4SC-202 as a Potential Treatment for the Pediatric Brain Tumor Medulloblastoma
Abstract
:1. Introduction
2. Materials and Methods
2.1. Cell Culture, Cell Viability Assays, and Immunocytochemistry
2.2. Spheroid Culture
2.3. Microarray Analysis
3. Results
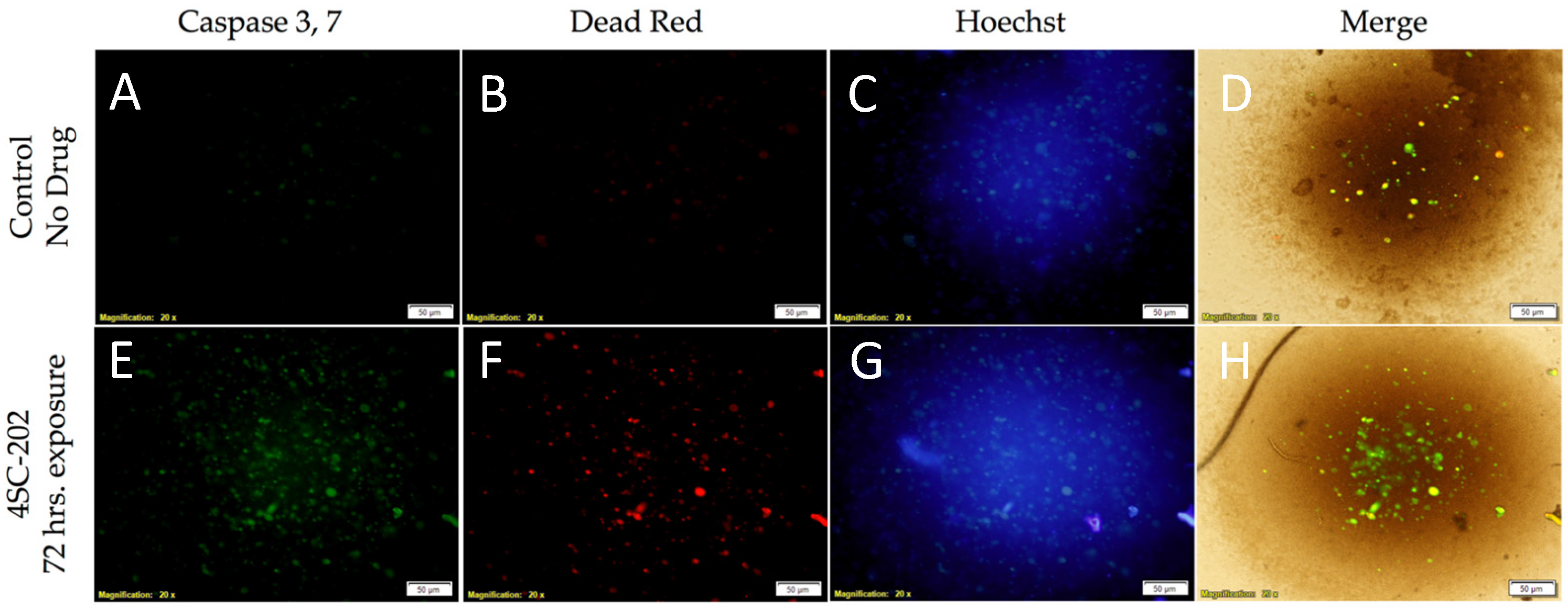
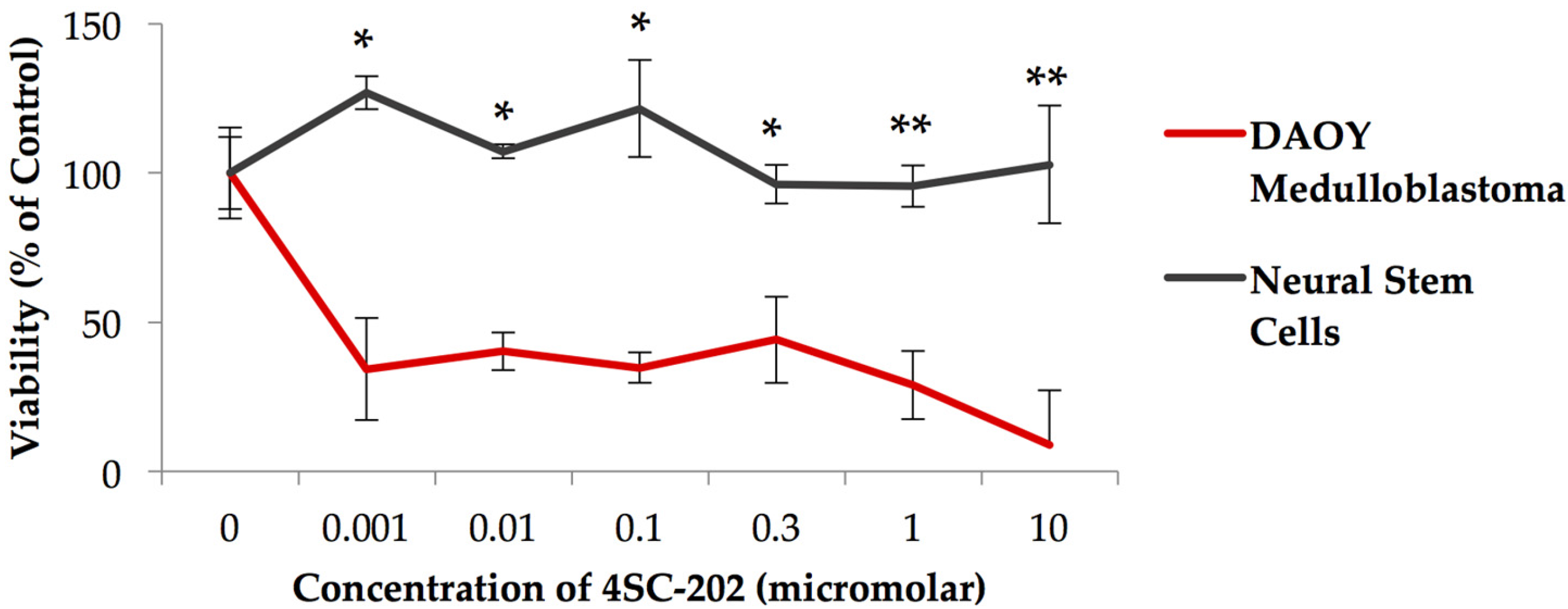
3.1. 4SC-202 Is Cytotoxic to Medulloblastoma in Cell Culture
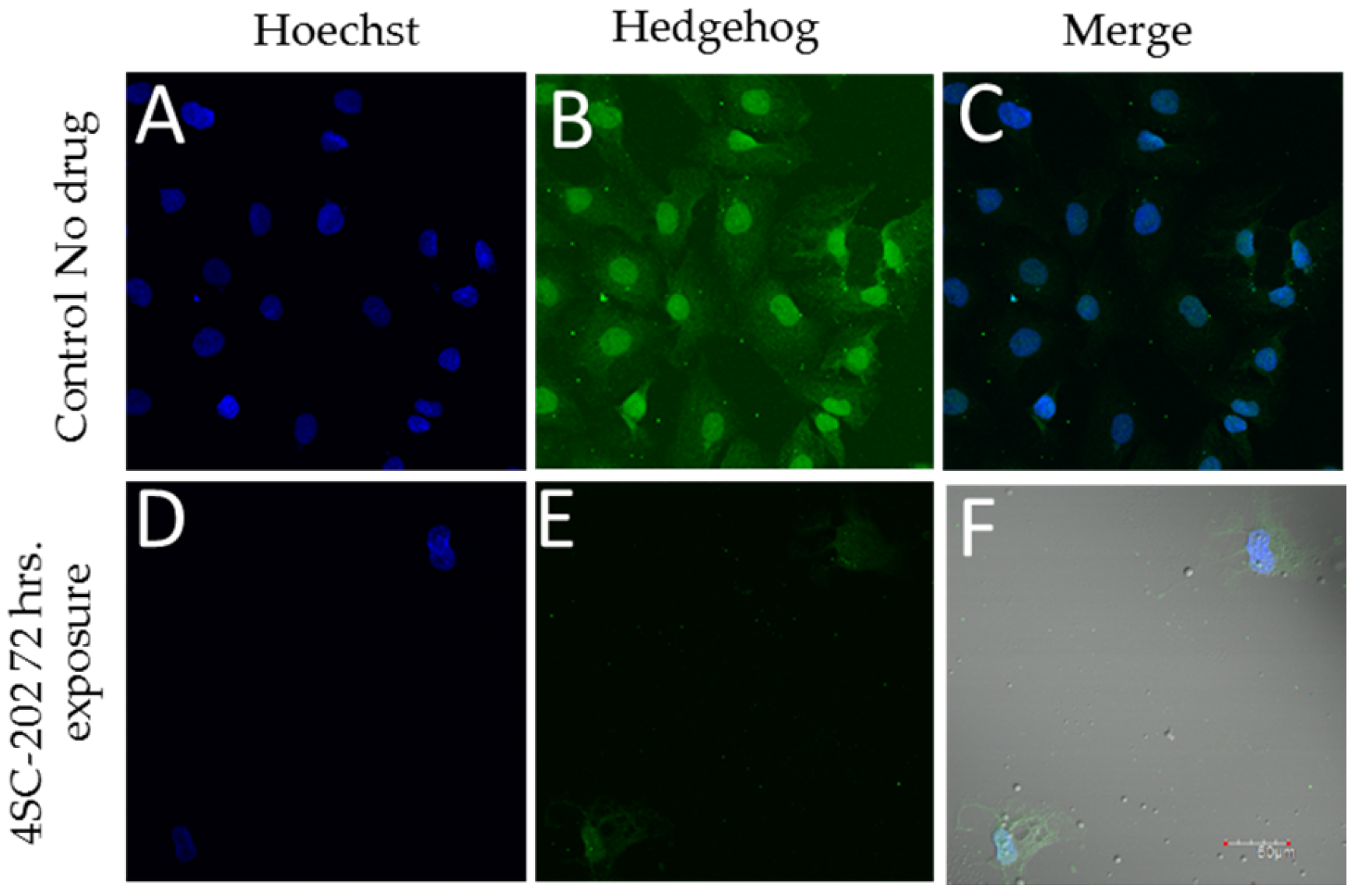
3.2. 4SC-202 Inhibits Hedgehog Immunoreactivity
3.3. Microarray Analysis of Human Medulloblastoma Tumors
4. Discussion
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Reith, W.; Bodea, S.; Mühl-Benninghaus, R. Pediatric brain tumors. Radiologe 2017. [Google Scholar] [CrossRef]
- Chakravadhanula, M.; Hampton, C.N.; Chodavadia, P.; Ozols, V.; Zhou, L.; Catchpoole, D.; Xu, J.; Erdreich-Epstein, A.; Bhardwaj, R.D. Wnt pathway in atypical teratoid rhabdoid tumors. Neuro. Oncol. 2014, 17, 526–535. [Google Scholar] [CrossRef] [PubMed]
- Messerli, S.M.; Hoffman, M.M.; Gnimpieba, E.Z.; Bhardwaj, R.D. Therapeutic Targeting of PTK7 is Cytotoxic in Atypical Teratoid Rhabdoid Tumors. Mol. Cancer Res. 2017, 15, 973–983. [Google Scholar] [CrossRef] [PubMed]
- Staedtke, V.; Dzaye, O.D.A.; Holdhoff, M. Actionable Molecular Biomarkers in Primary Brain Tumors. Trends Cancer 2016, 2, 338–349. [Google Scholar] [CrossRef] [PubMed]
