Activation Profile Analysis of CruCA4, an α-Carbonic Anhydrase Involved in Skeleton Formation of the Mediterranean Red Coral, Corallium rubrum
Abstract
:1. Introduction
2. Results and Discussion
2.1. CruCA4 Identification, Heterologous Expression, Purification, and Kinetic Analysis
2.2. CruCA4 Activation Profile
3. Materials and Methods
3.1. Chemistry
3.2. Gene Synthesis, Cloning, Expression, Purification
3.3. SDS-PAGE
3.4. CA Enzyme Activation Assay
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Supuran, C.T.; Capasso, C. An overview of the bacterial carbonic anhydrases. Metabolites 2017, 7, 56. [Google Scholar] [CrossRef] [PubMed]
- Supuran, C.T. Structure and function of carbonic anhydrases. Biochem. J. 2016, 473, 2023–2032. [Google Scholar] [CrossRef] [PubMed]
- Ozensoy Guler, O.; Capasso, C.; Supuran, C.T. A magnificent enzyme superfamily: Carbonic anhydrases, their purification and characterization. J. Enzyme Inhib. Med. Chem. 2016, 31, 689–694. [Google Scholar] [CrossRef] [PubMed]
- Supuran, C.T. Advances in structure-based drug discovery of carbonic anhydrase inhibitors. Expert Opin. Drug Discov. 2017, 12, 61–88. [Google Scholar] [CrossRef] [PubMed]
- Capasso, C.; Supuran, C.T. Inhibition of bacterial carbonic anhydrases as a novel approach to escape drug resistance. Curr. Top. Med. Chem. 2017, 17, 1237–1248. [Google Scholar] [CrossRef] [PubMed]
- Supuran, C.T. How many carbonic anhydrase inhibition mechanisms exist? J. Enzyme Inhib. Med. Chem. 2016, 31, 345–360. [Google Scholar] [CrossRef] [PubMed]
- Capasso, C.; Supuran, C.T. An overview of the carbonic anhydrases from two pathogens of the oral cavity: Streptococcus mutans and porphyromonas gingivalis. Curr. Top. Med. Chem. 2016, 16, 2359–2368. [Google Scholar] [CrossRef] [PubMed]
- Capasso, C.; Supuran, C.T. An overview of the alpha-, beta- and gamma-carbonic anhydrases from bacteria: Can bacterial carbonic anhydrases shed new light on evolution of bacteria? J. Enzyme Inhib. Med. Chem. 2015, 30, 325–332. [Google Scholar] [CrossRef] [PubMed]
- Capasso, C.; Supuran, C.T. An overview of the selectivity and efficiency of the bacterial carbonic anhydrase inhibitors. Curr. Med. Chem. 2015, 22, 2130–2139. [Google Scholar] [CrossRef] [PubMed]
