Assembly of DNA Architectures in a Non-Aqueous Solution
Abstract
:1. Introduction
2. Results and Discussion
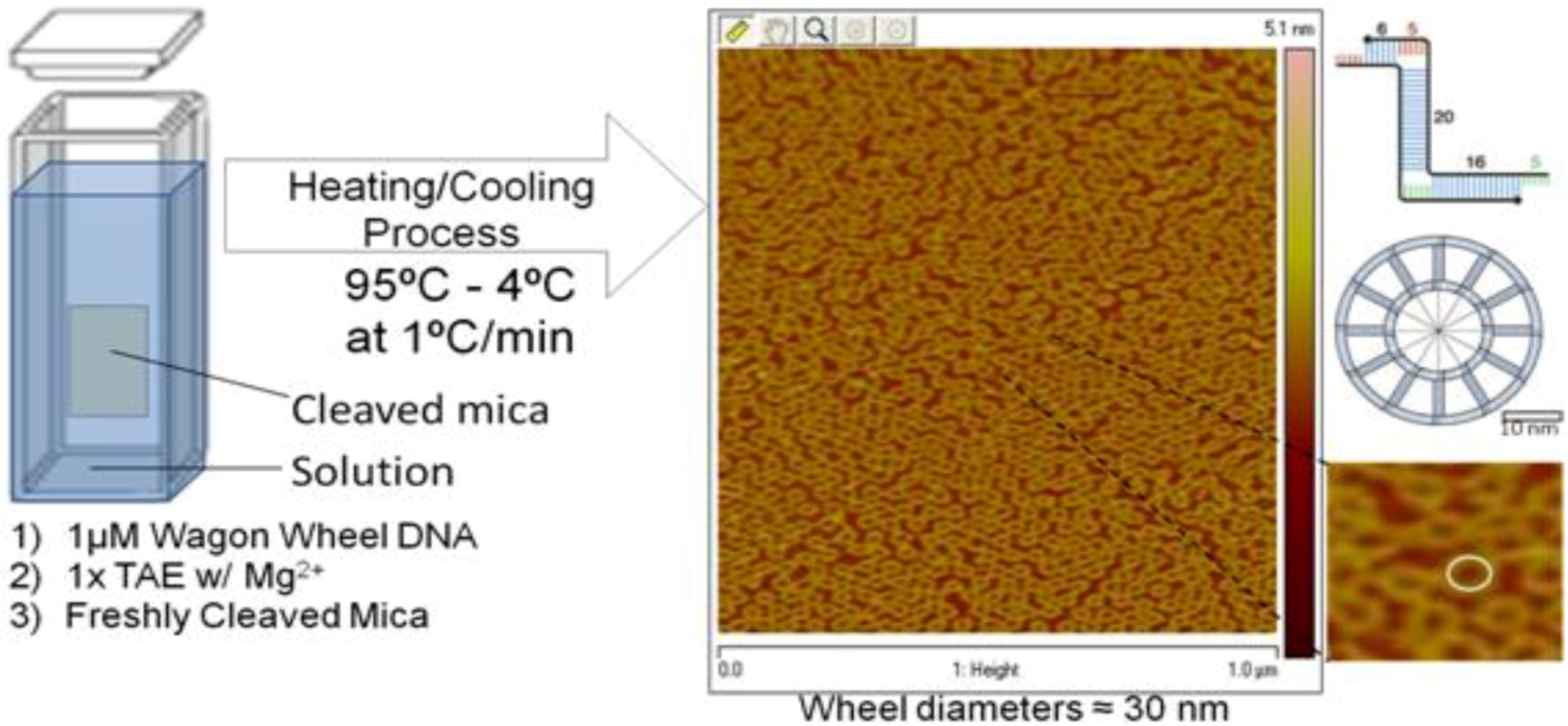
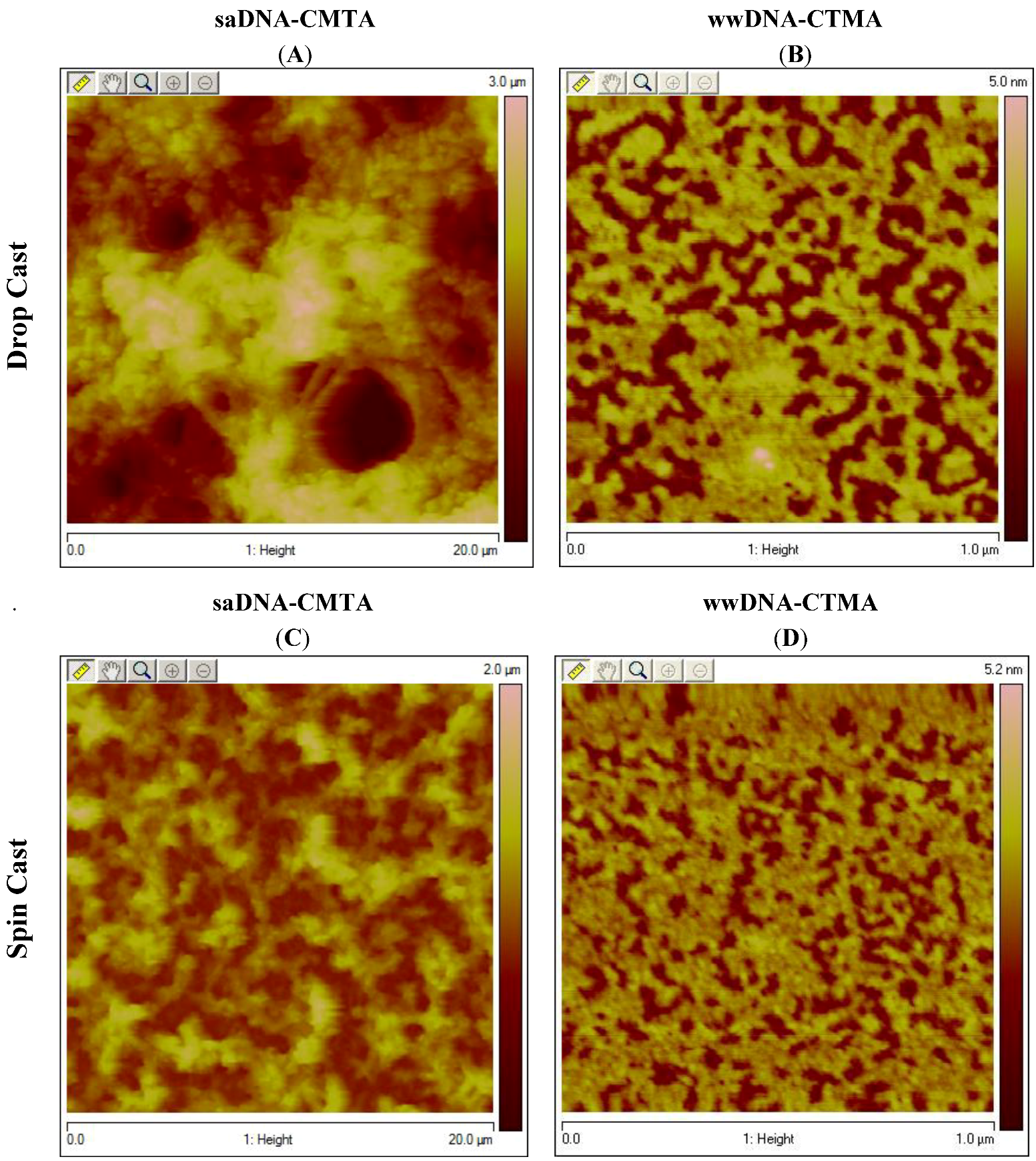

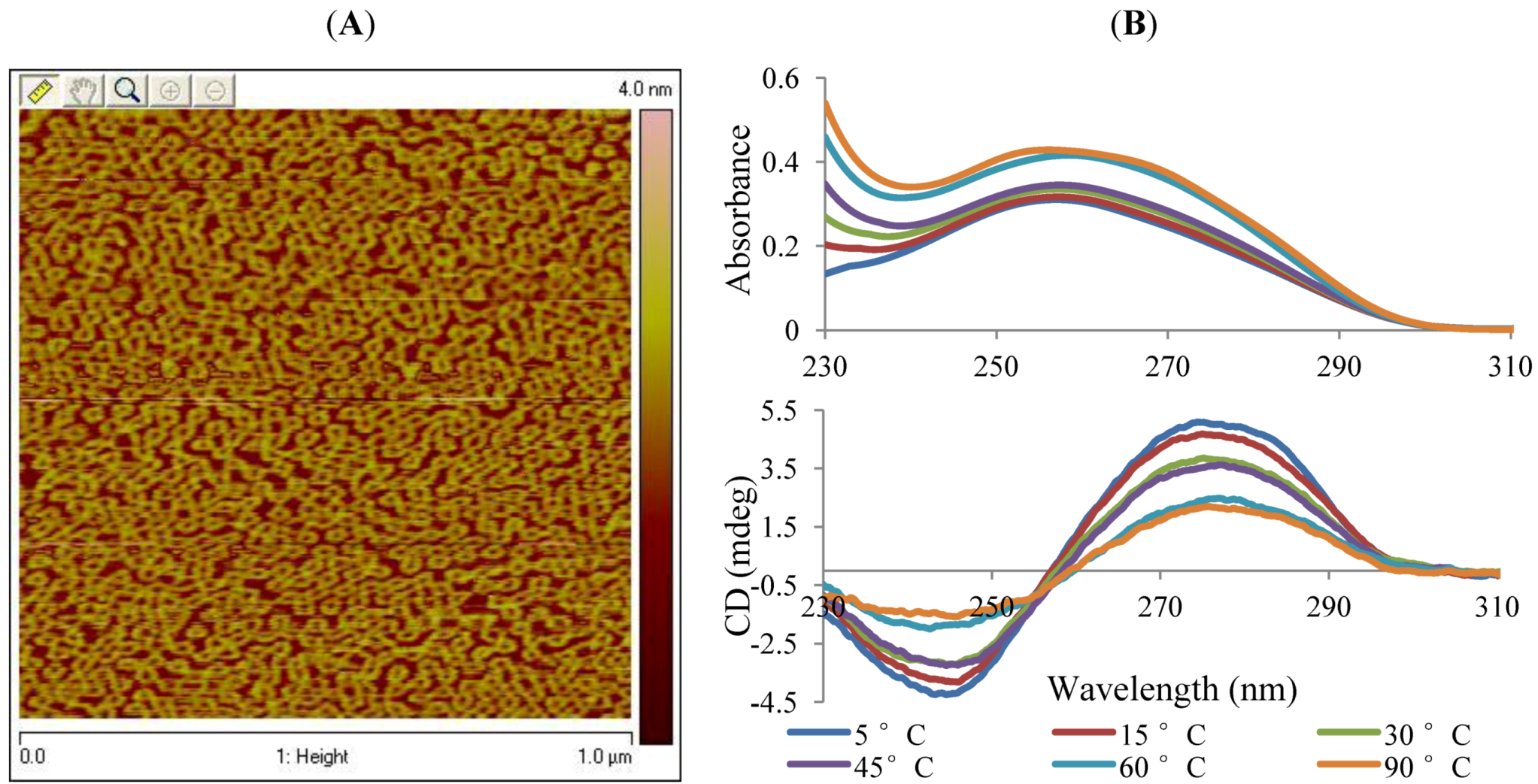
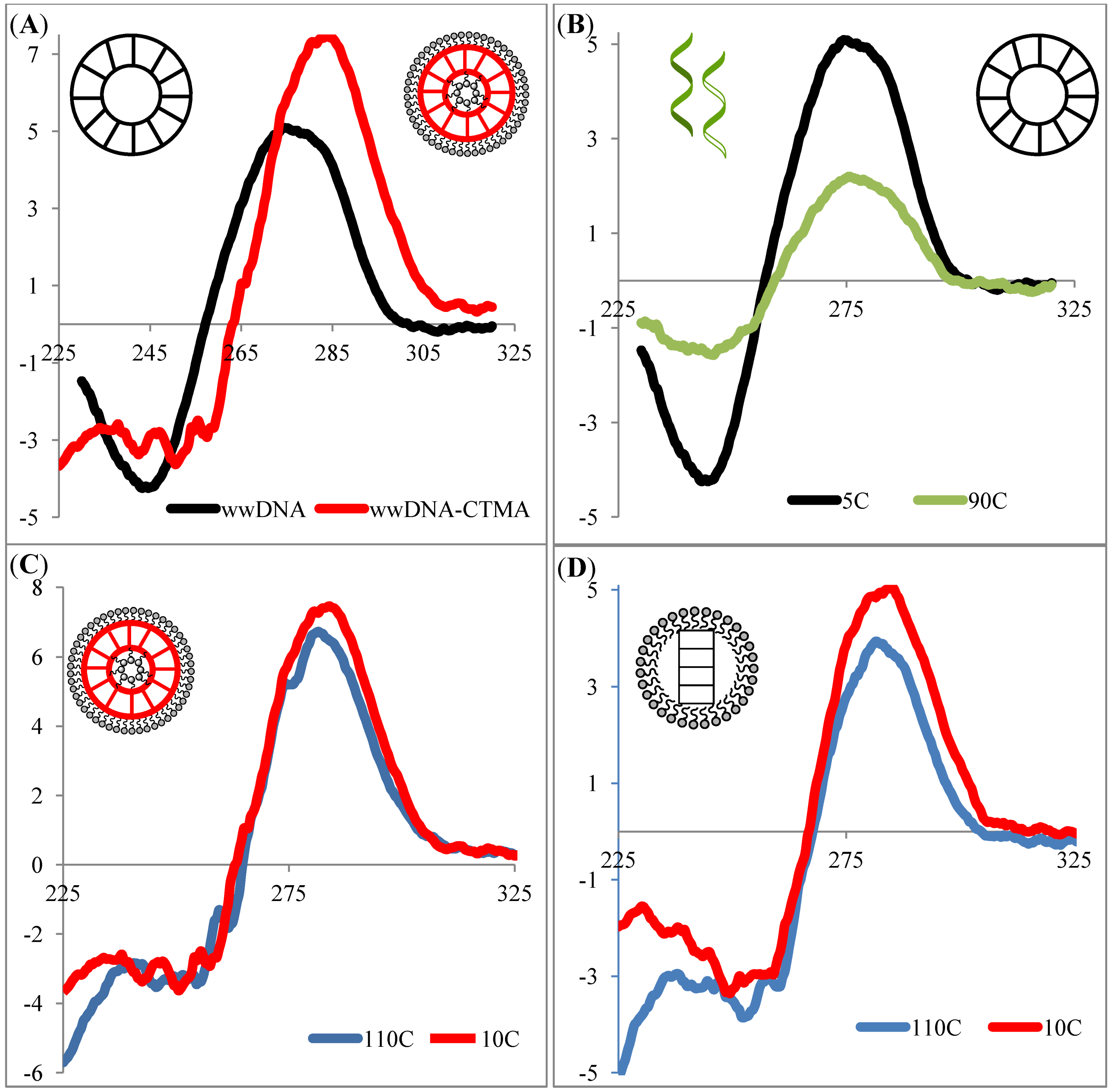

3. Experimental Section
3.1. Materials
3.2. Methods
4. Conclusions
Supplementary Materials
Supplementary File 1Acknowledgements
References
- Galatsis, K.; Wang, K.L.; Ozkan, M.; Ozkan, C.S.; Huang, Y.; Chang, J.P.; Monbouquette, H.G.; Chen, Y.; Nealey, P.; Botros, Y. Patterning and templating for nanoelectronics. Adv. Mater. 2010, 22, 769–778. [Google Scholar]
- Noy, A. Bionanoelectronics. Adv. Mater. 2011, 23, 807–820. [Google Scholar] [CrossRef]
- Sacca, B.; Niemeyer, C.M. DNA origami: The art of folding DNA. Angew. Chem. Int. Ed. 2012, 51, 58–66. [Google Scholar] [CrossRef]
- Seeman, N.C. Nanomaterials based on DNA. Annu. Rev. Biochem. 2010, 79, 65–87. [Google Scholar] [CrossRef]
- Andersen, E.S.; Dong, M.; Nielsen, M.M.; Jahn, K.; Subramani, R.; Mamdouh, W.; Golas, M.M.; Sander, B.; Stark, H.; Oliveira, C.L.P.; et al. Self-assembly of a nanoscale DNA box with a controllable lid. Nature 2009, 459, 73–76. [Google Scholar]
