1. Introduction
As an essential food for human beings, rice (
Oryza sativa L.) has been widely consumed and planted around the world. Particularly, 80% of rice production in the world comes from Asian countries [
1]. More than half of the world’s population constantly includes rice in their diet, and about 2808 calories are provided by rice per person per day [
2]. As world population growth and climate change are undeniable, demands for higher yield and better quality rice have become a critical issue [
3]. Therefore, breeding new rice varieties with improved quality and productivity is an unprecedented challenge [
4].
With assistance from biotechnology, rice breeding targets are not only improving crop productivity, but also upgrading quality characteristics through alterations in heredity [
5]. Many tremendous efforts and rapid progresses in rice breeding programs have been carried out and received remarkable achievements [
6]. As a result, new rice varieties have been developed and released with higher yield and quality.
Mutation, an effective method applied in many crops, plays a notable role in the development of desired varieties particularly in rice. Mutation may occur naturally due to alterations of base pairs in a DNA sequence [
5]. These alterations can be passed from generation to generation; thus, mutation is the primary source of genetic variation. In accordance with recombinant breeding and transgenic breeding, mutation is currently the most effective tool of plant breeding, especially throughout sexual production [
7]. As the major element of germplasm, genetic diversity is a natural source for rice breeding to encounter existing food requirements [
8]. The higher the level of genetic variation is in a population, the more valuable it is as a resource is used for enlarging the genetic base in breeding program [
9]. In addition, Haritha et al. [
10] postulated that genetic diversity obtained precious information for both basic studies and practical applications. DNA markers have revealed a promising approach toward detecting genetic diversity and assisting in management of plant genetic resources [
11]. The differentiations among accessions, individuals, and identification and characterization of novel germplasms at the molecular level, can be recognized by DNA markers and genetic engineering [
12]. In association with bioinformatic tools, DNA markers provide favorably successful and reliable technologies to facilitate evaluation of genetic variation in rice. Moreover, DNA markers offer a great promise for rice breeding because of their competence and accuracy [
13]. Numerous kinds of molecular markers have been practiced for calculating genetic dissimilarity in rice [
14]. Remarkably, among closely related rice cultivars, SSR markers are particularly prominent for evaluation of genetic diversity as they can detect a remarkable great level of polymorphism [
11,
15].
Chemical mutagens have been used for genetic screens crop breeding. Compounds such as EMS (ethyl methanesulfonate) and MNU (
N-Nitroso-
N-methylurea)-induced single nucleotide changes by alkylation of specific nucleotides [
16,
17], giving mutations that were high in density and essentially randomly distributed [
18]. Therefore, a relatively small population of rice can provide variety of missense changes with differing effects on protein function [
19]. In a previous study, Satoh et al. [
20] found that using of
N-methyl-
N-nitrosourea mutation caused effects to the
PHO1 gene in rice, which might play some crucial role in starch biosynthesis by forming a functional protein-protein complexed with carbohydrate metabolizing enzymes. The mutant rice lines were produced by treating fertilized egg cells of Japonica rice cultivars (
Oryza sativa cv. Kinmaze and T65) [
20], and when rice seeds mutated by MNU, the influence on the biosynthesis in rice endosperm, including amylopectin and amylose contents, has not been understood.
Rice grain quality, a persuasive factor for consumer preferences and global commerce, is a combination of physical and chemical characteristics [
3]. It is categorized in different major physical and chemical indicators [
6] such as appearance quality, milling quality, nutritional quality, cooking and eating quality. Amylose content (AC) is considered as one of the key characteristics for assessment of rice cooking and eating quality [
21], and it may also affect some physical traits [
3]. Hence, this study was conducted to evaluate the phenotypic variation of rice mutant lines achieved from MNU mutation, to determine the genetic divergence using SSR markers relevant to amylose content trait, and to investigate the alteration of rice germplasms after mutation.
