Conformational Changes in DNA upon Ligand Binding Monitored by Circular Dichroism
Abstract
:1. Introduction
2. Conformational Polymorphism of DNA and DNA-Drug Complexes Monitored by CD
2.1. DNA (B-form, A-form, Z-form and Quadruplexes)
2.2. Chromomycin A3(Chro)
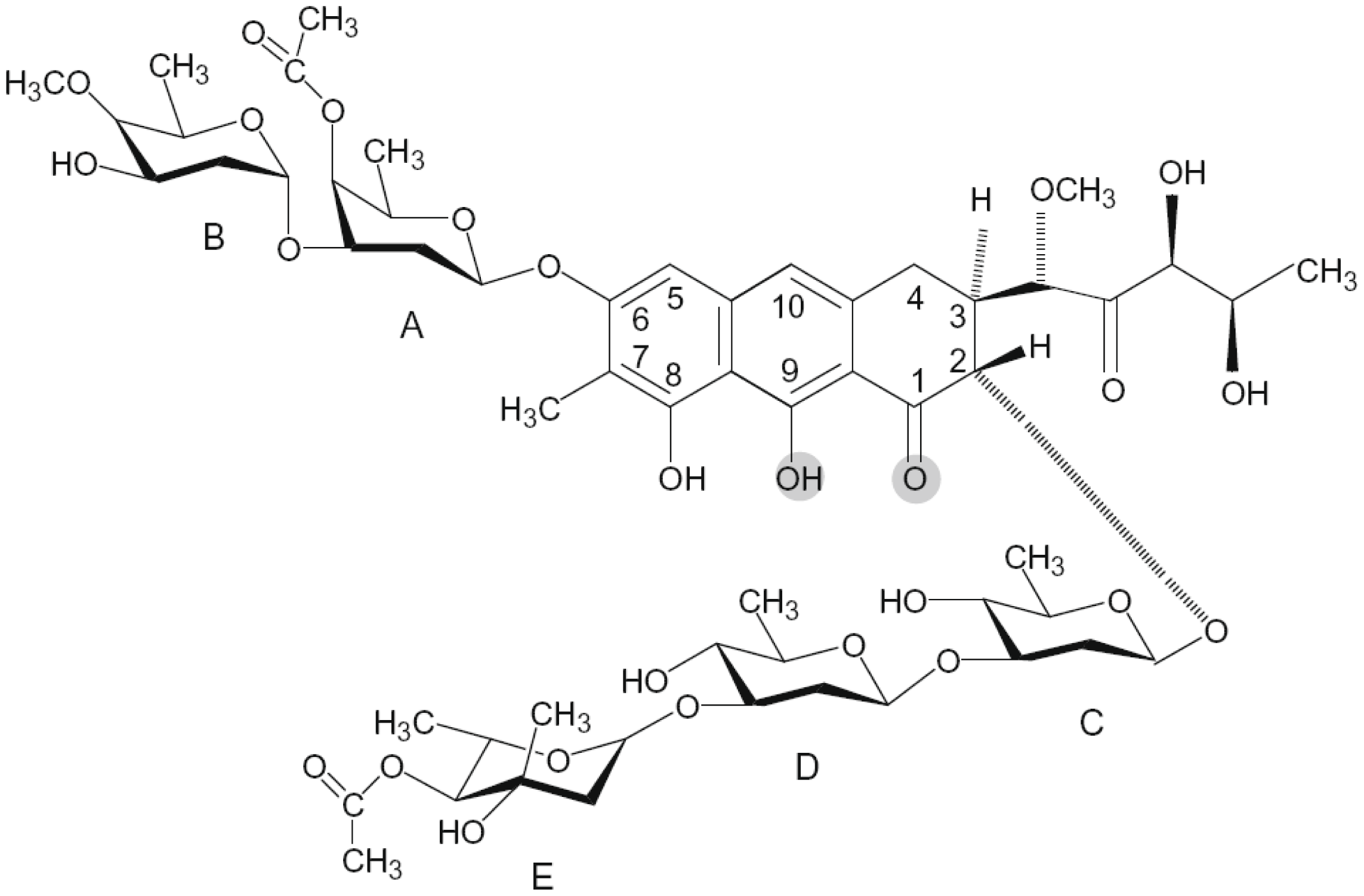
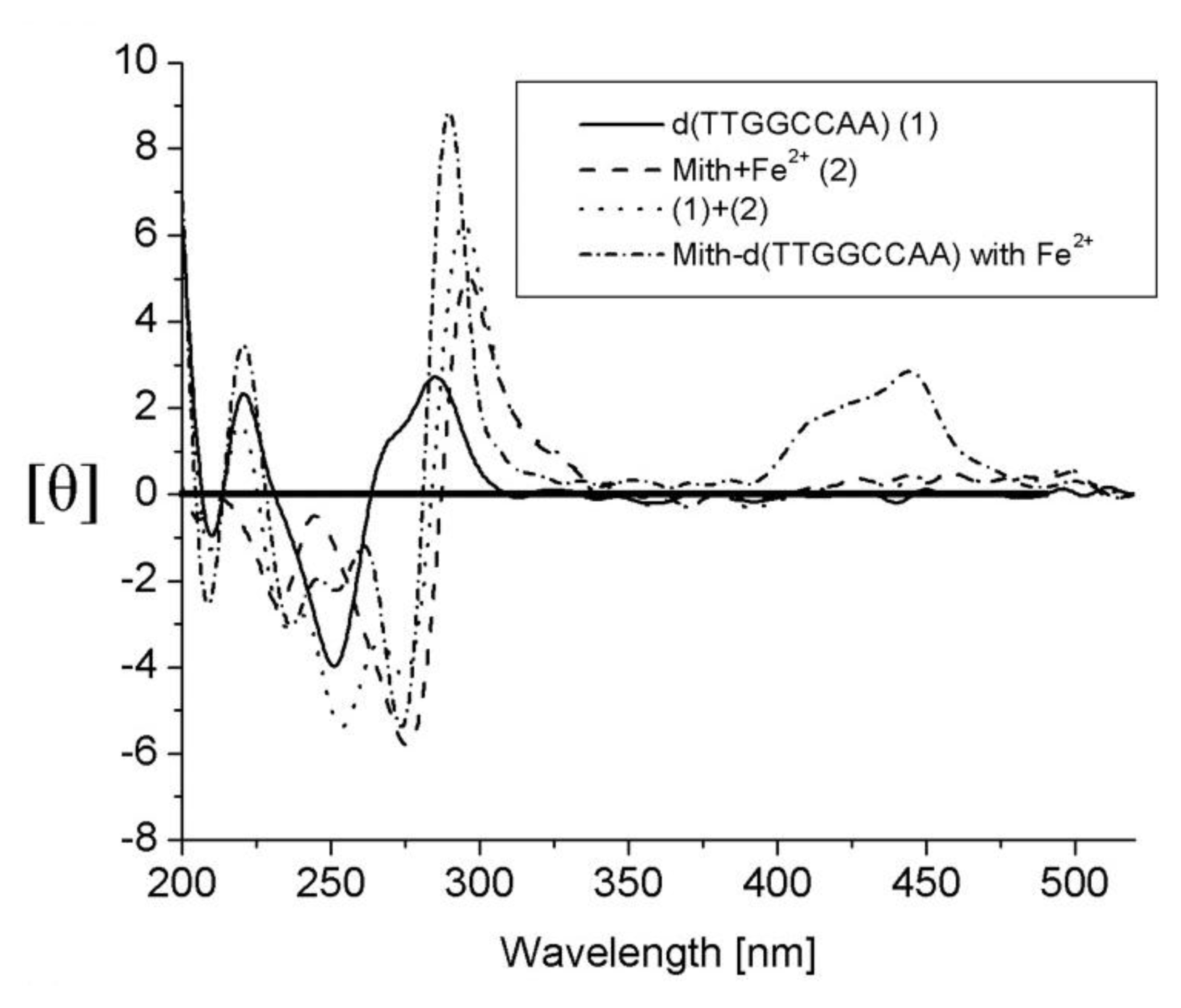
2.3. Mithramycin (Mith)
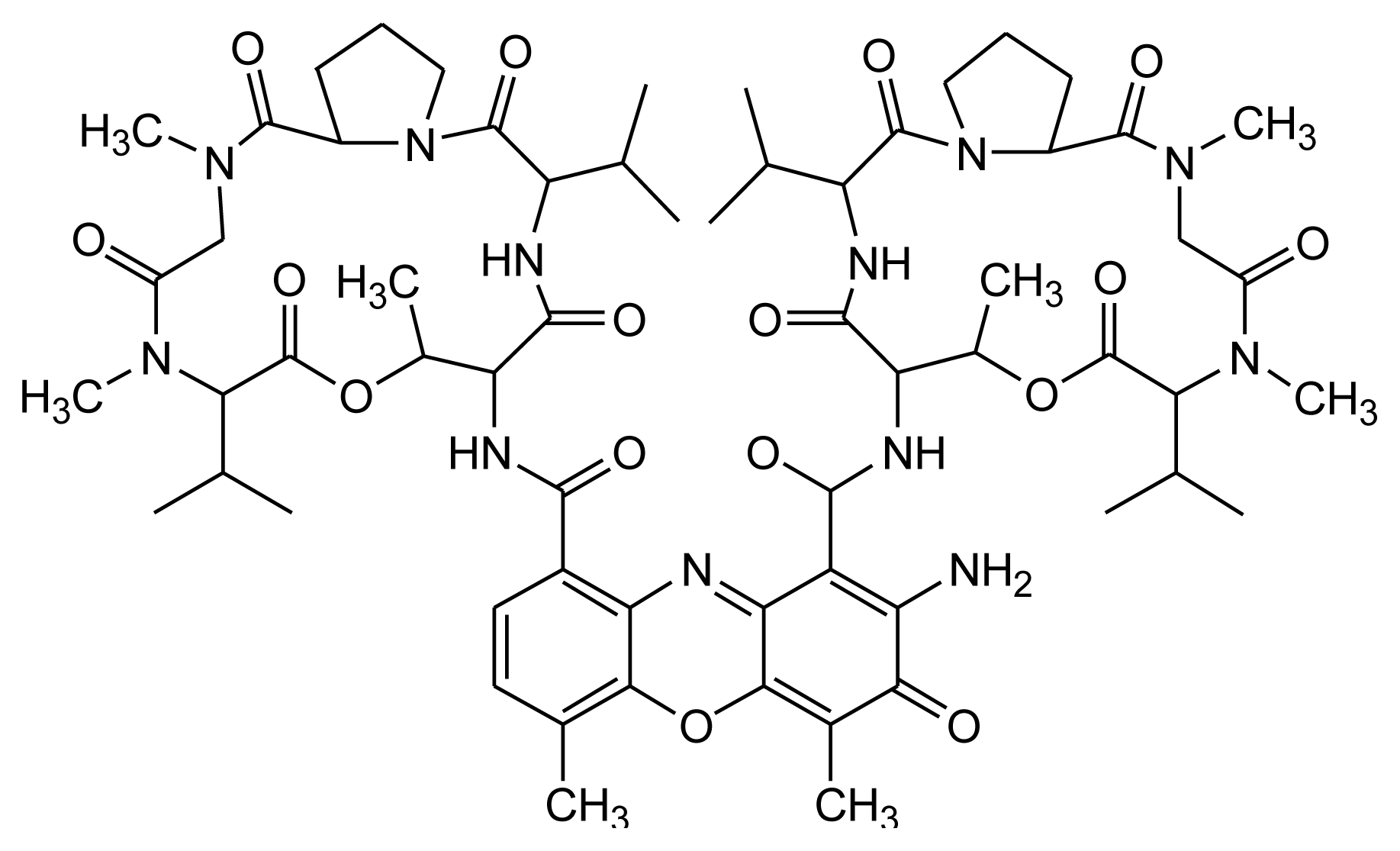
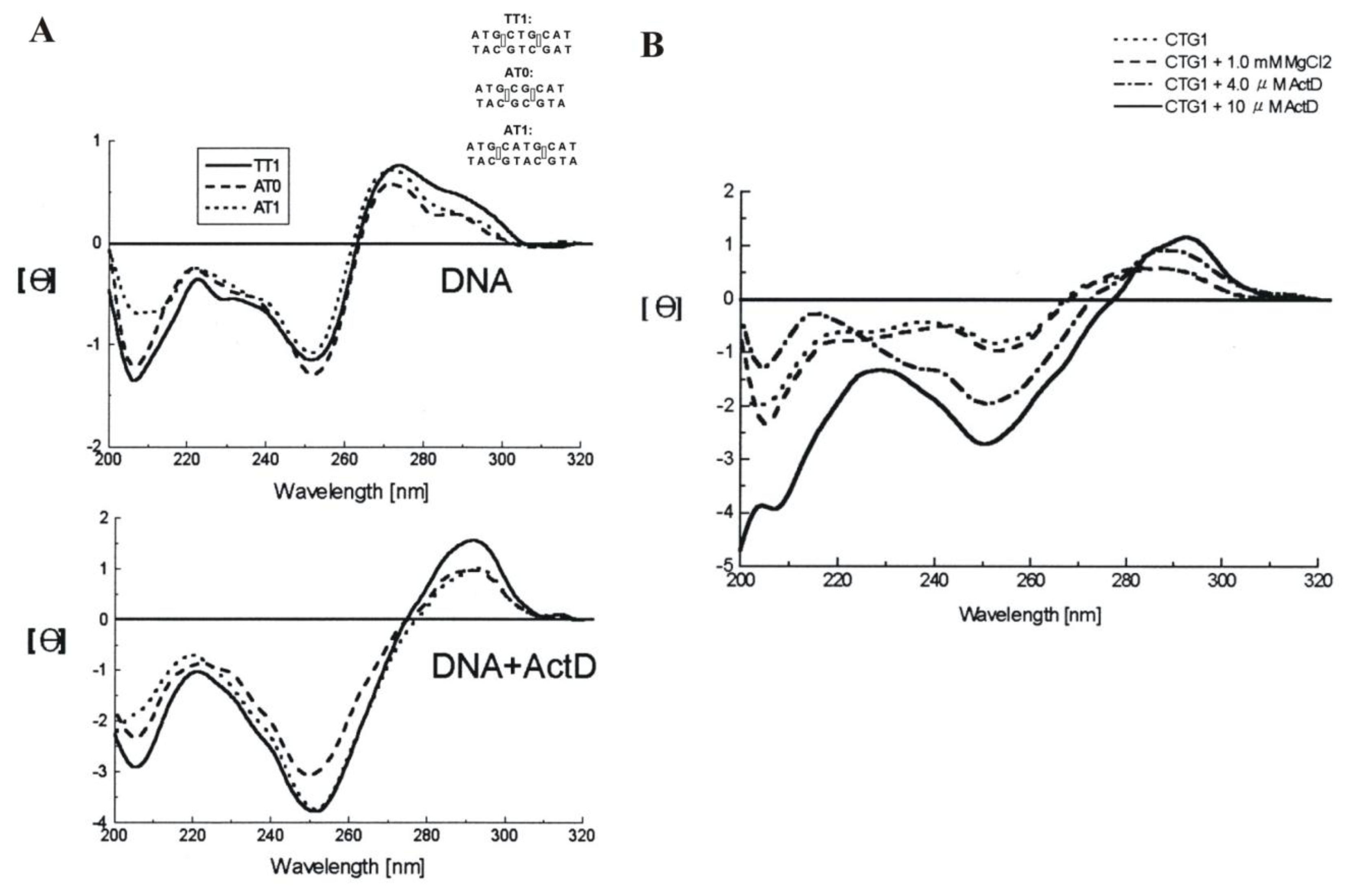
2.4. Actinomycin D (ActD)
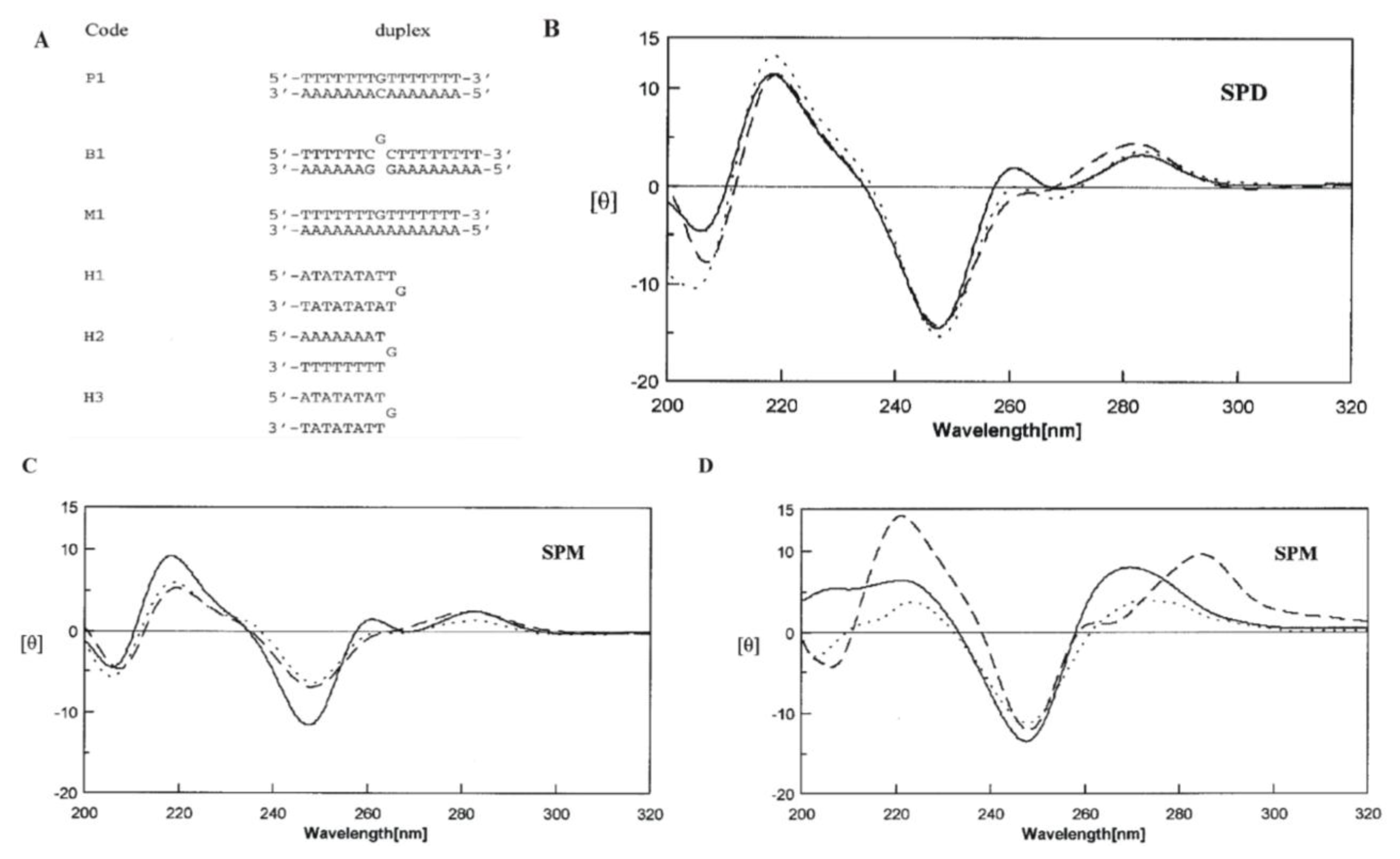
2.5. Polyamines
2.6. Neomycin
2.7. Cisplatin
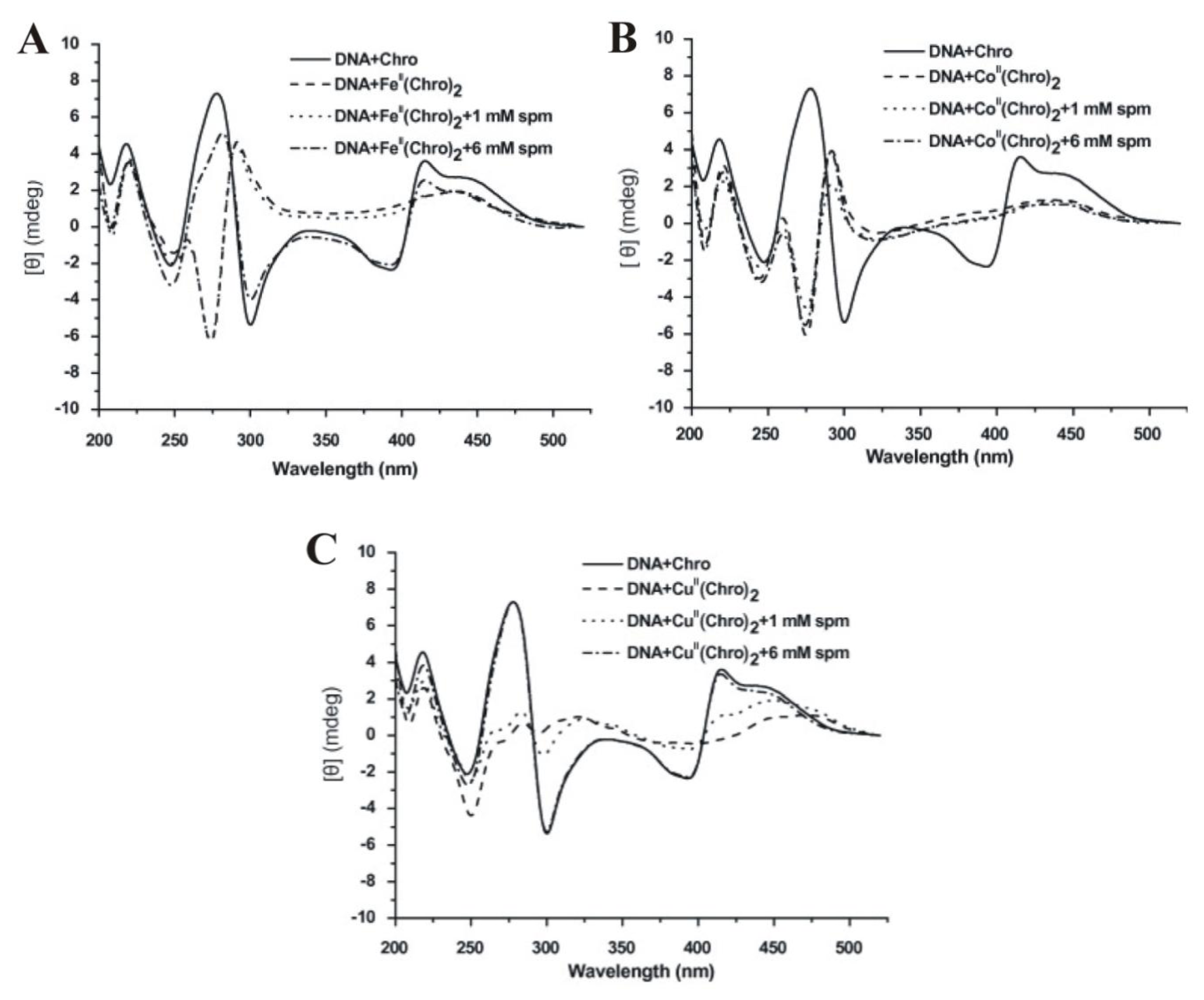
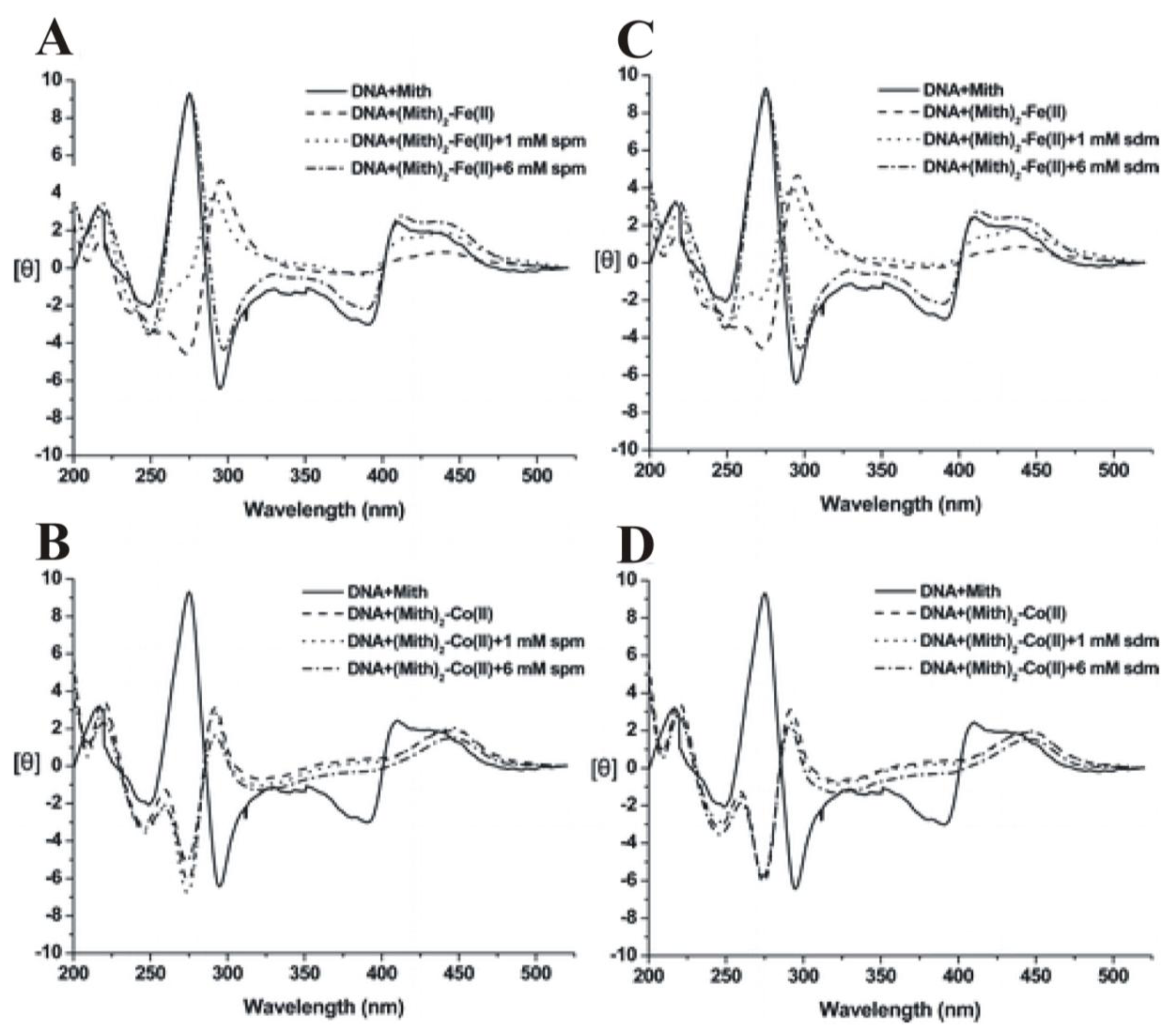
2.8. The Effect of Polyamines on the Metal Derivative Complex of Aureolic Family Drug-DNA Complexes
3. Conclusion
Acknowledgments
References
- Woody, R.W. Circular dichroism. Meth. Enzymology 1995, 246, 34–71. [Google Scholar]
- Lewis, D.G.; Johnson, W.C., Jr. Circular dichroism of DNA in the vacuum ultraviolet. J. Mol. Biol 1974, 86, 91–96. [Google Scholar]
- Holm, A.I.; Nielsen, L.M.; Hoffmann, S.V.; Nielsen, S.B. Vacuum-ultraviolet circular dichroism spectroscopy of DNA: A valuable tool to elucidate topology and electronic coupling in DNA. Phys. Chem. Chem. Phys 2010, 12, 9581–996. [Google Scholar]
- Matsuo, K.; Yonehara, R.; Gekko, K. Secondary-structure analysis of proteins by vacuum-ultraviolet circular dichroism spectroscopy. J. Biochem 2004, 135, 405–411. [Google Scholar]
- Sreerama, N.; Venyaminov, S.Y.; Woody, R.W. Estimation of protein secondary structure from circular dichroism spectra: Inclusion of denatured proteins with native proteins in the analysis. Anal. Biochem 2000, 287, 243–251. [Google Scholar]
- Porumb, H.; Monnot, M.; Fermandjian, S. Circular dichroism signatures of features simultaneously present in structured guanine-rich oligonucleotides: A combined spectroscopic and electrophoretic approach. Electrophoresis 2002, 23, 1013–1020. [Google Scholar]
- Zhou, J.; Yuan, G. Specific recognition of human telomeric G-quadruplex DNA with small molecules and the conformational analysis by ESI mass spectrometry and circular dichroism spectropolarimetry. Chemistry 2007, 13, 5018–5023. [Google Scholar]
- Andrushchenko, V.; Bour, P. Applications of the Cartesian coordinate tensor transfer technique in the simulations of vibrational circular dichroism spectra of oligonucleotides. Chirality 2010, 22, E96–E114. [Google Scholar]
- Daura, X.; Bakowies, D.; Seebach, D.; Fleischhauer, J.; van Gunsteren, W.F.; Kruger, P. Circular dichroism spectra of beta-peptides: Sensitivity to molecular structure and effects of motional averaging. Eur. Biophys. J 2003, 32, 661–670. [Google Scholar]
