A 2-Deoxyglucose-Resistant Mutant of Saccharomyces cerevisiae Shows Enhanced Maltose Fermentative Ability by the Activation of MAL Genes
Abstract
:1. Introduction
2. Materials and Methods
2.1. Yeast Strains
2.2. Investigation of 2-DOG Resistance in MCD4 Haploid Isolates
2.3. Maltose Fermentative Ability
2.4. DNA Sequencing and Identification of Amino Acid Substitutions/Deletions
2.5. Total RNA Extraction
2.6. Quantitative Reverse Transcription-Polymerase Chain Reaction (RT-PCR)
3. Results and Discussion
3.1. 2-DOG Resistance Improves Maltose Fermentative Ability
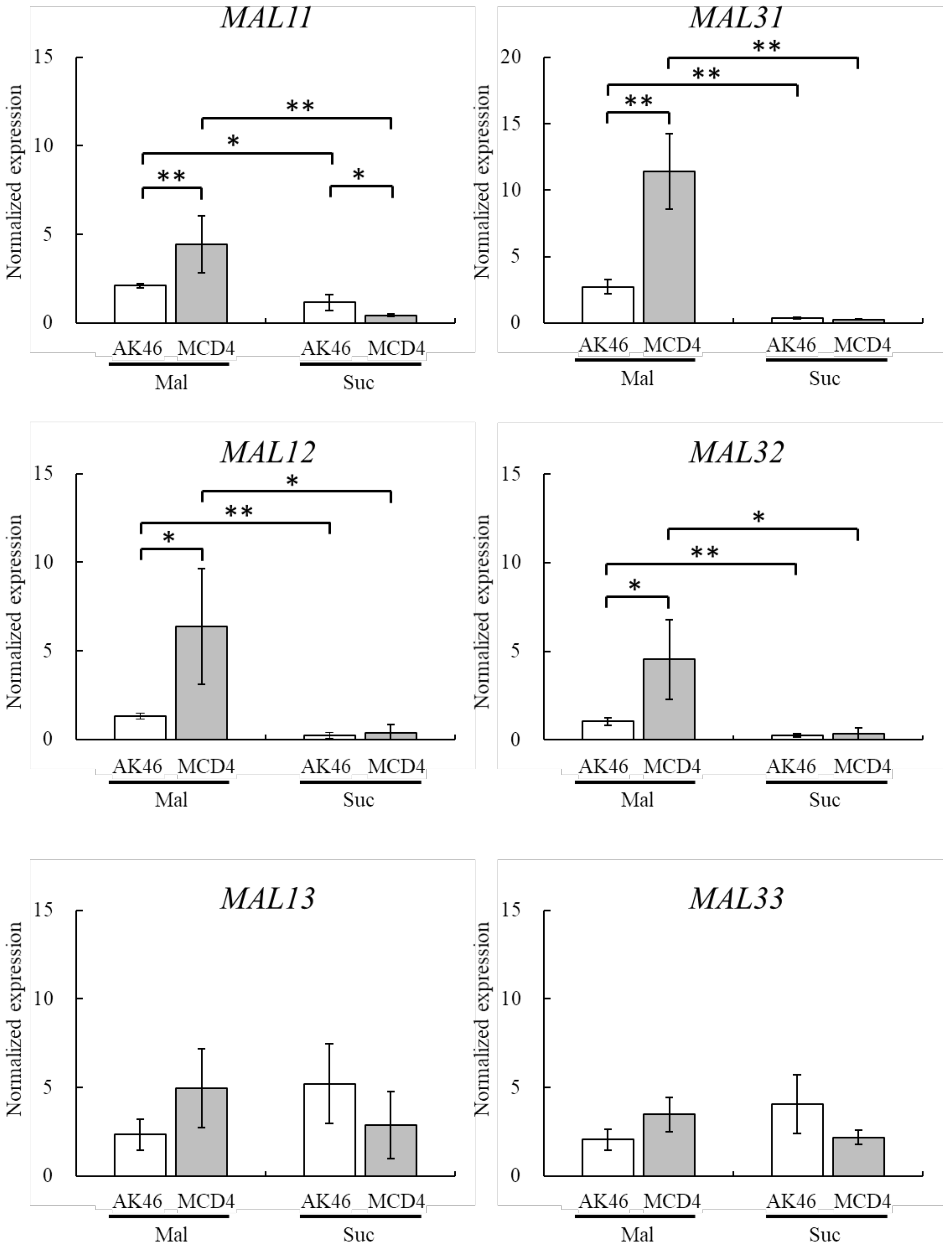
3.2. 2-DOG-Resistant Mutant MCD4 Activates MAL Genes
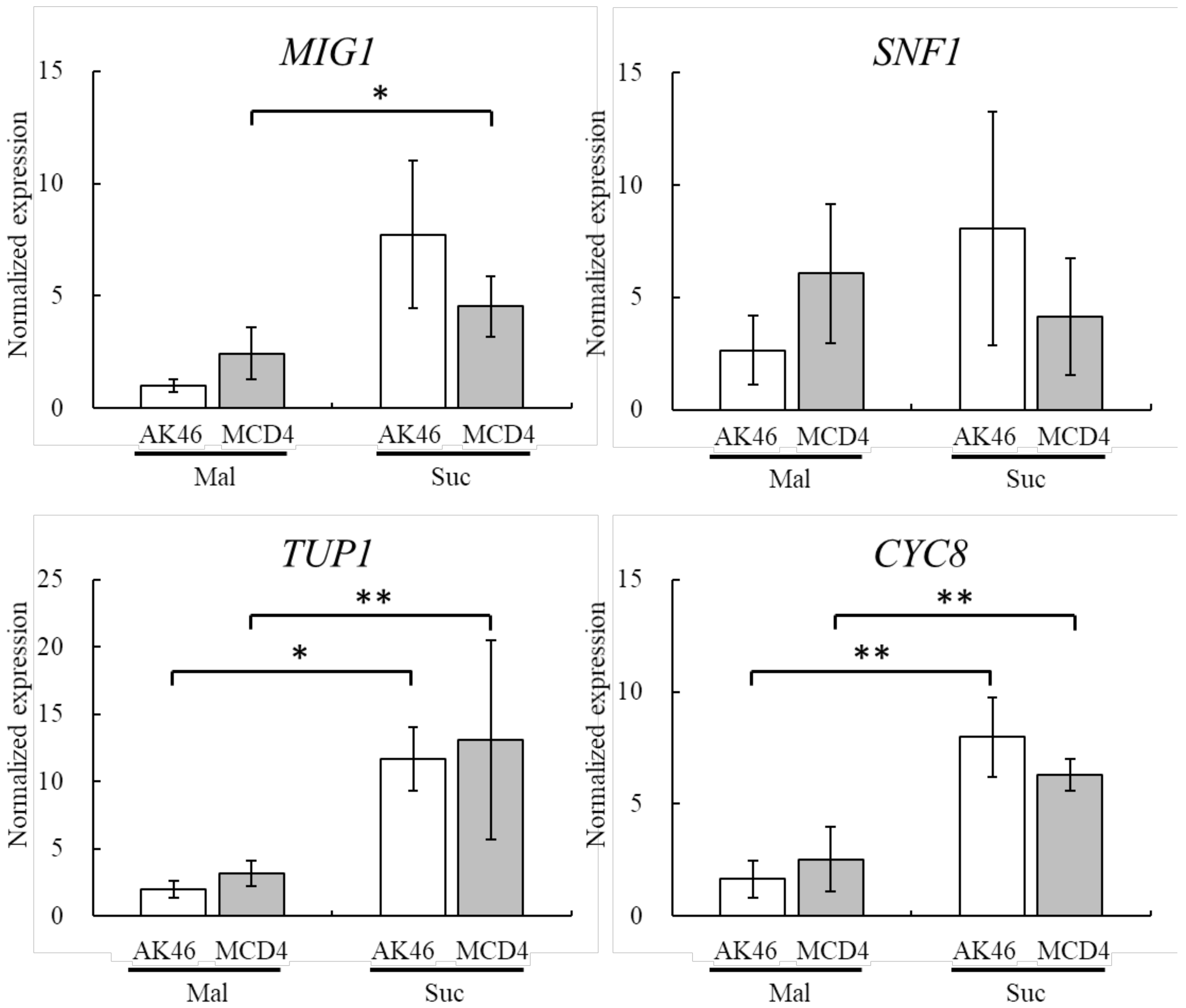
3.3. Involvement of Catabolite Repression-Related Gene in the Improved Maltose Fermentation of MCD4
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Sujaya, I.N.; Mikumo, D.; Orikasa, Y.; Urashima, T.; Oda, Y. Baking properties of Saccharomyces cerevisiae strains derived from brem, a traditional rice wine from Bali. Food Sci. Technol. Res. 2011, 17, 369–373. [Google Scholar] [CrossRef]
- Oda, Y.; Ouchi, K. Maltase gene and α-glucosidase activities: Their effects on dough-leavening. Yeast 1989, 5, 135–139. [Google Scholar]
- Oda, Y.; Ouchi, K. Principal-component analysis of the characteristics desirable in baker’s yeast. Appl. Environ. Microbiol. 1989, 55, 1495–1499. [Google Scholar] [PubMed]
- Randez-Gil, F.; Sanz, P. Construction of industrial baker’s yeast strains able to assimilate maltose under catabolite repression conditions. Appl. Microbiol. Biotechnol. 1994, 42, 581–586. [Google Scholar] [CrossRef]
- Charron, M.J.; Dubin, R.A.; Michels, C.A. Structural and functional analysis of the MALI Locus of Saccharomyces cerevisiae. Mol. Cell. Biol. 1986, 11, 3891–3899. [Google Scholar] [CrossRef]
- Charron, M.J.; Read, E.; Haut, S.R.; Michels, C.A. Molecular evolution of the telomere-associated MAL loci of Saccharomyces. Genetics 1989, 122, 307–316. [Google Scholar] [PubMed]
