Expression Analysis of Four Peroxiredoxin Genes from Tamarix hispida in Response to Different Abiotic Stresses and Exogenous Abscisic Acid (ABA)
Abstract
:1. Introduction
2. Results
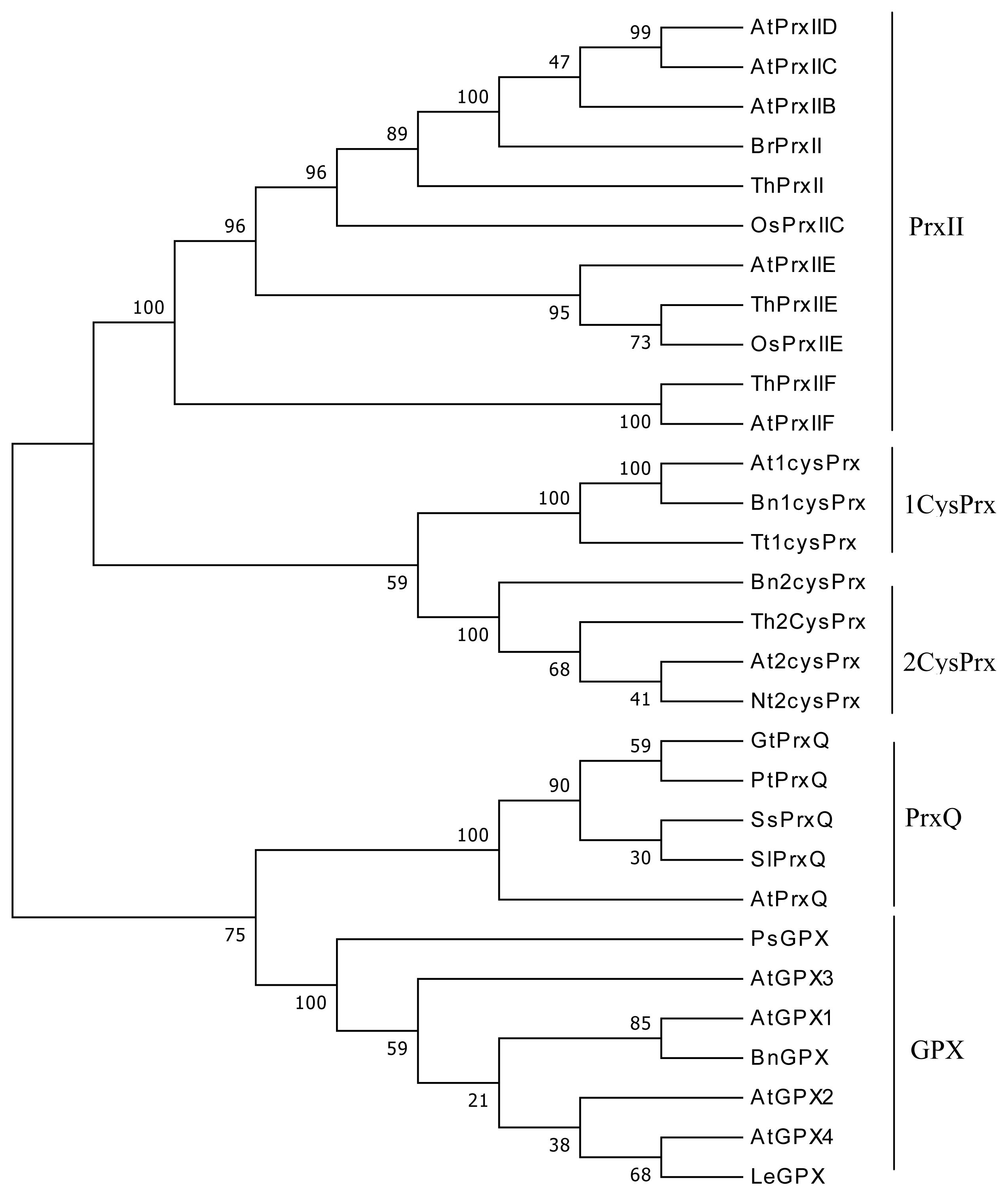
2.1. Cloning and Sequence Analysis of Four ThPrx Genes
2.2. Relative Expression Levels of Four ThPrx Genes in Roots, Stems and Leaves
2.3. Expression Profiles of ThPrx Genes in Response to Various Stresses
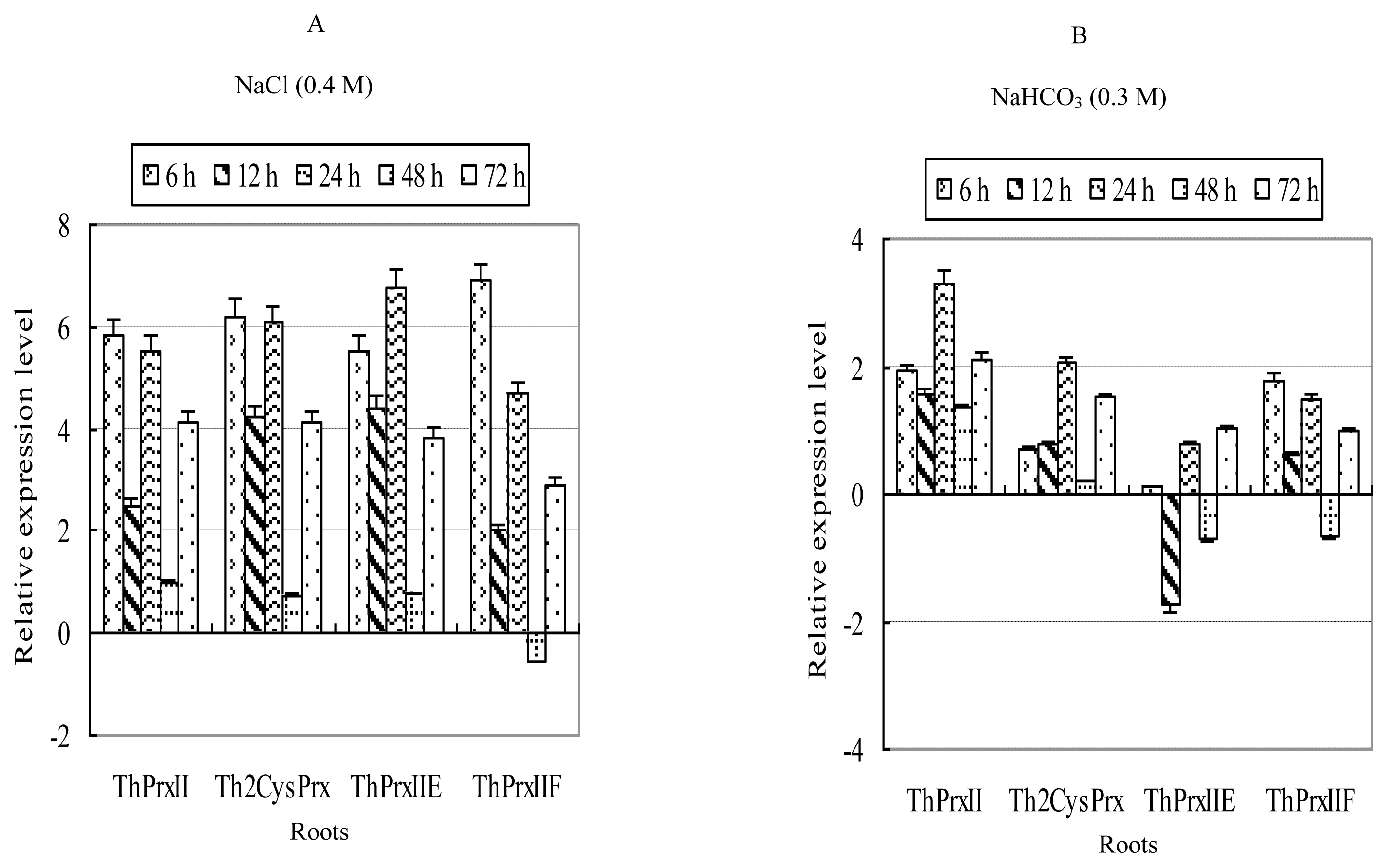
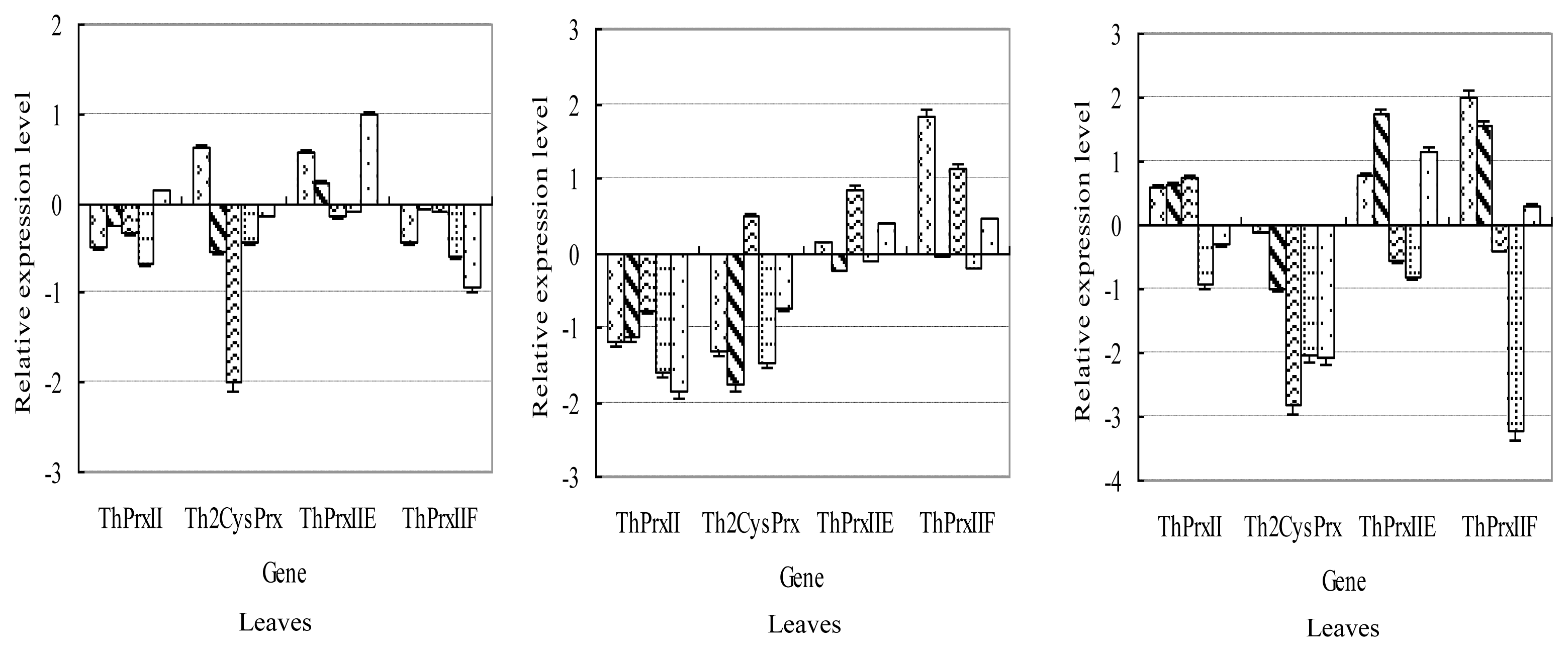
2.3.1. NaCl Stress
2.3.2. NaHCO3 Stress
2.3.3. PEG Stress
2.3.4. CdCl2 Stress
2.3.5. ABA Application
3. Discussion
4. Experimental Section
4.1. Plant Materials and Stress Treatments
4.2. Cloning and Identification of 4 ThPrx Genes
4.3. Sequence Alignments and Phylogenetic Analysis
4.4. RNA Extraction and Reverse Transcription (RT)
4.5. Real-time Quantitative RT-PCR
5. Conclusions
Acknowledgments
References
- Stacy, R.A.; Munthe, E.; Steinum, T.; Sharma, B.; Aalen, R.B. A peroxiredoxin antioxidant is encoded by a dormancy related gene, Per1, expressed during late development in the aleurone and embryo of barley grains. Plant Mol. Biol 1996, 31, 1205–1216. [Google Scholar]
- Baier, M.; Dietz, K.J. Primary structure and expression of plant homologues of animal and fungal thioredoxin-dependent peroxide reductases and bacterial alkyl hydroperoxide reductases. Plant Mol. Biol 1996, 31, 553–564. [Google Scholar]
- Pulido, P.; Spínola, M.C.; Kirchsteiger, K.; Guinea, M.; Pascual, M.B.; Sahrawy, M.; Sandalio, L.M.; Dietz, K.J.; González, M.; Cejudo, F.J. Functional analysis of the pathways for 2-Cys peroxiredoxin reduction in Arabidopsis thaliana chloroplasts. J. Exp. Bot 2010, 61, 4043–4054. [Google Scholar]
