Novel Signal-Enhancing Approaches for Optical Detection of Nucleic Acids—Going beyond Target Amplification
Abstract
:1. Introduction
2. Enzyme Labeling Strategies without Polymerase Target Amplification
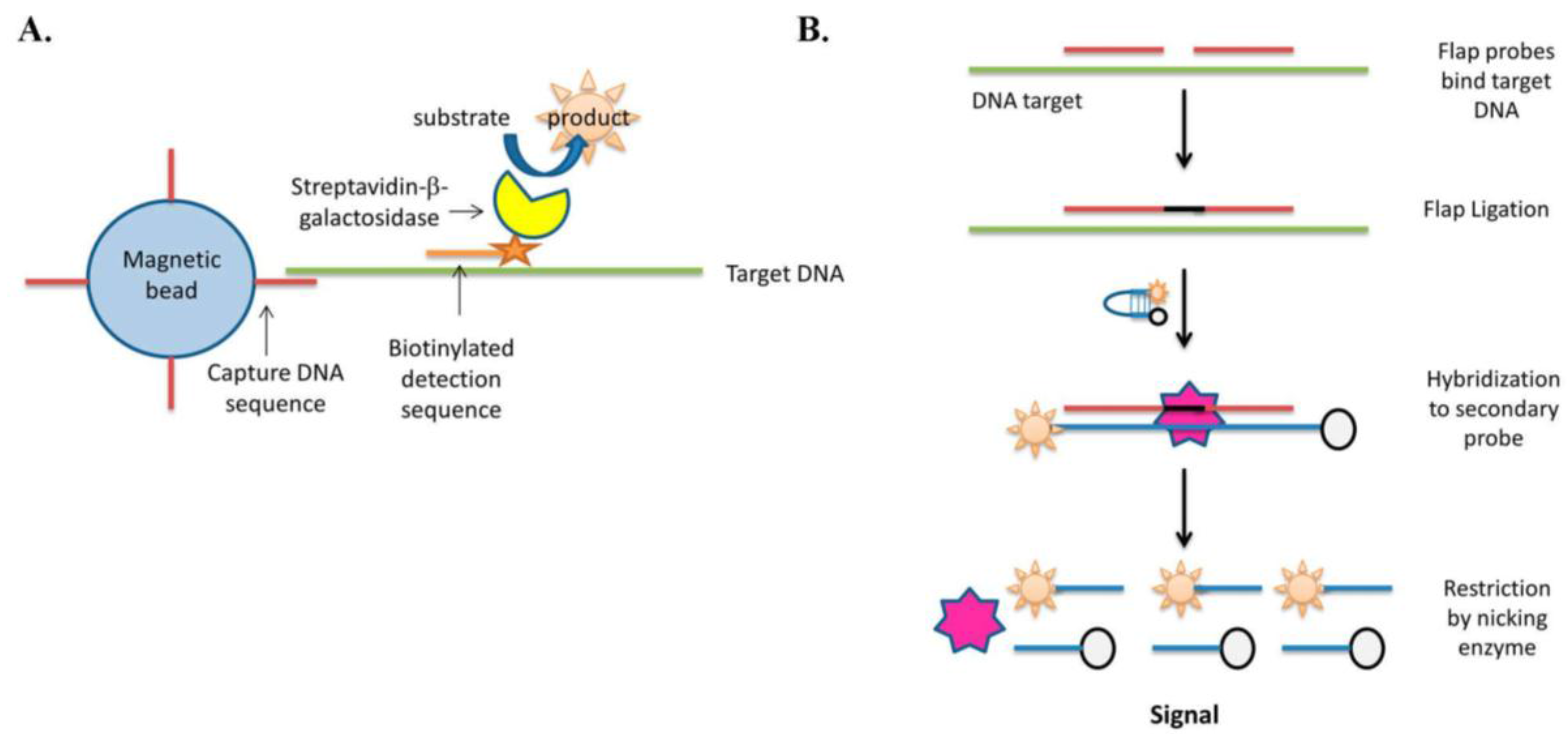
3. Enzyme-Free Hybridization Assays: Specificity in Focus
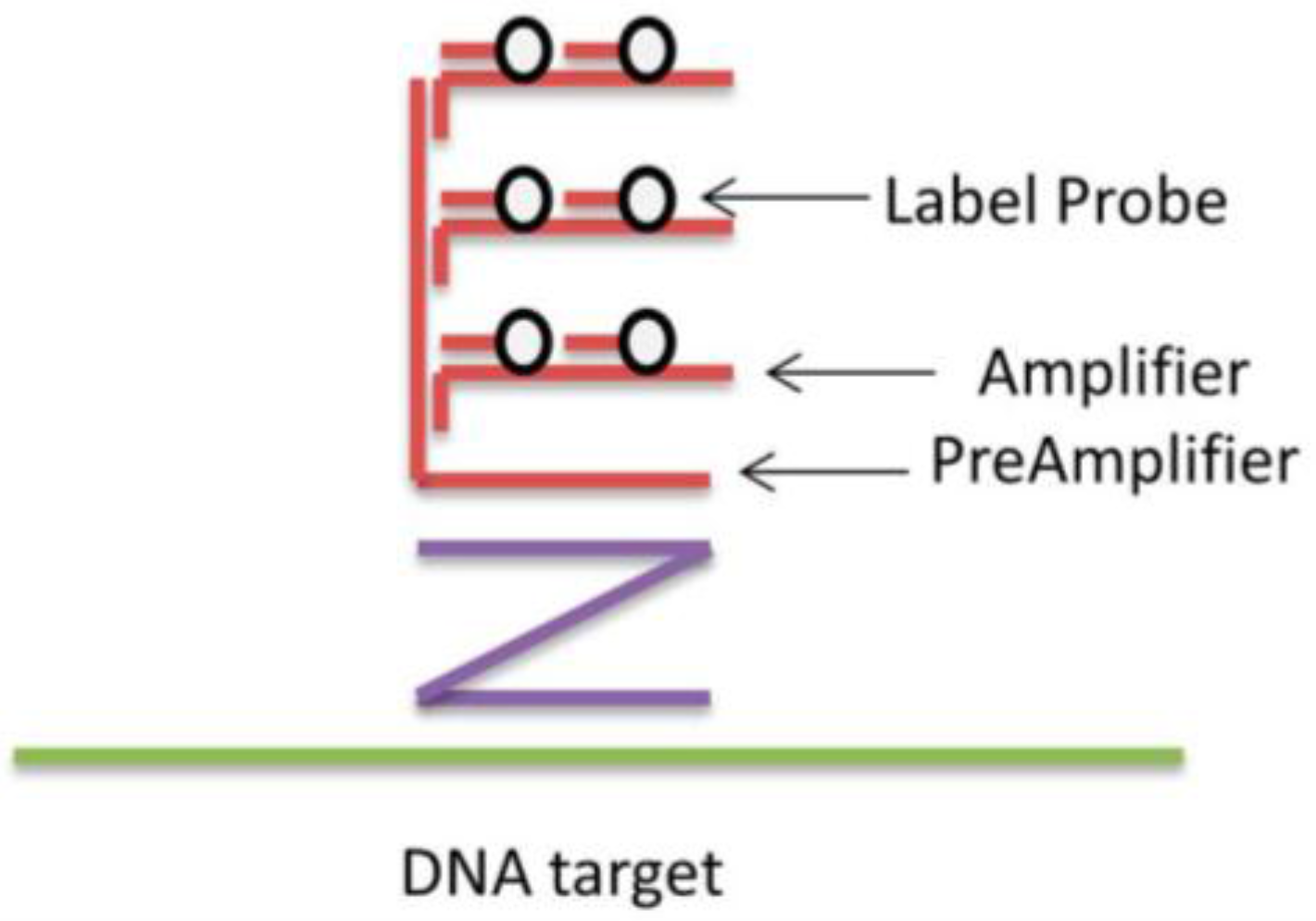
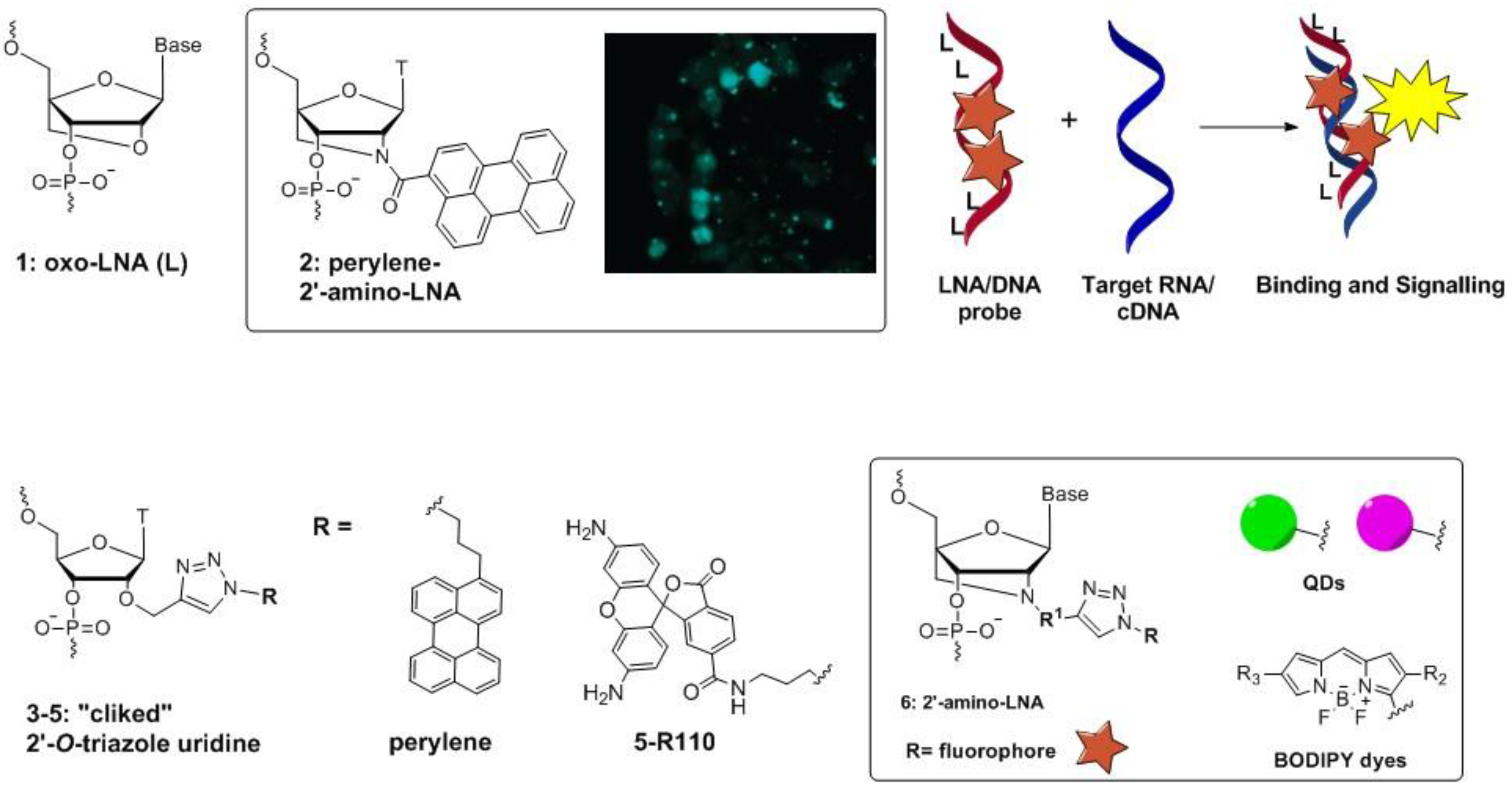
4. Ultra-Bright Fluorophores for Nucleic Acid Labeling: Macrofluorophores, Nucleic Acid Binding Dyes, and Nanoparticles

5. Signal Enhancement in Surface Plasmon Resonance Assays
6. Signal Enhancement with Optical Detectors
| Assay/Producer | Apparatus Specifications: | Compatible Labels; Absorbance-/Emission (abs-em) Range, nm; QY | LOD * | |||
|---|---|---|---|---|---|---|
| Light Source | Detector | Application | pmol | Ref. | ||
| Colorimetry: Konica Minolta/Hach | Filament lamp or LED | PR | Chromophore with abs at 380–780 nm | DNA | 5 × 10−3 | [50,57] |
| Fluorometry: Promega Glomax | Xe lamp or LED | PD/EPC + PMT | Fluorophore with abs-em at 260–750 nm, QY > 0.1 | Luciferase | 3 × 10−9 | [58] |
| Horiba spectrofluorometer | Xe lamp | PD/EPC + PMT | Fluorophore with abs-em at 185–1700 nm, QY > 0.1 | DNA | FLS: 9 × 10−4 LSC: 2 × 10−4 | [59] |
| TECAN multimode reader: Infinite F200 PRO | Xe lamp | PD/EPC + PMT | Fluorophore with abs-em at 230–1000 nm, QY > 0.1 | DNA | FLS: 3 × 10−4LSC: 1.2 × 10−5 | [60] |
| RT-qPCR: Roche Light Cycler 480 | HI LED | CCD | Fluorophore with abs-em at 488–660 nm, QY > 0.1 | DNA | 3 × 10−3 | [61] |
| Microscopy: Nikon/Leica/Olympus/Zeiss/Horiba | Lamp (Xe, MH)/LED | PD/EPC + PMT; CCD, CID, CMOS, EBCCD; GaAsP | Photostable fluorophores with abs-em at 240–940 nm, optimal at QY > 0.6 | DNA/RNA | <1 × 10−13 | [62] |
| Surface plasmon resonance: Biacore, BioNavis | Lamp, diode, laser or fiber | PD or CCD | n/a | DNA | 3 × 10−4 | [64] |
