Interaction of Proteins Identified in Human Thyroid Cells
Abstract
:1. Introduction
2. Results and Discussion
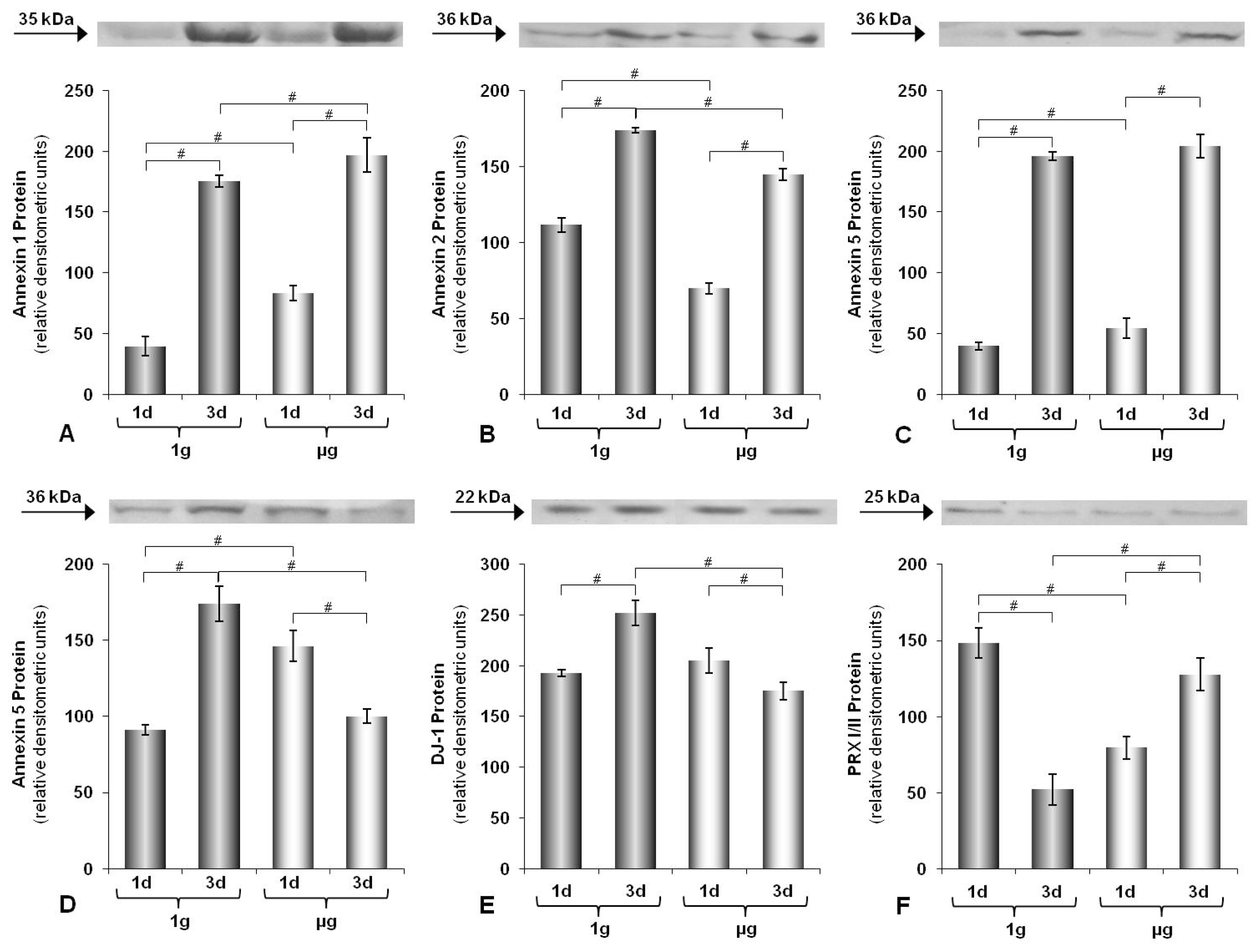
2.1. Time-Dependence of Protein Expression
2.2. STRING Network Analysis
2.3. Possible Ways of Signal Propagation
3. Experimental Section
3.1. Culture of Different Thyroid Cell Lines
3.2. Proteome Analysis—FF-IEF and Mass Spectrometry
3.3. STRING 9.0 Network Analysis
3.4. Western Blot Analysis
3.5. Statistical Analysis
4. Conclusions
Acknowledgments
References
- American Cancer Society. 2012. Available online: http://www.cancer.org/Research/CancerFactsFigures/CancerFactsFigures/cancer-facts-figures-2012 accessed on 17 August 2012.
