2.2. Best Models from Alignment 1
The best models
1B7 and
1B9 (1.0 Å grid cell) are described in
Table 1. Model
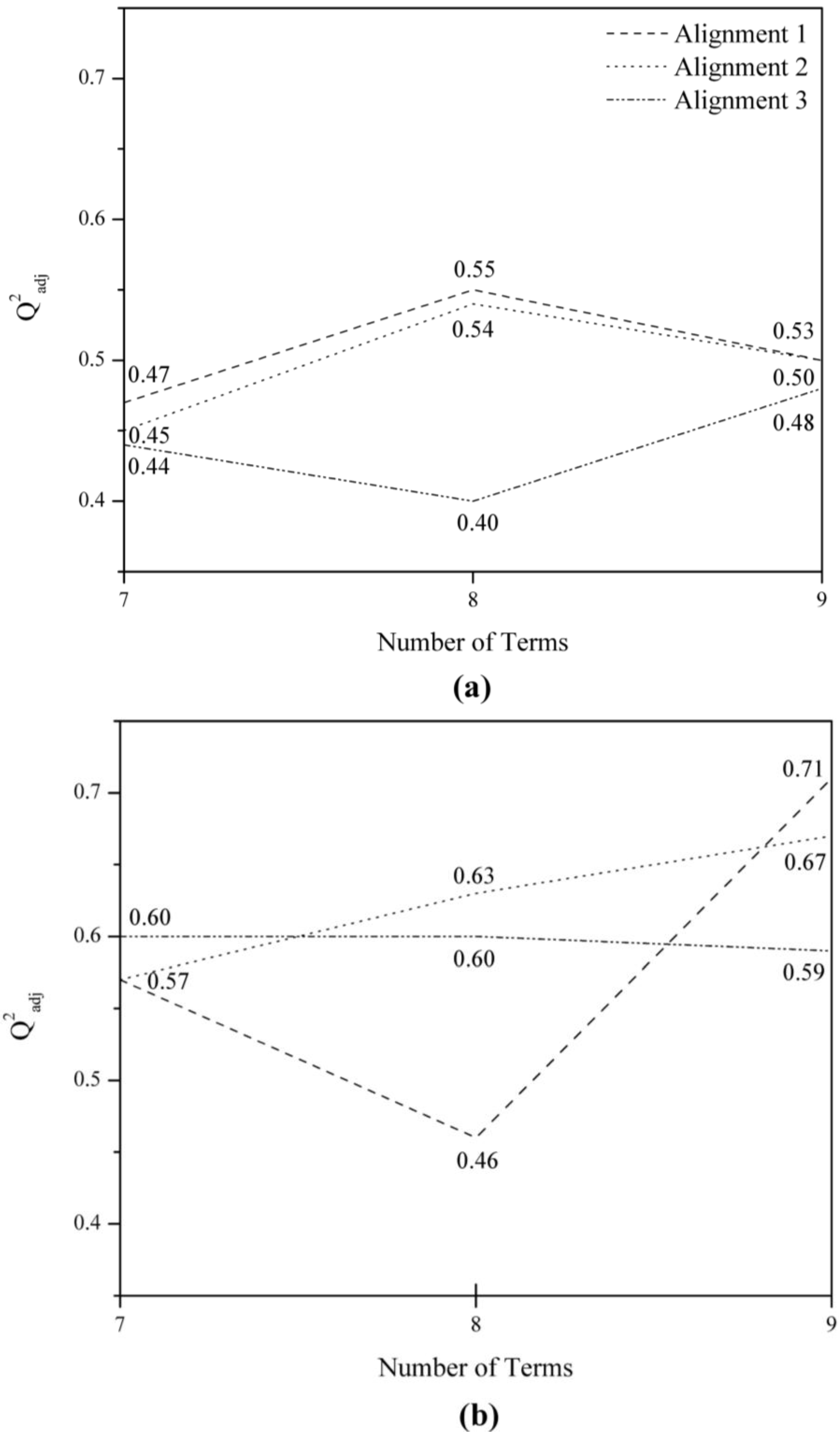
1B8 was eliminated from the analysis because it presented a low Q
2adj value (<0.5) (
Figure 3). In order to determine if the information in models
1B7 and
1B9 is redundant, the correlation coefficient (R) of their residuals was calculated (
i.e., pIC
50 experimental—pIC
50 calculated). Equivalent models typically have nearly the same distribution of residuals (
i.e., R ≈ 1) and independent models will have nearly uncorrelated residuals (
i.e., R ≈ 0) [
23]. The results show that models
1B7 and
1B9 have some degree of correlation (R = 0.59), probably due to the presence of spatially identical grid cells with the same IPE type (
Table 1), namely GCODs (
2,11,4)(
any),
(−1,8,2)(
any), and (
1,13,1)(
any).
Table 1.
Best models of Alignment 1 obtained from 1.0 Å grid cells.