- Kypr, J.; Kejnovska, I.; Renciuk, D.; Vorlickova, M. Circular dichroism and conformational polymorphism of DNA. Nucleic Acids Res 2009, 37, 1713–1725. [Google Scholar]
- Kankia, B.I.; Buckin, V.; Bloomfield, V.A. Hexamminecobalt(III)-induced condensation of calf thymus DNA: Circular dichroism and hydration measurements. Nucleic Acids Res 2001, 29, 2795–2801. [Google Scholar]
- Maestre, M.F. Circular dichroism of DNA films: Reversibility studies. J. Mol. Biol 1970, 52, 543–556. [Google Scholar]
- Boer, D.R.; Canals, A.; Coll, M. DNA-binding drugs caught in action: The latest 3D pictures of drug-DNA complexes. 2009, 399–414. [Google Scholar]
- Palchaudhuri, R.; Hergenrother, P.J. DNA as a target for anticancer compounds: Methods to determine the mode of binding and the mechanism of action. Curr. Opin. Biotechnol 2007, 18, 497–503. [Google Scholar]
- Watson, J.D.; Crick, F.H. Molecular structure of nucleic acids; a structure for deoxyribose nucleic acid. Nature 1953, 171, 737–738. [Google Scholar]
- Nelson, H.C.; Finch, J.T.; Luisi, B.F.; Klug, A. The structure of an oligo(dA)·oligo(dT) tract and its biological implications. Nature 1987, 330, 221–226. [Google Scholar]
- Alexeev, D.G.; Lipanov, A.A.; Skuratovskii, I. Poly(dA)·poly(dT) is a B-type double helix with a distinctively narrow minor groove. Nature 1987, 325, 821–823. [Google Scholar]
- Trantirek, L.; Stefl, R.; Vorlickova, M.; Koca, J.; Sklenar, V.; Kypr, J. An A-type double helix of DNA having B-type puckering of the deoxyribose rings. J. Mol. Biol 2000, 297, 907–922. [Google Scholar]
- Ivanov, V.I.; Minchenkova, L.E.; Minyat, E.E.; Frank-Kamenetskii, M.D.; Schyolkina, A.K. The B to A transition of DNA in solution. J. Mol. Biol 1974, 87, 817–833. [Google Scholar]
- Stefl, R.; Trantirek, L.; Vorlickova, M.; Koca, J.; Sklenar, V.; Kypr, J. A-like guanine-guanine stacking in the aqueous DNA duplex of d(GGGGCCCC). J. Mol. Biol 2001, 307, 513–524. [Google Scholar]
- Jaworski, A.; Hsieh, W.T.; Blaho, J.A.; Larson, J.E.; Wells, R.D. Left-handed DNA in vivo. Science 1987, 238, 773–777. [Google Scholar]
- Lezius, A.G.; Gottschalk, E.M. A reversible cooperative conformational change of a synthetic DNA under the influence of high salt concentrations. Hoppe-Seylers Z. Physiol. Chem 1970, 351, 413–416. [Google Scholar]
- Ivanov, V.I.; Minyat, E.E. The transitions between left- and right-handed forms of poly(dG-dC). Nucleic Acids Res 1981, 9, 4783–4798. [Google Scholar]
- Balagurumoorthy, P.; Brahmachari, S.K.; Mohanty, D.; Bansal, M.; Sasisekharan, V. Hairpin and parallel quartet structures for telomeric sequences. Nucleic Acids Res 1992, 20, 4061–4067. [Google Scholar]
