Discriminating Bacteria with Optical Sensors Based on Functionalized Nanoporous Xerogels
Abstract
:1. Introduction
2. Experimental Section
2.1. Synthesis of Nanoporous Pastilles Doped with DMACA
2.2. Characterization of the Xerogel Pastilles
2.3. Detection of Bacterial Indole
3. Results and Discussion
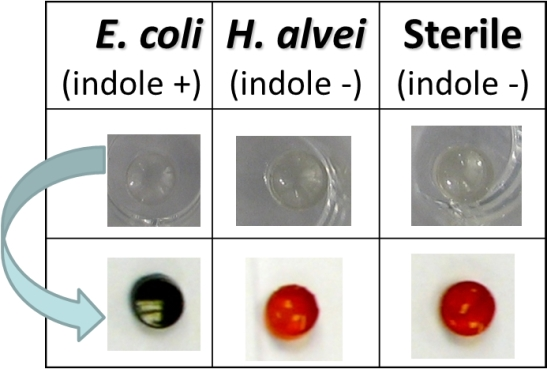
3.1. Probe Choice and Interferences
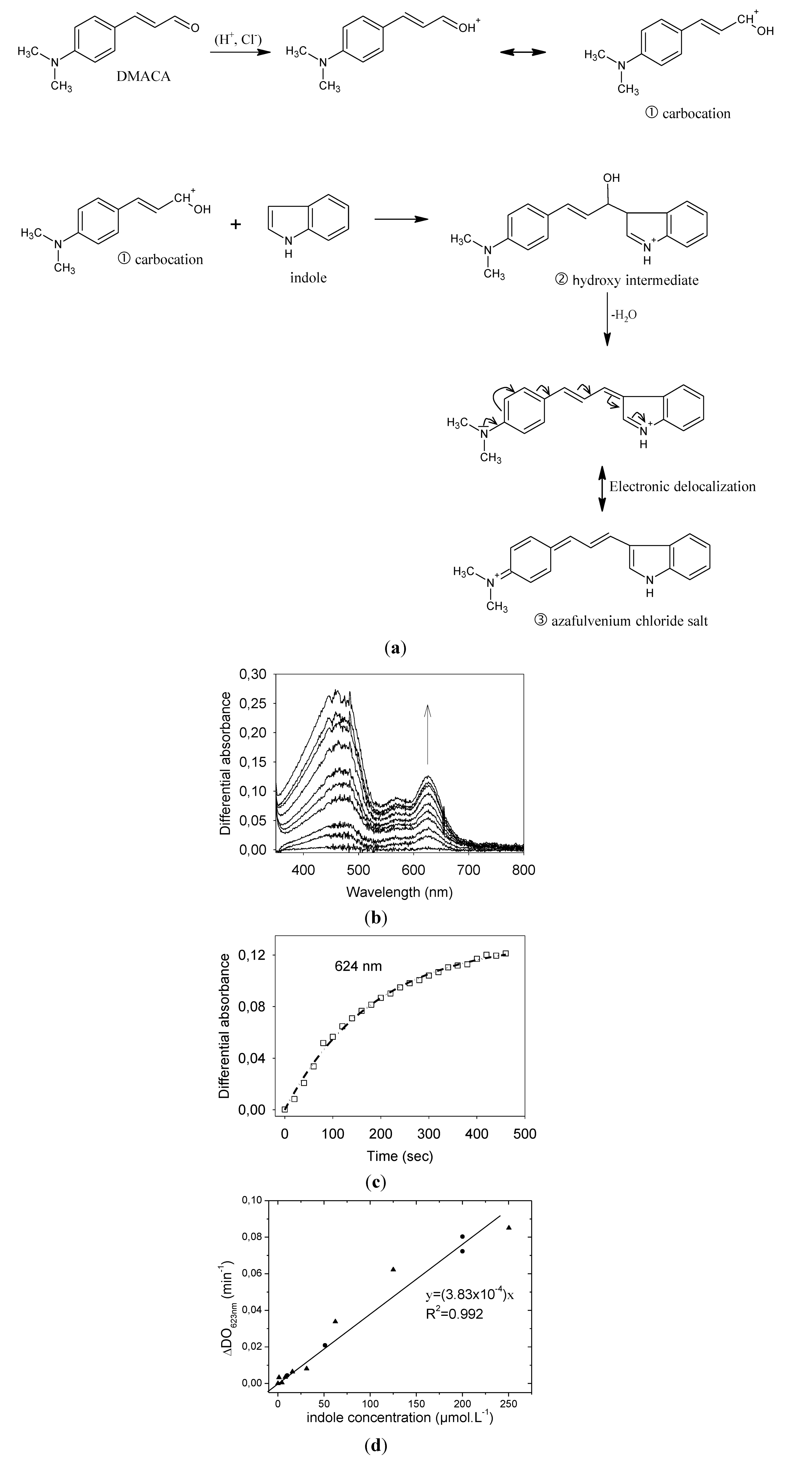
3.2. Preliminary Solution-phase Study of Indole Detection
3.3. Detection of Bacterial Indole in Liquid Phase

3.4. Detection of Bacterial Indole in the Gas Phase
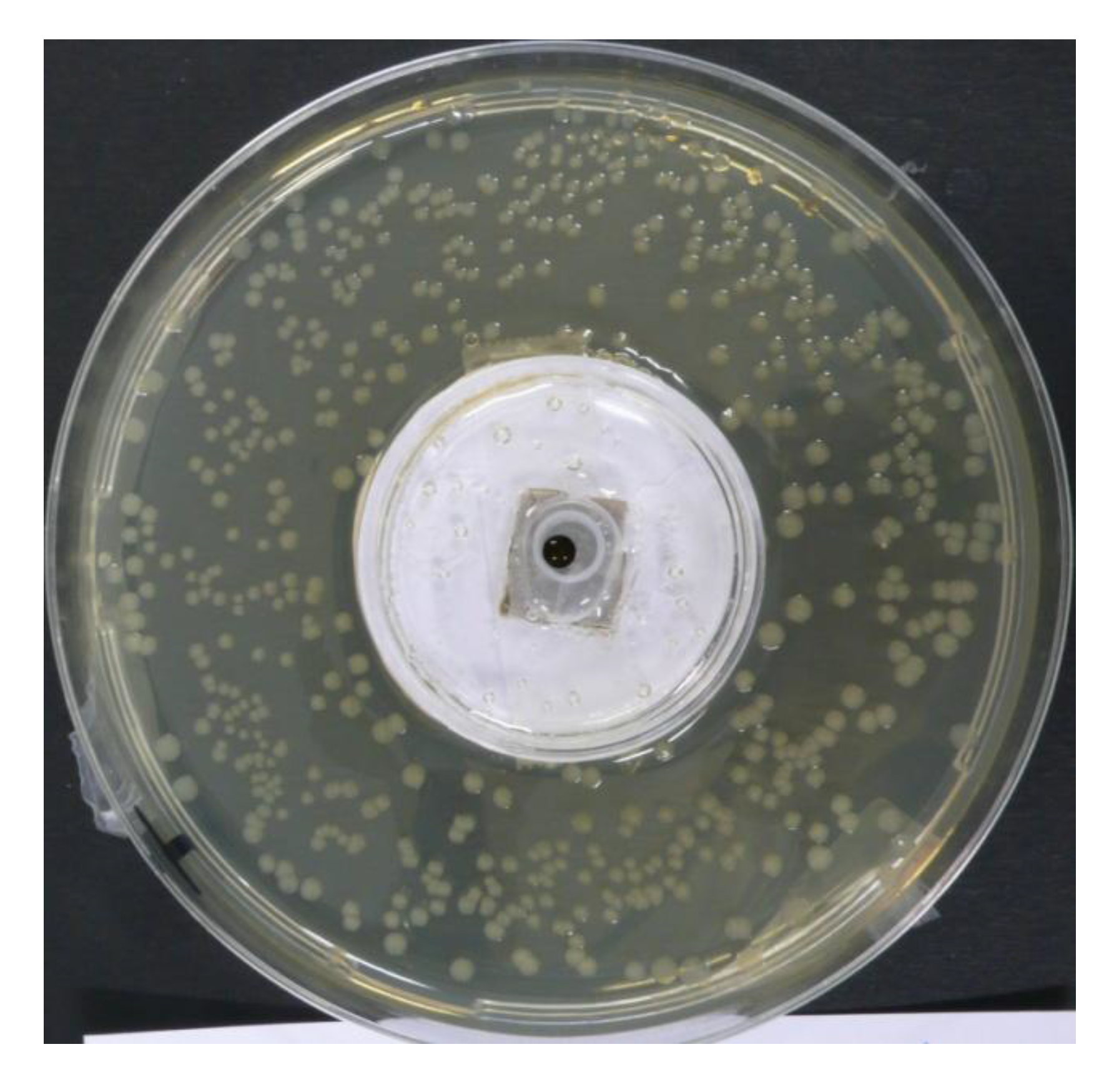
| Escherichia coli ATCC 11775 | Hafnia alvei ATCC 13337 | Control (no bacteria) | |
|---|---|---|---|
| tryptophan agar medium |  |  |  |
| LB agar medium |  |  |  |
4. Conclusions
Acknowledgments
Conflicts of Interest
References
- Fox, A. Mass spectrometry for species or strain identification after culture or without culture: Past, present and future. J. Clin. Microbiol. 2006, 44, 2677–2680. [Google Scholar] [CrossRef]
- Ammor, M.S. Recent advances in the use of intrinsic fluorescence for bacterial identification and characterization. J. Fluoresc. 2007, 17, 455–459. [Google Scholar] [CrossRef]
- Oust, A.; Moretro, T.; Kirschner, C.; Narvhus, J.A.; Kohler, A. FT-IR spectroscopy for identification of closely related lactobacilli. J. Microbiol. Methods 2004, 59, 149–162. [Google Scholar] [CrossRef]