- Perfetto, R.; Del Prete, S.; Vullo, D.; Sansone, G.; Barone, C.M.A.; Rossi, M.; Supuran, C.T.; Capasso, C. Production and covalent immobilisation of the recombinant bacterial carbonic anhydrase (SSPCA) onto magnetic nanoparticles. J. Enzyme Inhib. Med. Chem. 2017, 32, 759–766. [Google Scholar] [CrossRef] [PubMed]
- Supuran, C.T.; Capasso, C. New light on bacterial carbonic anhydrases phylogeny based on the analysis of signal peptide sequences. J. Enzyme Inhib. Med. Chem. 2016, 31, 1254–1260. [Google Scholar] [CrossRef] [PubMed]
- Del Prete, S.; De Luca, V.; Supuran, C.T.; Capasso, C. Protonography, a technique applicable for the analysis of eta-carbonic anhydrase activity. J. Enzyme Inhib. Med. Chem. 2015, 30, 920–924. [Google Scholar] [CrossRef] [PubMed]
- Del Prete, S.; De Luca, V.; Vullo, D.; Scozzafava, A.; Carginale, V.; Supuran, C.T.; Capasso, C. Biochemical characterization of the gamma-carbonic anhydrase from the oral pathogen porphyromonas gingivalis, pgica. J. Enzyme Inhib. Med. Chem. 2014, 29, 532–537. [Google Scholar] [CrossRef] [PubMed]
- Del Prete, S.; De Luca, V.; Scozzafava, A.; Carginale, V.; Supuran, C.T.; Capasso, C. Biochemical properties of a new alpha-carbonic anhydrase from the human pathogenic bacterium, vibrio cholerae. J. Enzyme Inhib. Med. Chem. 2014, 29, 23–27. [Google Scholar] [CrossRef] [PubMed]
- Clare, B.W.; Supuran, C.T. Carbonic anhydrase activators. 3: Structure-activity correlations for a series of isozyme II activators. J. Pharm. Sci. 1994, 83, 768–773. [Google Scholar] [CrossRef] [PubMed]
- Supuran, C.T.; Balaban, A.T.; Cabildo, P.; Claramunt, R.M.; Lavandera, J.L.; Elguero, J. Carbonic anhydrase activators. VII. Isozyme II activation by bisazolyl-methanes, -ethanes and related azoles. Biol. Pharm. Bull. 1993, 16, 1236–1239. [Google Scholar] [CrossRef] [PubMed]
- Vullo, D.; Del Prete, S.; Osman, S.M.; Alasmary, F.A.S.; AlOthman, Z.; Donald, W.A.; Capasso, C.; Supuran, C.T. Comparison of the amine/amino acid activation profiles of the beta- and gamma-carbonic anhydrases from the pathogenic bacterium burkholderia pseudomallei. J. Enzyme Inhib. Med. Chem. 2018, 33, 25–30. [Google Scholar] [CrossRef] [PubMed]
- Canto de Souza, L.; Provensi, G.; Vullo, D.; Carta, F.; Scozzafava, A.; Costa, A.; Schmidt, S.D.; Passani, M.B.; Supuran, C.T.; Blandina, P. Carbonic anhydrase activation enhances object recognition memory in mice through phosphorylation of the extracellular signal-regulated kinase in the cortex and the hippocampus. Neuropharmacology 2017, 118, 148–156. [Google Scholar] [CrossRef] [PubMed]