- Kohlhof, H.; Gruber, W.; Vitt, D.; Aberger, F.; Prenzel, T. Abstract C89: The small molecule inhibitor 4SC-202 controls aberrant HH signaling in cancer. Mol. Cancer Ther. 2015, 14. [Google Scholar] [CrossRef]
- Pinkerneil, M.; Hoffmann, M.J.; Kohlhof, H.; Schulz, W.A.; Niegisch, G. Evaluation of the Therapeutic Potential of the Novel Isotype Specific HDAC Inhibitor 4SC-202 in Urothelial Carcinoma Cell Lines. Target. Oncol. 2016, 11, 783–798. [Google Scholar] [CrossRef] [PubMed]
- Henning, S.W.; Doblhofer, R.; Kohlhof, H.; Jankowsky, R.; Maier, T.; Beckers, T.; Schmidt, M.; Hentsch, B. Preclinical characterization of 4SC-202, a novel isotype specific HDAC inhibitor. EJC Suppl. 2010, 8, 61. [Google Scholar] [CrossRef]
- Huang, Z.; Wang, S.; Min, H.; Li, J.; Qin, L.; Li, D. Pre-clinical characterization of 4SC-202, a novel class I HDAC inhibitor, against colorectal cancer cells. Tumor Biol. 2016, 37, 10257–10267. [Google Scholar]
- Von Tresckow, B.; Gundermann, S.; Eichenauer, D.A.; Aulitzky, W.E.; Göbeler, M.; Sayehli, C.; Bacchus, L.; Hauns, B.; Mais, A.; Hentsch, B.; et al. First-in-human study of 4SC-202, a novel oral HDAC inhibitor in advanced hematologic malignancies (TOPAS study). J. Clin. Oncol. 2014, 32, 8559. [Google Scholar]
- Elia, N.; Lippincott-Schwartz, J. Culturing MDCK cells in three dimensions for analyzing intracellular dynamics. Curr. Protoc. Cell Biol. 2009, 43, 1–18. [Google Scholar]
