The Ig CAM CAR is Implicated in Cardiac Development and Modulates Electrical Conduction in the Mature Heart
Abstract
:1. The CAR Subgroup of IgCAMs
2. Localization of CAR in the Heart
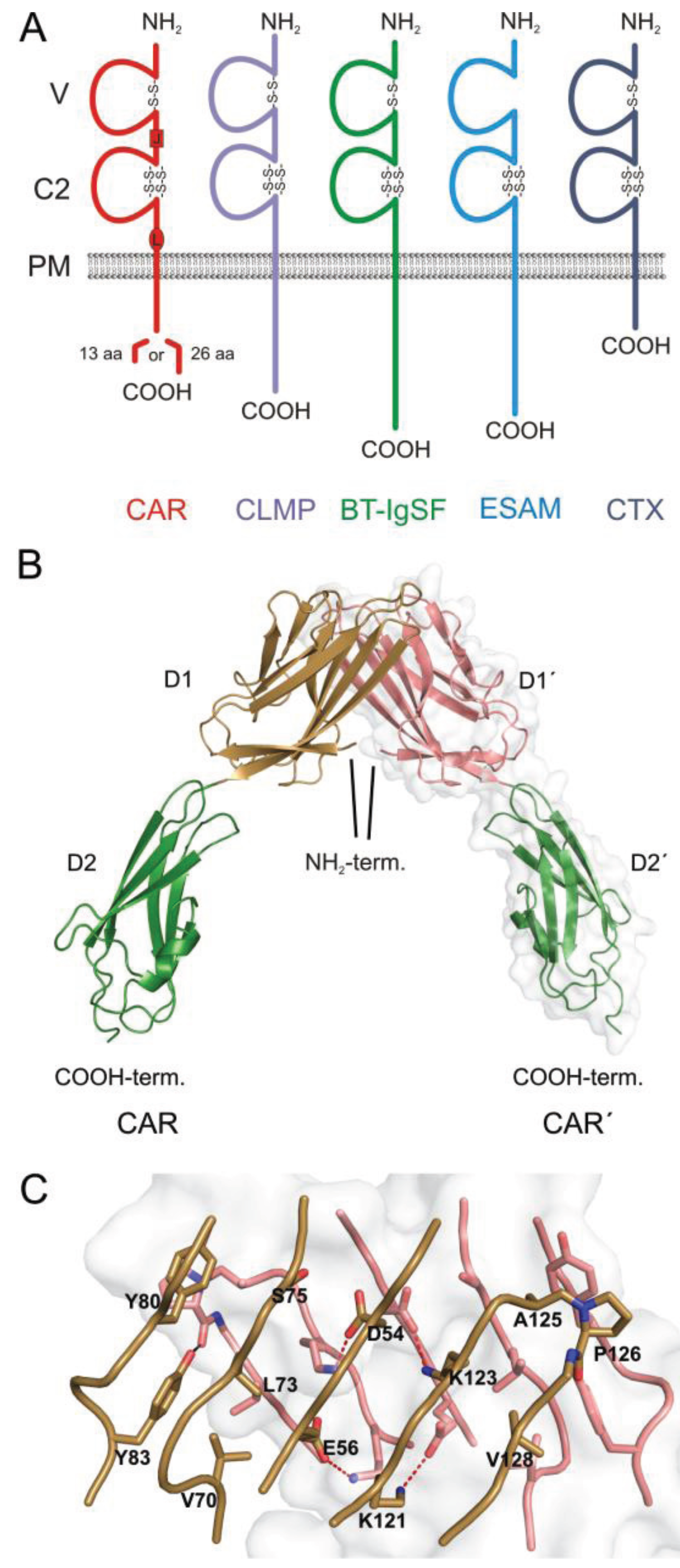
3. CAR Re-Expression in Diseased Cardiac Tissue
4. CAR is Essential for Embryonic Heart Development
5. CAR is Essential for Electrical Conduction in the Mature Heart
6. Outlook
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Bergelson, J.M.; Cunningham, J.A.; Droguett, G.; Kurt-Jones, E.A.; Krithivas, A.; Hong, J.S.; Horwitz, M.S.; Crowell, R.L.; Finberg, R.W. Isolation of a common receptor for Coxsackie B viruses and adenoviruses 2 and 5. Science 1997, 275, 1320–1323. [Google Scholar] [CrossRef]
- Tomko, R.P.; Xu, R.; Philipson, L. HCAR and MCAR: The human and mouse cellular receptors for subgroup C adenoviruses and group B coxsackieviruses. Proc. Natl. Acad. Sci. USA 1997, 94, 3352–3356. [Google Scholar] [CrossRef]
- Freimuth, P.; Philipson, L.; Carson, S.D. The coxsackievirus and adenovirus receptor. Curr. Top. Microbiol. Immunol. 2008, 323, 67–87. [Google Scholar]
- Coyne, C.B.; Bergelson, J.M. CAR: A virus receptor within the tight junction. Adv. Drug Deliv. Rev. 2005, 57, 869–882. [Google Scholar] [CrossRef]
- Schreiber, J.; Langhorst, H.; Jüttner, R.; Rathjen, F.G. The IgCAMs CAR, BT-IgSF and CLMP: Structure, function and diseases. In Cell Adhesion Molecules. Implications in Neurological Diseases; Berezin, V., Walmod, P.S., Eds.; Springer: New York, NY, USA, 2014; pp. 21–45. [Google Scholar]
- Hirata, K.; Ishida, T.; Penta, K.; Rezaee, M.; Yang, E.; Wohlgemuth, J.; Quertermous, T. Cloning of an immunoglobulin family adhesion molecule selectively expressed by endothelial cells. J. Biol. Chem. 2001, 276, 16223–16231. [Google Scholar]
- Nasdala, I.; Wolburg-Buchholz, K.; Wolburg, H.; Kuhn, A.; Ebnet, K.; Brachtendorf, G.; Samulowitz, U.; Kuster, B.; Engelhardt, B.; Vestweber, D.; et al. A transmembrane tight junction protein selectively expressed on endothelial cells and platelets. J. Biol. Chem. 2002, 277, 16294–16303. [Google Scholar] [CrossRef]
- Suzu, S.; Hayashi, Y.; Harumi, T.; Nomaguchi, K.; Yamada, M.; Hayasawa, H.; Motoyoshi, K. Molecular cloning of a novel immunoglobulin superfamily gene preferentially expressed by brain and testis. Biochem. Biophys. Res. Commun. 2002, 296, 1215–1221. [Google Scholar] [CrossRef]
- Raschperger, E.; Engstrom, U.; Pettersson, R.F.; Fuxe, J. CLMP, a novel member of the CTX family and a new component of epithelial tight junctions. J. Biol. Chem. 2004, 279, 796–804. [Google Scholar]
- Katoh, M.; Katoh, M. IGSF11 gene, frequently up-regulated in intestinal-type gastric cancer, encodes adhesion molecule homologous to CXADR, FLJ22415 and ESAM. Int. J. Oncol. 2003, 23, 525–531. [Google Scholar]