- Dapic, V.; Abdomerovic, V.; Marrington, R.; Peberdy, J.; Rodger, A.; Trent, J.O.; Bates, P.J. Biophysical and biological properties of quadruplex oligodeoxyribonucleotides. Nucleic Acids Res 2003, 31, 2097–2107. [Google Scholar]
- Paramasivan, S.; Rujan, I.; Bolton, P.H. Circular dichroism of quadruplex DNAs: Applications to structure, cation effects and ligand binding. Methods 2007, 43, 324–331. [Google Scholar]
- Foley, J.M. The size-distance relation and intrinsic geometry of visual space: Implications for processing. Vision Res 1972, 12, 323–332. [Google Scholar]
- Du Priest, R.W., Jr; Fletcher, W.S. Chemotherapy of testicular germinal tumors. Oncology 1973, 28, 147–163. [Google Scholar]
- Lu, W.J.; Wang, H.M.; Yuann, J.M.; Huang, C.Y.; Hou, M.H. The impact of spermine competition on the efficacy of DNA-binding Fe(II), Co(II), and Cu(II) complexes of dimeric chromomycin A(3). J. Inorg. Biochem 2009, 103, 1626–1633. [Google Scholar]
- Altaner, C.; Ban, J.; Zajac, V.; Rossler, H.; Rosenthal, S.; Kettmann, R.; Burny, A. Isolation and characterization of cell clones producing various amounts of bovine leukosis virus. Folia Biol. (Praha) 1985, 31, 107–114. [Google Scholar]
- Chakrabarti, S.; Aich, P.; Sarker, D.; Bhattacharyya, D.; Dasgupta, D. Role of Mg2+ in the interaction of anticancer antibiotic, chromomycin A3 with DNA: Does neutral antibiotic bind DNA in absence of the metal ion? J. Biomol. Struct. Dyn 2000, 18, 209–218. [Google Scholar]
- Aich, P.; Sen, R.; Dasgupta, D. Role of magnesium ion in the interaction between chromomycin A3 and DNA: Binding of chromomycin A3-Mg2+ complexes with DNA. Biochemistry 1992, 31, 2988–2997. [Google Scholar]
- Devi, P.G.; Pal, S.; Banerjee, R.; Dasgupta, D. Association of antitumor antibiotics, mithramycin and chromomycin, with Zn(II). J. Inorg. Biochem 2007, 101, 127–137. [Google Scholar]
- Hou, M.H.; Lu, W.J.; Lin, H.Y.; Yuann, J.M. Studies of sequence-specific DNA binding, DNA cleavage, and topoisomerase I inhibition by the dimeric chromomycin A3 complexed with Fe(II). Biochemistry 2008, 47, 5493–5502. [Google Scholar]
- Slavik, M.; Carter, S.K. Chromomycin A3, mithramycin, and olivomycin: Antitumor antibiotics of related structure. Adv. Pharmacol. Chemother 1975, 12, 1–30. [Google Scholar]
- Goldberg, I.H.; Friedman, P.A. Antibiotics and nucleic acids. Annu. Rev. Biochem 1971, 40, 775–810. [Google Scholar]
- Yang, X.L.; Wang, A.H. Structural studies of atom-specific anticancer drugs acting on DNA. Pharmacol. Ther 1999, 83, 181–215. [Google Scholar]
- Kennedy, B.J. Mithramycin therapy in testicular cancer. J. Urol 1972, 107, 429–432. [Google Scholar]
- Elias, E.G.; Evans, J.T. Mithramycin in the treatment of Paget’s disease of bone. J. Bone Joint Surg. Am 1972, 54, 1730–1736. [Google Scholar]
