Structure Elucidation of Procyanidins Isolated from Rhododendron formosanum and Their Anti-Oxidative and Anti-Bacterial Activities
Abstract
:1. Introduction
2. Results and Discussion
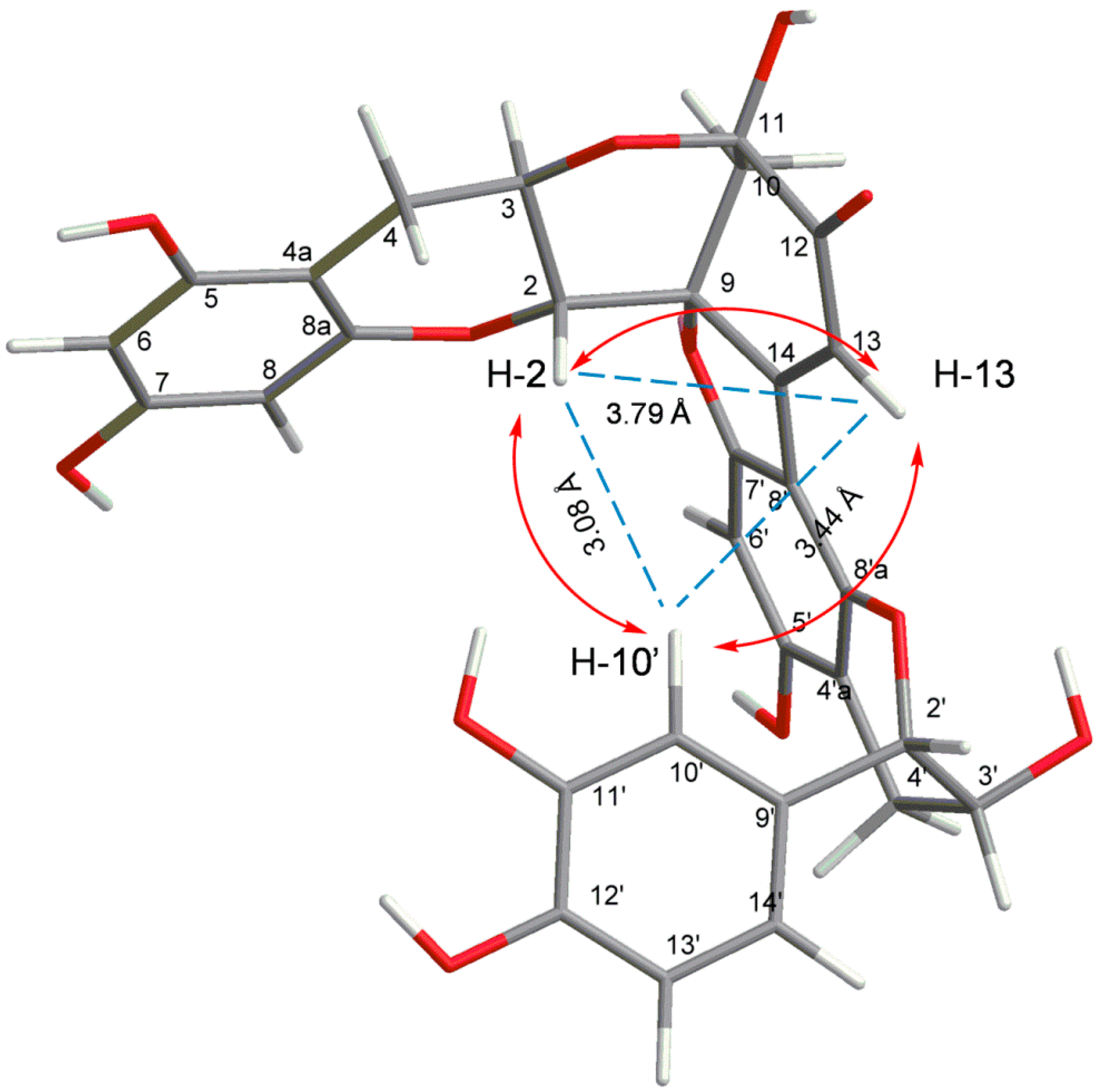
2.1. Identification of Isolated Procyanidins
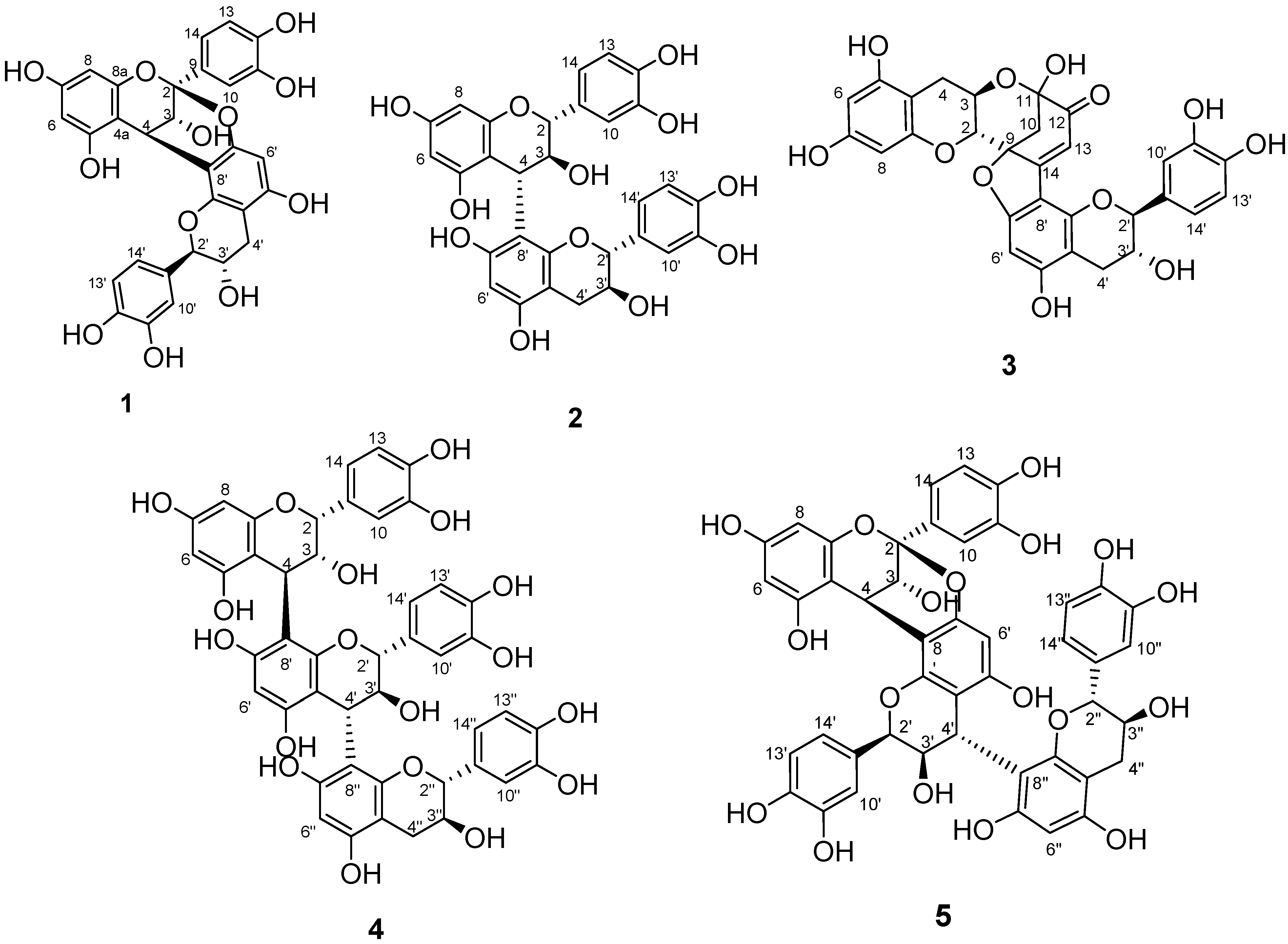

| Units | Position | 1H | 13C |
|---|---|---|---|
| I | 2 | 3.97 d (8.4) | 79.5 |
| 3 | 3.98 m | 66.8 | |
| 4β | 2.94 dd (5.4, 14.4) | 27.8 | |
| 4α | 2.52 dd (9.0, 14.4) | ||
| 4a | 100.4 | ||
| 5 | 157.6 | ||
| 6 | 5.90 (3.2) | 97.0 | |
| 7 | 157.9 | ||
| 8 | 5.54 (3.3) | 95.8 | |
| 8a | 156.3 | ||
| 9 | 89.8 | ||
| 10 | 2.68 d (11.4) | 45.9 | |
| 2.49 d (11.4) | |||
| 11 | 95.3 | ||
| 12 | 194.1 | ||
| 13 | 6.43 s | 112.8 | |
| 14 | 164.4 | ||
| II | 2′ | 4.92 d (7.2) | 83.4 |
| 3′ | 4.11 m | 67.8 | |
| 4′β | 2.86 dd (4.8, 16.2) | 28.3 | |
| 4′α | 2.60 dd (7.8, 16.2) | ||
| 4a′ | 103.9 | ||
| 5′ | 166.2 | ||
| 6′ | 6.13 s | 90.9 | |
| 7′ | 168.0 | ||
| 8′ | 105.6 | ||
| 8a′ | 155.1 | ||
| 9′ | 131.2 | ||
| 10′ | 6.85 d (2.4) | 114.8 | |
| 11′ | 146.4 | ||
| 12′ | 146.5 | ||
| 13′ | 6.79 d (8.4) | 116.3 | |
| 14′ | 6.74 dd (8.4, 2.4) | 119.7 |
2.2. Antibacterial Activity
| Pathogens | Minimum Inhibitory Concentration (μg/mL) | |||||||
|---|---|---|---|---|---|---|---|---|
| Antibiotics and Procyanidins | ||||||||
| Ap * | Tet | Met | 1 | 2 | 3 | 4 | 5 | |
| Staphylococcus aureus | 16 | 8 | +N.D. | 64 | 64 | 4 | >128 | >128 |
| Enterococcus faecalis | 2 | 4 | N.D. | >128 | >128 | >128 | >128 | >128 |
| Listeria monocytogenes | 1 | 2 | N.D. | 64 | >128 | >128 | >128 | >128 |
| Bacillus cereus | 128 | 4 | N.D. | 64 | >128 | >128 | >128 | >128 |
| Escherichia coli | 4 | 0.5 | N.D. | >128 | >128 | >128 | >128 | >128 |