4. Discussion
In this study, the rice materials were cultivated in the rice field to measure the variation in quality and yield characteristics. They were also genotyped to clarify the role of mutation. The results showed that both grain yield per plant and amylose content were varied among cultivars and mutant lines. The association among traits is a principal feature that may assist the selection of elite characteristics in breeding programs not only for rice but also for the other plants [
6,
30]. Discovered the role and the relationship of characteristics related to quality trait in rice is important to develop new varieties with expected quality [
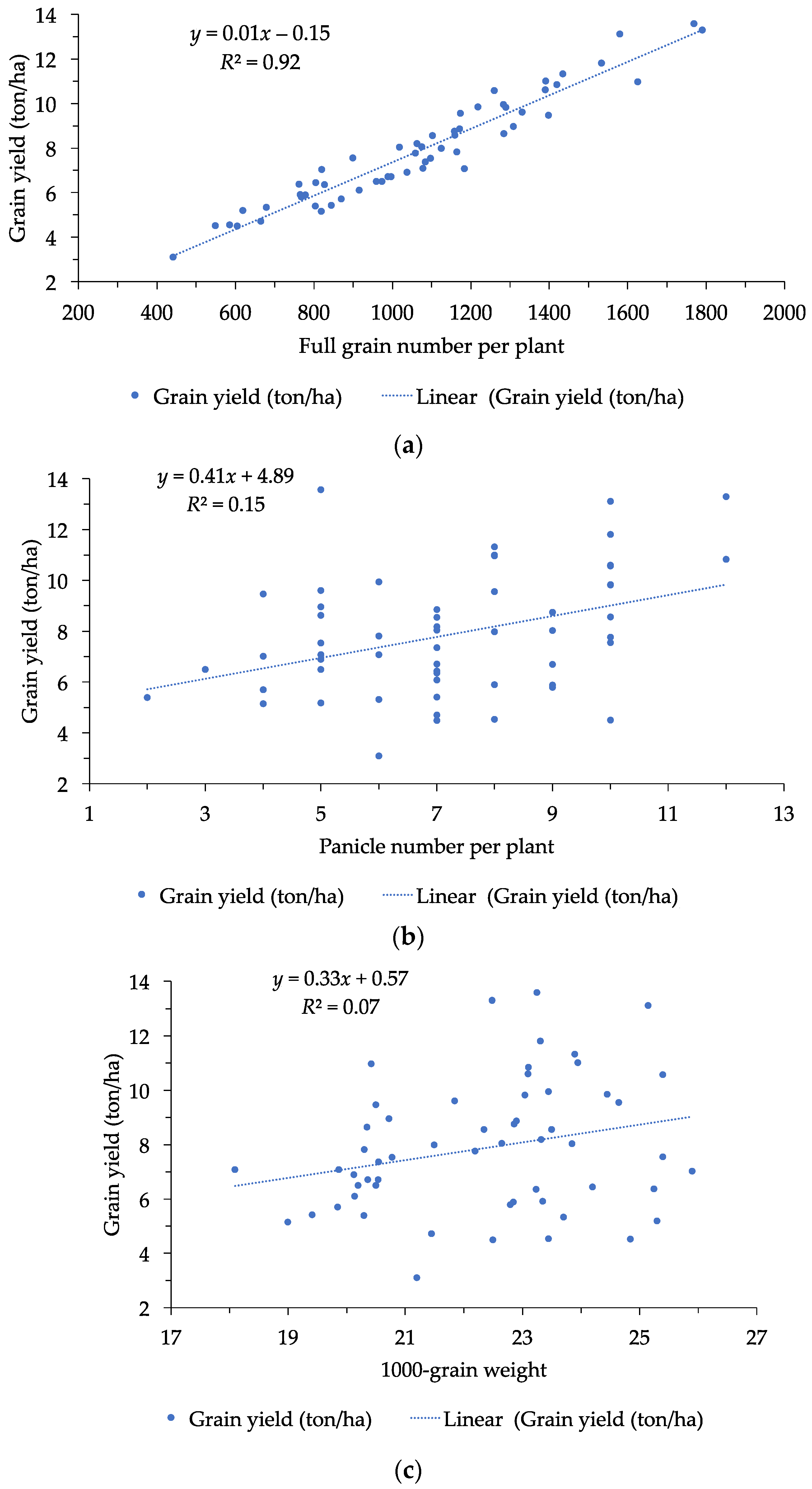
6]. Three main constituents of grain yield include number of panicles per plant, number of grains per panicle, and grain weight. Among them, grain weight, measured as 1000-grain weight, is a decisive trait to grain yield [
12,
31]. In this study, the highest significantly positive correlation was found between grain yield per plant and full grain number per plant, followed by number of panicles per plant and grain weight. This suggested that full grain number per plant has the highest contribution to grain yield, and an increase in full grain number per plant leads to an increase in grain yield.
Grain size is considered as a major determinant of grain appearance quality and grain weight, and it was contributed by grain length, grain width, and grain length to width ratio [
12]. Our study agreed with many findings showed that grain weight was significantly correlated with grain size [
12,
31,
32]. This result implied that the larger the grain, the heavier the grain. However, it is difficult for breeders to improve grain size efficiently by phenotyping, since this trait is quantitatively inherited although plants with large seed size were selected for breeding [
12].