- Fibach, E.; Bianchi, N.; Borgatti, M.; Prus, E.; Gambari, R. Mithramycin induces fetal hemoglobin production in normal and thalassemic human erythroid precursor cells. Blood 2003, 102, 1276–1281. [Google Scholar]
- Hou, M.H.; Wang, A.H. Mithramycin forms a stable dimeric complex by chelating with Fe(II): DNA-interacting characteristics, cellular permeation and cytotoxicity. Nucleic Acids Res 2005, 33, 1352–1361. [Google Scholar]
- Sobell, H.M.; Jain, S.C. Stereochemistry of actinomycin binding to DNA. II. Detailed molecular model of actinomycin-DNA complex and its implications. J. Mol. Biol 1972, 68, 21–34. [Google Scholar]
- Kamitori, S.; Takusagawa, F. Crystal structure of the 2:1 complex between d(GAAGCTTC) and the anticancer drug actinomycin D. J. Mol. Biol 1992, 225, 445–456. [Google Scholar]
- Hurwitz, J.; Furth, J.J.; Malamy, M.; Alexander, M. The role of deoxyribonucleic acid in ribonucleic acid synthesis. III. The inhibition of the enzymatic synthesis of ribonucleic acid and deoxyribonucleic acid by actinomycin D and proflavin. Proc. Natl. Acad. Sci. USA 1962, 48, 1222–1230. [Google Scholar]
- Aivasashvilli, V.A.; Beabealashvilli, R.S. Sequence-specific inhibition of RNA elongation by actinomycin D. FEBS Lett 1983, 160, 124–128. [Google Scholar]
- Chen, F.M. Kinetic and equilibrium binding studies of actinomycin D with some d(TGCA)-containing dodecamers. Biochemistry 1988, 27, 1843–1848. [Google Scholar]
- Margolis, R.L.; O’Hearn, E.; Rosenblatt, A.; Willour, V.; Holmes, S.E.; Franz, M.L.; Callahan, C.; Hwang, H.S.; Troncoso, J.C.; Ross, C.A. A disorder similar to Huntington’s disease is associated with a novel CAG repeat expansion. Ann. Neurol 2001, 50, 373–380. [Google Scholar]
- Seznec, H.; Agbulut, O.; Sergeant, N.; Savouret, C.; Ghestem, A.; Tabti, N.; Willer, J.C.; Ourth, L.; Duros, C.; Brisson, E.; et al. Mice transgenic for the human myotonic dystrophy region with expanded CTG repeats display muscular and brain abnormalities. Hum. Mol. Genet 2001, 10, 2717–2726. [Google Scholar]
- Paulson, H.L.; Fischbeck, K.H. Trinucleotide repeats in neurogenetic disorders. Annu. Rev. Neurosci 1996, 19, 79–107. [Google Scholar]
- Hou, M.H.; Robinson, H.; Gao, Y.G.; Wang, A.H. Crystal structure of actinomycin D bound to the CTG triplet repeat sequences linked to neurological diseases. Nucleic Acids Res 2002, 30, 4910–4917. [Google Scholar]
- Hou, M.H.; Lin, S.B.; Yuann, J.M.; Lin, W.C.; Wang, A.H.; Kan Ls, L. Effects of polyamines on the thermal stability and formation kinetics of DNA duplexes with abnormal structure. Nucleic Acids Res 2001, 29, 5121–5128. [Google Scholar]