- Chretien, I.; Robert, J.; Marcuz, A.; Garcia-Sanz, J.A.; Courtet, M.; Du, P.L. CTX, a novel molecule specifically expressed on the surface of cortical thymocytes in Xenopus. Eur. J. Immunol. 1996, 26, 780–791. [Google Scholar] [CrossRef]
- Chretien, I.; Marcuz, A.; Courtet, M.; Katevuo, K.; Vainio, O.; Heath, J.K.; White, S.J.; Du, P.L. CTX, a Xenopus thymocyte receptor, defines a molecular family conserved throughout vertebrates. Eur. J. Immunol. 1998, 28, 4094–4104. [Google Scholar] [CrossRef]
- Honda, T.; Saitoh, H.; Masuko, M.; Katagiri-Abe, T.; Tominaga, K.; Kozakai, I.; Kobayashi, K.; Kumanishi, T.; Watanabe, Y.G.; Odani, S.; et al. The coxsackievirus-adenovirus receptor protein as a cell adhesion molecule in the developing mouse brain. Brain Res. Mol. Brain Res. 2000, 77, 19–28. [Google Scholar] [CrossRef]
- Walters, R.W.; Freimuth, P.; Moninger, T.O.; Ganske, I.; Zabner, J.; Welsh, M.J. Adenovirus fiber disrupts CAR-mediated intercellular adhesion allowing virus escape. Cell 2002, 110, 789–799. [Google Scholar] [CrossRef]
- Patzke, C.; Max, K.E.; Behlke, J.; Schreiber, J.; Schmidt, H.; Dorner, A.A.; Kroger, S.; Henning, M.; Otto, A.; Heinemann, U.; et al. The coxsackievirus-adenovirus receptor reveals complex homophilic and heterophilic interactions on neural cells. J. Neurosci. 2010, 30, 2897–2910. [Google Scholar] [CrossRef]
- Excoffon, K.J.; Hruska-Hageman, A.; Klotz, M.; Traver, G.L.; Zabner, J. A role for the PDZ-binding domain of the coxsackie B virus and adenovirus receptor (CAR) in cell adhesion and growth. J. Cell Sci. 2004, 117, (Pt 19). 4401–4409. [Google Scholar] [CrossRef]
- Van Raaij, M.J.; Chouin, E.; van der Zandt, H.; Bergelson, J.M.; Cusack, S. Dimeric structure of the coxsackievirus and adenovirus receptor D1 domain at 1.7 A resolution. Structure 2000, 8, 1147–1155. [Google Scholar] [CrossRef]
- Verdino, P.; Witherden, D.A.; Havran, W.L.; Wilson, I.A. The molecular interaction of CAR and JAML recruits the central cell signal transducer PI3K. Science 2010, 329, 1210–1214. [Google Scholar] [CrossRef]
- Witherden, D.A.; Verdino, P.; Rieder, S.E.; Garijo, O.; Mills, R.E.; Teyton, L.; Fischer, W.H.; Wilson, I.A.; Havran, W.L. The junctional adhesion molecule JAML is a costimulatory receptor for epithelial gammadelta T cell activation. Science 2010, 329, 1205–1210. [Google Scholar] [CrossRef]
- Zen, K.; Liu, Y.; McCall, I.C.; Wu, T.; Lee, W.; Babbin, B.A.; Nusrat, A.; Parkos, C.A. Neutrophil migration across tight junctions is mediated by adhesive interactions between epithelial coxsackie and adenovirus receptor and a junctional adhesion molecule-like protein on neutrophils. Mol. Biol. Cell 2005, 16, 2694–2703. [Google Scholar] [CrossRef]