Table 1.
Best models of Alignment 1 obtained from 1.0 Å grid cells.
| Model | Terms | Equation a |
|---|
| 1B7 | 7 | pIC50 = 7.89 − 13.99 (2,11,4)(any)− 6.96 (−1,8,2)(any) − 2.04 (0,−2,9)(any) |
| + 22.68 (1,13,1)(any)+ 12.16 (1,−1,9)(any) |
| + 1.84 (1,2,2)(ar) + 19.60 (1,11,−2)(any) |
| 1B9 | 9 | pIC50 = 8.19 − 18.84 (2,11,4)(any) − 6.13 (0,−2,−1)(any) + 8.45 (0,1,−2)(any) |
| − 8.71 (0,11,5)(any) − 7.54 (−1,8,2)(any) − 20.98 (1,6,−2)(np) |
| + 15.96 (1,13,1)(any) + 3.93 (1,2,0)(any) + 1.61 (0,0,−2)(any) |
We also computed the cross-correlation matrix of the GCODs from models
1B7 and
1B9 (
Table 2) to determine if two or more highly correlated GCODs appear in the same 4D-QSAR model. According to
Table 2, except only for two pairs of cells, the other pairs of descriptors are poorly correlated (R < 0.5). This means that each of these descriptors contributes in different ways to the 4D-QSAR models [
22]. The highest correlations occur between the pair of GCODs (
−1,8,2)(
any) and (
1,13,1)(
any) (R = 0.52) and also between the pair of GCODs (
0,−2,−1)(
any) and (
0,1,−2)(
any) (R = 0.47). The first pair of GCODs may be found in both models (
1B7 and
1B9), while the second pair is found only in model
1B9.
Table 2.
Cross-correlation matrix of the GCODs and the experimental pIC50 values of models 1B7 and 1B9.
Table 2.
Cross-correlation matrix of the GCODs and the experimental pIC50 values of models 1B7 and 1B9.
| | Potency | (2,11,4) (any) | (0,−2,−1) (any) | (0,1,−2) (any) | (0,11,5) (any) | (−1,8,2) (any) | (1,6,−2) (np) |
|---|
| Potency | 1.00 | | | | | | |
| (2,11,4)(any) | −0.46 | 1.00 | | | | | |
| (0,−2,−1)(any) | −0.31 | 0.13 | 1.00 | | | | |
| (0,1,−2)(any) | −0.26 | 0.36 | 0.47 | 1.00 | | | |
| (0,11,5)(any) | −0.35 | 0.30 | 0.01 | 0.13 | 1.00 | | |
| (−1,8,2)(any) | −0.27 | 0.20 | 0.10 | 0.12 | −0.17 | 1.00 | |
| (1,6,−2)(np) | −0.21 | −0.21 | −0.07 | −0.13 | −0.21 | 0.06 | 1.00 |
| (1,13,1)(any) | 0.12 | 0.13 | 0.00 | −0.07 | −0.37 | 0.52 | 0.01 |
| (1,2,0)(any) | 0.36 | −0.01 | 0.18 | 0.18 | −0.16 | 0.22 | 0.11 |
| (0,0,−2)(any) | 0.43 | −0.31 | 0.26 | −0.19 | 0.01 | −0.11 | 0.07 |
| (0,−2,9)(any) | −0.30 | 0.05 | 0.20 | 0.16 | −0.29 | 0.18 | −0.05 |
| (1,−1,9)(any) | 0.36 | −0.26 | −0.11 | −0.08 | −0.11 | −0.09 | 0.06 |
| (1,2,2)(ar) | 0.33 | −0.18 | -0.27 | 0.21 | 0.14 | −0.16 | −0.09 |
| (1,11,−2)(any) | 0.60 | −0.24 | −0.10 | −0.17 | −0.41 | −0.06 | −0.27 |
| | (1,13,1)(any) | (1,2,0)(any) | (0,0,−2)(any) | (0,−2,9)(any) | (1,−1,9)(any) | (1,2,2)(ar) | (1,11,−2)(any) |
| (1,13,1)(any) | 1.00 | | | | | | |
| (1,2,0)(any) | 0.10 | 1.00 | | | | | |
| (0,0,−2)(any) | −0.13 | 0.28 | 1.00 | | | | |
| (0,−2,9)(any) | 0.12 | −0.11 | −0.35 | 1.00 | | | |
| (1,−1,9)(any) | −0.27 | 0.41 | 0.35 | −0.23 | 1.00 | | |
| (1,2,2)(ar) | −0.17 | 0.34 | 0.06 | −0.18 | 0.35 | 1.00 | |
| (1,11,−2)(any) | 0.06 | 0.08 | 0.31 | 0.07 | 0.02 | −0.08 | 1.00 |
The cross-correlation matrix of the experimental pIC
50 values and the frequency of grid cell occupancy of models
1B7 and
1B9 were calculated (
Table 2). It has been demonstrated that the highest individual correlation with activity, except only for GCOD (
1,11,−2)(
any), is shown by GCODs (
2,11,4)(
any) (R = −0.46), (
0,0,2)(
any) (R = 0.43), (
1,2,0)(
any) (R = 0.36), and (
0,11,5)(
any) (R = −0.35), which are present only in model
1B9.
