Glycobiology Aspects of the Periodontal Pathogen Tannerella forsythia
Abstract
:1. Introduction to Tannerella forsythia
1.1. Occurrence of T. forsythia
1.2. Description of T. forsythia
1.2.1. Taxonomic Affiliation
1.2.2. T. forsythia as a Periodontal Pathogen
2. Cellular Integrity
2.1. S-Layer Glycosylation of T. forsythia
2.1.1. S-Layers in General
2.1.2. S-Layer Ultrastructure of T. forsythia
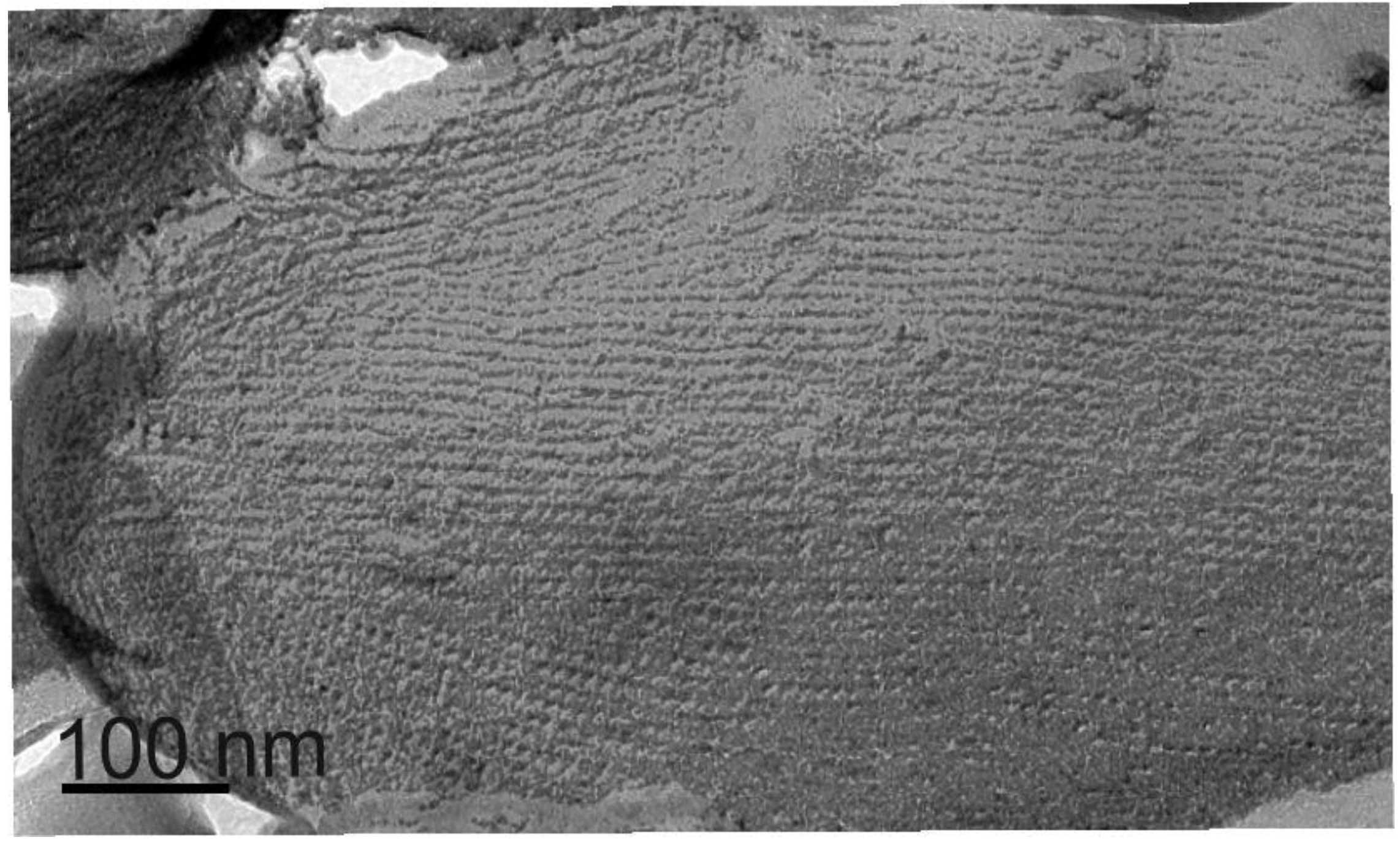

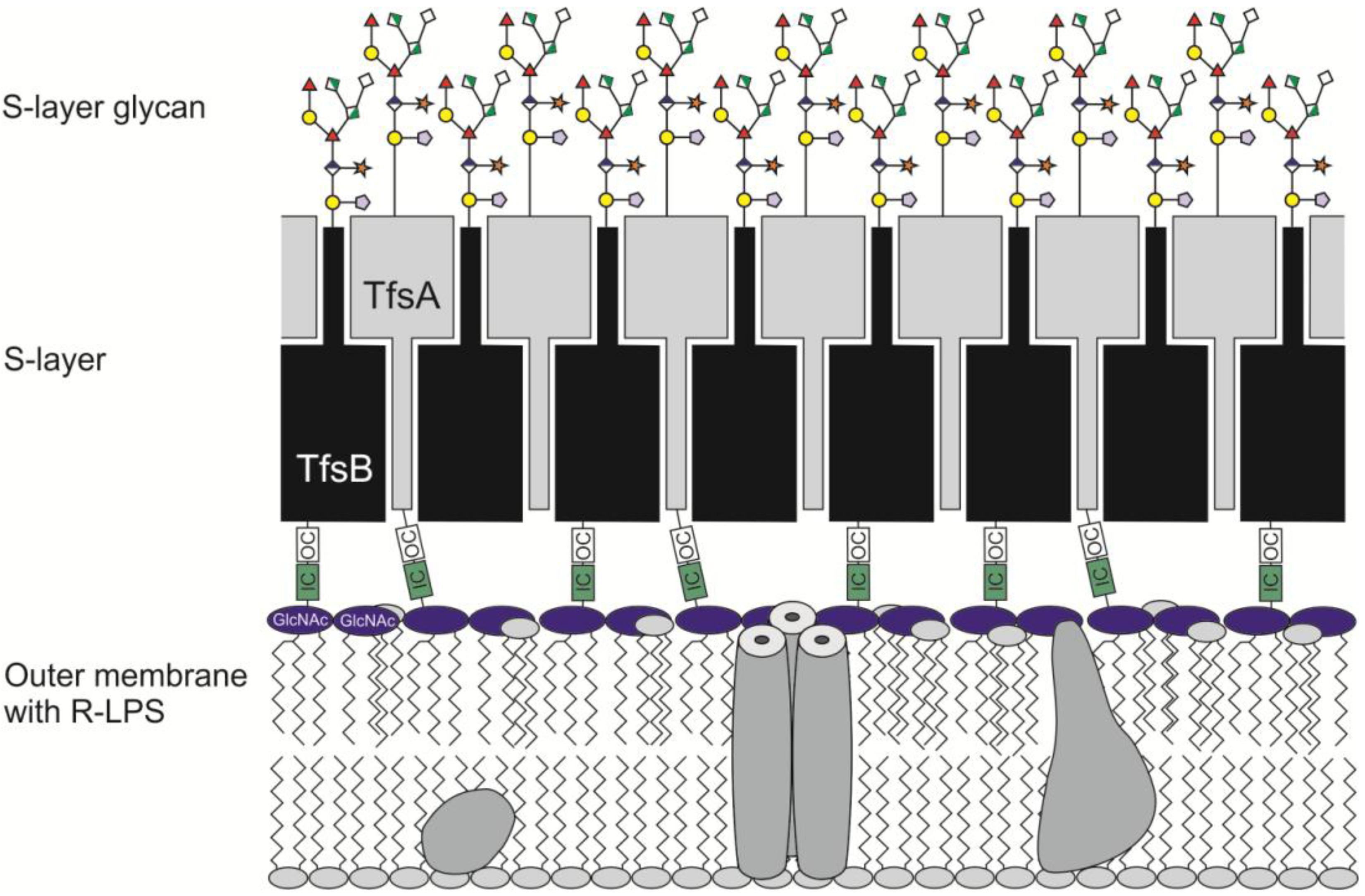
2.1.3. Glycosylation of the T. forsythia S-layer Proteins TfsA and TfsB
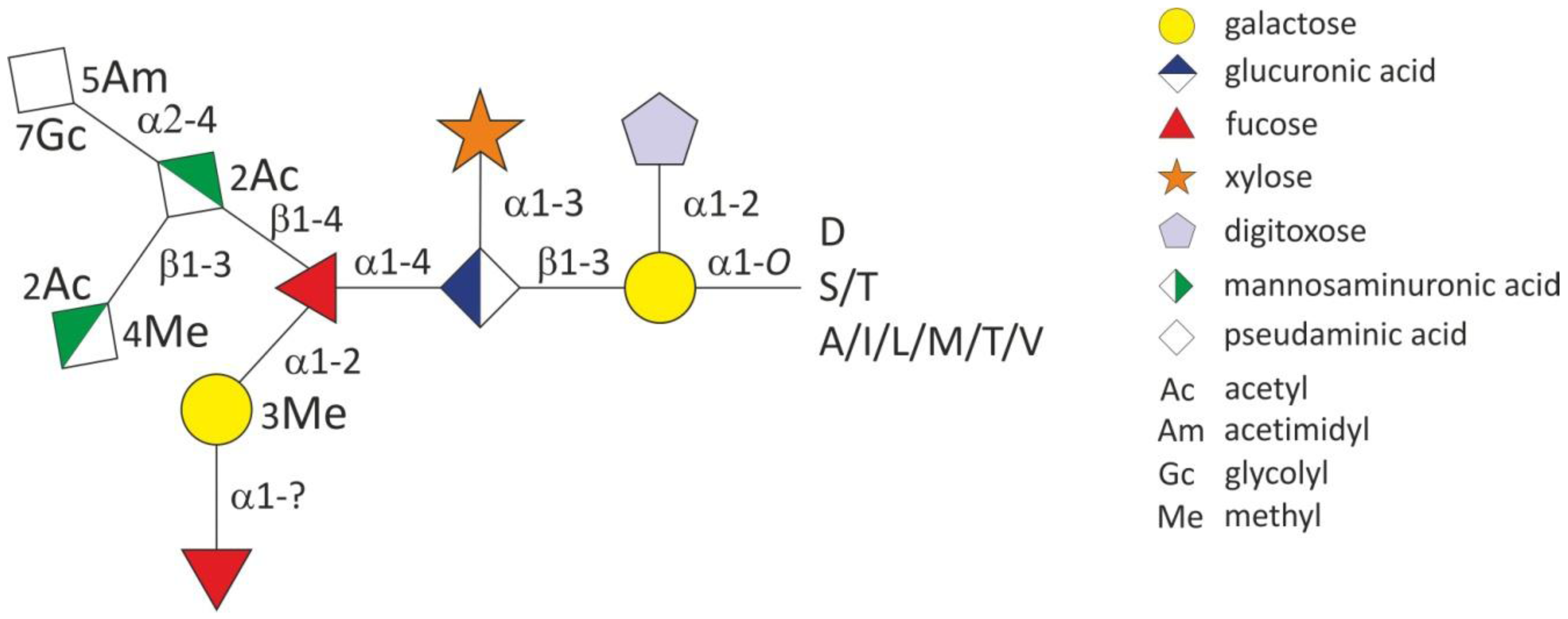
2.2. General O-Glycosylation System of T. forsythia
2.2.1. O-Glycoproteins of T. forsythia
2.2.2. Theoretical analysis of protein O‑glycosylation in T. forsythia
2.3. Lipopolysaccharide (LPS) of T. forsythia
3. Virulence Potential of the Glycosylated S‑Layer of T. forsythia
3.1. Virulence and Glycosylation in General
3.2. Immunological Data of T. forsythia
4. Cultivation and growth of T. forsythia
5. Glycosidic Activity of T. forsythia
6. Biofilm Life-Style of T. forsythia
6.1. General Remarks