- Higgins, V.J.; Braidwood, M.; Bell, P.; Bissinger, P.; Dawes, I.W.; Attfield, P.V. Genetic evidence that high noninduced maltase and maltose permease activities, governed by MALx3-encoded transcriptional regulators, determine efficiency of gas production by baker’s yeast in unsugared dough. Appl. Environ. Microbiol. 1999, 65, 680–685. [Google Scholar] [PubMed]
- Novak, S.; Zechner-Krpan, V.; Marić, V. Regulation of maltose transport and metabolism in Saccharomyces cerevisiae. Food Technol. Biotechnol. 2004, 42, 213–218. [Google Scholar]
- Naumov, G.I.; Naumova, E.S.; Michels, C.A. Genetic variation of the repeated MAL loci in natural populations of Saccharomyces cerevisiae and Saccharomyces paradoxus. Genetics 1994, 136, 803–812. [Google Scholar] [PubMed]
- Bell, P.J.; Higgins, V.J.; Attefield, P.V. Comparison of fermentative capacities of industrial baking and wild-type yeasts of the species Saccharomyces cerevisiae in different sugar media. Lett. Appl. Microbiol. 2001, 32, 224–229. [Google Scholar] [CrossRef] [PubMed]
- Oda, Y.; Mikumo, D.; Tajima, K.; Yamauchi, H. Characterization of an alternative baking strain of Saccharomyces cerevisiae isolated from fermented cherry fruits by the analysis of SUC2 gene. Food Sci. Technol. Res. 2010, 16, 45–50. [Google Scholar] [CrossRef]
- Rincón, A.M.; Codón, A.C.; Castrejón, F.; Benítez, T. Improved properties of baker’s yeast mutants resistant to 2-deoxy-d-glucose. Appl. Environ. Microbiol. 2001, 67, 4279–4285. [Google Scholar] [CrossRef] [PubMed]
- Oda, Y.; Ouchi, K. Hybridization of bakers’ yeast by the rare-mating method to improve leavening ability in dough. Enzym. Microb. Technol. 1990, 12, 989–993. [Google Scholar] [CrossRef]
- Oda, Y.; Ouchi, K. Construction of a sucrose-fermenting bakers’ yeast incapable of hydrolysing fructooligosaccharides. Enzym. Microb. Technol. 1991, 13, 495–498. [Google Scholar] [CrossRef]