Outliers were defined as those compounds whose residuals are higher than twice the standard deviation of the residual of fit (SDres). The standard deviations were computed for the residuals of all 41 compounds of the training set for models 1B7 (SDres = 0.53) and 1B9 (SDres = 0.38). The results show that model 1B9 presents lower SDres value and, consequently, only compound 9 was defined as an outlier. On the other hand, model 1B7 shows a higher SDres value and presents three outliers (namely, compounds 2, 6, and 13).
2.3. Graphical Analysis of the Representative Model of Alignment 1
Since Model
1B9, as described in Equation 1 (
Figure 4), exhibits higher R
2adj and Q
2adj values, and fewer number of outliers than Model
1B7, it was selected as the representative model. According to the cross-correlation matrix of the Model
1B9 GCODs (
Table 2), grid cells (
1,13,1)(
any) and (
−1,8,2)(
any) seem to partially supply (R = 0.52) the same type of structure-activity information, in spite of the fact that these cells are spatially distant (5.48 Å) and have opposite contributions. The GCOD (
1,13,1)(
any) occupation increases the compounds potency, while the GCOD (
−1,8,2)(
any) occupation decreases. Both descriptors are located close to the side chain of the arylbenzothiophenes, being related to the flexibility of this basic side chain. In fact, GCOD (
1,13,1)(
any) is close to the carbon atom (C4) of the piperidine ring, while GCOD (
−1,8,2)(
any) is close to the oxygen atom of the ethoxy-phenyl group (
Figure 3).
This correlation is probably due to an intramolecular hydrogen bonding (N∙∙∙O distance = 3.02 Å), reinforced by a strong attractive electrostatic interaction between the corresponding amino and ethoxyl groups. This interaction causes a preferential synclinal conformation of this lateral chain, involving the four atoms of the N-C-C-O moiety. Therefore, since GCOD (
1,13,1)(
any) is the descriptor that most contributes to increase the potency (coefficient value = 15.96, Equation 1,
Figure 4), this fact indicates the relevance of the synclinal conformation of the basic side chain for the structure-activity relationship (SAR) of this series of compounds.
Figure 4.
Graphical representation of compounds 1 (a) and 54 (b) according to the 4D-QSAR Model 1B9 (Alignment 1, grid cell size of 1.0 Å, and 9 terms). The postulated “bioactive” conformations of compounds 1 and 54 (stick models in yellow) were superposed (according Alignment 1) to the X-ray structure of raloxifene (stick models in blue) bound in the LBD of ERα (secondary structure and line models of selected amino acid residues in gray). GCODs of Model 1B9 are: (A) = (1,13,1)(any); (B) = (2,11,4)(any); (C) = (0,11,5)(any); (D) = (−1,8,2)(any); (E) = (1,6,−2)(np); (F) = (1,2,0)(any); (G) = (0,1,−2)(any); (H) = (0,0,−2)(any); and (I) = (0,−2,−1)(any). The white and black spheres represent the GCODs which occupation contribute to increase (GCODs A, F–H) or to decrease (GCODs B–E, and I) the potency of the compounds.
Figure 4.
Graphical representation of compounds 1 (a) and 54 (b) according to the 4D-QSAR Model 1B9 (Alignment 1, grid cell size of 1.0 Å, and 9 terms). The postulated “bioactive” conformations of compounds 1 and 54 (stick models in yellow) were superposed (according Alignment 1) to the X-ray structure of raloxifene (stick models in blue) bound in the LBD of ERα (secondary structure and line models of selected amino acid residues in gray). GCODs of Model 1B9 are: (A) = (1,13,1)(any); (B) = (2,11,4)(any); (C) = (0,11,5)(any); (D) = (−1,8,2)(any); (E) = (1,6,−2)(np); (F) = (1,2,0)(any); (G) = (0,1,−2)(any); (H) = (0,0,−2)(any); and (I) = (0,−2,−1)(any). The white and black spheres represent the GCODs which occupation contribute to increase (GCODs A, F–H) or to decrease (GCODs B–E, and I) the potency of the compounds.
The cells (
2,11,4)(
any) and (
0,11,5)(
any) are close to each other and present negative coefficients, being also related to the basic side chain, located near the carbon (C2) and nitrogen atoms of the piperidine ring, respectively (
Figure 4). Altogether, those four GCODs indicate a preferential orientation of the piperidine side chain. This group is involved in an intermolecular hydrogen bond, intensified by an electrostatic interaction with Asp351, which is corroborated by Wang and co-workers [
24]. The authors employed a series of compounds, structurally related to raloxifene, in construction of a 2D-QSAR model and proposed that hydrogen bonds are important, but not an unique feature to binding affinity. That orientation is essential to increase or decrease the potency of the raloxifene analogs. Additionally, the basic side chain of raloxifene also makes extensive hydrophobic contacts with the alpha helixes H3, H5/6, H11 and the loop between the alpha helixes H11 and H12 [
3], reinforcing the importance of the orientation and conformation of this basic side chain.