6.2. Biofilm Life-Style of T. forsythia and Glycosylation
7. Conclusions
Acknowledgments
Conflict of Interest
References
- Partridge, N.C.; Hillyard, C.J.; Nolan, R.D.; Martin, T.J. Regulation of prostaglandin production by osteoblast-rich calvarial cells. Prostaglandins 1985, 30, 527–539. [Google Scholar] [CrossRef]
- Kortsik, C.; Elmer, A.; Tamm, I. Pleural effusion due to Histoplasma capsulatum and idiopathic CD14 lymphocytopenia. Respiration 2003, 70, 118–122. [Google Scholar] [CrossRef]
- Smalley, J.W.; Birss, A.J.; Withnall, R.; Silver, J. Interactions of Porphyromonas gingivalis with oxyhaemoglobin and deoxyhaemoglobin. Biochem. J. 2002, 362, 239–245. [Google Scholar] [CrossRef]
- Holt, S.C.; Ebersole, J.L. Porphyromonas gingivalis, Treponema denticola, and Tannerella forsythia: the ‘red complex’, a prototype polybacterial pathogenic consortium in periodontitis. Periodontol. 2000 2005, 38, 72–122. [Google Scholar] [CrossRef]
- Wyss, C. Dependence of proliferation of Bacteroides forsythus on exogenous N‑acetylmuramic acid. Infect. Immun. 1989, 57, 1757–1759. [Google Scholar]