- Griesinger, A.M.; Josephson, R.J.; Donson, A.M.; Mulcahy Levy, J.M.; Amani, V.; Birks, D.K.; Hoffman, L.M.; Furtek, S.L.; Reigan, P.; Handler, M.H.; et al. Interleukin-6/STAT3 Pathway Signaling Drives an Inflammatory Phenotype in Group A Ependymoma. Cancer Immunol. Res. 2015, 3, 1165–1174. [Google Scholar] [CrossRef] [PubMed]
- Birks, D.K.; Donson, A.M.; Patel, P.R.; Sufit, A.; Algar, E.M.; Dunham, C.; Kleinschmidt-Demasters, B.K.; Handler, M.H.; Vibhakar, R.; Foreman, N.K. Pediatric rhabdoid tumors of kidney and brain show many differences in gene expression but share dysregulation of cell cycle and epigenetic effector genes. Pediatr. Blood Cancer 2013, 60, 1095–1102. [Google Scholar] [CrossRef] [PubMed]
- Smyth, G.K. Limma: Linear models for microarray data. In Bioinformatics and Computational Biology Solutions Using R and Bioconductor; Gentleman, R.C., Carey, V., Dudoit, S., Irizarry, R., Huber, W., Eds.; Springer: New York, NY, USA, 2005; pp. 397–420. [Google Scholar]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. B 1995, 57, 289–300. [Google Scholar]
- Pan, Y.; Bai, C.B.; Joyner, A.L.; Wang, B. Sonic hedgehog Signaling Regulates Gli2 Transcriptional Activity by Suppressing Its Processing and Degradation Sonic hedgehog Signaling Regulates Gli2 Transcriptional Activity by Suppressing Its Processing and Degradation. Mol. Cell. Biol. 2006, 26, 3365–3377. [Google Scholar] [CrossRef] [PubMed]