7. Conclusion and Perspective: Moving toward Sensitive, Specific and Detailed Nucleic Acid Analysis Using Non-PCR Assays
Acknowledgments
Conflicts of Interest
References
- Spengler, M.; Adler, M.; Niemeyer, C.M. Highly sensitive ligand-binding assays in pre-clinical and clinical applications: Immuno-PCR and other emerging techniques. Analyst 2015. [Google Scholar] [CrossRef] [PubMed]
- Jin, D.J.; Cagliero, C.; Martin, C.M.; Izard, J.; Zhou, Y.N. The dynamic nature and territory of transcriptional machinery in the bacterial chromosome. Front. Microbiol. 2015, 6, 1–14. [Google Scholar]
- The Cancer Genome Atlas Research Network. Comprehensive genomic characterization of squamous cell lung cancers. Nature 2012, 489, 519–525. [Google Scholar]
- Astakhova, K. Toward Non-Enzymatic Ultrasensitive Identification of Single Nucleotide Polymorphisms by Optical Methods. Chemosensors 2014, 2, 193–206. [Google Scholar] [CrossRef]
- Hu, R.; Liu, T.; Zhang, X.B.; Yang, Y.; Chen, T.; Wu, C.; Liu, Y.; Zhu, G.; Huan, S.; Fu, T.; et al. DLISA: A DNAzyme-based ELISA for protein enzyme-free immunoassay of multiple analytes. Anal. Chem. 2015, 87, 7746–7753. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Shan, D.; Zhao, M.; Pink, B.A.; Minnehan, K.A.; York, L.; Gardel, M.; Sullivan, S.; Phillips, A.F.; Hayman, R.B.; Walt, D.R.; Duffy, D.C. Direct detection of bacterial genomic DNA at sub-femtomolar concentrations using single molecule arrays. Anal. Chem. 2013, 85, 1932–1939. [Google Scholar] [CrossRef] [PubMed]
- Wabuyele, M.B.; Farquar, H.; Stryjewski, W.; Hammer, R.P.; Soper, S.A.; Cheng, Y.W.; Barany, F. Approaching real-time molecular diagnostics: Single-pair fluorescence resonance energy transfer (spFRET) detection for the analysis of low abundant point mutations in K-ras oncogenes. J. Am. Chem. Soc. 2003, 125, 6937–6945. [Google Scholar] [CrossRef] [PubMed]
