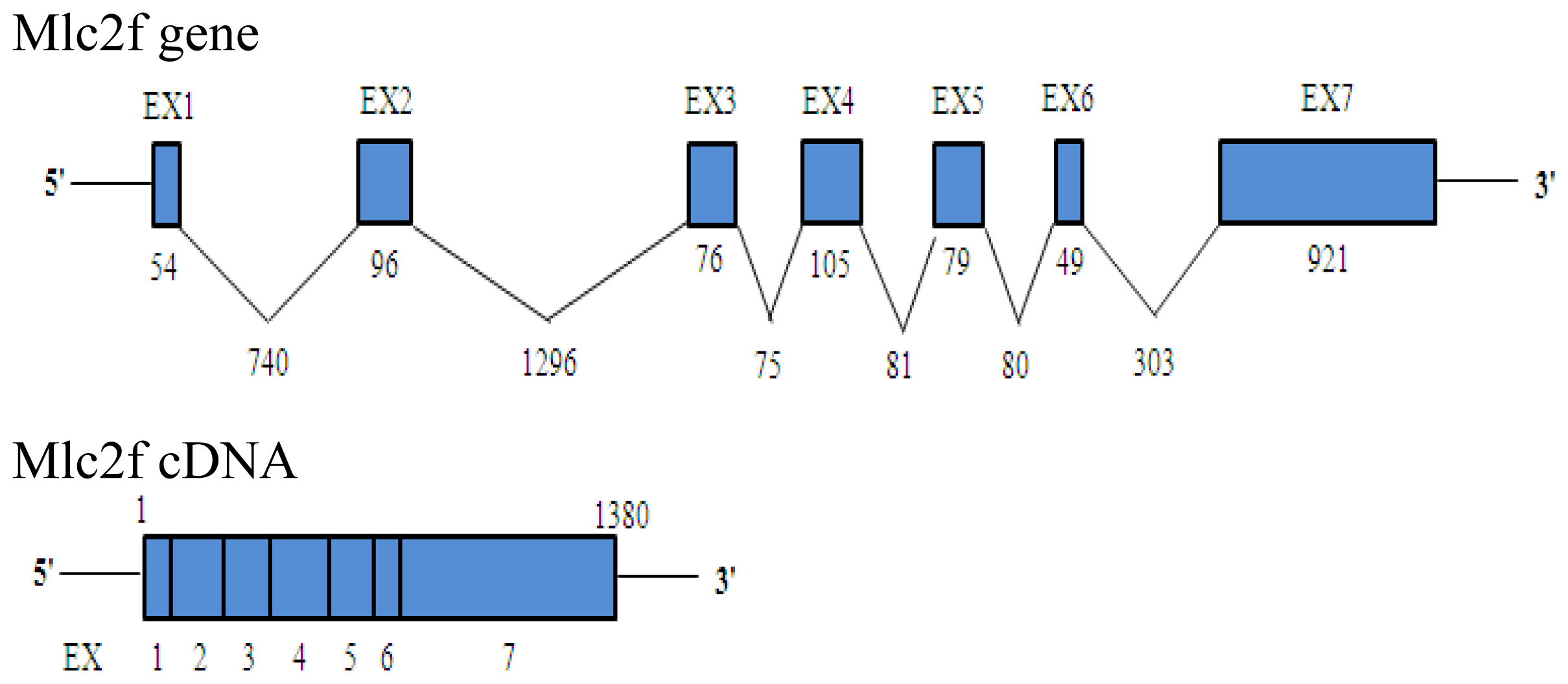
2.1. Characteristics of Akmlc2f cDNA, Genomic DNA and Deduced Amino Acid Sequences
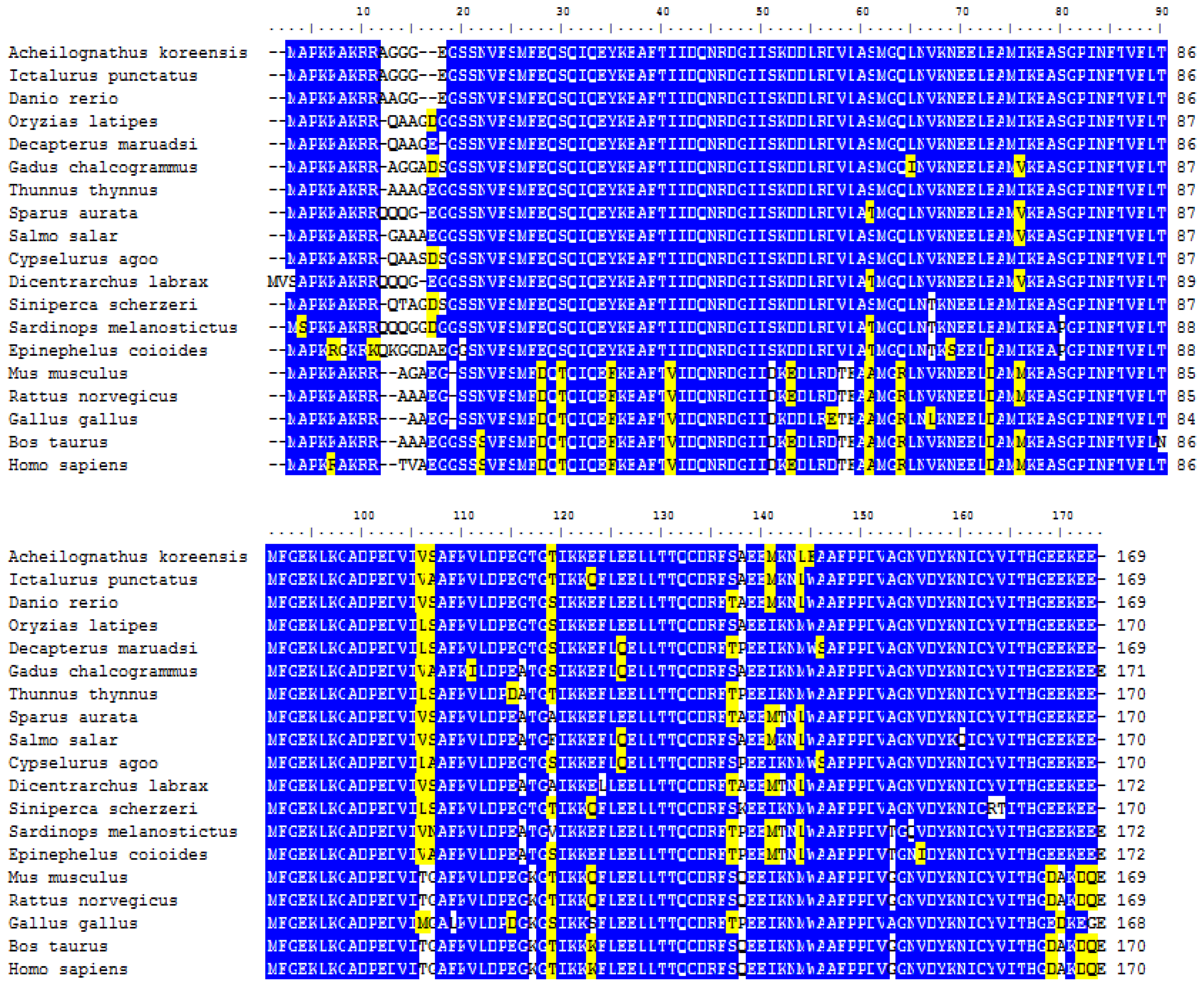
2.2. Pairwise Multiple Alignment of Akmlc2f
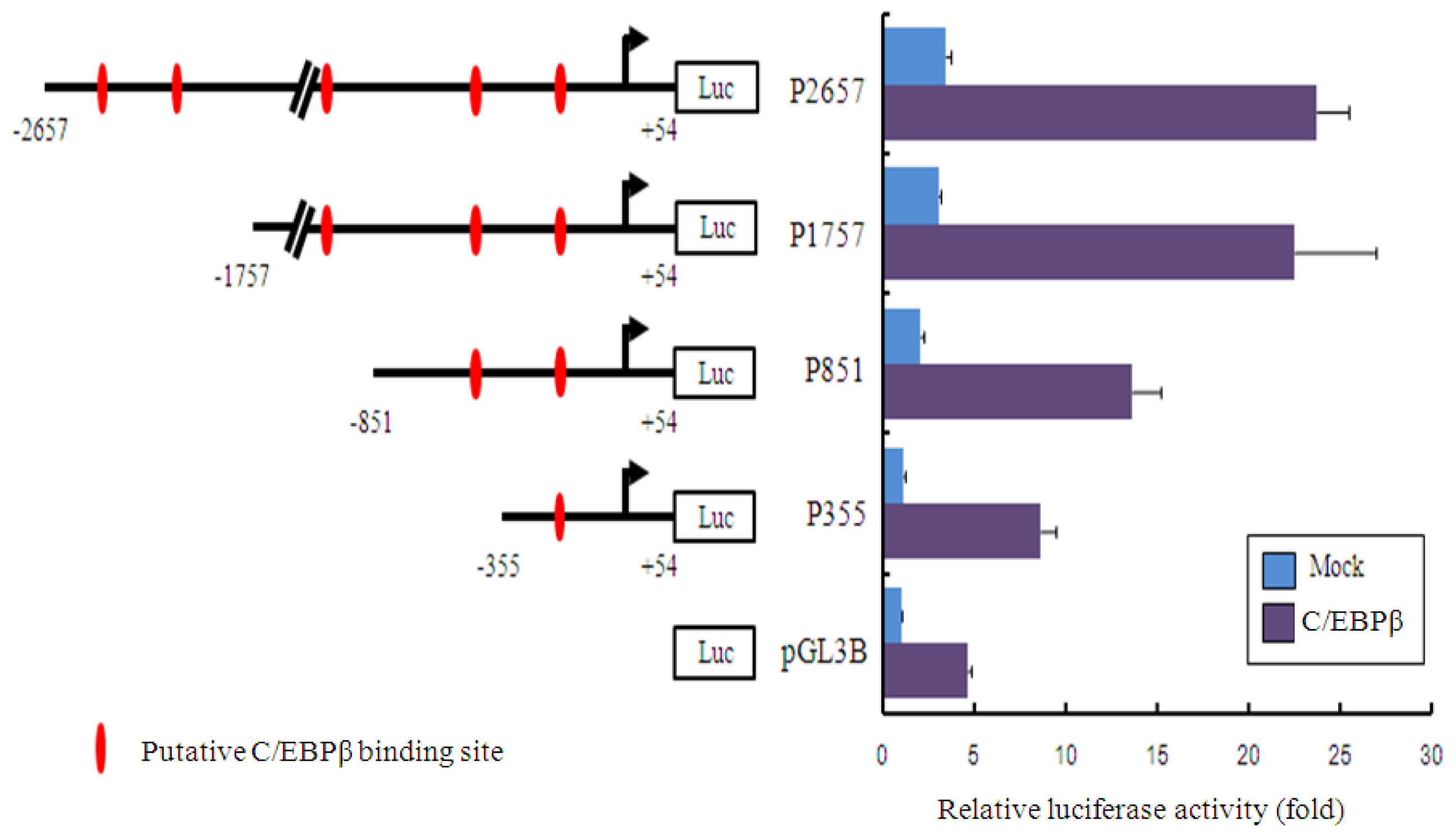
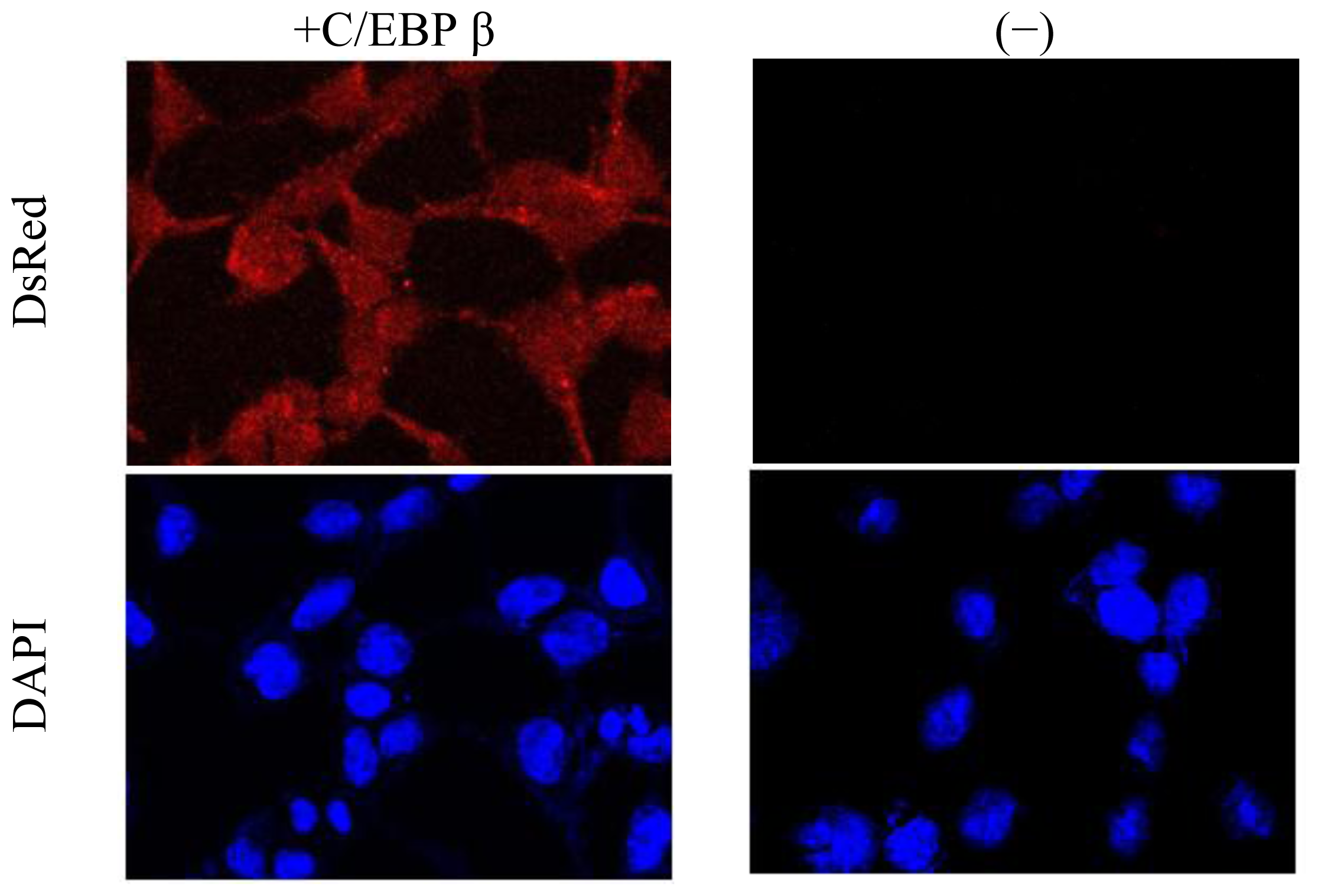
2.3. Promoter Activity of Akmlc2f
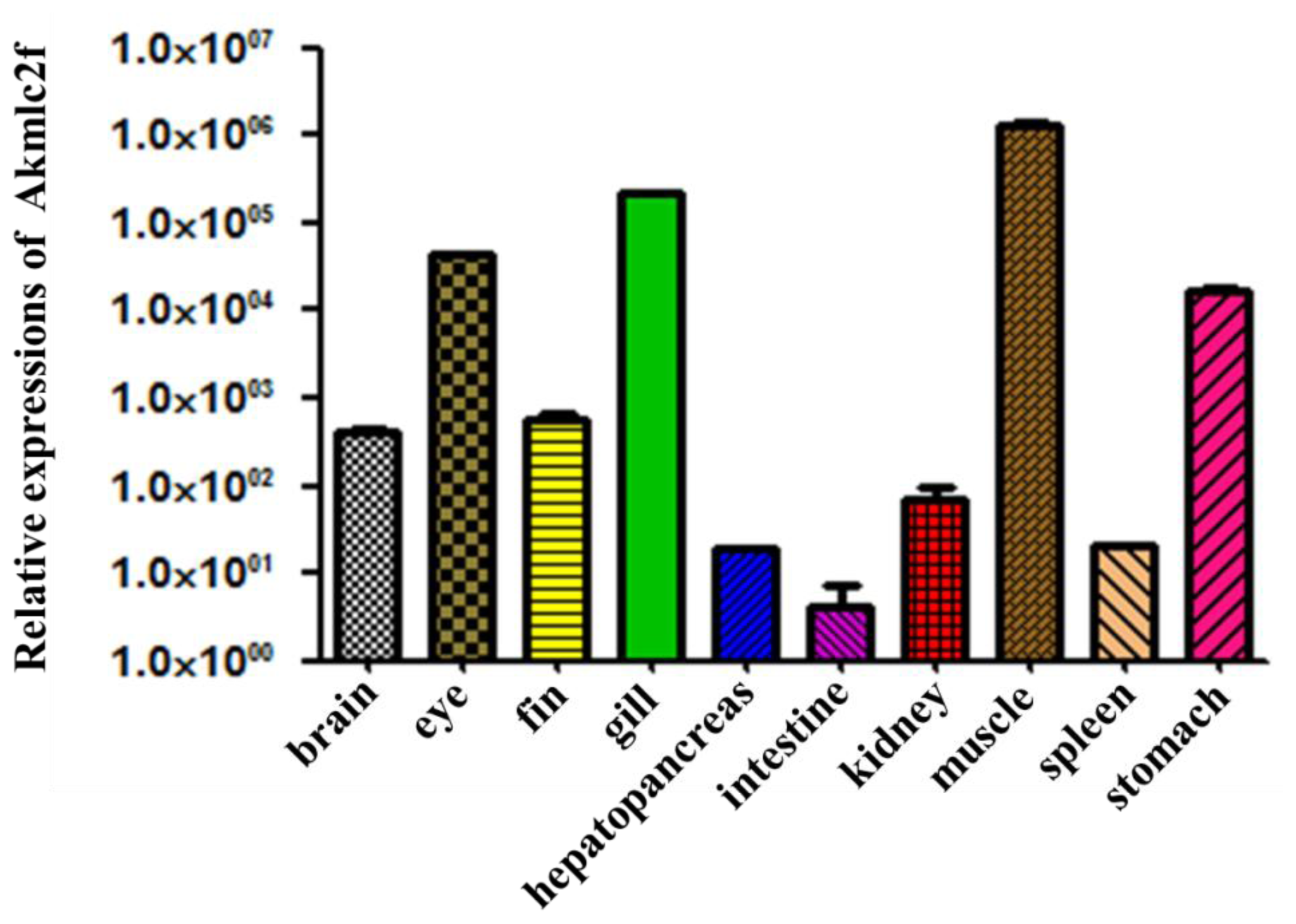
2.4. Expression of Akmlc2f mRNA
3. Materials and Methods
3.1. Isolation and Sequencing of A. koreensis Akmlc2f cDNA
3.2. Isolation and Sequence Analysis of A. koreensis mlc2f Gene
3.3. Sequence Alignments
3.4. Construction of Reporter Plasmids
3.5. Cell Culture, Transient Transfection and Luciferase Assay
3.6. Confocal Microscopy
3.7. Microinjection into Zebrafish Embryo
3.8. Fish and Tissue Samples
3.9. Quantitative Real-Time PCR
3.10. Statistical Analysis
4. Conclusions
Acknowledgements
Conflicts of Interest
References
- England, J.; Loughna, S. Heavy and light roles: Myosin in the morphogenesis of the heart. Cell Mol. Life Sci 2013, 70, 1221–1239. [Google Scholar]
- Harrington, W.F.; Rodgers, M.E. Myosin. Annu. Rev. Biochem 1984, 53, 35–73. [Google Scholar]