- Coni, S.; Mancuso, A.B.; Di Magno, L.; Sdruscia, G.; Manni, S.; Serrao, S.M.; Rotili, D.; Spiombi, E.; Bufalieri, F.; Petroni, M.; et al. Selective targeting of HDAC1/2 elicits anticancer effects through Gli1 acetylation in preclinical models of SHH Medulloblastoma. Sci. Rep. 2017, 7, 44079. [Google Scholar] [CrossRef] [PubMed]
- Mishra, V.K.; Wegwitz, F.; Kosinsky, R.L.; Sen, M.; Baumgartner, R.; Wulff, T.; Siveke, J.T.; Schildhaus, H.-U.; Najafova, Z.; Kari, V. Histone deacetylase class-I inhibition promotes epithelial gene expression in pancreatic cancer cells in a BRD4-and MYC-dependent manner. Nucleic Acids Res. 2017, 45, 6334–6349. [Google Scholar] [CrossRef] [PubMed]
- Javelaud, D.; Alexaki, V.I.; Dennler, S.; Mohammad, K.S.; Guise, T.A.; Mauviel, A. TGF-β/SMAD/GLI2 signaling axis in cancer progression and metastasis. Cancer Res. 2011, 71, 5606–5610. [Google Scholar] [CrossRef] [PubMed]
- Aref, D.; Moffatt, C.J.; Agnihotri, S.; Ramaswamy, V.; Dubuc, A.M.; Northcott, P.A.; Taylor, M.D.; Perry, A.; Olson, J.M.; Eberhart, C.G.; et al. Canonical TGF-β pathway activity is a predictor of SHH-driven medulloblastoma survival and delineates putative precursors in cerebellar development. Brain Pathol. 2013, 23, 178–191. [Google Scholar] [CrossRef] [PubMed]