- Edge, S.B.; Byrd, D.R.; Compton, C.C.; Fritz, A.G.; Greene, F.L.; Trotti, A. AJCC Cancer Staging Manual, 7th ed; Springer-Verlag: New York, NY, USA, 2010. [Google Scholar]
- Sakamoto, A.; Kasai, N.; Sugano, H. Poorly differentiated carcinoma of the thyroid. A clinicopathologic entity for a high-risk group of papillary and follicular carcinomas. Cancer 1983, 52, 1849–1855. [Google Scholar]
- Grimm, D.; Bauer, J.; Kossmehl, P.; Shakibaei, M.; Schönberger, J.; Pickenhahn, H.; Schulze-Tanzil, G.; Vetter, R.; Eilles, C.; Cogoli, A. Simulated microgravity alters differentiation and increases apoptosis in human follicular thyroid carcinoma cells. FASEB J 2002, 16, 604–616. [Google Scholar]
- Ulbrich, C.; Westphal, K.; Baatout, S.; Wehland, M.; Bauer, J.; Flick, B.; Infanger, M.; Kreutz, R.; Vadrucci, S.; Egli, M.; et al. Effects of basic fibroblast growth factor on endothelial cells under conditions of simulated microgravity. J. Cell Biochem 2008, 104, 1324–1341. [Google Scholar]
- Ulbrich, C.; Westphal, K.; Pietsch, J.; Winkler, H.D.; Leder, A.; Bauer, J.; Kossmehl, P.; Grosse, J.; Schöneberger, J.; Infanger, M.; et al. Characterization of human chondrocytes exposed to simulated microgravity. Cell Physiol. Biochem 2010, 25, 551–560. [Google Scholar]
- Ulbrich, C.; Leder, A.; Pietsch, J.; Flick, B.; Wehland, M.; Grimm, D. The impact of vascular endothelial growth factor and basic fibroblast growth factor on cardiac fibroblast grown under altered gravity conditions. Cell Physiol. Biochem 2010, 26, 1011–1022. [Google Scholar]
- Grimm, D.; Wise, P.; Lebert, M.; Richter, P.; Baatout, S. How and why does the proteome respond to microgravity? Expert Rev. Proteomics 2011, 8, 13–27. [Google Scholar]
- Pietsch, J.; Bauer, J.; Egli, M.; Infanger, M.; Wise, P.; Ulbrich, C.; Grimm, D. The effects of weightlessness on the human organism and mammalian cells. Curr. Mol. Med 2011, 11, 350–364. [Google Scholar]
- Infanger, M.; Ulbrich, C.; Baatout, S.; Wehland, M.; Kreutz, R.; Bauer, J.; Grosse, J.; Vadrucci, S.; Cogoli, A.; Derradji, H.; et al. Modeled gravitational unloading induced downregulation of endothelin-1 in human endothelial cells. J. Cell Biochem 2007, 101, 1439–1455. [Google Scholar]