The GCODs (
0,1,−2)(
any) and (
0,−2,−1)(
any) can also be considered to contain some degree of similarity (R = 0.47). In spite of presenting opposite contributions, these cells are located close to each other (3.16Å), partially leading to the same type of information. The GCODs (
1,2,0)(
any), (
0,1,−2)(
any), (
0,0,−2)(
any), and (
0,−2,−1)(
any) reflect the importance of the hydrogen-bonding network close to the benzothiophenyl moiety for the antagonist activity of the ERα ligands. In fact, the cell (
0,0,−2)(
any) is directly related to the hydrogen bonding interactions of the 6-OH group of the benzothiophene ring with Glu353 and Arg394, as described in previous SAR studies [
3,
11,
16,
25]. The occupation of this cell is drastically reduced when this position is non-substituted, or the substituents are unable to perform those hydrogen bond interactions (e.g.,
23,
30,
31,
35,
38, and
41). The same happens with GCOD (
0,1,−2)(
any), since compounds with hydrogen bond acceptor substituents at C6-position have a high frequency of occupation (e.g.,
37,
49, and
50).
The X-ray crystal structure of the raloxifene-ERα complex shows the benzothiophene ring of raloxifene surrounded by hydrophobic residues, such as Leu349, Ala350, Leu387, Leu391 and Phe404 [
3]. Therefore, the occupation by any atom types in grid cell
(0,−2,−1) decreases the potency of the compounds due to steric factors. This can be explained by the fact that compounds with bulky substituents, such as methoxyl or acetyl groups (e.g.,
21,
28,
37,
39,
43,
47,
49, and
52), have a greater occupation frequency than compounds with less bulky substituents (e.g.,
19,
23,
30, and
38). In addition, these substituents are not able to perform hydrogen bonding interactions with Glu353 and Arg394 or they can sterically impair these interactions.
Finally, the GCOD (
1,6,−2)(
np) (
Figure 4) is located in a 3D-box area that corresponds to the Ala350 residue. The negative coefficient of this GCOD (
Figure 4) indicates that the occupation of this cell by non-polar atoms reduces the compound potency, probably due to steric factors. Although
Figure 4 does not show clearly any atom of the ligands around this cell, some conformations and orientations adopted by the compounds during the MDS (data not shown) enable the carbon and hydrogen atoms of the piperidine ring to occupy this grid cell. It is important to notice that
Figure 4 shows only one conformation, selected as the “bioactive” one, among the 2,000 conformers from the MDS of each compound, which leads to maximum potency according to Model
1B9. In fact compounds with substituents at position 2' of the phenyl ring have low occupation frequency (e.g.,
3,
7,
8, and
16), as well as compounds with substituents at positions 4 and 5 of the benzothiophene ring (e.g.,
13,
29,
31,
35,
43,
44,
45,
47, and
49). This fact indicates that such substituents try to maintain a favorable conformation of the basic side chain of the compounds to the antagonism towards ERα. This additional characteristic has not been revealed by the LIV-3D-QSAR Model developed by Cunha and co-workers using this series of compounds [
18].
The absence of any descriptors around the phenyl ring, specifically related to the 4'-OH group, which is responsible for the hydrogen bond interaction with the backbone atoms of His524, corroborates what previous SAR studies already demonstrated [
11,
16],
i.e., the 6-OH group of the benzothiophenyl ring is more important for the biological activity than the 4'-OH group. However, the absence of any descriptors related to the 4'-position of the phenyl ring suggests some limitation of the model. Additionally, unlike observed in the LIV-3D-QSAR Model [
18], the postulated “bioactive” conformation obtained in the 4D-QSAR Model
1B9 (
Figure 4) is very similar to the one adopted by raloxifene in the X-ray co-crystal structure [
3]. Comparing the compounds conformations from Model
1B9 with the raloxifene-ERα X-ray structure, the nitrogen atom of the piperidine ring is in a very close position to that one observed in the crystal, being the distance among them of 1.33Å for
1 (the most potent), and of 0.29Å for
54 (the least potent). These distances were calculated after RMS superposition of these compounds conformations over the X-ray structure of raloxifene bound in the LBD of ERα, according Alignment 1.