- Gate, D.; Danielpour, M.; Rodriguez, J.; Kim, G.-B.; Levy, R.; Bannykh, S.; Breunig, J.J.; Kaech, S.M.; Flavell, R.A.; Town, T. T-cell TGF-signaling abrogation restricts medulloblastoma progression. Proc. Natl. Acad. Sci. USA 2014, 111, E3458–E3466. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.Y.; Yang, J.Y. Targeting the hedgehog pathway in pediatric medulloblastoma. Cancers 2015, 7, 2110–2123. [Google Scholar] [CrossRef] [PubMed] [Green Version]


© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Messerli, S.M.; Hoffman, M.M.; Gnimpieba, E.Z.; Kohlhof, H.; Bhardwaj, R.D. 4SC-202 as a Potential Treatment for the Pediatric Brain Tumor Medulloblastoma. Brain Sci. 2017, 7, 147. https://doi.org/10.3390/brainsci7110147
Messerli SM, Hoffman MM, Gnimpieba EZ, Kohlhof H, Bhardwaj RD. 4SC-202 as a Potential Treatment for the Pediatric Brain Tumor Medulloblastoma. Brain Sciences. 2017; 7(11):147. https://doi.org/10.3390/brainsci7110147
Chicago/Turabian StyleMesserli, Shanta M., Mariah M. Hoffman, Etienne Z. Gnimpieba, Hella Kohlhof, and Ratan D. Bhardwaj. 2017. "4SC-202 as a Potential Treatment for the Pediatric Brain Tumor Medulloblastoma" Brain Sciences 7, no. 11: 147. https://doi.org/10.3390/brainsci7110147
APA StyleMesserli, S. M., Hoffman, M. M., Gnimpieba, E. Z., Kohlhof, H., & Bhardwaj, R. D. (2017). 4SC-202 as a Potential Treatment for the Pediatric Brain Tumor Medulloblastoma. Brain Sciences, 7(11), 147. https://doi.org/10.3390/brainsci7110147