- Vullo, D.; De Luca, V.; Scozzafava, A.; Carginale, V.; Rossi, M.; Supuran, C.T.; Capasso, C. The first activation study of a bacterial carbonic anhydrase (CA). The thermostable alpha-CA from Sulfurihydrogenibium yellowstonense YO3AOP1 is highly activated by amino acids and amines. Bioorg. Med. Chem. Lett. 2012, 22, 6324–6327. [Google Scholar] [CrossRef] [PubMed]
- Supuran, C.T. Carbonic anhydrase inhibition/activation: Trip of a scientist around the world in the search of novel chemotypes and drug targets. Curr. Pharm. Des. 2010, 16, 3233–3245. [Google Scholar] [CrossRef] [PubMed]
- Supuran, C.T. Carbonic anhydrase inhibitors and activators for novel therapeutic applications. Future Med. Chem. 2011, 3, 1165–1180. [Google Scholar] [CrossRef] [PubMed]
- Supuran, C.T.; Vullo, D.; Manole, G.; Casini, A.; Scozzafava, A. Designing of novel carbonic anhydrase inhibitors and activators. Curr. Med. Chem. Cardiovasc. Hematol. Agents 2004, 2, 51–70. [Google Scholar] [CrossRef] [PubMed]
- Keller, H. Carbonic anhydrase, its inhibitors and activators. Z. Vitam. Horm. Fermentforsch. 1957, 9, 297–320. [Google Scholar] [PubMed]
- Supuran, C.T.; Claramunt, R.M.; Lavandera, J.L.; Elguero, J. Carbonic anhydrase activators. XV. A kinetic study of the interaction of bovine isozyme II with pyrazoles, bis- and tris-azolyl-methanes. Biol. Pharm. Bull. 1996, 19, 1417–1422. [Google Scholar] [CrossRef] [PubMed]
- Briganti, F.; Scozzafava, A.; Supuran, C.T. Carbonic anhydrase activators. Part 19 spectroscopic and kinetic investigations for the interaction of isozymes I and II with primary amines. Met. Based Drugs 1997, 4, 221–227. [Google Scholar] [CrossRef] [PubMed]
- Briganti, F.; Mangani, S.; Orioli, P.; Scozzafava, A.; Vernaglione, G.; Supuran, C.T. Carbonic anhydrase activators: X-ray crystallographic and spectroscopic investigations for the interaction of isozymes I and II with histamine. Biochemistry 1997, 36, 10384–10392. [Google Scholar] [CrossRef] [PubMed]
- Supuran, C.T.; Scozzafava, A. Carbonic anhydrase activators: Amino acyl/dipeptidyl histamine derivatives bind with high affinity to isozymes I, II and IV and act as efficient activators. Bioorg. Med. Chem. 1999, 7, 2915–2923. [Google Scholar] [CrossRef]