| Salmonella enterica | 1 | 8 | N.D. | >128 | >128 | >128 | >128 | >128 |
| Pseudomonas aeruginosa | 512 | 32 | N.D. | >128 | >128 | >128 | >128 | >128 |
| Helicobacter pylori ** | N.D. | N.D. | 2 | >256 | >256 | >256 | >256 | >256 |
2.3. Antioxidative Activity
| Compounds | Total OH Groups | Average OH/unit | Antioxidant Activity | |
|---|---|---|---|---|
| IC50/DPPH (μg/mL) | CUPRAC (TEACs) | |||
| Trolox | 2 | 2 | 61.12 | 1.00 |
| (−)-Catechin | 5 | 5 | 27.07 | 2.74 |
| 1 | 9 | 4.5 | 20.89 | 1.75 |
| 2 | 10 | 5 | 8.55 | 4.87 |
| 3 | 7 | 3.5 | 13.06 | 1.96 |
| 4 | 15 | 5 | 6.26 | 3.48 |
| 5 | 14 | 4.7 | 3.29 | 2.93 |

3. Experimental Section
3.1. General Information
3.2. Plant Material
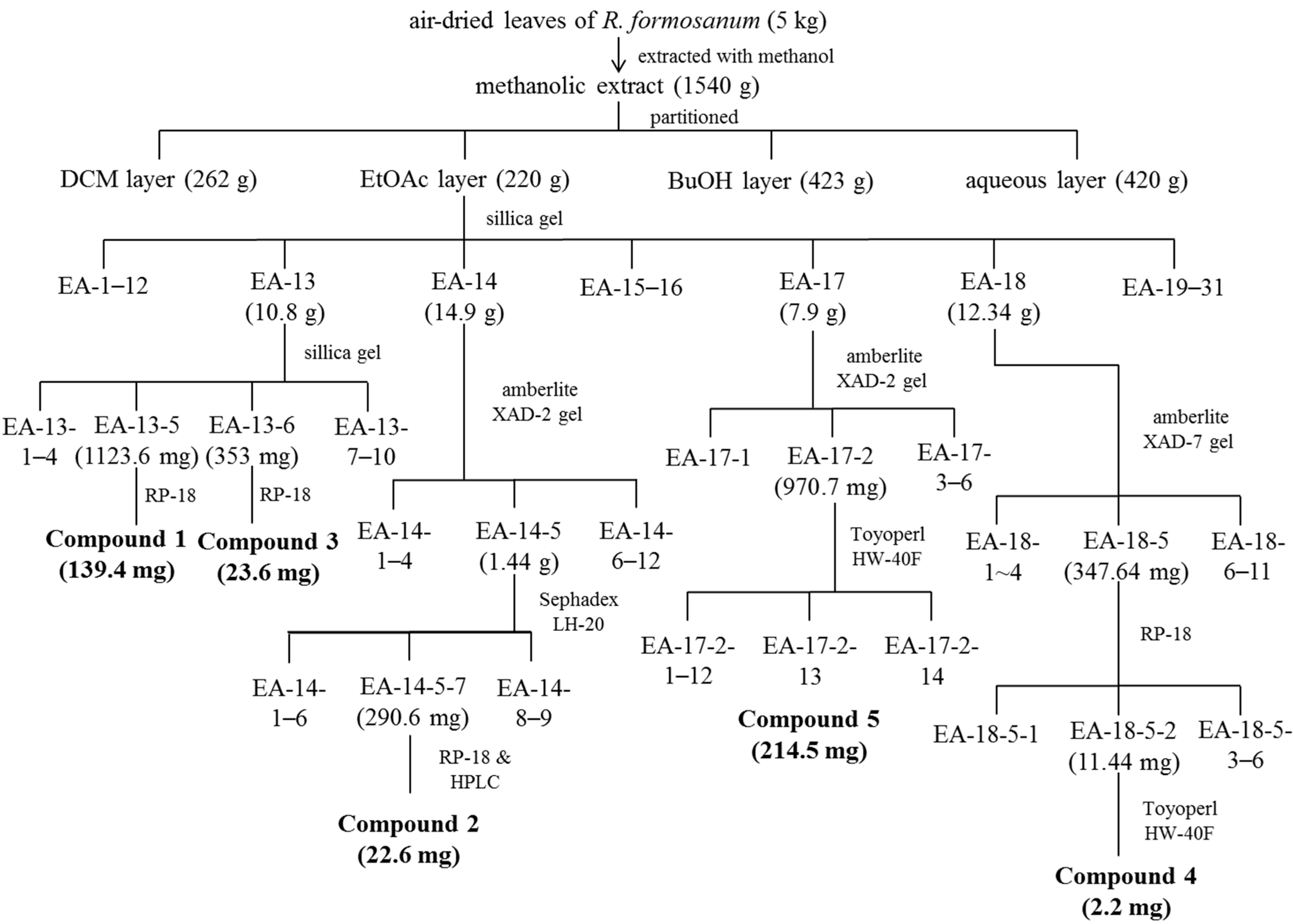
3.3. Isolation and Identification of Procyanidins
3.4. Total Antioxidant Capacity (TAC)
3.5. Free Radical Scavenging Activity
3.6. Antibacterial Activity
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References and Notes
- Maatta-Riihinen, K.R.; Kahkonen, M.P.; Torronen, A.R.; Heinonen, I.M. Catechins and procyanidins in berries of Vaccinium species and their antioxidant activity. J. Agric. Food Chem. 2005, 53, 8485–8491. [Google Scholar] [CrossRef] [PubMed]
- Fu, C.L.; Wang, H.Y.; Ng, W.L.; Song, L.X.; Huang, D.J. Antioxidant activity and proanthocyanidin profile of Selliguea feei rhizomes. Molecules 2013, 18, 4282–4292. [Google Scholar] [CrossRef] [PubMed]
- He, F.; Pan, Q.H.; Shi, Y.; Duan, C.Q. Biosynthesis and genetic regulation of proanthocyanidins in plants. Molecules 2008, 13, 2674–2703. [Google Scholar] [CrossRef] [PubMed]
- Zang, X.Y.; Shang, M.Y.; Xu, F.; Liang, J.; Wang, X.; Mikage, M.; Cai, S.Q. A-type proanthocyanidins from the stems of Ephedra sinica (Ephedraceae) and their antimicrobial activities. Molecules 2013, 18, 5172–5189. [Google Scholar] [CrossRef] [PubMed]