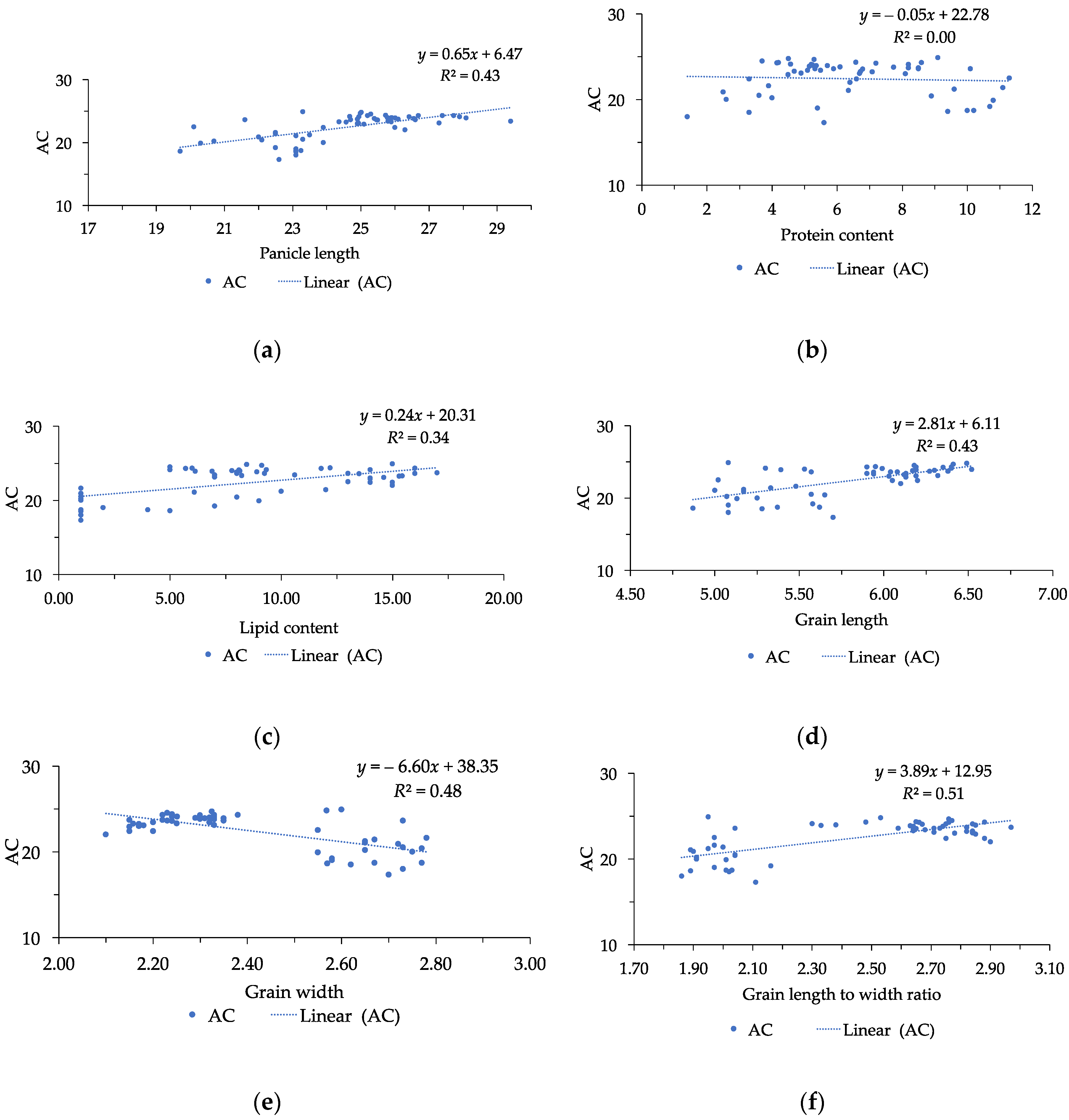
The correlation between appearance quality and nutritional quality was also found in this research. AC and protein content are two essential indicators for eating, cooking, and nutritional quality, respectively [
6]. With regards to AC and grain size, our results were consistent with that previously reported by Xu et al. [
33] and Abacar et al. [
6]. The significant correlation between AC and grain size suggested that selection of rice mutants with high values of grain length and grain length to width ratio could result in positive response of AC. The involvement of grain length to width ratio in AC was greater than the other traits, and grain length to width ratio was the most significant character influencing AC. Therefore, rice with high grain length to width ratio leads to an increase in AC in rice [
6]. Interestingly, this study also found that there was no correlation between PC and grain size. In a previous study, Lou et al. [
34] stated that there was no significant correlation between AC and other quality traits. It was reasoned that the principal element that regulates AC was a combination of the
Wx gene with the other minor QTLs. This finding proposed that marker-assisted selection can apply for AC. Furthermore, Abacar et al. [
6] highlighted that it was difficult to explain the relationship between AC and the other traits. Additionally, Huang et al. [
31] stated that there was a positive correlation between grain weight and grain size; however, the level of correlation of them was differed from independent studies.