- Cheney, R.E.; Riley, M.A.; Mooseker, M.S. Phylogenetic analysis of the myosin superfamily. Cell Motil. Cytoskeleton 1993, 24, 215–223. [Google Scholar]
- Warrick, H.M.; Spudich, J.A. Myosin structure and function in cell motility. Annu. Rev. Cell Biol 1987, 3, 379–421. [Google Scholar]
- Czosnek, H.; Nudel, U.; Shani, M.; Barker, P.E.; Pravtcheva, D.D.; Ruddle, F.H.; Yaffe, D. The genes coding for the muscle contractile proteins, myosin heavy chain, myosin light chain 2, and skeletal muscle actin are located on three different mouse chromosomes. EMBO J 1982, 1, 1299–1305. [Google Scholar]
- Moutou, K.A.; Canario, A.V.; Mamuris, Z.; Power, D.M. Molecular cloning and sequence of Sparus aurata skeletal myosin light chains expressed in white muscle: Developmental expression and thyroid regulation. J. Exp. Biol 2001, 204, 3009–3018. [Google Scholar]
- Ikebe, M.; Hartshorne, D.J. Phosphorylation of smooth muscle myosin at two distinct sites by myosin light chain kinase. J. Biol. Chem 1985, 260, 10027–10031. [Google Scholar]
- Venema, R.C.; Raynor, R.L.; Noland, T.A., Jr; Kuo, J.F. Role of protein kinase C in the phosphorylation of cardiac myosin light chain 2. Biochem. J. 1993, 294, 401–406. [Google Scholar]