- Kosman, J.; Juskowiak, B. Peroxidase-mimicking DNAzymes for biosensing applications: A review. Anal. Chim. Acta 2011, 707, 7–17. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Li, H.; Deng, M.; Weng, X.; Ma, H.; Feng, S.; Zhou, Y.; Zhou, X. Studies of the activity of peroxidase-like DNAzyme by modifying 3′- or 5′-end of aptamers. Chem. Biodivers. 2012, 9, 170–180. [Google Scholar] [CrossRef] [PubMed]
- Kong, D.M.; Cai, L.L.; Guo, J.H.; Wu, J.; Shen, H.X. Characterization of the G-quadruplex structure of a catalytic DNA with peroxidase activity. Biopolymers 2009, 91, 331–339. [Google Scholar] [CrossRef] [PubMed]
- Spittle, C.; Ward, M.R.; Nathanson, K.L.; Gimotty, P.A.; Rappaport, E.; Brose, M.S.; Medina, A.; Letrero, R.; Herlyn, M.; Edwards, R.H. Application of a BRAF pyrosequencing assay for mutation detection and copy number analysis in malignant melanoma. J. Mol. Diagn. 2007, 9, 464–471. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Wang, X.; Zhao, C.; Qu, K.; Ren, J.; Qu, X. Label-free colorimetric detection of single nucleotide polymorphism by using single-walled carbon nanotube intrinsic peroxidase-like activity. Chem. Eur. J. 2010, 16, 3617–3621. [Google Scholar] [CrossRef] [PubMed]
- Dirks, R.M.; Pierce, N.A. Triggered amplification by hybridization chain reaction. Proc. Natl. Acad. Sci. USA 2004, 101, 15275–15278. [Google Scholar] [CrossRef] [PubMed]
- Porichis, F.; Hart, M.G.; Griesbeck, M.; Everett, H.L.; Hassan, M.; Baxter, A.E.; Lindqvist, M.; Miller, S.M.; Soghoian, D.Z.; Kavanagh, D.G.; et al. High-throughput detection of miRNAs and gene-specific mRNA at the single-cell level by flow cytometry. Nat. Commun. 2014, 5, 5641–5654. [Google Scholar] [CrossRef] [PubMed]
- Bishop, R. Applications of fluorescence in situ hybridization (FISH) in detecting genetic aberrations of medical significance. Biosci. Horiz. 2010, 3, 85–92. [Google Scholar] [CrossRef]
- Bystricky, K. Chromosome dynamics and folding in eukaryotes: Insights from live cell microscopy. FEBS Lett. 2015. [Google Scholar] [CrossRef] [PubMed]