- Briganti, F.; Scozzafava, A.; Supuran, C.T. Novel carbonic anhydrase isozymes I, II and IV activators incorporating sulfonyl-histamino moieties. Bioorg. Med. Chem. Lett. 1999, 9, 2043–2048. [Google Scholar] [CrossRef]
- Licsandru, E.; Tanc, M.; Kocsis, I.; Barboiu, M.; Supuran, C.T. A class of carbonic anhydrase I—Selective activators. J. Enzyme Inhib. Med. Chem. 2017, 32, 37–46. [Google Scholar] [CrossRef] [PubMed]
- Vullo, D.; Del Prete, S.; Capasso, C.; Supuran, C.T. Carbonic anhydrase activators: Activation of the beta-carbonic anhydrase from Malassezia globosa with amines and amino acids. Bioorg. Med. Chem. Lett. 2016, 26, 1381–1385. [Google Scholar] [CrossRef] [PubMed]
- Supuran, C.T. Carbonic anhydrases: From biomedical applications of the inhibitors and activators to biotechnological use for CO2 capture. J. Enzyme Inhib. Med. Chem. 2013, 28, 229–230. [Google Scholar] [CrossRef] [PubMed]
- Akocak, S.; Lolak, N.; Vullo, D.; Durgun, M.; Supuran, C.T. Synthesis and biological evaluation of histamine schiff bases as carbonic anhydrase I, II, IV, VII, and IX activators. J. Enzyme Inhib. Med. Chem. 2017, 32, 1305–1312. [Google Scholar] [CrossRef] [PubMed]
- Scozzafava, A.; Supuran, C.T. Carbonic anhydrase activators: High affinity isozymes I, II, and IV activators, incorporating a beta-alanyl-histidine scaffold. J. Med. Chem. 2002, 45, 284–291. [Google Scholar] [CrossRef] [PubMed]
- Ilies, M.; Banciu, M.D.; Ilies, M.A.; Scozzafava, A.; Caproiu, M.T.; Supuran, C.T. Carbonic anhydrase activators: Design of high affinity isozymes I, II, and IV activators, incorporating tri-/tetrasubstituted-pyridinium-azole moieties. J. Med. Chem. 2002, 45, 504–510. [Google Scholar] [CrossRef] [PubMed]
- Green, D.W.; Ben-Nissan, B.; Yoon, K.S.; Milthorpe, B.; Jung, H.S. Natural and synthetic coral biomineralization for human bone revitalization. Trends Biotechnol. 2017, 35, 43–54. [Google Scholar] [CrossRef] [PubMed]
- Del Prete, S.; Vullo, D.; Zoccola, D.; Tambutte, S.; Capasso, C.; Supuran, C.T. Kinetic properties and affinities for sulfonamide inhibitors of an alpha-carbonic anhydrase (CRUCA4) involved in coral biomineralization in the mediterranean red coral Corallium rubrum. Bioorg. Med. Chem. 2017, 25, 3525–3530. [Google Scholar] [CrossRef] [PubMed]
- Le Goff, C.; Tambutte, E.; Venn, A.A.; Techer, N.; Allemand, D.; Tambutte, S. In vivo pH measurement at the site of calcification in an octocoral. Sci. Rep. 2017, 7, 11210. [Google Scholar] [CrossRef] [PubMed]