The SDres value for the training set was 0.38, which indicated compound 9 as an outlier (
Figure 5 and
Table 3). The only structural difference between this compound and raloxifene (1) is an additional substituent, 3'-Cl, at the phenyl ring of 9. According to Model 1B9, the predicted potency for compound 9 is lower than the experimental one, probably due to some limitation of the model, that does not show descriptors around the phenyl ring (as stated previously for the 4' position of the phenyl ring), or because of the presence of only few compounds with 3'-substituents, leading the model to underestimate the potency. Another possible explanation for the outlier behavior of compound 9 could be lipophilicity, which was not considered as a descriptor in this 4D-QSAR model. The calculated LogP value (cLogP) for the isomers containing one hydroxyl group in any position of the phenyl and benzothiophenyl rings is identical, e.g., the cLogP value for compounds
1,
16,
31,
35, and
41 is 5.96 [
26]. However, the insertion of a chlorine atom in the phenyl ring increases the lipophilicity, since the cLogP value of compound
9 is 6.63.
Wang and co-workers [
24] results indicated that LogP may not be essential to binding affinity on estrogen receptor α. However, the compounds have to be able to penetrate through cellular membranes. A 2D-QSAR [
26] was performed to the same data set of this study, including LogP as a potential descriptor into the analysis. The authors concluded that the benzothiophene moiety is lipophilic enough to pass through this stage, which corroborate this hypothesis, since there is a significant positive correlation between potency and lipophilicity [
25,
26].
Figure 5.
Graphical representation of compounds 9 (a) and 51 (b) according to the 4D-QSAR Model 1B9 (Alignment 1, grid cell size of 1.0 Å, and 9 terms). The postulated “bioactive” conformations of compounds 9 and 51 (stick models in yellow) are superposed (according Alignment 1) to the X-ray structure of raloxifene (stick models in blue) bound in the LBD of ERα (secondary structure and line models of selected amino acid residues in gray). GCODs of Model 1B9 are: (A) = (1,13,1)(any); (B) = (2,11,4)(any); (C) = (0,11,5)(any); (D) = (−1,8,2)(any); (E) = (1,6,−2)(np); (F) = (1,2,0)(any); (G) = (0,1,−2)(any); (H) = (0,0,−2)(any); and (I) = (0,−2,−1)(any). The white and black spheres represent the GCODs which occupation contributes to increase (GCODs A, F–H) or to decrease (GCODs B–E, and I) the potency of the compounds.
Figure 5.
Graphical representation of compounds 9 (a) and 51 (b) according to the 4D-QSAR Model 1B9 (Alignment 1, grid cell size of 1.0 Å, and 9 terms). The postulated “bioactive” conformations of compounds 9 and 51 (stick models in yellow) are superposed (according Alignment 1) to the X-ray structure of raloxifene (stick models in blue) bound in the LBD of ERα (secondary structure and line models of selected amino acid residues in gray). GCODs of Model 1B9 are: (A) = (1,13,1)(any); (B) = (2,11,4)(any); (C) = (0,11,5)(any); (D) = (−1,8,2)(any); (E) = (1,6,−2)(np); (F) = (1,2,0)(any); (G) = (0,1,−2)(any); (H) = (0,0,−2)(any); and (I) = (0,−2,−1)(any). The white and black spheres represent the GCODs which occupation contributes to increase (GCODs A, F–H) or to decrease (GCODs B–E, and I) the potency of the compounds.
Table 3.
Experimental (pIC50Exp) and calculated (pIC50Calc) potencies and residuals values (pIC50Calc—pIC50Exp) of Models 1B9 and 2B9 of 4D-QSAR.
Table 3.
Experimental (pIC50Exp) and calculated (pIC50Calc) potencies and residuals values (pIC50Calc—pIC50Exp) of Models 1B9 and 2B9 of 4D-QSAR.