- Luissint, A.C.; Lutz, P.G.; Calderwood, D.A.; Couraud, P.O.; Bourdoulous, S. JAM-L-mediated leukocyte adhesion to endothelial cells is regulated in cis by alpha4beta1 integrin activation. J. Cell Biol. 2008, 183, 1159–1173. [Google Scholar] [CrossRef]
- Guo, Y.L.; Bai, R.; Chen, C.X.; Liu, D.Q.; Liu, Y.; Zhang, C.Y.; Zen, K. Role of junctional adhesion molecule-like protein in mediating monocyte transendothelial migration. Arterioscler. Thromb. Vasc. Biol. 2009, 29, 75–83. [Google Scholar] [CrossRef]
- Mirza, M.; Hreinsson, J.; Strand, M.L.; Hovatta, O.; Soder, O.; Philipson, L.; Pettersson, R.F.; Sollerbrant, K. Coxsackievirus and adenovirus receptor (CAR) is expressed in male germ cells and forms a complex with the differentiation factor JAM-C in mouse testis. Exp. Cell Res. 2006, 312, 817–830. [Google Scholar] [CrossRef]
- Cohen, C.J.; Shieh, J.T.; Pickles, R.J.; Okegawa, T.; Hsieh, J.T.; Bergelson, J.M. The coxsackievirus and adenovirus receptor is a transmembrane component of the tight junction. Proc. Natl. Acad. Sci. USA 2001, 98, 15191–15196. [Google Scholar]
- Raschperger, E.; Thyberg, J.; Pettersson, S.; Philipson, L.; Fuxe, J.; Pettersson, R.F. The coxsackie- and adenovirus receptor (CAR) is an in vivo marker for epithelial tight junctions, with a potential role in regulating permeability and tissue homeostasis. Exp. Cell Res. 2006, 312, 1566–1580. [Google Scholar] [CrossRef]
- Coyne, C.B.; Voelker, T.; Pichla, S.L.; Bergelson, J.M. The coxsackievirus and adenovirus receptor interacts with the multi-PDZ domain protein-1 (MUPP-1) within the tight junction. J. Biol. Chem. 2004, 279, 48079–48084. [Google Scholar]
- Excoffon, K.J.; Gansemer, N.D.; Mobily, M.E.; Karp, P.H.; Parekh, K.R.; Zabner, J. Isoform-specific regulation and localization of the coxsackie and adenovirus receptor in human airway epithelia. PLoS ONE 2010, 5. [Google Scholar] [CrossRef]
- Mirza, M.; Raschperger, E.; Philipson, L.; Pettersson, R.F.; Sollerbrant, K. The cell surface protein coxsackie- and adenovirus receptor (CAR) directly associates with the Ligand-of-Numb Protein-X2 (LNX2). Exp. Cell Res. 2005, 309, 110–120. [Google Scholar] [CrossRef]
- Sollerbrant, K.; Raschperger, E.; Mirza, M.; Engstrom, U.; Philipson, L.; Ljungdahl, P.O.; Pettersson, R.F. The Coxsackievirus and adenovirus receptor (CAR) forms a complex with the PDZ domain-containing protein ligand-of-numb protein-X (LNX). J. Biol. Chem. 2003, 278, 7439–7444. [Google Scholar] [CrossRef]
- Bowles, K.R.; Gibson, J.; Wu, J.; Shaffer, L.G.; Towbin, J.A.; Bowles, N.E. Genomic organizationand chromosomal localization of the human Coxsackievirus B-adenovirus receptor gene. Hum. Genet. 1999, 105, 354–359. [Google Scholar] [CrossRef]