- Kresty, L.A.; Howell, A.B.; Baird, M. Cranberry proanthocyanidins mediate growth arrest of lung cancer cells through modulation of gene expression and rapid induction of apoptosis. Molecules 2011, 16, 2375–2390. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.Y.; Song, L.X.; Feng, S.B.; Liu, Y.C.; Zuo, G.; Lai, F.L.; He, G.Y.; Chen, M.J.; Huang, D.J. Characterization of proanthocyanidins in stems of Polygonum multiflorum thunb as strong starch hydrolase inhibitors. Molecules 2013, 18, 2255–2265. [Google Scholar] [CrossRef] [PubMed]
- Chou, S.C.; Krishna, V.; Chou, C.H. Hydrophobic Metabolites from Rhododendron formosanum and their Allelopathic Activities. Nat. Prod. Commun. 2009, 4, 1189–1192. [Google Scholar] [PubMed]
- Krishna, V.; Chang, C.I.; Chou, C.H. Two isomeric epoxysitosterols from Rhododendron formosanum: 1H- and 13C-NMR chemical shift assignments. Magn. Reson. Chem. 2006, 44, 817–819. [Google Scholar] [CrossRef] [PubMed]
- Way, T.D.; Tsai, S.J.; Wang, C.M.; Ho, C.T.; Chou, C.H. Chemical constituents of Rhododendron formosanum show pronounced growth inhibitory effect on non-small-cell lung carcinoma cells. J. Agric. Food Chem. 2014, 62, 875–884. [Google Scholar] [CrossRef] [PubMed]
- Chou, S.C.; Huang, C.H.; Hsu, T.W.; Wu, C.C.; Chou, C.H. Allelopathic potential of Rhododendron formosanum Hemsl in Taiwan. Allelopathy J. 2010, 25, 73–91. [Google Scholar]
- Wang, C.M.; Li, T.C.; Jhan, Y.L.; Weng, J.H.; Chou, C.H. The impact of microbial biotransformation of catechin in enhancing the allelopathic effects of Rhododendron formosanum. PLoS ONE 2013, 8, e85162. [Google Scholar] [CrossRef] [PubMed]