In general, there was a dissimilarity in grain yield and AC among the rice mutants. This can be reasoned that mutation may have affected these characteristics. However, the role of mutation involving these characteristics could not be studied. Therefore, it is necessary to do further study on these traits. In the previous reports, Kumbhar et al. [
27] and Salem et al. [
35] stated that it is difficult to discriminate close genotypes through phenotypic assessment. Furthermore, many agronomical characteristics such as quality and yield are highly affected by environmental conditions [
36]. In another report, Lo et al. [
37] indicated that in normal environmental conditions, rice mutants may not exhibit desirable phenotypes; subsequently, these mutants need to be recognized by screening under appropriate or inducible conditions.
Genetic variation in a population is a worthy source for enlarging the range of genetic materials in plant breeding, and molecular markers, such as SSRs and SNPs, are effective tools to discover genetic diversity [
9]. Compared to other genetic markers, SSR markers have remarkable potential to discriminate between rice genotypes [
35,
38]. Classification of genotypes based on polymorphism markers is a prevailing mean for assessment of genetic variance [
27,
38,
39]. PIC value is a principal factor in distinguishing the percentage of polymorphism of a marker at a specific locus; the higher PIC value is in SSR markers, the higher percentage of polymorphism is detected [
38]. In this research, most SSR markers amplified more than two alleles among all genotypes. The PIC values of the 56 polymorphism markers indicates moderate genetic diversity among rice mutants, and seven markers showed great PIC values (0.73 < PIC < 0.79). Salem et al. [
35] affirmed that markers with PIC values higher than 0.7 were very high PIC values, and they can be employed to enlarge the genetic foundation of the current genotypes. In addition, SSR markers with PIC value of 0.5 or higher considered as to be effective in discriminating the polymorphism rate [
10,
38], and they have a potential in evaluation of genetic divergence. Therefore, when the level of polymorphism of markers was high (50% highly informative markers followed with 43% moderately informative markers), it could indicate that the rice mutants used in this study were more diverse since only the amylose content trait-based DNA marker was used. The other explanation to this might be that the rice mutants have a related origin from quality rice. In previous study, Anumpam et al. [