- Hattori, M.; Fujiyama, A.; Taylor, T.D.; Watanabe, H.; Yada, T.; Park, H.S.; Toyoda, A.; Ishii, K.; Totoki, Y.; Choi, D.K.; et al. The DNA sequence of human chromosome 21. Nature 2000, 405, 311–319. [Google Scholar] [CrossRef]
- Chen, J.W.; Ghosh, R.; Finberg, R.W.; Bergelson, J.M. Structure and chromosomal localization of the murine coxsackievirus and adenovirus receptor gene. DNA Cell Biol. 2003, 22, 253–259. [Google Scholar] [CrossRef]
- Tomko, R.P.; Johansson, C.B.; Totrov, M.; Abagyan, R.; Frisen, J.; Philipson, L. Expression of the adenovirus receptor and its interaction with the fiber knob. Exp. Cell Res. 2000, 255, 47–55. [Google Scholar] [CrossRef]
- Fechner, H.; Noutsias, M.; Tschoepe, C.; Hinze, K.; Wang, X.; Escher, F.; Pauschinger, M.; Dekkers, D.; Vetter, R.; Paul, M.; et al. Induction of coxsackievirus-adenovirus-receptor expressionduring myocardial tissue formation and remodeling: Identification of a cell-to-cell contact-dependent regulatory mechanism. Circulation 2003, 107, 876–882. [Google Scholar] [CrossRef]
- Dorner, A.A.; Wegmann, F.; Butz, S.; Wolburg-Buchholz, K.; Wolburg, H.; Mack, A.; Nasdala, I.; August, B.; Westermann, J.; Rathjen, F.G.; et al. Coxsackievirus-adenovirus receptor (CAR) is essential for early embryonic cardiac development. J. Cell Sci. 2005, 118, (Pt 15). 3509–3521. [Google Scholar] [CrossRef]
- Hotta, Y.; Honda, T.; Naito, M.; Kuwano, R. Developmental distribution of coxsackie virus and adenovirus receptor localized in the nervous system. Brain Res. Dev. Brain Res. 2003, 143, 1–13. [Google Scholar] [CrossRef]
- Kashimura, T.; Kodama, M.; Hotta, Y.; Hosoya, J.; Yoshida, K.; Ozawa, T.; Watanabe, R.; Okura, Y.; Kato, K.; Hanawa, H.; et al. Spatiotemporal changes of coxsackievirus and adenovirus receptor in rat hearts during postnatal development and in cultured cardiomyocytes of neonatal rat. Virchows Arch. 2004, 444, 283–292. [Google Scholar] [CrossRef]
- Shaw, C.A.; Holland, P.C.; Sinnreich, M.; Allen, C.; Sollerbrant, K.; Karpati, G.; Nalbantoglu, J. Isoform-specific expression of the Coxsackie and adenovirus receptor (CAR) in neuromuscular junction and cardiac intercalated discs. BMC Cell Biol. 2004, 5. [Google Scholar] [CrossRef] [Green Version]
- Vigl, B.; Zgraggen, C.; Rehman, N.; Banziger-Tobler, N.E.; Detmar, M.; Halin, C. Coxsackie- and adenovirus receptor (CAR) is expressed in lymphatic vessels in human skin and affects lymphatic endothelial cell function in vitro. Exp. Cell Res. 2009, 315, 336–347. [Google Scholar] [CrossRef]
- Sasse, A.; Wallich, M.; Ding, Z.; Goedecke, A.; Schrader, J. Coxsackie-and-adenovirus receptor mRNA expression in human heart failure. J. Gene Med. 2003, 5, 876–882. [Google Scholar] [CrossRef]