- Mikumo, D.; Takaya, M.; Orikasa, Y.; Ohwada, T. Improved leavening ability of a wild yeast, Saccharomyces cerevisiae AK46 2-deoxyglucose resistant mutant. Food Sci. Technol. Res. 2015, 21, 623–630. [Google Scholar] [CrossRef]
- Oda, Y.; Mikumo, D.; Leo, F.; Urashima, T. Discrimination of Saccharomyces cerevisiae and Saccharomyces paradoxus strains by the SUC2 gene sequences. J. Gen. Appl. Microbiol. 2010, 56, 355–358. [Google Scholar]
- Milne, I.; Bayer, M.; Cardle, L.; Shaw, P.; Stephen, G.; Wright, F.; Marshall, D. Tablet-next generation sequence assembly visualization. Bioinformatics 2009, 26, 401–402. [Google Scholar] [CrossRef] [PubMed]
- Untergasser, A.; Nijveen, H.; Rao, X.; Bisseling, T.; Geurts, R.; Leunissen, J.A. Primer3Plus, an enhanced web interface to Primer3. Nucleic Acids Res. 2007, 35, W71–W74. [Google Scholar] [CrossRef] [PubMed]
- Hohjoh, H. Genri Kara Yoku Wakaru Riaru Taimu PCR Kanzen Jikken Gaido, 1st ed.; Yodosha Co., Ltd.: Tokyo, Japan, 2007; ISBN 9784758101721. (In Japanese) [Google Scholar]
- Masubuchi, T.; Tomizawa, Y.; Ueda, R.; Takashima, C.; Hayashi, H.; Ikenaga, Y.; Satoh, K.; Teshima, K.; Narumi, I.; Wada, S.; et al. Development of a New Sake Yeast Screening Method by Gene Function Analysis; Gunma Industrial Technology Center: Maebashi, Japan, 2012; pp. 20–24. [Google Scholar]
- Zhang, C.Y.; Lin, X.; Song, H.Y.; Xiao, D.G. Effects of MAL61 and MAL62 overexpression on maltose fermentation of baker’s yeast in lean dough. World J. Microbiol. Biotechnol. 2015, 31, 1241–1249. [Google Scholar] [CrossRef] [PubMed]
- Milkowski, C.; Krampe, S.; Weirich, J.; Hasse, V.; Boles, E.; Breunig, K.D. Feedback regulation of glucose transporter gene transcription in Kluyveromyces lactis by glucose uptake. J. Bacteriol. 2001, 183, 5223–5229. [Google Scholar] [CrossRef] [PubMed]
- Dunnen, J.T.; Antonarakis, S.E. Mutation nomenclature extensions and suggestions to describe complex mutations: A discussion. Hum. Mutat. 2000, 15, 7–12. [Google Scholar] [CrossRef]