- Wilkinson, S.; Paterson, H.F.; Marshall, C.J. Cdc42-MRCK and Rho-ROCK signalling cooperate in myosin phosphorylation and cell invasion. Nat. Cell Biol 2005, 7, 255–261. [Google Scholar]
- Xu, Y.; He, J.; Tian, H.L.; Chan, C.H.; Liao, J.; Yan, T.; Lam, T.J.; Gong, Z. Fast skeletal muscle-specific expression of a zebrafish myosin light chain 2 gene and characterization of its promoter by direct injection into skeletal muscle. DNA Cell Biol 1999, 18, 85–95. [Google Scholar]
- Krasnov, A.; Teerijoki, H.; Gorodilov, Y.; Molsa, H. Cloning of rainbow trout (Oncorhynchus mykiss) α-actin, myosin regulatory light chain genes and the 5′-flanking region of α-tropomyosin. Functional assessment of promoters. J. Exp. Biol 2003, 206, 601–608. [Google Scholar]
- Funkenstein, B.; Skopal, T.; Rapoport, B.; Rebhan, Y.; Du, S.J.; Radaelli, G. Characterization and functional analysis of the 5′ flanking region of myosin light chain-2 gene expressed in white muscle of the gilthead sea bream (Sparus aurata). Comp. Biochem. Physiol. Part D Genomics Proteomics 2007, 2, 187–199. [Google Scholar]