- Holst, E.; Larsson, L. Gas-chromatographic comparison of peptone yeast glucose and gas-liquid-chromatography growth media for anaerobic bacteria. Eur. J. Clin. Microbiol. 1987, 6, 724–728. [Google Scholar] [CrossRef]
- Tait, E.; Perry, J.D.; Stanforth, S.P.; Dean, J.R. Identification of volatile organic compounds produced by bacteria using HS-SPME-GC-MS. J. Chromatogr. Sci. 2014, 52, 363–373. [Google Scholar] [CrossRef]
- Bunge, M.; Araghipour, N.; Mikoviny, T.; Dunkl, J.; Schnitzhofer, R.; Hansel, A.; Schinner, F.; Wisthaler, A.; Margesin, R.; Märk, T.D. On-line monitoring of microbial volatile metabolites by proton transfer reaction-mass spectrometry. Appl. Environ. Microbiol. 2008, 74, 2179–2186. [Google Scholar] [CrossRef]
- Allardyce, R.A.; Hill, A.L.; Murdoch, D.R. The rapid evaluation of bacterial growth and antibiotic susceptibility in blood cultures by selected ion flow tube mass spectrometry. Diagn. Microbiol. Infect. Dis. 2006, 55, 255–261. [Google Scholar] [CrossRef]
- Allardyce, R.A.; Langford, V.S.; Hill, A.L.; Murdoch, D.R. Detection of volatile metabolites produced by bacterial growth in blood culture media by selected ion flow tube mass spectrometry (SIFT-MS). J. Microbiol. Methods 2006, 65, 361–365. [Google Scholar] [CrossRef]
- Scotter, J.M.; Langford, V.S.; Wilson, P.F.; McEwan, M.J.; Chambers, S.T. Real-time detection of common microbial volatile organic compounds from medically important fungi by Selected Ion Flow Tube-Mass Spectrometry (SIFT-MS). J. Microbiol. Methods 2005, 63, 127–134. [Google Scholar] [CrossRef]