- Haffajee, A.D.; Socransky, S.S. Microbial etiological agents of destructive periodontal diseases. Periodontol. 2000 1994, 5, 78–111. [Google Scholar] [CrossRef]
- Socransky, S.S.; Haffajee, A.D. Periodontal microbial ecology. Periodontol. 2000 2005, 38, 135–187. [Google Scholar] [CrossRef]
- Tanner, A.C.R.; Haffer, C.; Bratthall, G.T.; Visconti, R.A.; Socransky, S.S. A study of the bacteria associated with advancing periodontitis in man. J. Clin. Periodontol. 1979, 6, 278–307. [Google Scholar] [CrossRef]
- Tanner, A.C.R.; Izard, J. Tannerella forsythia, a periodontal pathogen entering the genomic era. Periodontol. 2000 2006, 42, 88–113. [Google Scholar] [CrossRef]
- Paster, B.J.; Dewhirst, F.E.; Olsen, I.; Fraser, G.J. Phylogeny of Bacteroides, Prevotella, and Porphyromonas spp. and related bacteria. J. Bacteriol. 1994, 176, 725–732. [Google Scholar]
- Sakamoto, M.; Suzuki, M.; Umeda, M.; Ishikawa, I.; Benno, Y. Reclassification of Bacteroides forsythus (Tanner et al. 1986) as Tannerella forsythensis corrig., gen. nov., comb. nov. Int. J. Syst. Evol. Microbiol. 2002, 52, 841–849. [Google Scholar] [CrossRef]
- Maiden, M.F.J.; Cohee, P.; Tanner, A.C.R. Proposal to conserve the adjectival form of the specific epithet in the reclassification of Bacteroides forsythus Tanner et al. 1986 to the genus Tannerella Sakamoto et al. 2002 as Tannerella forsythia corrig., gen. nov., comb. nov. Request for an Opinion. Int. J. Syst. Evol. Microbiol. 2003, 53, 2111–2112. [Google Scholar] [CrossRef]
- Sakamoto, M.; Benno, Y. Reclassification of Bacteroides distasonis, Bacteroides goldsteinii and Bacteroides merdae as Parabacteroides distasonis gen. nov., comb. nov., Parabacteroides goldsteinii comb. nov. and Parabacteroides merdae comb. nov. Int. J. Syst. Evol. Microbiol. 2006, 56, 1599–1605. [Google Scholar] [CrossRef]
- Braham, P.H.; Moncla, B.J. Rapid presumptive identification and further characterization of Bacteroides forsythus. J. Clin. Microbiol. 1992, 30, 649–654. [Google Scholar]
- Los Alamos National Laboratory Oral Pathogen Sequence Databases. Tannerella forsythensis. Available online: http://www.oralgen.lanl.gov/ (Accessed on 10 October 2012).