- Cho, Y.S.; Lee, S.Y.; Kim, D.S.; Nam, Y.K. Characterization of stable fluorescent transgenic marine medaka (Oryzias dancena) lines carrying red fluorescent protein gene driven by myosin light chain 2 promoter. Transgenic Res 2013, 22, 849–859. [Google Scholar]
- Lee, S.Y.; Kim, D.S.; Nam, Y.K. Genomic organization, intronic duplications, and promoter characteristics of the fast skeletal myosin light chain-2 gene (mlc2f) from javanese ricefish Oryzias javanicus. Fish Aquat. Sci 2012, 15, 325–335. [Google Scholar]
- Kim, I.S.; Choi, Y.; Lee, C.L.; Lee, Y.J.; Kim, B.Y.; Kim, J.H. Illustrated Book of Korean Fishes (in Korean); Kyo-Hak Publishing Co. Ltd.: Seoul, Korea, 2005; p. 613. [Google Scholar]
- Hwang, D.S.; Lee, W.O.; Lee, J.S. Complete mitochondrial genome of the Korean bitterling Acheilognathus koreensis(Cypriniformes; Cyprinidae). Mitochondrial DNA 2013. [Google Scholar] [CrossRef]
- Ogut, O.; Brozovich, F.V. Regulation of force in vascular smooth muscle. J. Mol. Cell Cardiol 2003, 35, 347–355. [Google Scholar]