- Santini, M.T.; Rainaldi, F. Three-dimensional spheroid model in tumor biology. Pathobiology 1999, 67, 148–157. [Google Scholar]
- Infanger, M.; Kossmehl, P.; Shakibaei, S.; Baatout, S.; Witzing, A.; Grosse, J.; Bauer, J.; Cogoli, A.; Faramarzi, S.; Derradji, H.; et al. Induction of three-dimensional assembly and increase in apoptosis of human endothelial cells by simulated microgravity: Impact of vascular endothelial growth factor. Apoptosis 2006, 11, 749–764. [Google Scholar]
- Grimm, D.; Bauer, J.; Schönberger, J. Blockade of neoangiogenesis, a new and promising technique to control the growth of malignant tumors and their metastases. Curr. Vasc. Pharmacol 2009, 7, 374–357. [Google Scholar]
- Hirschhaeuser, F.; Menne, H.; Dittfeld, C.; West, J.; Mueller-Klieser, W.; Kunz-Schughart, L.A. Multicellular tumor spheroids: An underestimated tool is catching up again. J. Biotechnol 2010, 148, 3–15. [Google Scholar]
- Ulbrich, C.; Pietsch, J.; Grosse, J.; Wehland, M.; Schulz, H.; Saar, K.; Hubner, N.; Hauslage, J.; Hemmersbach, R.; Braun, M.; et al. Differential gene regulation under altered gravity conditions in follicular thyroid cancer cells: Relationship between the extracellular matrix and the cytoskeleton. Cell Physiol. Biochem 2011, 28, 185–198. [Google Scholar]
- Obermaier, C.; Jankowski, V.; Schmutzler, C.; Bauer, J.; Wildgruber, R.; Infanger, M.; Köhrle, J.; Krause, E.; Weber, G.; Grimm, D. Free-flow isoelectric focusing of proteins remaining in cell fragments following sonication of thyroid carcinoma cells. Electrophoresis 2005, 26, 2109–2116. [Google Scholar]
- Pietsch, J.; Kussian, R.; Sickmann, A.; Bauer, J.; Weber, G.; Nissum, M.; Westphal, K.; Egli, M.; Grosse, J.; Schönberger, J.; et al. Application of free-flow IEF to identify protein candidates changing under microgravity conditions. Proteomics 2010, 10, 904–913. [Google Scholar]
- Pietsch, J.; Sickmann, A.; Weber, G.; Bauer, J.; Egli, M.; Wildgruber, R.; Infanger, M.; Grimm, D. A proteomic approach to analyzing spheroid formation of two human thyroid cell lines cultured on a random positioning machine. Protemics 2011, 11, 2095–2104. [Google Scholar]