- Scotter, J.M.; Allardyce, R.A.; Langford, V.S.; Hill, A.; Murdoch, D.R. The rapid evaluation of bacterial growth in blood cultures by selected ion flow tube-mass spectrometry (SIFT-MS) and comparison with the BacT/ALERT automated blood culture system. J. Microbiol. Methods 2006, 65, 628–631. [Google Scholar] [CrossRef]
- Siripatrawan, U. Rapid differentiation between E. coli and Salmonella typhimurium using metal oxide sensors integrated with pattern recognition. Sens. Actuators B 2008, 133, 414–419. [Google Scholar] [CrossRef]
- Carey, J.R.; Suslick, K.S.; Hulkower, K.I.; Imlay, J.A.; Imlay, K.R.C.; Ingison, C.K.; Ponder, J.B.; Sen, A.; Wittrig, A.E. Rapid identification of bacteria with a disposable colorimetric sensing array. J. Am. Chem. Soc. 2011, 133, 7571–7576. [Google Scholar] [CrossRef]
- McDonald, J.G.; Borders, R.A. Technique for Detecting Microorganisms US Pat. U.S. Patent 2006/0223052 A1, 2006. [Google Scholar]
- Lee, J.-H.; Lee, J. Indole as an intercellular signal in microbial communities. FEMS Microbiol. Rev. 2010, 34, 426–444. [Google Scholar]
- Lee, H.H.; Molla, M.N.; Cantor, C.R.; Collins, J.J. Bacterial charity work leads to population-wide resistance. Nature 2010, 467, 82–86. [Google Scholar] [CrossRef]
- Di Martino, P.; Fursy, R.; Bret, L.; Sundararaju, B.; Phillips, R.S. Indole can act as an extracellular signal to regulate biofilm formation of Escherichia coli and other indole-producing bacteria. Can. J. Microbiol. 2003, 49, 443–449. [Google Scholar] [CrossRef]