- Botha, J.J.; Ferreira, D.; Roux, D.G. Condensed tannins-circular-dichroism method of assessing absolute-configuration at C-4 of 4-arylflavan-3-ols, and stereochemistry of their formation from flavan-3,4-diols. J. Chem. Soc. Chem. Comm. 1978, 698–700. [Google Scholar] [CrossRef]
- Barrett, M.W.; Klyne, W.; Scopes, P.M.; Fletcher, A.C.; Porter, L.J.; Haslam, E. Plant proanthocyanidins. Part 6. chiroptical studies. Part 95. Circular-Dichroism of procyanidins. J. Chem. Soc. Perkin Trans. 1 1979, 2375–2377. [Google Scholar] [CrossRef]
- Lou, H.X.; Yamazaki, Y.; Sasaki, T.; Uchida, M.; Tanaka, H.; Oka, S. A-type proanthocyanidins from peanut skins. Phytochemistry 1999, 51, 297–308. [Google Scholar] [CrossRef]
- Oizumi, Y.; Mohri, Y.; Hirota, M.; Makabe, H. Synthesis of procyanidin B3 and its anti-inflammatory activity. The effect of 4-alkoxy group of catechin electrophile in the Yb(OTf)(3)-catalyzed condensation with catechin nucleophile. J. Org. Chem. 2010, 75, 4884–4886. [Google Scholar] [CrossRef] [PubMed]
- Guyot, S.; Vercauteren, J.; Cheynier, V. Structural determination of colourless and yellow dimers resulting from (+)-catechin coupling catalysed by grape polyphenoloxidase. Phytochemistry 1996, 42, 1279–1288. [Google Scholar] [CrossRef]
- Yan, G.Q.; Li, S.P.; Hu, J.; Zhai, X.Y.; Ma, W.; Li, N.; Wang, K.J. Phenolic constituents from the roots of Rosa laevigata (Rosaceae). Biochem. Syst. Ecol. 2014, 52, 23–26. [Google Scholar] [CrossRef]
- Karioti, A.; Bilia, A.R.; Gabbiani, C.; Messori, L.; Skaltsa, H. Proanthocyanidin glycosides from the leaves of Quercus ilex L. (Fagaceae). Tetrahedron Lett. 2009, 50, 1771–1776. [Google Scholar] [CrossRef]
- Slade, D.; Ferreira, D.; Marais, J.P.J. Circular dichroism, a powerful tool for the assessment of absolute configuration of flavonoids. Phytochemistry 2005, 66, 2177–2215. [Google Scholar] [CrossRef] [PubMed]