- Douglas, S.M.; Dietz, H.; Liedl, T.; Hogberg, B.; Graf, F.; Shih, W.M. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 2009, 459, 414–418. [Google Scholar]
- Ke, Y.; Douglas, S.M.; Liu, M.; Sharma, J.; Cheng, A.; Leung, A.; Liu, Y.; Shih, W.M.; Yan, H. Multilayer DNA origami packed on a square lattice. J. Am. Chem. Soc. 2009, 131, 15903–15908. [Google Scholar]
- Ke, Y.; Sharma, J.; Liu, M.; Jahn, K.; Liu, Y.; Yan, H. Scaffolded DNA origami of a DNA tetrahedron molecular container. Nano Lett. 2009, 9, 2445–2447. [Google Scholar]
- Kuzuya, A.; Komiyama, M. Design and construction of a box-shaped 3D-DNA origami. Chem. Commun. 2009, 28, 4182–4184. [Google Scholar] [CrossRef]
- Kuzuya, A.; Komiyama, M. DNA origami: Fold, stick, and beyond. Nanoscale 2010, 2, 310–322. [Google Scholar] [CrossRef]
- Rothemund, P.W. Folding DNA to create nanoscale shapes and patterns. Nature 2006, 440, 297–302. [Google Scholar] [CrossRef]
- Tang, H.; Chen, L.; Xing, C.; Guo, Y.-G.; Wang, S. DNA-templated synthesis of cationic poly(3,4-ethylenedioxythiophene) derivative for supercapacitor electrodes. Macromol. Rapid Commun. 2010, 31, 1892–1896. [Google Scholar] [CrossRef]