- Ito, M.; Kodama, M.; Masuko, M.; Yamaura, M.; Fuse, K.; Uesugi, Y.; Hirono, S.; Okura, Y.; Kato, K.; Hotta, Y.; et al. Expression of coxsackievirus and adenovirus receptor in hearts of rats with experimental autoimmune myocarditis. Circ. Res. 2000, 86, 275–280. [Google Scholar] [CrossRef]
- Noutsias, M.; Fechner, H.; de Jonge, H.; Wang, X.; Dekkers, D.; Houtsmuller, A.B.; Pauschinger, M.; Bergelson, J.; Warraich, R.; Yacoub, M.; et al. Human coxsackie-adenovirus receptor is colocalized with integrins alpha(v)beta(3) and alpha(v)beta(5) on the cardiomyocyte sarcolemma and upregulated in dilated cardiomyopathy: Implications for cardiotropic viral infections. Circulation 2001, 104, 275–280. [Google Scholar] [CrossRef]
- Tatrai, E.; Bedi, K.; Kovalszky, I.; Hartyanszky, I.; Laszik, A.; Acsady, G.; Sotonyi, P.; Hubay, M. No mutation but high mRNA expression of Coxsackie-Adenovirus Receptor was observed in both dilated and ischemic cardiomyopathy. Forensic Sci. Int. 2011, 212, 47–50. [Google Scholar] [CrossRef]
- Yuen, S.; Smith, J.; Caruso, L.; Balan, M.; Opavsky, M.A. The coxsackie-adenovirus receptor induces an inflammatory cardiomyopathy independent of viral infection. J. Mol. Cell. Cardiol. 2011, 50, 826–840. [Google Scholar] [CrossRef]
- Caruso, L.; Yuen, S.; Smith, J.; Husain, M.; Opavsky, M.A. Cardiomyocyte-targeted overexpression of the coxsackie-adenovirus receptor causes a cardiomyopathy in association with beta-catenin signaling. J. Mol. Cell. Cardiol. 2010, 48, 1194–1205. [Google Scholar] [CrossRef]
- Bowles, N.E.; Javier Fuentes-Garcia, F.; Makar, K.A.; Li, H.; Gibson, J.; Soto, F.; Schwimmbeck, P.L.; Schultheiss, H.P.; Pauschinger, M. Analysis of the coxsackievirus B-adenovirus receptor gene in patients with myocarditis or dilated cardiomyopathy. Mol. Genet. Metab. 2002, 77, 257–259. [Google Scholar] [CrossRef]
- Excoffon, K.J.; Avenarius, M.R.; Hansen, M.R.; Kimberling, W.J.; Najmabadi, H.; Smith, R.J.; Zabner, J. The Coxsackievirus and Adenovirus Receptor: A new adhesion protein in cochlear development. Hear. Res. 2006, 215, 1–9. [Google Scholar] [CrossRef]
- Asher, D.R.; Cerny, A.M.; Weiler, S.R.; Horner, J.W.; Keeler, M.L.; Neptune, M.A.; Jones, S.N.; Bronson, R.T.; DePinho, R.A.; Finberg, R.W. Coxsackievirus and adenovirus receptor is essential for cardiomyocyte development. Genesis 2005, 42, 77–85. [Google Scholar] [CrossRef]
- Chen, J.W.; Zhou, B.; Yu, Q.C.; Shin, S.J.; Jiao, K.; Schneider, M.D.; Baldwin, H.S.; Bergelson, J.M. Cardiomyocyte-specific deletion of the coxsackievirus and adenovirus receptor results in hyperplasia of the embryonic left ventricle and abnormalities of sinuatrial valves. Circ. Res. 2006, 98, 923–930. [Google Scholar] [CrossRef]