- Socransky, S.S. Criteria for the infectious agents in dental caries and periodontal disease. J. Clin. Periodontol. 1979, 6, 16–21. [Google Scholar] [CrossRef]
- Socransky, S.S.; Haffajee, A.D.; Cugini, M.A.; Smith, C.; Kent, R.L., Jr. Microbial complexes in subgingival plaque. J. Clin. Periodontol. 1998, 25, 134–144. [Google Scholar] [CrossRef]
- Sharma, A.; Sojar, H.T.; Glurich, I.; Honma, K.; Kuramitsu, H.K.; Genco, R.J. Cloning, expression, and sequencing of a cell surface antigen containing a leucine-rich repeat motif from Bacteroides forsythus ATCC 43037. Infect. Immun. 1998, 66, 5703–5710. [Google Scholar]
- Bird, P.S.; Shakibaie, F.; Gemmell, E.; Polak, B.; Seymour, G.J. Immune response to Bacteroides forsythus in a murine model. Oral Microbiol. Immunol. 2001, 16, 311–315. [Google Scholar] [CrossRef]
- Yoo, J.Y.; Kim, H.C.; Zhu, W.; Kim, S.M.; Sabet, M.; Handfield, M.; Hillman, J.; Progulske-Fox, A.; Lee, S.W. Identification of Tannerella forsythia antigens specifically expressed in patients with periodontal disease. FEMS Microbiol. Lett. 2007, 275, 344–352. [Google Scholar] [CrossRef]
- Takemoto, T.; Kurihara, H.; Dahlen, G. Characterization of Bacteroides forsythus isolates. J. Clin. Microbiol. 1997, 35, 1378–1381. [Google Scholar]
- Sharma, A.; Inagaki, S.; Honma, K.; Sfintescu, C.; Baker, P.J.; Evans, R.T. Tannerella forsythia-induced alveolar bone loss in mice involves leucine-rich-repeat BspA protein. J. Dent. Res. 2005, 84, 462–467. [Google Scholar] [CrossRef]
- Kesavalu, L.; Sathishkumar, S.; Bakthavatchalu, V.; Matthews, C.; Dawson, D.; Steffen, M.; Ebersole, J.L. Rat model of polymicrobial infection, immunity, and alveolar bone resorption in periodontal disease. Infect. Immun. 2007, 75, 1704–1712. [Google Scholar] [CrossRef]
- Sharma, A. Virulence mechanisms of Tannerella forsythia. Periodontol. 2000 2010, 54, 106–116. [Google Scholar] [CrossRef]
- Ishikura, H.; Arakawa, S.; Nakajima, T.; Tsuchida, N.; Ishikawa, I. Cloning of the Tannerella forsythensis (Bacteroides forsythus) siaHI gene and purification of the sialidase enzyme. J. Med. Microbiol. 2003, 52, 1101–1107. [Google Scholar] [CrossRef]
- Hughes, C.V.; Malki, G.; Loo, C.Y.; Tanner, A.C.R.; Ganeshkumar, N. Cloning and expression of a-D-glucosidase and N-acetyl-b-glucosaminidase from the periodontal pathogen, Tannerella forsythensis (Bacteroides forsythus.). Oral Microbiol. Immunol. 2003, 18, 309–312. [Google Scholar] [CrossRef]
- Sabet, M.; Lee, S.-W.; Nauman, R.K.; Sims, T.; Um, H.-S. The surface (S-) layer is a virulence factor of Bacteroides forsythus. Microbiology 2003, 149, 3617–3627. [Google Scholar] [CrossRef]
- Sekot, G.; Posch, G.; Messner, P.; Matejka, M.; Rausch-Fan, X.; Andrukhov, O.; Schäffer, C. Potential of the Tannerella forsythia S-layer to delay the immune response. J. Dent. Res. 2011, 90, 109–114. [Google Scholar] [CrossRef]
- Posch, G.; Pabst, M.; Brecker, L.; Altmann, F.; Messner, P.; Schäffer, C. Characterization and scope of S-layer protein O-glycosylation in Tannerella forsythia. J. Biol. Chem. 2011, 286, 38714–38724. [Google Scholar]
- Sleytr, U.B.; Messner, P. Crystalline surface layers on bacteria. Annu. Rev. Microbiol. 1983, 37, 311–339. [Google Scholar] [CrossRef]
- Messner, P.; Schäffer, C.; Egelseer, E.M.; Sleytr, U.B. Occurrence, structure, chemistry, genetics, morphogenesis, and functions of S-layers. In Prokaryotic Cell Wall Compounds-Structure and Biochemistry; König, H., Claus, H., Varma, A., Eds.; Springer-Verlag: Berlin, Germany, 2010; pp. 53–109. [Google Scholar]