- Johnston, M.; Carlson, M. Regulation of carbon and phosphate utilization: The molecular and cellular biology of the yeast Saccharomyces. In The Molecular and Cellular Biology of the Yeast Saccharomyces: Gene Expression; Jones, E.W., Pringle, J.R., Broach, J.R., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 1992; Volume II, pp. 193–281. [Google Scholar]
- Hong, S.H.; Marmuri, J. Upstream regulatory regions controlling the expression of the yeast maltase gene. Mol. Cell. Biol. 1987, 7, 2477–2483. [Google Scholar] [CrossRef] [PubMed]
- Laity, J.H.; Lee, B.M.; Wright, P.E. Zinc finger proteins: New insights into structural and functional diversity. Curr. Opin. Struct. Biol. 2001, 11, 39–46. [Google Scholar] [CrossRef]
- Hu, Z.; Gibson, A.W.; Kim, J.H.; Wojciechowicz, L.A.; Zhang, B.; Michels, C.A. Functional domain analysis of the Saccharomyces MAL-activator. Curr. Genet. 1999, 36, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Gibson, A.W.; Wojciechowicz, L.A.; Danzi, S.E.; Zhang, B.; Kim, J.H.; Hu, Z.; Michels, C.A. Constitutive mutations of the Saccharomyces cerevisiae MAL-activator genes MAL23, MAL43, MAL63, and MAL64. Genetics 1997, 146, 1287–1298. [Google Scholar]
- Varanasi, U.S.; Klis, M.; Mikesell, P.B.; Trumbly, R.J. The Cyc8 (Ssn6)-Tup1 corepressor complex is composed of one Cyc8 and four Tup1 subunits. Mol. Cell. Biol. 1996, 16, 6707–6714. [Google Scholar] [CrossRef] [PubMed]
- Klein, C.J.; Olsson, L.; Rønnow, B.; Mikkelsen, J.D.; Nielsen, J. Alleviation of glucose repression of maltose metabolism by MIG1 disruption in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 1996, 62, 4441–4449. [Google Scholar] [PubMed]
- Treitel, M.A.; Carlson, M. Repression by SSN6-TUP1 is directed by MIG1, a repressor/activator protein. Proc. Natl. Acad. Sci. USA 1995, 92, 3132–3136. [Google Scholar] [CrossRef]
- Gancedo, J.M. Yeast carbon catabolite repression. Microbiol. Mol. Biol. Rev. 1998, 62, 334–361. [Google Scholar] [PubMed]