- Asher, D.; Finberg, R. CAR might provide a survival signal for myocardial cells. J. Cell Sci. 2005, 118, (Pt 24). 5679–5680. [Google Scholar] [CrossRef]
- Lim, B.K.; Xiong, D.; Dorner, A.; Youn, T.J.; Yung, A.; Liu, T.I.; Gu, Y.; Dalton, N.D.; Wright, A.T.; Evans, S.M.; et al. Coxsackievirus and adenovirus receptor (CAR) mediates atrioventricular-node function and connexin 45 localization in the murine heart. J. Clin. Investig. 2008, 118, 2758–2770. [Google Scholar] [CrossRef]
- Lisewski, U.; Shi, Y.; Wrackmeyer, U.; Fischer, R.; Chen, C.; Schirdewan, A.; Juttner, R.; Rathjen, F.; Poller, W.; Radke, M.H.; et al. The tight junction protein CAR regulates cardiac conduction and cell-cell communication. J. Exp. Med. 2008, 205, 2369–2379. [Google Scholar] [CrossRef]
- Nishii, K.; Kumai, M.; Egashira, K.; Miwa, T.; Hashizume, K.; Miyano, Y.; Shibata, Y. Mice lacking connexin45 conditionally in cardiac myocytes display embryonic lethality similar to that of germline knockout mice without endocardial cushion defect. Cell Commun. Adhes. 2003, 10, 365–369. [Google Scholar] [CrossRef]
- Marsman, R.F.; Bezzina, C.R.; Freiberg, F.; Verkerk, A.O.; Adriaens, M.E.; Podliesna, S.; Chen, C.; Purfurst, B.; Spallek, B.; Koopmann, T.T.; et al. Coxsackie and adenovirus receptor (CAR) is a modifier of cardiac conduction and arrhythmia vulnerability in the setting of myocardial ischemia. J. Am. Coll. Cardiol. 2014, 63, 549–559. [Google Scholar] [CrossRef] [Green Version]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Matthäus, C.; Schreiber, J.; Jüttner, R.; Rathjen, F.G. The Ig CAM CAR is Implicated in Cardiac Development and Modulates Electrical Conduction in the Mature Heart. J. Cardiovasc. Dev. Dis. 2014, 1, 111-120. https://doi.org/10.3390/jcdd1010111
Matthäus C, Schreiber J, Jüttner R, Rathjen FG. The Ig CAM CAR is Implicated in Cardiac Development and Modulates Electrical Conduction in the Mature Heart. Journal of Cardiovascular Development and Disease. 2014; 1(1):111-120. https://doi.org/10.3390/jcdd1010111
Chicago/Turabian StyleMatthäus, Claudia, Jadwiga Schreiber, René Jüttner, and Fritz G. Rathjen. 2014. "The Ig CAM CAR is Implicated in Cardiac Development and Modulates Electrical Conduction in the Mature Heart" Journal of Cardiovascular Development and Disease 1, no. 1: 111-120. https://doi.org/10.3390/jcdd1010111
APA StyleMatthäus, C., Schreiber, J., Jüttner, R., & Rathjen, F. G. (2014). The Ig CAM CAR is Implicated in Cardiac Development and Modulates Electrical Conduction in the Mature Heart. Journal of Cardiovascular Development and Disease, 1(1), 111-120. https://doi.org/10.3390/jcdd1010111