- Lin, Y.; Tao, Y.; Ren, J.; Pu, F.; Qu, X. Highly sensitive and selective detection of thiol-containing biomolecules using DNA-templated silver deposition. Biosens. Bioelectron. 2011, 28, 339–343. [Google Scholar] [CrossRef]
- Shukla, S.; Sastry, M. Probing differential Ag+-nucleobase interactions with isothermal titration calorimetry (ITC): Towards patterned DNA metallization. Nanoscale 2009, 1, 122–127. [Google Scholar] [CrossRef]
- Boeneman, K.; Prasuhn, D.E.; Blanco-Canosa, J.B.; Dawson, P.E.; Melinger, J.S.; Ancona, M.; Stewart, M.H.; Susumu, K.; Huston, A.; Medintz, I.L. Self-assembled quantum dot-sensitized multivalent DNA photonic wires. J. Am. Chem. Soc. 2010, 132, 18177–18190. [Google Scholar]
- Kim, H.J.; Roh, Y.; Hong, B. Selective formation of a latticed nanostructure with the precise alignment of DNA-templated gold nanowires. Langmuir 2010, 26, 18315–18319. [Google Scholar] [CrossRef]
- Heckman, E.M.; Aga, R.S.; Rossbach, A.T.; Telek, B.A.; Bartsch, C.M.; Grote, J.G. DNA biopolymer conductive cladding for polymer electro-optic waveguide modulators. Appl. Phys. Lett. 2011, 98, 103304:1–103304:3. [Google Scholar]