- Umate, P. Genome-wide analysis of thioredoxin fold superfamily peroxiredoxins in Arabidopsis and rice. Plant Signal Behav 2010, 5, 1543–1546. [Google Scholar]
- Horling, F.; Baier, M.; Dietz, K.J. Redox-regulation of the expression of the peroxide- detoxifying chloroplast 2-cys peroxiredoxin in the liverwort Riccia. fluitans. Planta 2001, 214, 304–313. [Google Scholar]
- Baier, M.; Dietz, K.J. The plant 2-Cys peroxiredoxin BAS1 is a nuclear-encoded chloroplast protein: its expressional regulation, phylogenetic origin, and implications for its specific physiological function in plants. Plant J 1997, 12, 179–190. [Google Scholar]
- Rouhier, N.; Gelhaye, E.; Sautiere, P.E.; Brun, A.; Laurent, P.; Tagu, D.; Gerard, J.; de Fay, E.; Meyer, Y.; Jacquot, J.P. Isolation and characterization of a new peroxiredoxin from poplar sieve tubes that uses either glutaredoxin or thioredoxin as a proton donor. Plant Physiol 2001, 127, 1299–1309. [Google Scholar]
- Kiba, A.; Nishihara, M.; Tsukatani, N.; Nakatsuka, T.; Kato, Y.; Yamamura, S. A peroxiredoxin Q homolog from gentians is involved in both resistance against fungal disease and oxidative stress. Plant Cell Physiol 2005, 46, 1007–1015. [Google Scholar]
- Berberich, T.; Uebler, M.; Feierabend, J. Cloning of a cDNA encoding a thioredoxin peroxidase (TPx) homolog from winter rye (Secale cereale L.) (Accession No. AF076920) (PGR98-167). Plant Physiol 1998, 118, 711. [Google Scholar]
- Bréhélin, C.; Meyer, E.H.; de Souris, J.P.; Bonnard, G.; Meyer, Y. Resemblance and dissemblance of Arabidopsis type II peroxiredoxins: similar sequences for divergent gene expression, protein localization, and activity. Plant Physiol 2003, 132, 2045–2057. [Google Scholar]