- Lin, X.; Zhang, C.Y.; Bai, X.W.; Liu, X.E.; Xiao, D.G. Effects of MIG1, TUP1 and SSN6 deletion on maltose metabolism and leavening ability of baker’s yeast in lean dough. Microb. Cell Fact. 2014, 13, 93–101. [Google Scholar] [CrossRef] [PubMed]
- Flick, J.S.; Johnston, M. GRR1 of Saccharomyces cerevisiae is required for glucose repression and encodes a protein with leucine-rich repeats. Mol. Cell. Biol. 1991, 11, 5101–5112. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Zhang, C.Y.; Bai, X.W.; Xiao, D.G. Effects of GLC7 and REG1 deletion on maltose metabolism and leavening ability of baker’s yeast in lean dough. J. Biotechnol. 2015, 209, 1–6. [Google Scholar] [CrossRef]
- Trumbly, R.J. Glucose repression in the yeast Saccharomyces cerevisiae. Mol. Microbiol. 1992, 6, 15–21. [Google Scholar] [CrossRef] [PubMed]

| Gene | Orientation | Sequence (5′→3′) |
|---|---|---|
| Maltose metabolism | ||
| MAL11 | Forward | GTCTTGGGTTAGCGGGTACA |
| Reverse | CAACTCCGCTGATGGAATTT | |
| MAL12 | Forward | AAGGTATCACTTCCAAGTTGC |
| Reverse | AGTCCTCATTGGTACCTATG | |
| MAL13 | Forward | GCAACCGTCGAGAAAAAGAG |
| Reverse | ATAGAGCCGCAAGCACTGAT | |
| MAL31 | Forward | GTAGCCATGGGGTTGTTTC |
| Reverse | CAGATCCACTGCAAAGCAAA | |
| MAL32 | Forward | ACATACGGTACCAACGAGGA |
| Reverse | GTTTGCGAGTCGTCAAGTTG | |
| MAL33 | Forward | ATGAAGTTGGAGGCTTGGAA |
| Reverse | ATCATTTAGGCGCAGTGGTC | |
| Catabolite repression | ||
| MIG1 | Forward | GGTTGTGGGCTCTCCAATAA |
| Reverse | CCATCGTTTTGGGAGAAGAA | |
| SNF1 | Forward | TACCACTACGGGCCAAAAAG |
| Reverse | CCCGGCGTACTCTATAACCA | |
| TUP1 | Forward | AAGGACGCGTACGAAGAAGA |
| Reverse | GCAACTGGAACAGATGCAGA | |
| CYC8 | Forward | GCCAAAGTTTTGGAATTGGA |
| Reverse | CATGCTCGTAGGCTTCCTTC | |
| Normalization | ||
| TDH1 | Forward | CTCTACCGGTGCTGCTAAGC |
| Reverse | AACGGCATCTTCGGTGTAAC |
| Protein | Description | Amino Acid substitutions * | Reference |
|---|---|---|---|
| Maltose metabolism | |||
| Mal11 | sugar transporter | H591L/D592I/S593R/I594X ** | [15] |
| Mal12 | α-glucosidase | none | [15] |