- Popescu, R.; Moldoveanu, M.; Rau, I. Biopolymer thin films for photonics applications. Key Eng. Mater. 2009, 415, 33–36. [Google Scholar] [CrossRef]
- Heckman, E.; Bartsch, C.; Yaney, P.; Subramanyam, G.; Ouchen, F.; Grote, J.G. DNA-Surfactant Thin-Film Processing and Characterization. In Materials Science of DNA; CRC Press: Boca Raton, FL, USA, 2011; pp. 179–230. [Google Scholar]
- Gao, L.; Ma, N. DNA-templated semiconductor nanocrystal growth for controlled DNA packing and gene delivery. ACS Nano 2012, 6, 689–695. [Google Scholar]
- Zang, D.Y.; Grote, J. DNA-based nanoparticle composite materials for EMI shielding. Proc. SPIE 2012. [Google Scholar]
- Singh, T.B.; Sariciftci, N.S.; Grote, J.G. Bio-Organic Optoelectronic Devices Using DNA. In Organic Electronics; Meller, G., Grasser, T., Eds.; Springer: New York, NY, USA, 2010; Volume 223, pp. 189–212. [Google Scholar]
- Séverac, F.; Alphonse, P.; Estève, A.; Bancaud, A.; Rossi, C. High-energy Al/Cuo nanocomposites obtained by DNA-directed assembly. Adv. Funct. Mater. 2012, 22, 323–329. [Google Scholar]