- N’Soukpoe-Kossi, C.N.; Ouameur, A.A.; Thomas, T.; Shirahata, A.; Thomas, T.J.; Tajmir-Riahi, H.A. DNA interaction with antitumor polyamine analogues: A comparison with biogenic polyamines. Biomacromolecules 2008, 9, 2712–2718. [Google Scholar]
- Ouameur, A.A.; Bourassa, P.; Tajmir-Riahi, H.A. Probing tRNA interaction with biogenic polyamines. RNA 2010, 16, 1968–1979. [Google Scholar]
- N’Soukpoe-Kossi, C.N.; Ahmed Ouameur, A.; Thomas, T.; Thomas, T.J.; Tajmir-Riahi, H.A. Interaction of tRNA with antitumor polyamine analogues. Biochem. Cell Biol 2009, 87, 621–630. [Google Scholar]
- Marty, R.; N’Soukpoe-Kossi, C.N.; Charbonneau, D.M.; Kreplak, L.; Tajmir-Riahi, H.A. Structural characterization of cationic lipid-tRNA complexes. Nucleic Acids Res 2009, 37, 5197–5207. [Google Scholar]
- Igarashi, K.; Kashiwagi, K. Polyamines: Mysterious modulators of cellular functions. Biochem. Biophys. Res. Commun 2000, 271, 559–564. [Google Scholar]
- Patel, A.R.; Wang, J.Y. Polyamine depletion is associated with an increase in JunD/AP-1 activity in small intestinal crypt cells. Am. J. Physiol 1999, 276, G441–G450. [Google Scholar]
- Panagiotidis, C.A.; Artandi, S.; Calame, K.; Silverstein, S.J. Polyamines alter sequence-specific DNA-protein interactions. Nucleic Acids Res 1995, 23, 1800–1809. [Google Scholar]
- Frugier, M.; Florentz, C.; Hosseini, M.W.; Lehn, J.M.; Giege, R. Synthetic polyamines stimulate in vitro transcription by T7 RNA polymerase. Nucleic Acids Res 1994, 22, 2784–2790. [Google Scholar]
- Wang, Y.; Devereux, W.; Stewart, T.M.; Casero, R.A., Jr. Characterization of the interaction between the transcription factors human polyamine modulated factor (PMF-1) and NF-E2-related factor 2 (Nrf-2) in the transcriptional regulation of the spermidine/spermine N1-acetyltransferase (SSAT) gene. Biochem. J 2001, 355, 45–49. [Google Scholar]
- Westerhoff, H.V.; O’Dea, M.H.; Maxwell, A.; Gellert, M. DNA supercoiling by DNA gyrase. A static head analysis. Cell Biophys 1988, 12, 157–181. [Google Scholar]
- Pommier, Y.; Kerrigan, D.; Kohn, K. Topological complexes between DNA and topoisomerase II and effects of polyamines. Biochemistry 1989, 28, 995–1002. [Google Scholar]
- Gao, Y.G.; Robinson, H.; Wang, A.H. High-resolution A-DNA crystal structures of d(AGGGGCCCCT). An A-DNA model of poly(dG)·poly(dC). Eur. J. Biochem 1999, 261, 413–420. [Google Scholar]
- Ohishi, H.; Terasoma, N.; Nakanishi, I.; van der Marel, G.; van Boom, J.H.; Rich, A.; Wang, A.H.; Hakoshima, T.; Tomita, K. Interaction between left-handed Z-DNA and polyamine-3. The crystal structure of the d(CG)3 and thermospermine complex. FEBS Lett 1996, 398, 291–296. [Google Scholar]