| # a | pIC50Exp | Model 1B9 | Model 2B9 |
|---|
| pIC50Calc | Residue | pIC50Calc | Residue |
|---|
| 1 | 9.70 | 9.84 | 0.14 | 9.27 | −0.43 |
| 2 | 9.52 | 9.09 | −0.43 | 8.11 | −1.41 * |
| 3 | 9.15 | 9.30 | 0.15 | 9.48 | 0.33 |
| 4 | 9.10 | 8.77 | −0.33 | 9.04 | −0.06 |
| 5 | 9.00 | 9.16 | 0.16 | 8.97 | −0.03 |
| 6 | 9.00 | 8.49 | −0.51 | 8.81 | −0.19 |
| 7 | 8.70 | 8.57 | −0.13 | 8.89 | 0.19 |
| 8 | 8.70 | 8.24 | −0.46 | 8.32 | −0.38 |
| 9 | 8.64 | 7.83 | −0.81 * | 8.83 | 0.19 |
| 10 | 8.64 | 8.45 | −0.19 | 8.97 | 0.33 |
| 11 | 8.60 | 8.01 | −0.59 | 8.32 | −0.28 |
| 12 | 8.60 | 8.08 | −0.52 | 8.59 | −0.01 |
| 13 | 8.52 | 7.86 | −0.66 | 8.47 | −0.05 |
| 14 | 8.30 | 8.38 | 0.08 | 7.88 | −0.42 |
| 15 | 8.15 | 8.09 | −0.06 | 7.97 | −0.18 |
| 16 | 8.00 | 8.39 | 0.39 | 7.86 | −0.14 |
| 17 | 8.00 | 8.25 | 0.25 | 7.45 | −0.55 |
| 18 | 7.70 | 7.85 | 0.15 | 7.76 | 0.06 |
| 19 | 7.70 | 7.41 | −0.29 | 6.92 | −0.78 |
| 20 | 7.52 | 7.81 | 0.29 | 7.58 | 0.06 |
| 21 | 7.52 | 7.62 | 0.10 | 7.41 | −0.11 |
| 22 | 7.49 | 7.73 | 0.24 | 7.14 | −0.35 |
| 23 | 7.46 | 7.10 | −0.36 | 6.87 | −0.59 |
| 24 | 7.40 | 7.46 | 0.06 | 7.86 | 0.46 |
| 25 | 7.30 | 7.89 | 0.59 | 7.28 | −0.02 |
| 26 | 7.30 | 7.68 | 0.38 | 7.11 | −0.19 |
| 27 | 7.30 | 6.93 | −0.37 | 7.19 | −0.11 |
| 28 | 7.22 | 6.76 | −0.46 | 7.01 | −0.21 |
| 29 | 7.00 | 7.08 | 0.08 | 6.45 | −0.55 |
| 30 | 7.00 | 6.73 | −0.27 | 6.59 | −0.41 |
| 31 | 7.00 | 7.31 | 0.31 | 7.46 | 0.46 |
| 32 | 7.00 | 7.64 | 0.64 | 7.47 | 0.47 |
| 33 | 7.00 | 7.71 | 0.71 | 7.60 | 0.60 |
| 34 | 7.00 | 6.37 | −0.63 | 6.86 | −0.14 |
| 35 | 6.72 | 6.08 | −0.64 | 6.27 | −0.45 |
| 36 | 6.70 | 6.84 | 0.14 | 6.33 | −0.37 |
| 37 | 6.60 | 6.53 | −0.07 | 7.01 | 0.41 |
| 38 | 6.52 | 6.39 | −0.13 | 6.91 | 0.39 |
| 39 | 6.52 | 6.02 | −0.50 | 6.40 | −0.12 |
| 40 | 6.52 | 6.63 | 0.11 | 6.32 | −0.20 |
| 41 | 6.52 | 6.97 | 0.45 | 6.57 | 0.05 |
| 42 | 6.49 | 6.77 | 0.28 | 6.58 | 0.09 |
| 43 | 6.46 | 6.09 | −0.37 | 6.39 | −0.07 |
| 44 | 6.46 | 6.56 | 0.10 | 6.76 | 0.30 |
| 45 | 6.40 | 7.01 | 0.61 | 6.26 | −0.14 |
| 46 | 6.30 | 6.15 | −0.15 | 6.07 | −0.23 |
| 47 | 6.30 | 6.18 | −0.12 | 6.45 | 0.15 |
| 48 | 6.30 | 6.65 | 0.35 | 6.52 | 0.22 |
| 49 | 6.30 | 6.27 | −0.03 | 7.44 | 1.14 * |
| 50 | 6.22 | 6.43 | 0.21 | 6.40 | 0.18 |
| 51 | 6.00 | 7.12 | 1.12* | 7.14 | 1.14 * |
| 52 | 6.00 | 6.46 | 0.46 | 7.05 | 1.05 * |
| 53 | 6.00 | 6.17 | 0.17 | 6.28 | 0.28 |
| 54 | 6.00 | 6.39 | 0.39 | 6.61 | 0.61 |
As described in the previous section, the test data set were used to accomplish a “real” prediction using the 4D-QSAR Model
1B9. The value of SD
res found for the test data set was 0.38, indicating compound
51 as an outlier (
Table 3). The only structural difference between this compound and raloxifene (1) is the substitution of the 4'-OH of
1 by the 4'-OMe in
51. The potency of compound
51 was overestimated by Model
1B9. Again, this fact can be due to a limitation of the model, which does not present descriptors around the phenyl ring. Thus, it would not distinguish some putative unfavorable interaction of the methoxyl group with the neighboring residues (
Figure 5). Besides, classic QSAR studies demonstrated that there is a negative steric effect for 4'-substituents in the phenyl ring [
26].