- Bowling, A.J.; Pence, H.E.; Church, J.B. Application of a novel and automated branched DNA in situ hybridization method for the rapid and sensitive localization of mRNA molecules in plant tissues. Appl. Plant Sci. 2014, 2. [Google Scholar] [CrossRef] [PubMed]
- Baumeister, M.A.; Zhang, N.; Beas, H.; Brooks, J.R.; Canchola, J.A.; Cosenza, C.; Kleshik, F.; Rampersad, V.; Surtihadi, J.; Battersby, T. A sensitive branched DNA HIV-1 signal amplification viral load assay with single day turnaround. PLoS ONE 2012, 7, e33295. [Google Scholar] [CrossRef] [PubMed]
- Patel, J.A.; Anderson, E.; Dong, J. False Positive Ultrasensitive HIV bDNA Viral load results in diagnosis of perinatal HIV-Infection in the era of low transmission. Lab. Med. 2009, 40, 611–614. [Google Scholar] [CrossRef]
- Briley, W.E.; Bondy, M.H.; Randeria, P.S.; Dupperc, T.J.; Mirkin, C.A. Quantification and real-time tracking of RNA in live cells using Sticky-flares. Proc. Natl. Acad. Sci. USA 2015, 112, 9591–9595. [Google Scholar] [CrossRef] [PubMed]
- Rino, J.; Martin, R.M.; Carvalho, C.; de Jesus, A.C.; Carmo-Fonseca, M. Single-Molecule Imaging of RNA Splicing in Live Cells. Method Enzymol. 2015, 558, 571–585. [Google Scholar]
- Viswanathan, S.; Williams, M.E.; Bloss, E.B.; Stasevich, T.J.; Speer, C. M.; Nern, A.; Pfeiffer, B.D.; Hooks, B.M.; Li, W.P.; English, B.P.; et al. High-performance probes for light and electron microscopy. Nat. Methods 2015, 12, 568–576. [Google Scholar] [CrossRef] [PubMed]
- Gerling, T.; Wagenbauer, K.F.; Neuner, A.M.; Dietz, H. Dynamic DNA devices and assemblies formed by shape-complementary, non-base pairing 3D components. Science 2015, 347, 1446–1452. [Google Scholar] [CrossRef] [PubMed]