- Kobayashi, N. Bioled with DNA/conducting polymer complex as active layer. Nonlinear Opt. Quantum Opt. 2011, 43, 233–251. [Google Scholar]
- Sun, Q.; Chang, D.W.; Dai, L.; Grote, J.; Naik, R. Multilayer white polymer light-emitting diodes with deoxyribonucleic acid-cetyltrimetylammonium complex as a hole-transporting/electron-blocking layer. Appl. Phys. Lett. 2008, 92, 251108:1–251108:3. [Google Scholar]
- Singh, T.B.; Sariciftci, N.S.; Grote, J.G. Bio-organic optoelectronic devices using DNA. Adv. Polym. Sci. 2010, 223, 189–212. [Google Scholar]
- Horowitz, G. Interfaces in Organic Field-Effect Transistors. In Organic Electronics; Meller, G., Grasser, T., Eds.; Springer: New York, NY, USA, 2010; Volume 223, pp. 113–153. [Google Scholar]
- Finch, A.S.; Jacob, C.M.; Sumner, J.J. DNA architectures for templated material growth. Proc. SPIE 2011. [Google Scholar]
- Hamada, S.; Murata, S. Substrate-assisted assembly of interconnected single-duplex DNA nanostructures. Angew. Chem. Int. Ed. 2009, 48, 6820–6823. [Google Scholar] [CrossRef]
- Sun, X.; Hyeon Ko, S.; Zhang, C.; Ribbe, A.E.; Mao, C. Surface-mediated DNA self-assembly. J. Am. Chem. Soc. 2009, 131, 13248–13249. [Google Scholar] [CrossRef]