- Le Goff, C.; Ganot, P.; Zoccola, D.; Caminiti-Segonds, N.; Allemand, D.; Tambutté, S. Carbonic anhydrases in cnidarians: Novel perspectives from the octocorallian Corallium rubrum. PLoS ONE 2016, 11, e0160368. [Google Scholar] [CrossRef] [PubMed]
- Debreuil, J.; Tambutte, E.; Zoccola, D.; Deleury, E.; Guigonis, J.M.; Samson, M.; Allemand, D.; Tambutte, S. Molecular cloning and characterization of first organic matrix protein from sclerites of red coral, Corallium rubrum. J. Biol. Chem. 2012, 287, 19367–19376. [Google Scholar] [CrossRef] [PubMed]
- Wilkins, M.R.; Gasteiger, E.; Bairoch, A.; Sanchez, J.C.; Williams, K.L.; Appel, R.D.; Hochstrasser, D.F. Protein identification and analysis tools in the expasy server. Methods Mol. Biol. 1999, 112, 531–552. [Google Scholar] [PubMed]
- Perfetto, R.; Del Prete, S.; Vullo, D.; Sansone, G.; Barone, C.; Rossi, M.; Supuran, C.T.; Capasso, C. Biochemical characterization of the native alpha-carbonic anhydrase purified from the mantle of the Mediterranean mussel, Mytilus galloprovincialis. J. Enzyme Inhib. Med. Chem. 2017, 32, 632–639. [Google Scholar] [CrossRef] [PubMed]
- Perfetto, R.; Del Prete, S.; Vullo, D.; Carginale, V.; Sansone, G.; Barone, C.M.A.; Rossi, M.; Alasmary, F.A.S.; Osman, S.M.; AlOthman, Z.; et al. Cloning, expression and purification of the alpha-carbonic anhydrase from the mantle of the Mediterranean mussel, Mytilus galloprovincialis. J. Enzyme Inhib. Med. Chem. 2017, 32, 1029–1035. [Google Scholar] [CrossRef] [PubMed]
- Di Cesare Mannelli, L.; Micheli, L.; Carta, F.; Cozzi, A.; Ghelardini, C.; Supuran, C.T. Carbonic anhydrase inhibition for the management of Cerebral ischemia: In vivo evaluation of sulfonamide and coumarin inhibitors. J. Enzyme Inhib. Med. Chem. 2016, 31, 894–899. [Google Scholar] [CrossRef] [PubMed]
- Temperini, C.; Innocenti, A.; Scozzafava, A.; Supuran, C.T. Carbonic anhydrase activators: Kinetic and x-ray crystallographic study for the interaction of d- and l-tryptophan with the mammalian isoforms I-XIV. Bioorg. Med. Chem. 2008, 16, 8373–8378. [Google Scholar] [CrossRef] [PubMed]
- Temperini, C.; Scozzafava, A.; Vullo, D.; Supuran, C.T. Carbonic anhydrase activators. Activation of isozymes I, II, IV, VA, VII, and XIV with l- and d-histidine and crystallographic analysis of their adducts with isoform II: Engineering proton-transfer processes within the active site of an enzyme. Chemistry 2006, 12, 7057–7066. [Google Scholar] [CrossRef] [PubMed]