- Astakhova, I.K.; Samokhina, E.; Babu, B.R.; Wengel, J. Novel (Phenylethynyl)pyrene–LNA Constructs for Fluorescence SNP Sensing in Polymorphic Nucleic Acid Targets. ChemBioChem 2012, 13, 1509–1519. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Astakhova, I.K.; Korshun, V.A.; Jahn, K.; Kjems, J.; Wengel, J. Perylene Attached to 2′-Amino-LNA: Synthesis, Incorporation into Oligonucleotides, and Remarkable Fluorescence Properties in vitro and in Cell Culture. Bioconjug. Chem. 2008, 19, 1995–2007. [Google Scholar] [CrossRef] [PubMed]
- Okholm, A.; Kjemsa, J.; Astakhova, K. Fluorescence detection of natural RNA using rationally designed “clickable” oligonucleotide probes. RSC Adv. 2014, 4, 45653–45656. [Google Scholar] [CrossRef]
- Jørgensen, A.S.; Gupta, P.; Wengel, J.; Astakhova, I.K. “Clickable” LNA/DNA probes for fluorescence sensing of nucleic acids and autoimmune antibodies. Chem. Commun. 2013, 49, 10751–10753. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dziuba, D.; Pohl, R.; Hocek, M. Bodipy-labeled nucleoside triphosphates for polymerase synthesis of fluorescent DNA. Bioconjug. Chem. 2014, 25, 1984–1995. [Google Scholar] [CrossRef] [PubMed]
- Yan, Z.; Gan, N.; Zhang, H.; Wang, D.; Qiao, L.; Cao, Y.; Li, T.; Hu, F. A sandwich-hybridization assay for simultaneous determination of HIV and tuberculosis DNA targets based on signal amplification by quantum dots-PowerVision (™) polymer coding nanotracers. Biosens. Bioelectron. 2015, 71, 207–213. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Z.; Chen, Y.C.; Li, H.W.; Chang, H.T. Fluorescent silver nanoclusters stabilized by DNA scaffolds. Chem. Commun. 2014, 50, 9800–9815. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; You, J.; Zhuang, Y.; Han, C.; Hu, J.; Wang, A.; Xu, K.; Zhu, J.J. A “light-up” and “spectrum-shift” response of aptamer-functionalized silver nanoclusters for intracellular mRNA imaging. Chem. Commun. 2014, 50, 7107–7110. [Google Scholar] [CrossRef] [PubMed]