- Kim, M.D.; Kim, Y.H.; Kwon, S.Y.; Jang, B.Y.; Lee, S.Y.; Yun, D.J.; Cho, J.H.; Kwak, S.S.; Lee, H.S. Overexpression of 2-cysteine peroxiredoxin enhances tolerance to methyl viologen-mediated oxidative stress and high temperature in potato plants. Plant Physiol. Biochem 2011, 49, 891–897. [Google Scholar]
- Finkemeier, I.; Goodman, M.; Lamkemeyer, P.; Kandlbinder, A.; Sweetlove, L.J.; Dietz, K.J. The mitochondrial type II peroxiredoxin F is essential for redox homeostasis and root growth of Arabidopsis thaliana under stress. J. Biol. Chem 2005, 280, 12168–12180. [Google Scholar]
- Haslekås, C.; Viken, M.K.; Grini, P.E.; Nygaard, V.; Nordgard, S.H.; Meza, T.J.; Aalen, R.B. Seed 1-cysteine peroxiredoxin antioxidants are not involved in dormancy, but contribute to inhibition of germination during stress. Plant Physiol 2003, 133, 1148–1157. [Google Scholar]
- Lee, K.O.; Jang, H.H.; Jung, B.G.; Chi, Y.H.; Lee, J.Y.; Choi, Y.O.; Lee, J.R.; Lim, C.O.; Cho, M.J.; Lee, S.Y. Rice 1Cys-peroxiredoxin over-expressed in transgenic tobacco does not maintain dormancy but enhances antioxidant activity. FEBS Lett. 2000, 486, 103–106. [Google Scholar]
- Kim, S.Y.; Jang, H.H.; Lee, J.R.; Sung, N.R.; Lee, H.B.; Lee, D.H.; Park, D.J.; Kang, C.H.; Chung, W.S.; Lim, C.O.; et al. Oligomerization and chaperone activity of a plant 2-Cys peroxiredoxin in response to oxidative stress. Plant Sci 2009, 177, 227–232. [Google Scholar]
- Kim, S.Y.; Paeng, S.K.; Nawkar, G.M.; Maibam, P.; Lee, E.S.; Kim, K.S.; Lee, D.H.; Park, D.J.; Kang, S.B.; Kim, M.R.; et al. The 1-Cys peroxiredoxin, a regulator of seed dormancy, functions as a molecular chaperone under oxidative stress conditions. Plant Sci 2011, 181, 119–124. [Google Scholar]