- Saito, A.; Doi, Y.; Tanaka, A.; Matsuura, N.; Ubukata, M.; Nakajima, N. Systematic synthesis of four epicatechin series procyanidin trimers and their inhibitory activity on the Maillard reaction and antioxidant activity. Bioorg. Med. Chem. 2004, 12, 4783–4790. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.C.; Lee, S.S. Proanthocyanidins from the leaves of Machilus philippinensis. J. Nat. Prod. 2010, 73, 1375–1380. [Google Scholar] [CrossRef] [PubMed]
- Killday, K.B.; Davey, M.H.; Glinski, J.A.; Duan, P.G.; Veluri, R.; Proni, G.; Daugherty, F.J.; Tempesta, M.S. Bioactive A-type proanthocyanidins from Cinnamomum cassia. J. Nat. Prod. 2011, 74, 1833–1841. [Google Scholar] [CrossRef] [PubMed]
- Kamiya, K.; Ohno, A.; Horii, Y.; Endang, H.; Umar, M.; Satake, T. A-type proanthocyanidins from the bark of Parameria laevigata. Heterocycles 2003, 60, 1697–1708. [Google Scholar]
- De Bruyne, T.; Pieters, L.; Witvrouw, M.; de Clercq, E.; vanden Berghe, D.; Vlietinck, A.J. Biological evaluation of proanthocyanidin dimers and related polyphenols. J. Nat. Prod. 1999, 62, 954–958. [Google Scholar] [CrossRef] [PubMed]
- Ming, D.S.; Lopez, A.; Hillhouse, B.J.; French, C.J.; Hudson, J.B.; Towers, G.H.N. Bioactive constituents from Iryanthera megistophylla. J. Nat. Prod. 2002, 65, 1412–1416. [Google Scholar] [CrossRef] [PubMed]
- Ricardo-Da-Silva, J.M.; Darmon, N.; Fernandez, Y.; Mitjavila, S. Oxygen free-radical scavenger capacity in aqueous models of different procyanidins from grape seeds. J. Agric. Food Chem. 1991, 39, 1549–1552. [Google Scholar] [CrossRef]
- Wang, C.M.; Chen, H.T.; Li, T.C.; Weng, J.H.; Jhan, Y.L.; Lin, S.X.; Chou, C.H. The role of pentacyclic triterpenoids in the allelopathic effects of Alstonia scholaris. J. Chem. Ecol. 2014, 40, 90–98. [Google Scholar] [CrossRef] [PubMed]