- Temperini, C.; Scozzafava, A.; Vullo, D.; Supuran, C.T. Carbonic anhydrase activators. Activation of isoforms I, II, IV, VA, VII, and XIV with l- and d-phenylalanine and crystallographic analysis of their adducts with isozyme II: Stereospecific recognition within the active site of an enzyme and its consequences for the drug design. J. Med. Chem. 2006, 49, 3019–3027. [Google Scholar] [PubMed]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef] [PubMed]
- Khalifah, R.G. The carbon dioxide hydration activity of carbonic anhydrase. I. Stop-flow kinetic studies on the native human isoenzymes B and C. J. Biol. Chem. 1971, 246, 2561–2573. [Google Scholar] [PubMed]
- Alber, B.E.; Ferry, J.G. A carbonic anhydrase from the archaeon Methanosarcina thermophila. Proc. Natl. Acad. Sci. USA 1994, 91, 6909–6913. [Google Scholar] [CrossRef] [PubMed]
- Alber, B.E.; Ferry, J.G. Characterization of heterologously produced carbonic anhydrase from Methanosarcina thermophila. J. Bacteriol. 1996, 178, 3270–3274. [Google Scholar] [CrossRef] [PubMed]
- Kisker, C.; Schindelin, H.; Alber, B.E.; Ferry, J.G.; Rees, D.C. A left-hand beta-helix revealed by the crystal structure of a carbonic anhydrase from the archaeon Methanosarcina thermophila. EMBO J. 1996, 15, 2323–2330. [Google Scholar] [PubMed]
- Wang, X.; Schröder, H.C.; Schlossmacher, U.; Neufurth, M.; Feng, Q.; Diehl-Seifert, B.; Müller, W.E. Modulation of the initial mineralization process of SaOS-2 cells by carbonic anhydrase activators and polyphosphate. Calcif. Tissue Int. 2014, 94, 495–509. [Google Scholar] [CrossRef] [PubMed]
- Müller, W.E.; Schröder, H.C.; Schlossmacher, U.; Grebenjuk, V.A.; Ushijima, H.; Wang, X. Induction of carbonic anhydrase in SaOS-2 cells, exposed to bicarbonate and consequences for calcium phosphate crystal formation. Biomaterials 2013, 34, 8671–8680. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of all the compounds are available from the authors. |
| Acronym | GenBank Accession Number | Amino Acid Number | M.W. (kDa) | Calcifying Fraction 1 | Non-Calcifying Fraction 1 |
|---|---|---|---|---|---|
| CruCA1 | KU557743.1 | 356 | 40.89 | − | + |
| CruCA2 | KU557744.1 | 322 | 36.90 | − | + |