- Jing, L.W.; Chen, S.H.; Guo, X.L.; Zhang, H.; Zhao, Y.X. Overexpression of a chloroplast-located peroxiredoxin Q gene, SsPrxQ, increases the salt and low-temperature tolerance of Arabidopsis. J. Integr. Plant Biol 2006, 48, 1244–1249. [Google Scholar]
- Kim, K.H.; Alam, I.; Lee, K.W.; Sharmin, S.A.; Kwak, S.S.; Lee, S.Y.; Lee, B.H. Enhanced tolerance of transgenic tall fescue plants overexpressing 2-Cys peroxiredoxin against methyl viologen and heat stresses. Biotechnol Lett 2010, 32, 571–576. [Google Scholar]
- Muthuramalingam, M.; Seidel, T.; Laxa, M.; Nunes de Miranda, S.M.; Gärtner, F.; Ströher, E.; Kandlbinder, A.; Dietz, K.J. Multiple redox and non-redox interactions define 2-Cys peroxiredoxin as a regulatory hub in the chloroplast. Mol. Plant. 2009, 2, 1273–1288. [Google Scholar]
- Dietz, K.J. Peroxiredoxins in Plants and Cyanobacteria. Antioxid. Redox Signal 2011, 15, 1129–1159. [Google Scholar]
- Mowla, S.B.; Thomson, J.A.; Farrant, J.M.; Mundree, S.G. A novel stress-inducible antioxidant enzyme identified from the resurrection plant Xerophyta viscosa Baker. Planta 2002, 215, 716–726. [Google Scholar]
- Horling, F.; Lamkemeyer, P.; König, J.; Finkemeier, I.; Kandlbinder, A.; Baier, M.; Dietz, K.J. Divergent light-, ascorbate-, and oxidative stress-dependent regulation of expression of the peroxiredoxin gene family in Arabidopsis. Plant Physiol 2003, 131, 317–325. [Google Scholar]
- Barranco-Medina, S.; Krell, T.; Bernier-Villamor, L.; Sevilla, F.; Lazaro, J.J.; Dietz, K.J. Hexameric oligomerization of mitochondrial peroxiredoxin PrxIIF and formation of an ultrahigh affinity complex with its electron donor thioredoxin Trx-o. J. Exp. Bot 2008, 59, 3259–3269. [Google Scholar]