- Ishiguro, E.E.; Kay, W.W.; Ainsworth, T.; Chamberlain, J.B.; Austen, R.A.; Buckley, J.T.; Trust, T.J. Loss of virulence during culture of Aeromonas salmonicidaat high temperature. J. Bacteriol. 1981, 148, 333–340. [Google Scholar]
- Borinski, R.; Holt, S.C. Studies on the surface of Wolinella recta ATCC 33238 and human clinical isolates: correlation of structure with function. Infect. Immun. 1990, 58, 2770–2776. [Google Scholar]
- Garduño, R.; Moore, A.; Olivier, G.; Lizama, A.L.; Garduño, E.; Kay, W.W. Host cell invasion and intracellular residence by Aeromonas salmonicida: role of the S-layer. Can. J. Microbiol. 2000, 46, 660–668. [Google Scholar]
- Kerosuo, E. Ultrastructure of the cell envelope of Bacteroides forsythus strain ATCC 43037. Oral Microbiol. Immunol. 1988, 3, 134–137. [Google Scholar] [CrossRef]
- Higuchi, N.; Murakami, Y.; Moriguchi, K.; Ohno, N.; Nakamura, H.; Yoshimura, F. Localization of major, high molecular weight proteins in Bacteroides forsythus. Microbiol. Immunol. 2000, 44, 777–780. [Google Scholar]
- Yoneda, M.; Hirofuji, T.; Motooka, N.; Nozoe, K.; Shigenaga, K.; Anan, H.; Miura, M.; Kabashima, H.; Matsumoto, A.; Maeda, K. 2003. Humoral immune responses to S‑layer-like proteins of Bacteroides forsythus. Clin. Diagn. Lab. Immunol. 2003, 10, 383–387. [Google Scholar]
- Lee, S.-W.; Sabet, M.; Um, H.-S.; Yang, J.; Kim, H.C.; Zhu, W. Identification and characterization of the genes encoding a unique surface (S-) layer of Tannerella forsythia. Gene 2006, 371, 102–111. [Google Scholar] [CrossRef]
- Sekot, G.; Posch, G.; Oh, Y.J.; Zayni, S.; Mayer, H.F.; Pum, D.; Messner, P.; Hinterdorfer, P.; Schäffer, C. Analysis of the cell surface layer ultrastructure of the oral pathogen Tannerella forsythia. Arch. Microbiol. 2012, 194, 525–539. [Google Scholar] [CrossRef]
- Fletcher, C.M.; Coyne, M.J.; Bentley, D.L.; Villa, O.F.; Comstock, L.E. Phase-variable expression of a family of glycoproteins imparts a dynamic surface to a symbiont in its human intestinal ecosystem. Proc. Natl. Acad. Sci. USA 2007, 104, 2413–2418. [Google Scholar]
- Nguyen, K.A.; Travis, J.; Potempa, J. Does the importance of the C-terminal residues in the maturation of RgpB from Porphyromonas gingivalis reveal a novel mechanism for protein export in a subgroup of Gram-negative bacteria? J. Bacteriol. 2007, 189, 833–843. [Google Scholar] [CrossRef]
- Chen, Y.-Y.; Peng, B.; Yang, Q.; Glew, M.D.; Veith, P.D.; Cross, K.J.; Goldie, K.N.; Chen, D.; O’Brien-Simpson, N.; Daspher, S.G.; Reynolds, E.C. The outer membrane protein LptO is essential for the O-deacetylation of LPS and the co-ordinated secretion and attachment of A-LPS and CTD proteins in Porphyromonas gingivalis. Mol. Microbiol. 2011, 79, 1380–1401. [Google Scholar] [CrossRef]
- Wang, B.N.; Kraig, E.; Kolodrubetz, D. Use of defined mutants to assess the role of the Campylobacter rectus S-layer in bacterium-epithelial cell interactions. Infect. Immun. 2000, 68, 1465–1473. [Google Scholar] [CrossRef]
- Tanner, A.C.R.; Listgarten, M.A.; Ebersole, J.L.; Strzempko, M.N. Bacteroides forsythus sp. nov., a slow-growing, fusiform Bacteroides sp. from the human oral cavity. Int. J. Syst. Bacteriol. 1986, 36, 213–221. [Google Scholar] [CrossRef]
- Veith, P.D.; O’Brien-Simpson, N.M.; Tan, Y.; Djatmiko, D.C.; Dashper, S.G.; Reynolds, E.C. Outer membrane proteome and antigens of Tannerella forsythia. J. Proteome Res. 2009, 8, 4279–4292. [Google Scholar]
- Fletcher, C.M.; Coyne, M.J.; Villa, O.F.; Chatzidaki-Livanis, M.; Comstock, L.E. A general O‑glycosylation system important to the physiology of a major human intestinal symbiont. Cell 2009, 137, 321–331. [Google Scholar] [CrossRef]