- Ouameur, A.A.; Tajmir-Riahi, H.A. Structural analysis of DNA interactions with biogenic polyamines and cobalt(III)hexamine studied by Fourier transform infrared and capillary electrophoresis. J. Biol. Chem 2004, 279, 42041–42054. [Google Scholar]
- Marky, L.A.; Snyder, J.G.; Breslauer, K.J. Calorimetric and spectroscopic investigation of drug—DNA interactions: II. Dipyrandium binding to poly d(AT). Nucleic Acids Res 1983, 11, 5701–5715. [Google Scholar]
- Antony, T.; Thomas, T.; Shirahata, A.; Thomas, T.J. Selectivity of polyamines on the stability of RNA-DNA hybrids containing phosphodiester and phosphorothioate oligodeoxyribonucleotides. Biochemistry 1999, 38, 10775–10784. [Google Scholar]
- Gyi, J.I.; Conn, G.L.; Lane, A.N.; Brown, T. Comparison of the thermodynamic stabilities and solution conformations of DNA·RNA hybrids containing purine-rich and pyrimidine-rich strands with DNA and RNA duplexes. Biochemistry 1996, 35, 12538–12548. [Google Scholar]
- Arya, D.P.; Coffee, R.L., Jr; Willis, B.; Abramovitch, A.I. Aminoglycoside-nucleic acid interactions: Remarkable stabilization of DNA and RNA triple helices by neomycin. J. Am. Chem. Soc 2001, 123, 5385–5395. [Google Scholar]
- Arya, D.P.; Xue, L.; Willis, B. Aminoglycoside (neomycin) preference is for A-form nucleic acids, not just RNA: Results from a competition dialysis study. J. Am. Chem. Soc 2003, 125, 10148–10149. [Google Scholar]
- Hamilton, P.L.; Arya, D.P. Natural product DNA major groove binders. Nat. Prod. Rep 2012, 29, 134–143. [Google Scholar]
- Xi, H.; Davis, E.; Ranjan, N.; Xue, L.; Hyde-Volpe, D.; Arya, D.P. Thermodynamics of nucleic acid “shape readout” by an aminosugar. Biochemistry 2011, 50, 9088–9113. [Google Scholar]
- Xi, H.; Gray, D.; Kumar, S.; Arya, D.P. Molecular recognition of single-stranded RNA: Neomycin binding to poly(A). FEBS Lett 2009, 583, 2269–2275. [Google Scholar]
- Kumar, S.; Xue, L.; Arya, D.P. Neomycin-neomycin dimer: An all-carbohydrate scaffold with high affinity for AT-rich DNA duplexes. J. Am. Chem. Soc 2011, 133, 7361–7375. [Google Scholar]
- Shaw, N.N.; Xi, H.; Arya, D.P. Molecular recognition of a DNA:RNA hybrid: Sub-nanomolar binding by a neomycin-methidium conjugate. Bioorg. Med. Chem. Lett 2008, 18, 4142–4145. [Google Scholar]
- Xue, L.; Ranjan, N.; Arya, D.P. Synthesis and spectroscopic studies of the aminoglycoside (neomycin)-perylene conjugate binding to human telomeric DNA. Biochemistry 2011, 50, 2838–2849. [Google Scholar]
- Willis, B.; Arya, D.P. Recognition of B-DNA by neomycin-Hoechst 33258 conjugates. Biochemistry 2006, 45, 10217–10232. [Google Scholar]
- Arya, D.P.; Xue, L.; Tennant, P. Combining the best in triplex recognition: Synthesis and nucleic acid binding of a BQQ-neomycin conjugate. J. Am. Chem. Soc 2003, 125, 8070–8071. [Google Scholar]