- Dziuba, D.; Pohl, R.; Hocek, M. Polymerase synthesis of DNA labelled with benzylidene cyanoacetamide-based fluorescent molecular rotors: Fluorescent light-up probes for DNA-binding proteins. Chem. Commun. 2015, 51, 4880–4882. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, A.; Saito, M.; Katoh, R.; Saito, Y. Synthesis of 8-aza-3,7-dideaza-2′-deoxyadenosines possessing a new adenosine skeleton as an environmentally sensitive fluorescent nucleoside for monitoring the DNA minor groove. Org. Biomol. Chem. 2015, 13, 7459–7468. [Google Scholar] [CrossRef] [PubMed]
- Kanamori, T.; Ohzeki, H.; Masaki, Y.; Ohkubo, A.; Takahashi, M.; Tsuda, K.; Ito, T.; Shirouzu, M.; Kuwasako, K.; Muto, Y.; et al. Controlling the fluorescence of benzofuran-modified uracil residues in oligonucleotides by triple-helix formation. ChemBioChem 2015, 16, 167–176. [Google Scholar] [CrossRef] [PubMed]
- Hopkins, P.A.; Sinkeldam, R.W.; Tor, Y. Visibly emissive and responsive extended 6-aza-uridines. Org. Lett. 2014, 16, 5290–5293. [Google Scholar] [CrossRef] [PubMed]
- Riedl, J.; Ménová, P.; Pohl, R.; Orság, P.; Fojta, M.; Hocek, M. GFP-like fluorophores as DNA labels for studying DNA-protein interactions. J. Org. Chem. 2012, 77, 8287–8293. [Google Scholar] [CrossRef] [PubMed]
- Çamli, T.; Tuncel, M.; Şenel, S.; Tuncel, A. Functional, uniform and macroporous latex particles: Preparation, electron microscopic characterisation, and nonspecific protein adsorption properties. J. Appl. Polym. Sci. 2002, 84, 414–429. [Google Scholar] [CrossRef]
- Zhu, X.; Li, J.; He, H.; Huang, M.; Zhang, X.; Wang, S. Application of nanomaterials in the bioanalytical detection of disease-related genes. Biosens. Bioelectron. 2015, 74, 113–133. [Google Scholar] [CrossRef] [PubMed]
- Introduction to Signal Amplification—Section 6.1. Available online: https://www.lifetechnologies.com/dk/en/home/references/molecular-probes-the-handbook/ultrasensitive-detection-technology/introduction-to-detection-methods.html (accessed on 5 August 2015).