- Pietsch, J.; Sickmann, A.; Weber, G.; Bauer, J.; Egli, M.; Wildgruber, R.; Infanger, M.; Grimm, D. Metabolic enzyme diversity in different human thyroid cell lines and their sensitivity to gravitational forces. Proteomics 2012, 12, 1–8. [Google Scholar]
- Pietsch, J.; Bauer, J.; Weber, G.; Nissum, M.; Westphal, K.; Egli, M.; Grosse, J.; Schönberger, J.; Eilles, C.; Infanger, M.; et al. Proteome analysis of thyroid cancer cell after long-term exposure to a random positioning machine. Microgravity Sci. Technol 2011, 23, 381–390. [Google Scholar]
- Jensen, L.J.; Kuhn, M.; Stark, M.; Chaffron, S.; Muller, J.; Doerks, T.; Julien, P.; Roth, A.; Simonovic, M.; Bork, P.; et al. STRING 8—A global view on proteins and their functional interactions in 630 organisms. Nucleic Acids Res 2009, 37, D412–D416. [Google Scholar]
- Ma, X.; Wildgruber, R.; Bauer, J.; Weber, G.; Infanger, M.; Grosse, J.; Grimm, D. The use of sigmoid pH gradients in free-flow isoelectric focusing of human endothelial cell proteins. Electrophoresis 2012, 33, 1349–1355. [Google Scholar]
- Arpin, M.; Chirivino, D.; Naba, A.; Zwaenepoel, I. Emerging role for ERM proteins in cell adhesion and migration. Cell Adh. Migr 2011, 5, 199–206. [Google Scholar]
- Niggli, V.; Rossy, J. Ezrin/radixin/moesin: Versatile controllers of signaling molecules and of the cortical cytoskeleton. Int. J. Biochem. Cell Biol 2008, 40, 344–349. [Google Scholar]
- Mackay, D.J.G.; Esch, F.; Furthmayr, H.; Hall, A. Rho- and Rac-dependent assembly of focal adhesion complexes and actin filaments in permeabilized fibroblasts: An essential role for ezrin/radixin/moesin proteins. J. Cell Biol 1997, 138, 927–938. [Google Scholar]
- Hall, A.; Nobes, C.D. Rho GTPases: Molecular switches that control the organization and dynamics of the actin cytoskeleton. Philos. Trans. R Soc. B 2000, 355, 965–970. [Google Scholar]
- Wilson, M.A. The role of cysteine oxidation in DJ-1 function and dysfunction. Antioxid. Redox Signal 2011, 15, 111–122. [Google Scholar]