- Haslekås, C.; Grini, P.E.; Nordgard, S.H.; Thorstensen, T.; Viken, M.K.; Nygaard, V.; Aalen, R.B. ABI3 mediates expression of the peroxiredoxin antioxidant AtPER1 gene and induction by oxidative stress. Plant Mol. Biol 2003, 53, 313–326. [Google Scholar]
- Baier, M.; Ströher, E.; Dietz, K.J. The acceptor availability at photosystem I and ABA control nuclear expression of 2-Cys peroxiredoxin-A in Arabidopsis thaliana. Plant Cell Physiol 2004, 45, 997–1006. [Google Scholar]
- Gao, C.; Wang, Y.; Liu, G.; Yang, C.; Jiang, J.; Li, H. Expression profiling of salinity-alkali stress responses by large-scale expressed sequence tag analysis in Tamarix hispid. Plant Mol. Biol. 2008, 66, 245–258. [Google Scholar]
- Li, H.; Wang, Y.; Jiang, J.; Liu, G.; Gao, C.; Yang, C. Identification of genes responsive to salt stress on Tamarix hispida roots. Gene 2009, 433, 65–71. [Google Scholar]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar]



| Gene | GenBank accession number | Type | Deduced number of amino acid | Isoelectric point | Molecular mass (kDa) |
|---|---|---|---|---|---|
| ThPrxII | JQ341201 | type II peroxiredoxin | 162 | 5.79 | 17.3 |
| Th2CysPrx | JQ341202 | 2Cys peroxiredoxin | 274 | 6.9 | 29.8 |
| ThPrxIIE | JQ341203 | type II peroxiredoxin | 227 | 8.57 | 24.0 |
| ThPrxIIF | JQ341204 | type II peroxiredoxin | 196 | 8.37 | 21.0 |
| Gene | Library | |||||
|---|---|---|---|---|---|---|
| Root libraries | Leaf libraries | |||||
| 0 h | 24 h | 48 h | 0 h | 24 h | 52 h | |
| ThPrxII | 1 | 0 | 0 | 0 | 1 | 2 |
| Th2CysPrx | 0 | 0 | 0 | 0 | 0 | 1 |
| ThPrxIIE | 0 | 0 | 0 | 2 | 1 | 0 |
| ThPrxIIF | 0 | 0 | 1 | 1 | 1 | 3 |