- Ihara, T.; Tanaka, S.; Chikaura, Y.; Jyo, A. Preparation of DNA-modified nanoparticles and preliminary study for colorimetric SNP analysis using their selective aggregations. Nucleic Acids Res. 2004, 32, e105. [Google Scholar] [CrossRef] [PubMed]
- Thomson, D.A.C.; Tee, E.H.-L.; Tran, N.T.D.; Monteiro, M.J.; Cooper, M.A. Oligonucleotide and polymer functionalized nanoparticles for amplification-free detection of DNA. Biomacromolecules 2012, 13, 1981–1989. [Google Scholar] [CrossRef] [PubMed]
- Mao, F.; Leung, W.Y.; Xin, X. Characterization of EvaGreen and the implication of its physicochemical properties for qPCR applications. BMC Biotechnol. 2007, 7, 76. [Google Scholar] [CrossRef] [PubMed]
- Nafisi, S.; Saboury, A.A.; Keramat, N.; Neault, J.F.; Tajmir-Riahi, H.A. Stability and structural features of DNA intercalation with ethidium bromide, acridine orange and methylene blue. J. Mol. Struct. 2007, 827, 35–43. [Google Scholar] [CrossRef]
- QuantiFluor® dsDNA System. Available online: https://dk.promega.com/products/dna-and-rna-purification/dna-and-rna-quantitation/sensitive-easy-to-use-quantifluor-dsdna-quantitation/quantifluor-dsdna-system/ (accessed on 5 August 2015).
- Bhunia, S.K.; Saha, A.; Maity, A.R.; Ray, S.C.; Jana, N.R. Carbon nanoparticle-based fluorescent bioimaging probes. Sci. Rep. 2013. [Google Scholar] [CrossRef] [PubMed]
- Misra, S.K.; Ohoka, A.; Kolmodin, N.J.; Pan, D. Next generation carbon nanoparticles for efficient gene therapy. Mol. Pharm. 2015, 12, 375–385. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Cheng, Y.; Wang, H.; Li, Z. A homogeneous fluorescence sensing platform with water-soluble carbon nanoparticles for detection of microRNA and nuclease activity. Analyst 2012, 137, 3667–3672. [Google Scholar] [CrossRef] [PubMed]
- Mirkin, C.A.; Letsinger, R.L.; Mucic, R.C.; Storhoff, J.J. A DNA-based method for rationally assembling nanoparticles into macroscopic materials. Nature 1996, 382, 607–609. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.H.; Asami, Y.; Murata, M.; Kitazaki, H.; Sadanaga, N.; Tokunaga, E.; Shiotani, S.; Okada, S.; Maehara, Y.; Niidome, T.; et al. Gold nanoparticle-based colorimetric assay for cancer diagnosis. Biosens. Bioelectron. 2010, 25, 1869–1874. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Yang, X.; Sun, S.; Wang, Q.; Wang, K.; Huang, J.; Liu, J.; He, L. Enzyme-Free Colorimetric Detection of DNA by Using Gold Nanoparticles and Hybridization Chain Reaction Amplification. Anal. Chem. 2013, 85, 7689–7695. [Google Scholar] [CrossRef] [PubMed]
- Su, X.; Teh, H.F.; Aung, K.M.; Zong, Y.; Gao, Z. Femtomol SPR detection of DNA-PNA hybridization with the assistance of DNA-guided polyaniline deposition. Biosens. Bioelectron. 2008, 23, 1715–1720. [Google Scholar] [CrossRef] [PubMed]
- Cao, D.; Huang, H.; Hu, M.; Cui, L.; Geng, F.; Rao, Z.; Niu, H.; Cai, Y.; Kang, Y. Comprehensive characterization of natural organic matter by MALDI- and ESI-Fourier transform ion cyclotron resonance mass spectrometry. Anal. Chim. Acta 2015, 866, 48–58. [Google Scholar] [CrossRef] [PubMed]
- Halpern, A.R.; Wood, J.B.; Wang, Y.; Corn, R.M. Previous Article Next Article Table of Contents Single-Nanoparticle Near-Infrared Surface Plasmon Resonance Microscopy for Real-Time Measurements of DNA Hybridization Adsorption. ACS Nano 2014, 8, 1022–1030. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Wang, Y.; Wang, L.; Wei, Q. A surface plasmon resonance assay coupled with a hybridization chain reaction for amplified detection of DNA and small molecules. Chem. Commun. 2014, 50, 5049–5052. [Google Scholar] [CrossRef] [PubMed]
- Qiu, X.; Liu, X.; Zhang, W.; Zhang, H.; Jiang, T.; Fan, D.; Luo, Y. Dynamic Monitoring of MicroRNA-DNA Hybridization Using DNAase-Triggered Signal Amplification. Anal. Chem. 2015, 87, 6303–6310. [Google Scholar] [CrossRef] [PubMed]
- Zagorodko, O.; Spadavecchia, J.; Serrano, A.Y.; Larroulet, I.; Pesquera, A.; Zurutuza, A.; Boukherroub, R.; Szunerits, S. Highly Sensitive Detection of DNA Hybridization on Commercialized Graphene-Coated Surface Plasmon Resonance Interfaces. Anal. Chem. 2014, 86, 11211–11216. [Google Scholar] [CrossRef] [PubMed]
- Konica Minolta. Available online: http://www.konicaminolta.com/instruments/ (accessed on 5 August 2015).