- He, X.; Zheng, Z.; Li, J.; Ben, Q.; Liu, J.; Zhang, J.; Ji, J.; Yu, B.; Chen, X.; Su, L.; et al. DJ-1 promotes invasion and metastasis of pancreatic cancer cells by activating SRC/ERK/uPA. Carcinogenesis 2012, 33, 555–562. [Google Scholar]
- Mhawech, P. 14-3-3 proteins—an update. Cell Res 2005, 15, 228–236. [Google Scholar]
- Sorimachi, H.; Hata, S.; Ono, Y. Calpain chronicle-an enzyme family under multidisciplinary characterization. Proc. Jpn. Acad. Ser. B 2011, 87, 287–327. [Google Scholar]
- Humphries, J.D.; Byron, A.; Bass, M.D.; Craig, S.E.; Pinney, J.W.; Knight, D.; Humphries, M.J. Proteomic analysis of integrin-associated complexes identifies RCC2 as a Dual regulator of Rac1 and Arf6. Sci. Signal. 2009, 2. [Google Scholar] [CrossRef]
- Su, L.K.; Qui, Y. Characterization of human MAPRE genes and their proteins. Genomics 2001, 71, 142–149. [Google Scholar]
- Hammond, A.T.; Glick, B.S. Dynamics of transitional endoplasmic reticulum sites in vertebrate cells. Mol. Biol. Cell 2000, 11, 3013–3030. [Google Scholar]
- Riera, J.; Lazo, P.S. The mammalian NudC-like genes: A family with functions other than regulating nuclear distribution. Cell. Mol. Life Sci 2009, 66, 2383–2390. [Google Scholar]
- Papaseit, C.; Pochon, N.; Tabony, J. Microtubuli self-organization is gravity dependent. Proc. Natl. Acad. Sci. USA 2000, 97, 8364–8368. [Google Scholar]
- Meloni, M.A.; Galleri, G.; Pippia, P.; Cogoli-Greuter, M. Cytoskeleton changes and impaired motility of monocytes at modeled low gravity. Protoplasma 2006, 229, 243–249. [Google Scholar]
- Infanger, M.; Kossmehl, P.; Shakibaei, M.; Bauer, J.; Kossmehl-Zorn, S.; Cogoli, A.; Curcio, F.; Oksche, A.; Wehland, M.; Kreutz, R.; et al. Simulated weightlessness changes the cytoskeleton and extracellular matrix proteins in papillary thyroid carcinoma cells. Cell Tissue Res 2006, 324, 267–277. [Google Scholar]
- Schneider, G.; Nieznanski, K.; Jozwiak, J.; Slomnicki, L.P.; Redowicz, M.J.; Filipek, A. Tubulin binding protein, CacyBP/SIP, induces actin polymerization and may link actin and tubulin cytoskeletons. Biochim. Biophys. Acta 2010, 1803, 1308–1317. [Google Scholar]