- Steinbacher, P.; Haslett, J.R.; Six, M.; Gollmann, H.P.; Sänger, A.M.; Stoiber, W. Phases of myogenic cell activation and possible role of dermomyotome cells in teleost muscle formation. Dev. Dynam 2006, 235, 3132–3143. [Google Scholar]
- Latinkic, B.V.; Cooper, B.; Smith, S.; Kotecha, S.; Towers, N.; Sparrow, D.; Mohun, T.J. Transcriptional regulation of the cardiac specific Mlc2 gene during Xenopus embryonic development. Development 2004, 131, 669–679. [Google Scholar]
- Gove, C.; Walmsley, M.; Nijjar, S.; Bertwistle, D.; Guille, M.; Partington, G.; Bomford, A.; Patient, R. Over-expression of GATA-6 in Xenopus embryos blocks differentiation of heartprecursors. EMBO J 1997, 16, 355–368. [Google Scholar]
- Ramji, D.P.; Foka, P. CCAAT/enhancer-binding proteins: Structure, function and regulation. Biochem. J 2002, 365, 561–575. [Google Scholar]
- Harries, L.W.; Pilling, L.C.; Hernandez, L.D.; Bradley-Smith, R.; Henley, W.; Singleton, A.B.; Guralnik, J.M.; Bandinelli, S.; Ferrucci, L.; Melzer, D. CCAAT-enhancer-binding protein-beta expression in vivo is associated with muscle strength. Aging Cell 2012, 11, 262–268. [Google Scholar]
- Lyons, S.E.; Shue, B.C.; Lei, L.; Oates, A.C.; Zon, L.I.; Liu, P.P. Molecular cloning, genetic mapping, and expression analysis of four zebrafish c/ebp genes. Gene 2001, 281, 43–51. [Google Scholar]