- Han, T.H.; Lee, J.-H.; Cho, M.H.; Wood, T.K.; Lee, J. Environmental factors affecting indole production in Escherichia coli. Res. Microbiol. 2011, 162, 108–116. [Google Scholar] [CrossRef]
- Ahmad, A.; Qureshi, M.; Khan, I.A.; Alam, K.Z. Spectrophotometric determination of diphenylamine, pyrrole, and indole—A kinetic study. Anal. Lett. 1990, 23, 1139–1157. [Google Scholar]
- Lowrance, B.L.; Reich, P.; Traub, W.H. Evaluation of two spot-indole reagents. Appl. Microbiol. 1969, 17, 923–924. [Google Scholar]
- Pastan, I.; Perlman, R.L. Stimulation of Tryptophanase Synthesis in Escherichia coli by Cyclic 3',5'-Adenosine Monophosphate. J. Biol. Chem. 1969, 244, 2226–2232. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Crunaire, S.; Marcoux, P.R.; Ngo, K.-Q.; Moy, J.-P.; Mallard, F.; Tran-Thi, T.-H. Discriminating Bacteria with Optical Sensors Based on Functionalized Nanoporous Xerogels. Chemosensors 2014, 2, 171-181. https://doi.org/10.3390/chemosensors2020171
Crunaire S, Marcoux PR, Ngo K-Q, Moy J-P, Mallard F, Tran-Thi T-H. Discriminating Bacteria with Optical Sensors Based on Functionalized Nanoporous Xerogels. Chemosensors. 2014; 2(2):171-181. https://doi.org/10.3390/chemosensors2020171
Chicago/Turabian StyleCrunaire, Sabine, Pierre R. Marcoux, Khanh-Quyen Ngo, Jean-Pierre Moy, Frédéric Mallard, and Thu-Hoa Tran-Thi. 2014. "Discriminating Bacteria with Optical Sensors Based on Functionalized Nanoporous Xerogels" Chemosensors 2, no. 2: 171-181. https://doi.org/10.3390/chemosensors2020171