- Kypr, J.; Kejnovska, I.; Renciuk, D.; Vorlickova, M. Circular dichroism and conformational polymorphism of DNA. Nucleic Acids Res. 2009, 37, 1713–1725. [Google Scholar] [CrossRef]
- Tanaka, K.; Okahata, Y. A DNA-lipid complex in organic media and formation of an aligned cast film. J. Am. Chem. Soc. 1996, 118, 10679–10683. [Google Scholar] [CrossRef]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Finch, A.S.; Anton, C.M.; Jacob, C.M.; Proctor, T.J.; Stratis-Cullum, D.N. Assembly of DNA Architectures in a Non-Aqueous Solution. Nanomaterials 2012, 2, 275-285. https://doi.org/10.3390/nano2030275
Finch AS, Anton CM, Jacob CM, Proctor TJ, Stratis-Cullum DN. Assembly of DNA Architectures in a Non-Aqueous Solution. Nanomaterials. 2012; 2(3):275-285. https://doi.org/10.3390/nano2030275
Chicago/Turabian StyleFinch, Amethist S., Christopher M. Anton, Christina M. Jacob, Thomas J. Proctor, and Dimitra N. Stratis-Cullum. 2012. "Assembly of DNA Architectures in a Non-Aqueous Solution" Nanomaterials 2, no. 3: 275-285. https://doi.org/10.3390/nano2030275