- GloMax®-Multi Detection System. Available online: https://dk.promega.com/products/instruments/multimode-readers/glomax_multi-detection-system/ (accessed on 5 August 2015).
- Fluoromax-4. Available online: http://www.horiba.com/fileadmin/uploads/Scientific/Documents/Fluorescence/Fluoromax-4.pdf (accessed on 5 August 2015).
- TECAN. Available online: http://www.tecan.com/page/content/index.asp?MenuID=23&ID=10&Menu=1&Item=21.2 (accessed on 5 August 2015).
- LightCycler® 480 System. Available online: http://www.roche.com/products/product-details.htm?type=product&id=64 (accessed on 5 August 2015).
- Murphy, D.B.; Davidson, M.W. Fundamentals of Light Microscopy and Electronic Imaging, 1st ed.; Wiley-Liss: New York, NY, USA, 2001. [Google Scholar]
- Microscopy Resource Center. Available online: http://www.olympusmicro.com/primer/techniques/fluorescence/fluorointrohome.html (accessed on 5 August 2015).
- Homola, J. Surface plasmon resonance sensors for detection of chemical and biological species. Chem. Rev. 2008, 108, 462–493. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.Y.; Chen, S.X.; Yin, P. Optimizing the specificity of nucleic acid hybridization. Nat. Chem. 2012, 4, 208–214. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.S.; Zhang, D.Y. Simulation-guided DNA probe design for consistently ultraspecific hybridization. Nat. Chem. 2015, 7, 545–553. [Google Scholar] [CrossRef] [PubMed]
- Ramos, E.; Levinson, B.T.; Chasnoff, S.; Hughes, A.; Young, A.L.; Thornton, K.; Li, A.; Vallania, F.L.M.; Province, M.; Druley, T.E. Population-based rare variant detection via pooled exome or custom hybridization capture with or without individual indexing. BMC Genomics 2012. [Google Scholar] [CrossRef] [PubMed]
- Cui, M.; Fan, M.; Jing, R.; Wang, H.; Qin, J.; Sheng, H.; Wang, Y.; Wu, X.; Zhang, L.; Zhu, J.; et al. Cell-Free Circulating DNA: A New Biomarker for the Acute Coronary Syndrome. Cardiology 2013, 124, 76–84. [Google Scholar] [CrossRef] [PubMed]
- Choi, N.W.; Kim, J.; Chapin, S.C.; Duong, T.; Donohue, E.; Pandey, P.; Broom, W.; Hill, W.A.; Doyle, P.S. Multiplexed Detection of mRNA Using Porosity-Tuned Hydrogel Microparticles. Anal. Chem. 2012, 84, 9370–9378. [Google Scholar] [CrossRef] [PubMed]
- Blin, A.; Cissé, I.; Bockelmann, U. Electronic hybridization detection in microarray format and DNA genotyping. Sci. Rep. 2014. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Miotke, L.; Barducci, M.C.; Astakhova, K. Novel Signal-Enhancing Approaches for Optical Detection of Nucleic Acids—Going beyond Target Amplification. Chemosensors 2015, 3, 224-240. https://doi.org/10.3390/chemosensors3030224
Miotke L, Barducci MC, Astakhova K. Novel Signal-Enhancing Approaches for Optical Detection of Nucleic Acids—Going beyond Target Amplification. Chemosensors. 2015; 3(3):224-240. https://doi.org/10.3390/chemosensors3030224
Chicago/Turabian StyleMiotke, Laura, Maria Carla Barducci, and Kira Astakhova. 2015. "Novel Signal-Enhancing Approaches for Optical Detection of Nucleic Acids—Going beyond Target Amplification" Chemosensors 3, no. 3: 224-240. https://doi.org/10.3390/chemosensors3030224