- Bernstein, B.W.; Bamburg, J.R. ADF/Cofilin: A functional node in cell biology. Trends Cell Biol 2010, 20, 187–195. [Google Scholar]
- Jong, J.H.; Yoon, T.; Choi, E.C.; Lee, K. Interaction of cofilin with triose-phosphate isomerase contributes glycolytic fuel for Na,K-ATPase via Rho-mediated signaling pathway machine. J. Biol. Chem 2002, 277, 48931–48937. [Google Scholar]
- Burkard, T.R.; Planyavsky, M.; Kaupe, I.; Breitwieser, F.P.; Burckstummer, T.; Bennett, K.L.; Superti-Furga, G.; Colinge, J. Initial characterization of the human central proteome. BMC Syst. Biol 2011, 5, 17. [Google Scholar]
- Zahid, S.; Oellerich, M.; Asif, A.R.; Ahmed, N. Phosphoproteome profiling of substantia nigra and cortex regions of Alzheimer’s disease patients. J. Neurochem 2012, 121, 954–963. [Google Scholar]
- Liu, Y.S.; Teng, X.H.; Yang, X.X.; Song, Q.; Lu, R.; Xiong, J.X.; Liu, B.; Zeng, N.J.; Zeng, Y.; Long, J.; et al. Shotgun proteomics and network analysis between plasma membrane and extracellular matrix proteins from rat olfactory ensheathing cells. Cell Transplantat 2010, 19, 133–146. [Google Scholar]
- Muller, T.; Schrotter, A.; Loosse, C.; Helling, S.; Stephan, C.; Ahrens, M.; Uszkoreit, J.; Eisenacher, M.; Meyer, H.E.; Marcus, K. Sense and nonsense of pathway analysis software in proteomics. J. Proteome Res 2011, 10, 5398–5408. [Google Scholar]
- Curcio, F.; Ambesi-Impiombato, F.S.; Perrella, G.; Coon, H.G. Long-term culture and functional characterization of follicular cells from adult normal thyroids. Proc. Natl. Acad. Sci. USA 1994, 91, 9004–9008. [Google Scholar]
- Goretzki, P.E.; Frilling, A.; Simon, D.; Roeher, H.D. Growth regulation of normal thyroids and thyroid tumors in man. Recent Results Cancer Res 1990, 118, 48–63. [Google Scholar]
- Lin, J.-D.; Chao, T.-C.; Weng, H.-F.; Huang, H.-S.; Ho, Y.-S. Establishment of xenografts and cell lines from well-differentiated human thyroid carcinoma. J. Surg. Oncol 1996, 63, 112–118. [Google Scholar]
- Perkins, D.N.; Pappin, D.J.; Creasy, D.M.; Cottrell, J.S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 1999, 20, 3551–3567. [Google Scholar]