- Braun, T.; Tannich, E.; Buschhausen-Denker, G.; Arnold, H.H. Promoter upstream elements of the chicken cardiac myosin light-chain 2-A gene interact with trans-actin regulatory factors for muscle-specific transcription. Mol. Cell. Biol 1989, 9, 2513–2525. [Google Scholar]
- Ju, B.; Chong, S.W.; He, J.; Wang, X.; Xu, Y.; Wan, H.; Tong, Y.; Yan, T.; Korzh, V.; Gong, Z. Recapitulation of fast skeletal muscle development in zebrafish by transgenic expression of GFP under the mylz2 promoter. Dev. Dyn 2003, 227, 14–26. [Google Scholar]
- Chae, S.H.; Kim, J.W.; Cho, J.M.; Larkin, D.M.; Everts-van der Wind, A.; Park, H.S.; Yeo, J.S.; Choi, I. Chromosomal localization of Korean cattle (Hanwoo) BAC clones via BAC end sequence analysis. Asian Australas. J. Anim. Sci 2007, 20, 316–327. [Google Scholar]
- Westfield, M. The Zebrafish Book; University of Oregon: Eugene, OR, USA; p. 1995.
- Kikuchi, Y.; Segawa, H.; Tokumoto, M.; Tsubokawa, T.; Hotta, Y.; Uyemura, K.; Okamoto, H. Ocular and cerebellar defects in zebrafish induced by overexpression of the LIM domains of the islet-3 LIM/homeodomain protein. Neuron 1997, 18, 369–382. [Google Scholar]


© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Kong, H.J.; Lee, Y.-J.; Kim, W.-J.; Kim, H.S.; Kim, B.-S.; An, C.M.; Yeo, S.-Y.; Cho, H.K. Molecular and Functional Analyses of the Fast Skeletal Myosin Light Chain2 Gene of the Korean Oily Bitterling, Acheilognathus koreensis. Int. J. Mol. Sci. 2013, 14, 16672-16684. https://doi.org/10.3390/ijms140816672
Kong HJ, Lee Y-J, Kim W-J, Kim HS, Kim B-S, An CM, Yeo S-Y, Cho HK. Molecular and Functional Analyses of the Fast Skeletal Myosin Light Chain2 Gene of the Korean Oily Bitterling, Acheilognathus koreensis. International Journal of Molecular Sciences. 2013; 14(8):16672-16684. https://doi.org/10.3390/ijms140816672
Chicago/Turabian StyleKong, Hee Jeong, Ye-Ji Lee, Woo-Jin Kim, Hyung Soo Kim, Bong-Seok Kim, Cheul Min An, Sang-Yeob Yeo, and Hyun Kook Cho. 2013. "Molecular and Functional Analyses of the Fast Skeletal Myosin Light Chain2 Gene of the Korean Oily Bitterling, Acheilognathus koreensis" International Journal of Molecular Sciences 14, no. 8: 16672-16684. https://doi.org/10.3390/ijms140816672