- Xue, L.; Charles, I.; Arya, D.P. Pyrene-neomycin conjugate: Dual recognition of a DNA triple helix. Chem. Commun (Camb) 2002, 1, 70–71. [Google Scholar]
- Xi, H.; Kumar, S.; Dosen-Micovic, L.; Arya, D.P. Calorimetric and spectroscopic studies of aminoglycoside binding to AT-rich DNA triple helices. Biochimie 2010, 92, 514–529. [Google Scholar]
- Willis, B.; Arya, D.P. Triple recognition of B-DNA by a neomycin-Hoechst 33258-pyrene conjugate. Biochemistry 2010, 49, 452–469. [Google Scholar]
- Rosenberg, B.; Vancamp, L.; Krigas, T. Inhibition of cell division in escherichia coli by electrolysis products from a platinum electrode. Nature 1965, 205, 698–699. [Google Scholar]
- Pruefer, F.G.; Lizarraga, F.; Maldonado, V.; Melendez-Zajgla, J. Participation of Omi Htra2 serine-protease activity in the apoptosis induced by cisplatin on SW480 colon cancer cells. J. Chemother 2008, 20, 348–354. [Google Scholar]
- Vrana, O.; Brabec, V.; Kleinwachter, V. Polarographic studies on the conformation of some platinum complexes: Relations to anti-tumour activity. Anticancer Drug Des 1986, 1, 95–109. [Google Scholar]
- Brabec, V.; Kleinwachter, V.; Butour, J.L.; Johnson, N.P. Biophysical studies of the modification of DNA by antitumour platinum coordination complexes. Biophys. Chem 1990, 35, 129–141. [Google Scholar]
- Heringova, P.; Kasparkova, J.; Brabec, V. DNA adducts of antitumor cisplatin preclude telomeric sequences from forming G quadruplexes. J. Biol. Inorg. Chem 2009, 14, 959–968. [Google Scholar]
- Ming, L.J. Structure and function of “metalloantibiotics”. Med. Res. Rev 2003, 23, 697–762. [Google Scholar]
- Itzhaki, L.; Weinberger, S.; Livnah, N.; Berman, E. A unique binding cavity for divalent cations in the DNA-metal-chromomycin A3 complex. Biopolymers 1990, 29, 481–489. [Google Scholar]
- Gaboriau, F.; Kreder, A.; Clavreul, N.; Moulinoux, J.P.; Delcros, J.G.; Lescoat, G. Polyamine modulation of iron uptake in CHO cells. Biochem. Pharmacol 2004, 67, 1629–1637. [Google Scholar]
- Hou, M.H.; Lu, W.J.; Huang, C.Y.; Fan, R.J.; Yuann, J.M. Effects of polyamines on the DNA-reactive properties of dimeric mithramycin complexed with cobalt(II): Implications for anticancer therapy. Biochemistry 2009, 48, 4691–4698. [Google Scholar]

| Drugs | Sequence Specificity | Target Sequence |
|---|---|---|
| Anticancer drug | ||
| Chromomycin A3 | G/C-rich sequence | d(TTGGCCAA)2 |
| G/C-rich sequence | d(TTGGCCAA)2 | |
| Mithramycin | G/C-rich sequence | d(TTGGCCAA)2 |
| Blenomycin | G/C-rich sequence | d(CCCCAGCTGGGG) |
| Actinomycin D | CTG triplet repeat sequence | d[(CAG)4·(CTG)16] |
| telomeric G-quadruplex | d[GGG(TTAGGG)3] | |
| CC-1065 | AT-rich sequence | d(GAATT) |