- Koebnik, R. Ton-B dependent trans-envelope signalling: the exception of the rule? Trends Microbiol. 2005, 13, 343–347. [Google Scholar] [CrossRef]
- Honma, K.; Inagaki, S.; Okuda, K.; Kuramitsu, H.K.; Sharma, A. Role of a Tannerella forsythia exopolysaccharide synthesis operon in biofilm development. Microb. Pathog. 2007, 42, 156–166. [Google Scholar] [CrossRef]
- Raetz, C.R.H.; Whitfield, C. Lipopolysaccharide endotoxins. Annu. Rev. Biochem. 2002, 71, 635–700. [Google Scholar] [CrossRef]
- Settem, R.P.; Honma, K.; Nakajima, T.; Phansopa, C.; Roy, S.; Stafford, G.P.; Sharma, A. A bacterial glycan core linked to surface (S)-layer proteins modulates host immunity through Th17 suppression. Mucosal Immunol. 2012. [Google Scholar] [CrossRef]
- Griffiths, S.G.; Lynch, W.H. Characterization of Aeromonas salmonicida variants with altered cell surfaces and their use in studying surface protein assembly. Arch. Microbiol. 1990, 154, 308–312. [Google Scholar] [CrossRef]
- Schmidt, M.A.; Riley, L.W.; Benz, I. Sweet new world: glycoproteins in bacterial pathogens. Trends Microbiol. 2003, 11, 554–561. [Google Scholar] [CrossRef]
- Lebeer, S.; Vanderleyden, J.; De Keersmaecker, S.C. Host interactions of probiotic bacterial surface molecules: comparison with commensals and pathogens. Nat. Rev. Microbiol. 2010, 8, 171–184. [Google Scholar] [CrossRef]
- Szymanski, C.M.; Wren, B.W. Protein glycosylation in bacterial mucosal pathogens. Nat. Rev. Microbiol. 2005, 3, 225–237. [Google Scholar] [CrossRef]
- Sakakibara, J.; Nagano, K.; Murakami, Y.; Higuchi, N.; Nakamura, H.; Shimozato, K.; Yoshimura, F. Loss of adherence ability to human gingival epithelial cells in S-layer protein-deficient mutants of Tannerella forsythensis. Microbiology 2007, 153, 866–876. [Google Scholar] [CrossRef]
- Roy, S.; Douglas, C.W.; Stafford, G.P. A novel sialic acid utilization and uptake system in the periodontal pathogen Tannerella forsythia. J. Bacteriol. 2010, 192, 2285–2293. [Google Scholar] [CrossRef]
- Gabriel, M.O.; Grunheid, T.; Zentner, A. Glycosylation pattern and cell attachment-inhibiting property of human salivary mucins. J. Periodontol. 2005, 76, 1175–1181. [Google Scholar] [CrossRef]
- Prakobphol, A.; Tangemann, K.; Rosen, S.D.; Hoover, C.I.; Leffler, H.; Fisher, S.J. Separate oligosaccharide determinants mediate interactions of the low molecular weight salivary mucin with neutrophils and bacteria. Biochemistry 1999, 38, 6817–6825. [Google Scholar] [CrossRef]
- Murakami, Y.; Higuchi, N.; Nakamura, H.; Yoshimura, F.; Oppenheim, F.G. Bacteroides forsythus hemagglutinin is inhibited by N-acetylneuraminyllactose. Oral Microbiol. Immunol. 2002, 17, 125–128. [Google Scholar] [CrossRef]
- Stafford, G.; Roy, S.; Honma, K.; Sharma, A. Sialic acid, periodontal pathogens and Tannerella forsythia: stick around and enjoy the feast. Mol. Oral Microbiol. 2012, 27, 11–22. [Google Scholar] [CrossRef]
- Honma, K.; Mishima, E.; Sharma, A. Role of Tannerella forsythia NanH sialidase in epithelial cell attachment. Infect. Immun. 2011, 79, 393–401. [Google Scholar] [CrossRef]
- Roy, S.; Honma, K.; Douglas, I.; Sharma, A.; Stafford, G.P. Role of sialidase in glycoprotein utilisation by Tannerella forsythia. Microbiology 2011, 157, 3195–3202. [Google Scholar] [CrossRef]
- Marsh, P.D.; Moter, A.; Devine, D.A. Dental plaque biofilms: communities, conflict and control. Periodontol. 2000 2011, 55, 16–35. [Google Scholar] [CrossRef]