2.5. Graphical Analysis of the Representative Model of Alignment 2
Model
2B9 (described in Equation 2) was selected as the most representative model of Alignment 2, as previous reported in the selection criteria outlined above. According to the cross-correlation matrix of grid cell occupancy of model
2B9 (
Table 5), the descriptors are nearly orthogonal and contribute in a different way to the 4D-QSAR models, except for only two pairs of GCODs. The GCODs (
0,10,−2)(
any) and
(0,12,−1)(
any) shows high correlation (R = 0.73). Although the GCOD (
0,10,−2)(
any) occupation increases the compounds potency and the GCOD (
0,12,−1)(
any) occupation decreases, these cells are close in space (distance of 2.24 Å), what would justify the correlation between them. The GCOD (
0,10,−2)(
any) shows an ambiguity, because it is located in an area of the 3D grid cell close to Asp351 (
Figure 6). Therefore, it is not expected that this GCOD occupation increases the compounds potency. It demonstrates that model
2B9 is unable to “predict” the presence of Asp351 and the attractive electrostatic interaction between this residue and the piperidine group of raloxifene, as can be noticed by visual inspection of the 3D structure of the raloxifene-ERα complex (PDB code 1ERR [
3]). This data may be used to rationalize the underestimation of the potency of compound
1, the most potent compound of the series under study.
The second pair of GCODs that shows medium correlation (R = 0.44) corresponds to GCODs (
0,2,2)(
ar) and
(1,3,6)(
hba). Unlike the previous case, those descriptors are distant in space (distance of 4.24Å) and both contribute to increase the potency. The GCOD (
0,2,2)(
ar) is located close to the carbon and hydrogen atoms at position 4 of the benzothiophene ring, while the GCOD (
1,3,6)(
hba) is located close to the oxygen atom of the carbonyl group. Those GCODs are related to the dihedral angle formed by the carbonyl and benzothiophenyl planes, indicating the importance of the coplanar orientation of the side chain, as described by Grese and coworkers [
11]. The molecules that occupy these cells most frequently are those that present substituents at position 2' of the phenyl ring, e.g., compounds
3,
7,
8, and
16. Substituents at these positions generate steric repulsion with the oxygen atom of the carbonyl group, leading the side chain to adopt a non-coplanar orientation.
GCODs (−1,12,6)(any) and (0,11,3)(p−) are located close to the piperidine ring, corroborating the importance of the side chain orientation. The occupation of GCOD (−1,12,6) by any atom type decreases the potency of the compounds, displacing the nitrogen atom of the piperidine group from the favorable position, related to the hydrogen bonding with residue Asp351. The GCOD (0,11,3)(p−) occupation is also related to the nitrogen atom of the side chain, since substituents at position 4 of the benzothiophene ring or at position 3' of the phenyl group have a high occupation frequency at this cell, e.g., compounds 5, 9, 29, 35, and 44. This indicates that these substituents are able to maintain a favorable conformation of the basic side chain of the compounds to the antagonism on ERα.
The GCOD (
2,5,0)(
any) (
Figure 6), located close to the residue Ala350, has the same behavior of grid cell (
1,6,−
2)(
np) from Model
1B9.
GCOD (0,0,−2)(any), located close to Glu353, indicates the importance of hydrogen bonding of this residue with the substituent at position 6 of the benzothiophene ring. This grid cell has a low frequency of occupation when there are no substituents at this position (e.g., compounds 23, 30 and 38) or the substituents are unable to perform this type of interaction (e.g., compound 19).
Figure 6.
Graphical representation of compounds 1 (a) and 54 (b) according to the 4D-QSAR Model 2B9 (Alignment 2, grid cell size of 1.0 Å, and 9 terms). The postulated “bioactive” conformations of compounds 1 and 54 (stick models in yellow) are superposed (according Alignment 2) to the X-ray structure of raloxifene (stick models in blue) bound in the LBD of ERα (secondary structure and line models of selected amino acid residues in gray). GCODs of Model 2B9 are (A) = (0,10,−2)(any); (B) = (0,12,−1)(any); (C) = (0,11,3)(p−); (D) = (−1,12,6)(any); (E) = (2,5,0)(any); (F) = (1,3,6)(hba); (G) = (0,2,2)(ar); (H) = (0,0,−2)(any); and (I) = (−1,−2,11)(any). The white and black spheres represent the GCODs which occupation contributes to increase (GCODs A, C, F, G, and H) or to decrease (GCODs B, D, E, and I) the potency of the compounds.
Figure 6.
Graphical representation of compounds 1 (a) and 54 (b) according to the 4D-QSAR Model 2B9 (Alignment 2, grid cell size of 1.0 Å, and 9 terms). The postulated “bioactive” conformations of compounds 1 and 54 (stick models in yellow) are superposed (according Alignment 2) to the X-ray structure of raloxifene (stick models in blue) bound in the LBD of ERα (secondary structure and line models of selected amino acid residues in gray). GCODs of Model 2B9 are (A) = (0,10,−2)(any); (B) = (0,12,−1)(any); (C) = (0,11,3)(p−); (D) = (−1,12,6)(any); (E) = (2,5,0)(any); (F) = (1,3,6)(hba); (G) = (0,2,2)(ar); (H) = (0,0,−2)(any); and (I) = (−1,−2,11)(any). The white and black spheres represent the GCODs which occupation contributes to increase (GCODs A, C, F, G, and H) or to decrease (GCODs B, D, E, and I) the potency of the compounds.