38] found that the genetic diversity value was low when using only SSR markers related to drought trait.
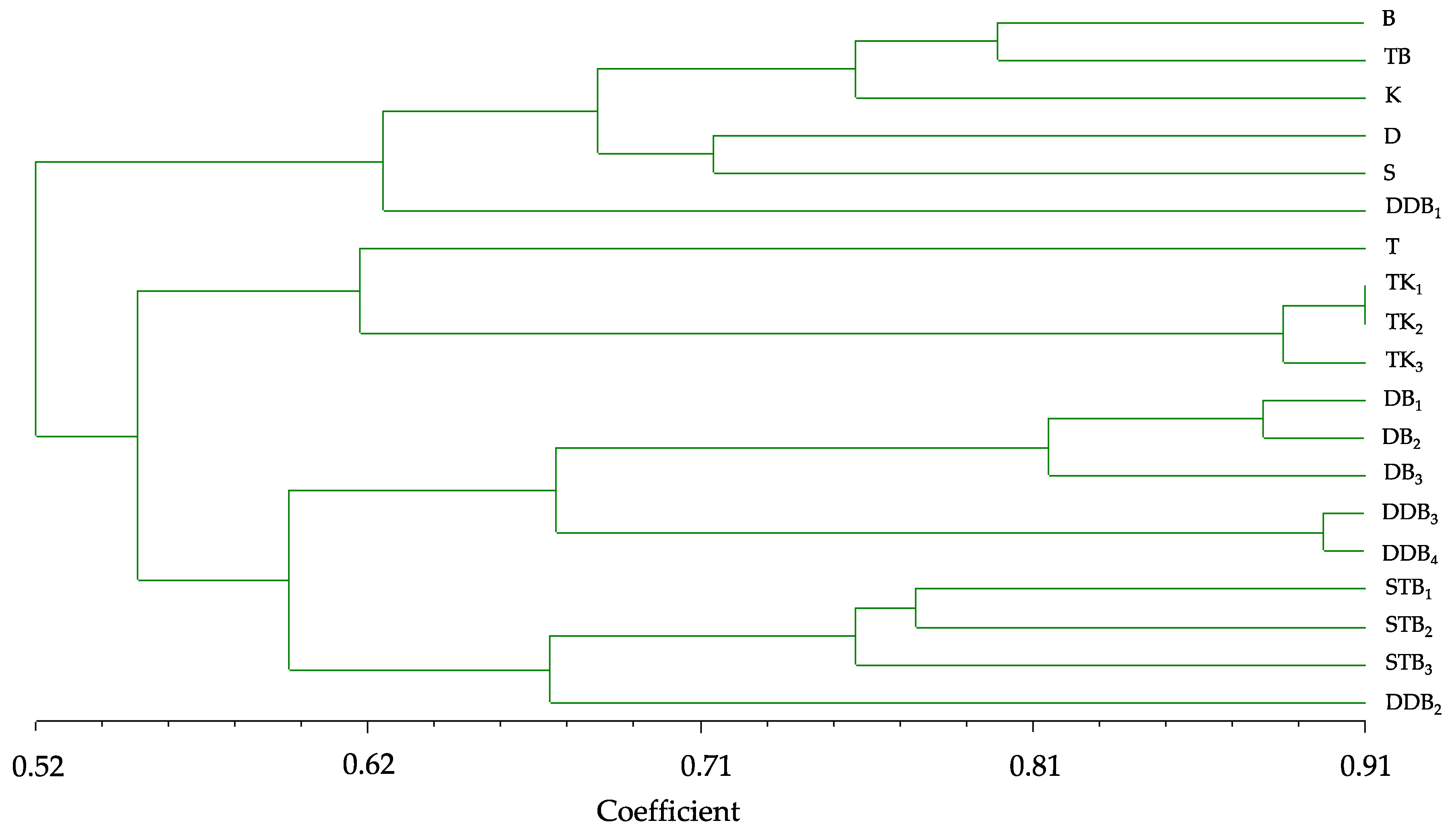
The UPGMA clustering analysis delineated all 19 genotypes in four groups with difference genotypes indicating the greatness of the genetic variation among the rice mutant genotypes. Cluster I represented six mutant genotypes originated from varieties and mutant lines, and the genetic similarity of this cluster was the lowest. It was significant that these genotypes were resolved into this cluster because they were originated from different rice cultivars. Some lines were generated from a common genotype, and they obviously grouped together, such as cluster II. TK1, TK2, and TK3 were originated from T and K varieties, and they grouped with T into this cluster. Interestingly, although T and K were parents of these lines, these lines clustered together with T (female parent) and separated from K (male parent); K was in cluster I, and the highest genetic similarity was found in this cluster. Although generated from a common genotype, some rice genotypes did not cluster together for instance DDB1, DDB2, DDB3, and DDB4; they clustered into different clusters while they have the same origin. Originated from D and B, DB1, DB2, and DB3 were belong to cluster II; however, their parents belong to cluster I. This result was similar STB1, STB2, and STB3. These results could be reasoned that, mutation may change genetic materials, and these changes lead to the genetic differentiation among rice mutants.
Aside from alleles obtained from polymorphism SSR markers, the differentiation within groups and among groups of nineteen rice genotypes was estimated. Based on the result of UPGMA clustering method, mutant genotypes and their parents were grouped into four groups with significant divergence among group (
p > 0.001). By using AMOVA analysis, high genetic diversity within groups was reveal (68%), whereas, among groups, genetic diversity was only 30%. There was a significant genetic variation among rice mutants. Mao-bai et al. [
40] previously referred that it may be due to farming system alteration. This can be explained that extensive cultivation may have contributed to the occurrence of rice mutants. Although the rice mutants used in this study were selected randomly, further investigation should be conducted on the other mutant lines to understand the influence of MNU mutation on the phenotypes and genotypes correlated to amylose content in rice. Findings of this study are helpful for controlling the amylose content in rice breeding, and for widening the understanding of MNU mutation to rice quality.