| Total | 1 | 0 | 1 | 3 | 3 | 6 |
| Gene | Relative abundance | ||
|---|---|---|---|
| Roots | Stems | Leaves | |
| ThPrxII | 83.4 | 155.6 | 226.9 |
| Th2CysPrx | 17.8 | 42.3 | 187.9 |
| ThPrxIIE | 5.6 | 6.3 | 30.4 |
| ThPrxIIF | 20.1 | 15.0 | 38.4 |
| Actin | 100 | 100 | 100 |
| Gene | Forward Primers (5′–3′) | Reverse Primers (5′–3′) |
|---|---|---|
| ThPrxII | TCAGCAGGTTTCAGTTCACT | CAGAGCCATCAGCAAGGA |
| Th2CysPrx | GAGAAGGCTTGGACTGAG | GAGGAACGGCAAGATGAG |
| ThPrxIIE | CCCTCTCCTATTTTGACTCC | TCAGCAGCAGGACTTCATC |
| ThPrxIIF | CTTCCTCATCGGAAATATGTCG | AACAAACACCTGTGTACGCACC |
| Actin | AAACAATGGCTGATGCTG | ACAATACCGTGCTCAATAGG |
| α-tubulin | CACCCACCGTTGTTCCAG | ACCGTCGTCATCTTCACC |
| β-tubulin | GGAAGCCATAGAAAGACC | CAACAAATGTGGGATGCT |
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Gao, C.; Zhang, K.; Yang, G.; Wang, Y. Expression Analysis of Four Peroxiredoxin Genes from Tamarix hispida in Response to Different Abiotic Stresses and Exogenous Abscisic Acid (ABA). Int. J. Mol. Sci. 2012, 13, 3751-3764. https://doi.org/10.3390/ijms13033751
Gao C, Zhang K, Yang G, Wang Y. Expression Analysis of Four Peroxiredoxin Genes from Tamarix hispida in Response to Different Abiotic Stresses and Exogenous Abscisic Acid (ABA). International Journal of Molecular Sciences. 2012; 13(3):3751-3764. https://doi.org/10.3390/ijms13033751
Chicago/Turabian StyleGao, Caiqiu, Kaimin Zhang, Guiyan Yang, and Yucheng Wang. 2012. "Expression Analysis of Four Peroxiredoxin Genes from Tamarix hispida in Response to Different Abiotic Stresses and Exogenous Abscisic Acid (ABA)" International Journal of Molecular Sciences 13, no. 3: 3751-3764. https://doi.org/10.3390/ijms13033751