The GCOD (−1,−2,11)(any), located close to position 4' of the phenyl group, is related to the nonpolar interactions around this ring. As this area is wrapped up by hydrophobic residues (e.g., Ile424, Gly521, and Leu525), compounds with bulky substituents at this position (e.g., compounds 22, 33, 36, 39, 43 and 51) show steric hindrance and, consequently, lower potency.
Three compounds of the training set were identified as outliers:
2,
49 and
52 (
Table 3). The predicted activity for compound
2 is lower than the experimental one. The structural difference between this compound and raloxifene (
1) is an additional substituent, 3'-F, at the phenyl ring of
2. Like Model
1B9, this fact can be due to the existence of few compounds with substituents at position 3' of the phenyl ring. Therefore, the model does not reveal the importance of the substitution pattern at this position. Compounds
49 and
52 show higher predicted potencies than the experimental ones. The chemical difference between raloxifene (
1) and
49 is the presence of two additional substituents, 5,7-Me, at the phenyl ring. Model
2B9 does not show any descriptors around those substituents, which turns the model unable to recognize how they may influence the conformation of the side chain, leading to an unfavorable orientation for potency (
Figure 7). Compound
52 has an amide substituent at position 6 of the benzothiophene ring that has a high frequency of occupation of GCOD (
0,0,−
2)(
any). However, this cell presents a low coefficient (Equation 2), which tends to have a small contribution to Model
2B9.
Figure 7.
Graphical representation of compounds 2 (a), 49 (b) and 52 (c) according to the 4D-QSAR Model 2B9 (Alignment 2, grid cell size of 1.0 Å, and 9 terms). The postulated “bioactive” conformations of compounds 2, 49 and 52 (stick models in yellow) are superposed (according Alignment 2) to the X-ray structure of raloxifene (stick models in blue) bound in the LBD of ERα (secondary structure and line models of selected amino acid residues in gray). GCODs of Model 2B9 are (A) = (0,10,−2)(any); (B) = (0,12,−1)(any); (C) = (0,13,3)(p−); (D) = (−1,12,6)(any); (E) = (2,5,0)(any); (F) = (1,3,6)(hba); (G) = (0,2,2)(ar); (H) = (0,0,−2)(any); and (I) = (−1,−2,11)(any). The white and black spheres represent the GCODs which occupation contributes to increase (GCODs A, C, F, G, and H) or to decrease (GCODs B, D, E, and I) the potency of the compounds.
Figure 7.
Graphical representation of compounds 2 (a), 49 (b) and 52 (c) according to the 4D-QSAR Model 2B9 (Alignment 2, grid cell size of 1.0 Å, and 9 terms). The postulated “bioactive” conformations of compounds 2, 49 and 52 (stick models in yellow) are superposed (according Alignment 2) to the X-ray structure of raloxifene (stick models in blue) bound in the LBD of ERα (secondary structure and line models of selected amino acid residues in gray). GCODs of Model 2B9 are (A) = (0,10,−2)(any); (B) = (0,12,−1)(any); (C) = (0,13,3)(p−); (D) = (−1,12,6)(any); (E) = (2,5,0)(any); (F) = (1,3,6)(hba); (G) = (0,2,2)(ar); (H) = (0,0,−2)(any); and (I) = (−1,−2,11)(any). The white and black spheres represent the GCODs which occupation contributes to increase (GCODs A, C, F, G, and H) or to decrease (GCODs B, D, E, and I) the potency of the compounds.
Compound
51, from test data set, was identified as an outlier and its potency was overestimated by Model
2B9 (
Table 3). The same behavior was observed in Model
1B9. In a different fashion as observed with the representative model of alignment 1, Model
2B9 has a close descriptor at position 4' of the phenyl ring,
i.e., GCOD
(−1,−2,11)(any). This fact supports 2D-QSAR data about the negative steric effect of substituents at this position [
26]. This GCOD is also occupied by compound
51, but its contribution to the model is not significant, due to its small coefficient value when compared to other GCODs.
Although alignment 2 has considered atoms belonging to more rigid regions of the molecules and shows descriptors better distributed in space, alignment 1 has superior statistical indices and is more consistent with the considered raloxifene mechanism of action. Thus, based in the results of the 4D-QSAR Model
1B9 and in a previous LIV-3D-QSAR model from our group [
18], the synthesis of new SERMs candidates has been suggested (
Figure 8).
Figure 8.
Structures of the raloxifene IV (pIC50Exp = 9.70 M) and the proposed compounds V and VI (and calculated potencies based on the 4D-QSAR Model 1B9).
Figure 8.
Structures of the raloxifene IV (pIC50Exp = 9.70 M) and the proposed compounds V and VI (and calculated potencies based on the 4D-QSAR Model 1B9).