- Ozyurek, M.; Guclu, K.; Apak, R. The main and modified CUPRAC methods of antioxidant measurement. TrAC Trend. Anal. Chem. 2011, 30, 652–664. [Google Scholar] [CrossRef]
- Prior, R.L.; Wu, X.L.; Schaich, K. Standardized methods for the determination of antioxidant capacity and phenolics in foods and dietary supplements. J. Agric. Food Chem. 2005, 53, 4290–4302. [Google Scholar] [CrossRef] [PubMed]
- CLSI. Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria that Grow Aerobically, 10th ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2015. [Google Scholar]
- Sample Availability: Samples of the compounds 1, 2, and 5 are available from the authors.
© 2015 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, C.-M.; Hsu, Y.-M.; Jhan, Y.-L.; Tsai, S.-J.; Lin, S.-X.; Su, C.-H.; Chou, C.-H. Structure Elucidation of Procyanidins Isolated from Rhododendron formosanum and Their Anti-Oxidative and Anti-Bacterial Activities. Molecules 2015, 20, 12787-12803. https://doi.org/10.3390/molecules200712787
Wang C-M, Hsu Y-M, Jhan Y-L, Tsai S-J, Lin S-X, Su C-H, Chou C-H. Structure Elucidation of Procyanidins Isolated from Rhododendron formosanum and Their Anti-Oxidative and Anti-Bacterial Activities. Molecules. 2015; 20(7):12787-12803. https://doi.org/10.3390/molecules200712787
Chicago/Turabian StyleWang, Chao-Min, Yuan-Man Hsu, Yun-Lian Jhan, Shang-Jie Tsai, Shi-Xun Lin, Chiu-Hsian Su, and Chang-Hung Chou. 2015. "Structure Elucidation of Procyanidins Isolated from Rhododendron formosanum and Their Anti-Oxidative and Anti-Bacterial Activities" Molecules 20, no. 7: 12787-12803. https://doi.org/10.3390/molecules200712787