| Doxorubicin | telomeric G-quadruplex | d[GGG(TTAGGG)3] |
| Sabarubicin | telomeric G-quadruplex | d[GGG(TTAGGG)3] |
| Streptonigrin | little sequence specificity | d(GCATGC)2 |
| cis-Platin | GC-rich sequence | poly(dG)·poly(dC) |
| SN-6999 | AT-rich sequence | poly(dA)·poly(dT) |
| SN-18071 | AT-rich sequence | poly(dA)·poly(dT) |
| Rebeccam ycin | sequences containing GpT (ApC) and TpG (CpA) steps | d(TGTTACGTT)2 |
| Antiviral drug | ||
| Lamivudine | little sequence specificity | Calf thymus DNA |
| Valacyclovir | little sequence specificity | Calf thymus DNA |
| isatin-β-thiosemicarbazone | little sequence specificity | Calf thymus DNA |
| Tartrazine | little sequence specificity | Calf thymus DNA |
| bis-netropsin | AT-rich sequence | poly(dA)·poly(dT) |
| Distamycin | AT-rich sequence | poly(dA)·poly(dT) |
| SN-7167 | AT-rich sequence | d(CGCGAATTCGCG) 2 |
| Berenil | AT-rich sequence | d(G4A4G4-[T4]*-C4T4C4-[T4]*-G4T4G4) |
| Antimicrobial drug | ||
| Neomycin | A-form nucleic acids | 16S A-site rRNA |
| Neomycin dimer | AT-rich sequence | d[5′-A12-x-T12-3′] |
| NHP | AT-rich sequence | poly(dA)·poly(dT) |
| NM | DNA:RNA hybrids | poly(dA)·poly(rU) |
| perylene-neomycin conjugate | G-quadruplex DNA | 5′-d[AG3(T2AG3)3] |
| Mitomycin C | GC-rich sequence | poly(dG) · poly(dC) |
| Phenanthrolin | little sequence specificity | Calf thymus DNA |
| Nogalamycin | little sequence specificity | Calf thymus DNA |
| Enrofloxacin | little sequence specificity | Calf thymus DNA |
| Neocarzinostatin | sequence-specific bulged DNA | |
| Others | ||
| Polyamine | Major groove of GC-rich duplexes | poly(dG)·poly(dC) |
| Minor groove of AT-rich duplexes | poly(dA)·poly(dT) | |
| Pentamidine | AT-rich sequence | poly(dA)·poly(dT) |
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Chang, Y.-M.; Chen, C.K.-M.; Hou, M.-H. Conformational Changes in DNA upon Ligand Binding Monitored by Circular Dichroism. Int. J. Mol. Sci. 2012, 13, 3394-3413. https://doi.org/10.3390/ijms13033394
Chang Y-M, Chen CK-M, Hou M-H. Conformational Changes in DNA upon Ligand Binding Monitored by Circular Dichroism. International Journal of Molecular Sciences. 2012; 13(3):3394-3413. https://doi.org/10.3390/ijms13033394
Chicago/Turabian StyleChang, Yu-Ming, Cammy K.-M. Chen, and Ming-Hon Hou. 2012. "Conformational Changes in DNA upon Ligand Binding Monitored by Circular Dichroism" International Journal of Molecular Sciences 13, no. 3: 3394-3413. https://doi.org/10.3390/ijms13033394
APA StyleChang, Y.-M., Chen, C. K.-M., & Hou, M.-H. (2012). Conformational Changes in DNA upon Ligand Binding Monitored by Circular Dichroism. International Journal of Molecular Sciences, 13(3), 3394-3413. https://doi.org/10.3390/ijms13033394