| Mal13 | activator |
T299I/T318A/S320X/N327Y/T330I,V/S333A/R336W/R337H /I341V/N361R/G362A/Q363H/I364V/R370S/E381D,K,N /D385E,G/V391I/V393A,I,M,T/T395A/L396I/I398V/T400N | [15] |
| Mal31 | sugar transporter |
H49R/A122S/S146P/Q166H/M175L/Q261T/A265P /E268N/E339K/T349S/V354L/G357S/I358V/C374I,S /S375T/A378T/S379P,Q,X/S394G/V508A,I,T/T509R,S /K526L/F534L/L536F/A540V/V544I | [15] |
| Mal32 | α-glucosidase | none | [15] |
| Mal33 | activator |
S240A/V243I/H244D,K,Q/Q257L/F260V/D269E/F272L /M274V/F286Y/E292V/K305R/K308N/A313T/L315H /E316D/I327F/F329C/S330F,L/H332P/A336T/F343L /Q344H/N346K/K365R/D369E,G/I371M,T/S390A/V393I /K403Q/Y404H/H406K,N,Q | [15] |
| Catabolite repression | |||
| Mig1 | transcription factor | none | [15] |
| Snf1 | carbon catabolite derepressing protein kinase | none | [15] |
| Tup1 | glucose repression regulatory protein | none | [15] |
| Cyc8 | general transcriptional co-repressor | Q26R,Q/Q28R,Q | This study |
| Grr1 | ubiquitin-ligase | N44N,-/N45N,-/N46N,- | This study |
| Reg1 | protein phosphatase | none | This study |
| Hxt2 | high-affinity glucose transporter | none | This study |
| Hxt4 | high-affinity glucose transporter | none | This study |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Orikasa, Y.; Mikumo, D.; Ohwada, T. A 2-Deoxyglucose-Resistant Mutant of Saccharomyces cerevisiae Shows Enhanced Maltose Fermentative Ability by the Activation of MAL Genes. Foods 2018, 7, 52. https://doi.org/10.3390/foods7040052
Orikasa Y, Mikumo D, Ohwada T. A 2-Deoxyglucose-Resistant Mutant of Saccharomyces cerevisiae Shows Enhanced Maltose Fermentative Ability by the Activation of MAL Genes. Foods. 2018; 7(4):52. https://doi.org/10.3390/foods7040052
Chicago/Turabian StyleOrikasa, Yoshitake, Dai Mikumo, and Takuji Ohwada. 2018. "A 2-Deoxyglucose-Resistant Mutant of Saccharomyces cerevisiae Shows Enhanced Maltose Fermentative Ability by the Activation of MAL Genes" Foods 7, no. 4: 52. https://doi.org/10.3390/foods7040052