| CruCA3 | KU557745.1 | 262 | 29.04 | − | + |
| CruCA4 | KU557746.1 | 284 | 32.86 | + | − |
| CruCA5 | KU557747.1 | 335 | 36.06 | − | + |
| CruCA5 | KU557748.1 | 281 | 32.06 | − | + |
| Isoenzyme | kcat * (s−1) | KM * (mM) | (kcat)l-Tyr ** (s−1) | KA *** l-Tyr (μM) |
|---|---|---|---|---|
| hCA I a | 2.0 × 105 | 4.0 | 13.9 × 105 | 0.020 |
| hCA II a | 1.4 × 106 | 9.3 | 12.8 × 106 | 0.011 |
| CruCA4 b | 2.4 × 105 | 4.6 | 18.7 × 105 | 0.73 |
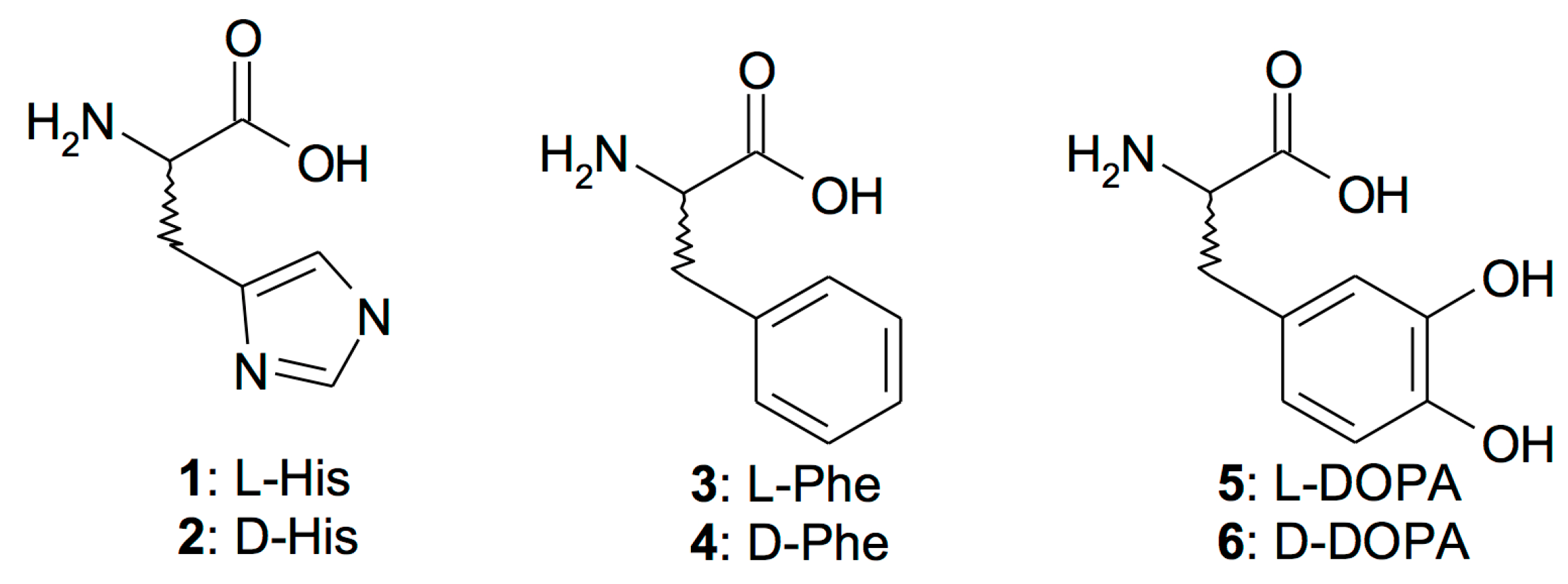
| No. | Compound | KA (μM) * | ||
|---|---|---|---|---|
| hCA I a | hCA II a | CruCA4 b | ||
| 1 | l-His | 0.03 | 10.9 | 36.9 |
| 2 | d-His | 0.09 | 43 | 0.098 |
| 3 | l-Phe | 0.07 | 0.013 | 15.4 |
| 4 | d-Phe | 86 | 0.035 | 1.01 |
| 5 | l-DOPA | 3.1 | 11.4 | 13.7 |
| 6 | d-DOPA | 4.9 | 7.8 | 0.93 |
| 7 | l-Trp | 44 | 27 | 9.48 |
| 8 | d-Trp | 41 | 12 | 8.35 |
| 9 | l-Tyr | 0.02 | 0.011 | 0.73 |
| 10 | d-Tyr | 0.04 | 0.013 | 18.9 |
| 11 | 4-H2N-l-Phe | 0.24 | 0.15 | 0.074 |
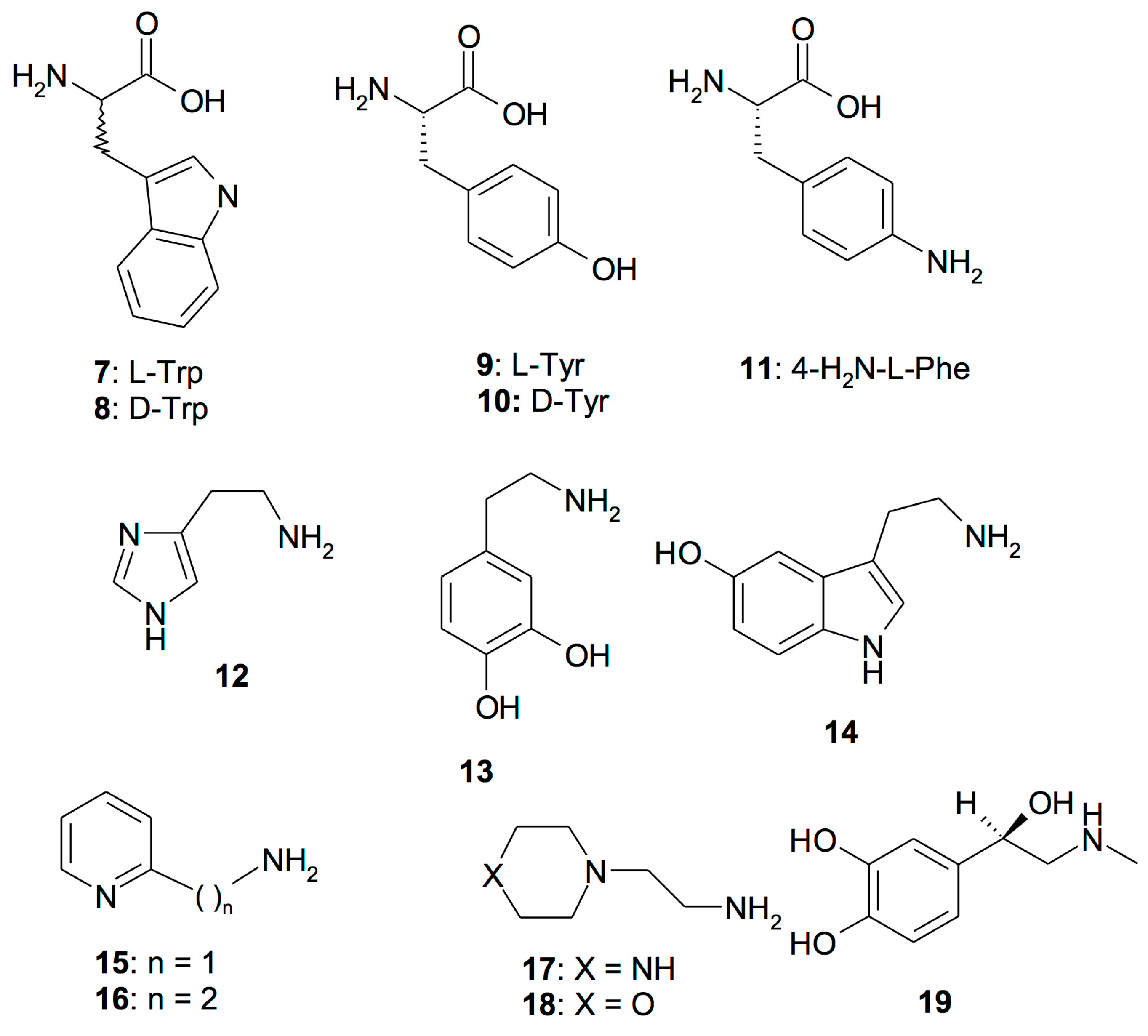
| 12 | Histamine | 2.1 | 125 | 0.007 |
| 13 | Dopamine | 13.5 | 9.2 | 0.005 |
| 14 | Serotonin | 45 | 50 | 0.006 |
| 15 | 2-Pyridyl-methylamine | 26 | 34 | 0.41 |
| 16 | 2-(2-Aminoethyl)pyridine | 13 | 15 | 0.26 |
| 17 | 1-(2-Aminoethyl)-piperazine | 7.4 | 2.3 | 0.004 |
| 18 | 4-(2-Aminoethyl)-morpholine | 0.14 | 0.19 | 0.15 |
| 19 | l-Adrenaline | 0.09 | 96 | 0.009 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Del Prete, S.; Vullo, D.; Zoccola, D.; Tambutté, S.; Supuran, C.T.; Capasso, C. Activation Profile Analysis of CruCA4, an α-Carbonic Anhydrase Involved in Skeleton Formation of the Mediterranean Red Coral, Corallium rubrum. Molecules 2018, 23, 66. https://doi.org/10.3390/molecules23010066
Del Prete S, Vullo D, Zoccola D, Tambutté S, Supuran CT, Capasso C. Activation Profile Analysis of CruCA4, an α-Carbonic Anhydrase Involved in Skeleton Formation of the Mediterranean Red Coral, Corallium rubrum. Molecules. 2018; 23(1):66. https://doi.org/10.3390/molecules23010066
Chicago/Turabian StyleDel Prete, Sonia, Daniela Vullo, Didier Zoccola, Sylvie Tambutté, Claudiu T. Supuran, and Clemente Capasso. 2018. "Activation Profile Analysis of CruCA4, an α-Carbonic Anhydrase Involved in Skeleton Formation of the Mediterranean Red Coral, Corallium rubrum" Molecules 23, no. 1: 66. https://doi.org/10.3390/molecules23010066