- Kossmehl, P.; Kurth, E.; Faramarzi, S.; Habighorst, B.; Shakibaei, M.; Wehland, M.; Kreutz, R.; Infanger, M.; Danser, A.H.J.; Grosse, J.; et al. Mechanisms of apoptosis after ischemia and reperfusion: Role of the renin-angiotensin system. Apoptosis 2006, 11, 347–358. [Google Scholar]
- Infanger, M.; Faramarzi, S.; Grosse, J.; Kurth, E.; Ulbrich, C.; Bauer, J.; Wehland, M.; Kreutz, R.; Kossmehl, P.; Paul, M.; et al. Expression of vascular endothelial growth factor and receptor tyrosine kinases in cardiac ischemia/reperfusion injury. Cardovasc. Pathol 2007, 16, 291–299. [Google Scholar]
- Grimm, D.; Jabusch, H.C.; Kossmehl, P.; Huber, M.; Fredersdorf, S.; Griese, D.P.; Krämer, B.K.; Kromer, E.P. Experimental diabetes and left ventricular hypertrophy: Effects of beta-receptor blockade. Cardovasc. Pathol 2002, 11, 229–237. [Google Scholar]
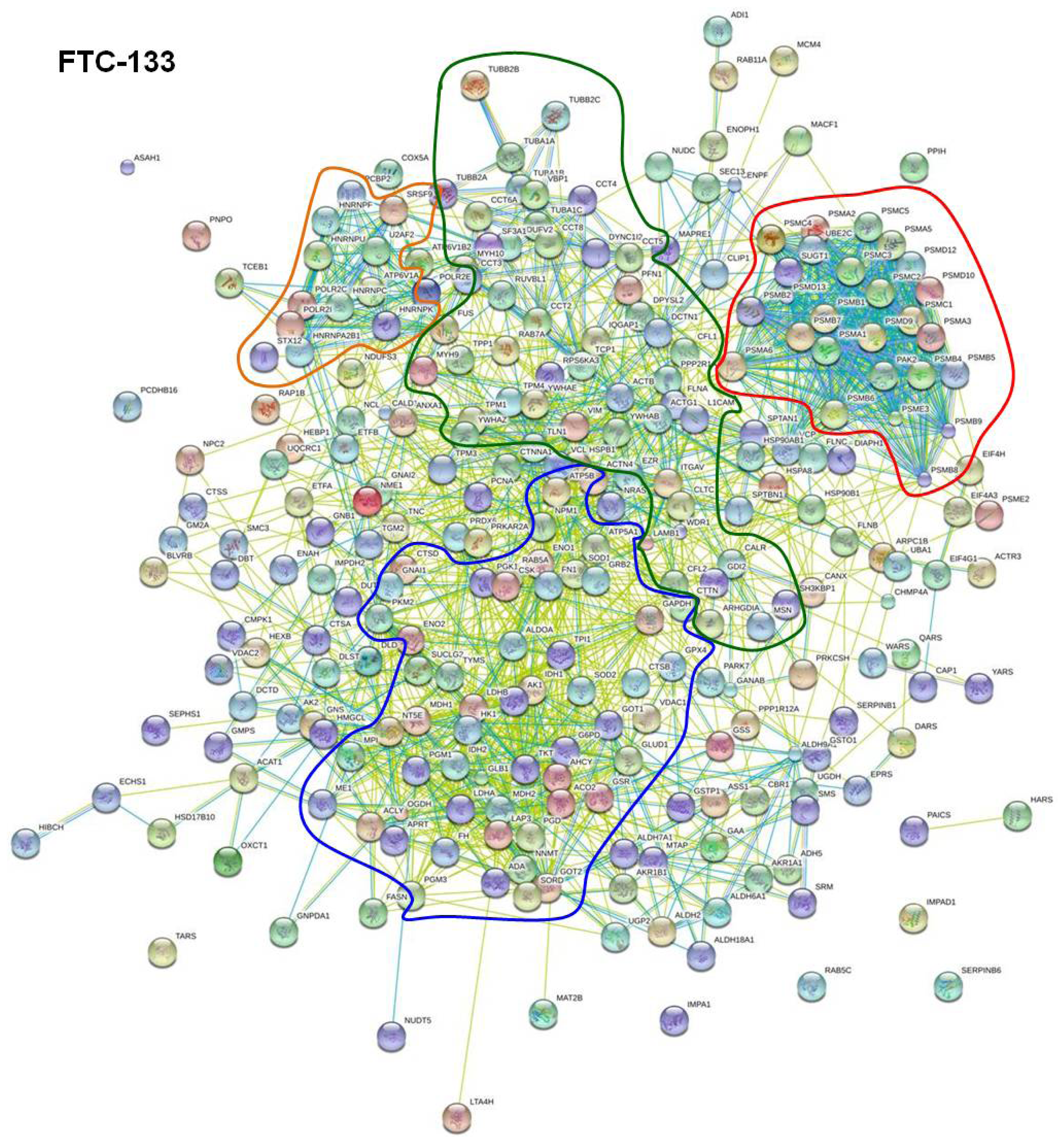
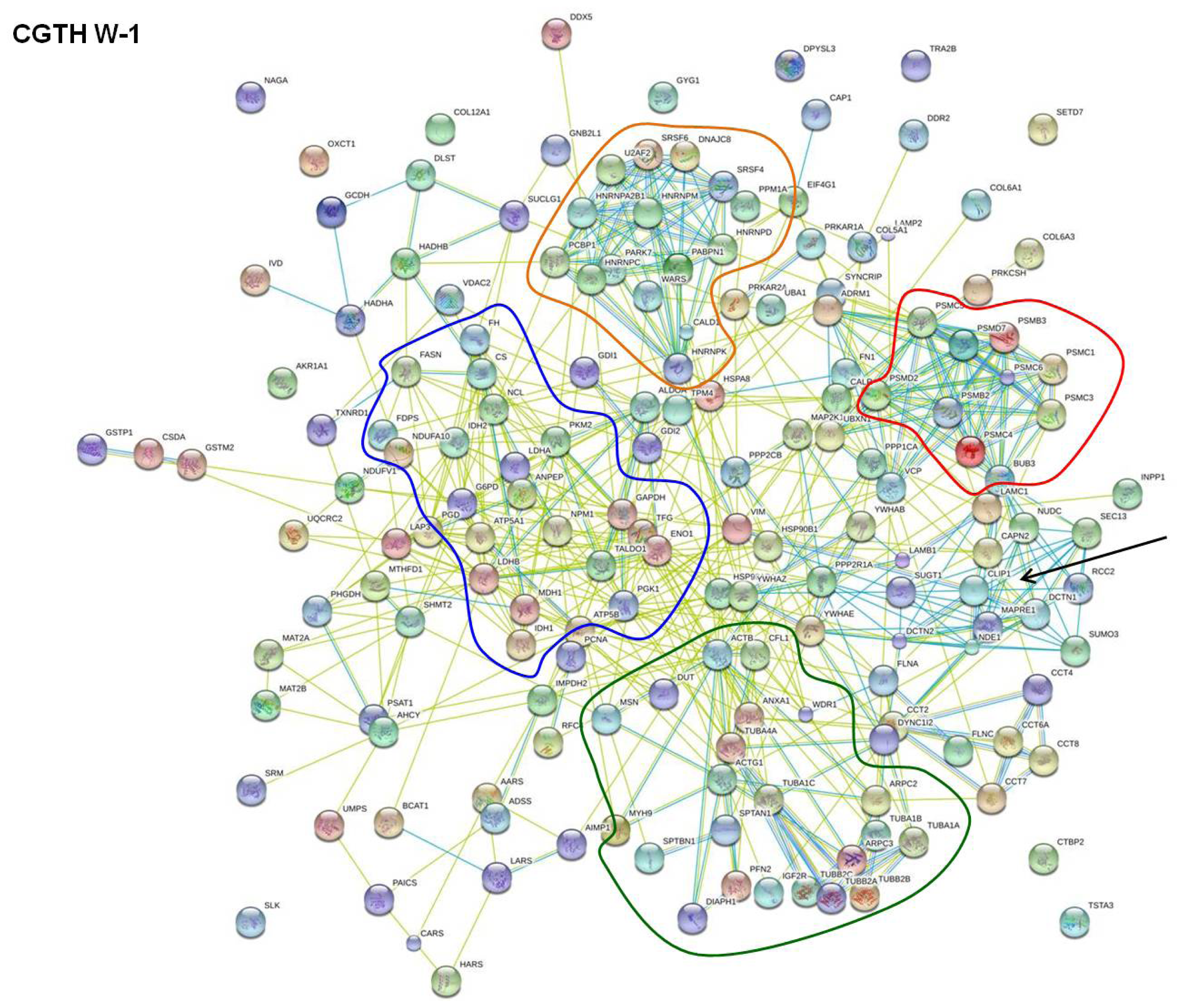

| Functionality of the interacting proteins | Found in FTC-133 cells | Found in CGTH W-1 cells | Found in HUT-5 cells |
|---|---|---|---|
| Carbohydrate metabolism | 106 | 53 | 63 |
| Maintaining cell structure | 73 | 43 | 58 |
| Protein synthesis/transcription | 38 | 32 | 14 |
| Degradation | 28 | 13 | 8 |
| Regulation | 109 | 70 | 64 |
© 2013 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Pietsch, J.; Riwaldt, S.; Bauer, J.; Sickmann, A.; Weber, G.; Grosse, J.; Infanger, M.; Eilles, C.; Grimm, D. Interaction of Proteins Identified in Human Thyroid Cells. Int. J. Mol. Sci. 2013, 14, 1164-1178. https://doi.org/10.3390/ijms14011164
Pietsch J, Riwaldt S, Bauer J, Sickmann A, Weber G, Grosse J, Infanger M, Eilles C, Grimm D. Interaction of Proteins Identified in Human Thyroid Cells. International Journal of Molecular Sciences. 2013; 14(1):1164-1178. https://doi.org/10.3390/ijms14011164
Chicago/Turabian StylePietsch, Jessica, Stefan Riwaldt, Johann Bauer, Albert Sickmann, Gerhard Weber, Jirka Grosse, Manfred Infanger, Christoph Eilles, and Daniela Grimm. 2013. "Interaction of Proteins Identified in Human Thyroid Cells" International Journal of Molecular Sciences 14, no. 1: 1164-1178. https://doi.org/10.3390/ijms14011164