- Yang, H.W.; Huang, Y.F.; Chou, M.Y. Occurrence of Porphyromonas gingivalis and Tannerella forsythensis in periodontally diseased and healthy subjects. J. Periodontol. 2004, 75, 1077–1083. [Google Scholar] [CrossRef]
- Pham, T.K.; Roy, S.; Noirel, J.; Douglas, I.; Wright, P.C.; Stafford, G.P. A quantitative proteomic analysis of biofilm adaptation by the periodontal pathogen Tannerella forsythia. Proteomics 2010, 10, 3130–3141. [Google Scholar] [CrossRef]
- Ristl, R.; Steiner, K.; Zarschler, K.; Zayni, S.; Messner, P.; Schäffer, C. The S-layer glycome – adding to the sugar coat of bacteria. Int. J. Microbiol. 2011, 2011, 127870:1–127870:16. [Google Scholar]
- Messner, P.; Steiner, K.; Zarschler, K.; Schäffer, C. S-layer nanoglycobiology of bacteria. Carbohydr. Res. 2008, 343, 1934–1951. [Google Scholar] [CrossRef]
- Calo, D.; Eilam, Y.; Lichtenstein, R.G.; Eichler, J. Towards glycoengineering in Archaea: replacement of Haloferax volcanii AglD with homologous glycosyltransferases from other halophilic archaea. Appl. Environ. Microbiol. 2010, 76, 5684–5692. [Google Scholar] [CrossRef]
- Knirel, Y.A.; Shashkov, A.S.; Tsvetkov, Y.E.; Jansson, P.-E.; Zähringer, U. 5,7-Diamino-3,5,7,9-tetradeoxynon-2-ulosonic acids in bacterial glycopolymers: chemistry and biochemistry. Adv. Carbohydr. Chem. Biochem. 2003, 58, 371–417. [Google Scholar] [CrossRef]
- Kiss, E.; Kereszt, A.; Barta, F.; Stephens, S.; Reuhs, B.L.; Kondorosi, A.; Putnoky, P. The rkp-3 gene region of Sinorhizobium meliloti Rm41 contains strain-specific genes that determine K antigen structure. Mol. Plant-Microbe Interact. 2001, 14, 1395–1403. [Google Scholar] [CrossRef]
- Castric, P.; Cassels, F.J.; Carlson, R.W. Structural characterization of the Pseudomonas aeruginosa 1244 pilin glycan. J. Biol. Chem. 2001, 276, 26479–26485. [Google Scholar] [CrossRef]
- Thibault, P.; Logan, S.M.; Kelly, J.F.; Brisson, J.-R.; Ewing, C.P.; Trust, T.J.; Guerry, P. Identification of the carbohydrate moieties and glycosylation motifs in Campylobacter jejuni flagellin. J. Biol. Chem. 2001, 276, 34862–34870. [Google Scholar]
- Schoenhofen, I.C.; McNally, D.J.; Brisson, J.-R.; Logan, S.M. Elucidation of the CMP-pseudaminic acid pathway in Helicobacter pylori: synthesis from UDP-N-acetylglucosamine by a single enzymatic reaction. Glycobiology 2006, 16, 8–14. [Google Scholar] [CrossRef]
- Hsu, K.-L.; Pilobello, K.T.; Mahal, L.K. Analyzing the dynamic bacterial glycome with a lectin microarray approach. Nat. Chem. Biol. 2006, 2, 153–157. [Google Scholar]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Posch, G.; Sekot, G.; Friedrich, V.; Megson, Z.A.; Koerdt, A.; Messner, P.; Schäffer, C. Glycobiology Aspects of the Periodontal Pathogen Tannerella forsythia. Biomolecules 2012, 2, 467-482. https://doi.org/10.3390/biom2040467
Posch G, Sekot G, Friedrich V, Megson ZA, Koerdt A, Messner P, Schäffer C. Glycobiology Aspects of the Periodontal Pathogen Tannerella forsythia. Biomolecules. 2012; 2(4):467-482. https://doi.org/10.3390/biom2040467
Chicago/Turabian StylePosch, Gerald, Gerhard Sekot, Valentin Friedrich, Zoë A. Megson, Andrea Koerdt, Paul Messner, and Christina Schäffer. 2012. "Glycobiology Aspects of the Periodontal Pathogen Tannerella forsythia" Biomolecules 2, no. 4: 467-482. https://doi.org/10.3390/biom2040467
APA StylePosch, G., Sekot, G., Friedrich, V., Megson, Z. A., Koerdt, A., Messner, P., & Schäffer, C. (2012). Glycobiology Aspects of the Periodontal Pathogen Tannerella forsythia. Biomolecules, 2(4), 467-482. https://doi.org/10.3390/biom2040467




