Constructive Approaches for Understanding the Origin of Self-Replication and Evolution
Abstract
:1. Introduction
2. What Is the Constructive Approach?
3. Construction of Self-Replicating Systems
4. Construction of the Ability to Evolve
- Replication
- Inheritable variation
- Selection
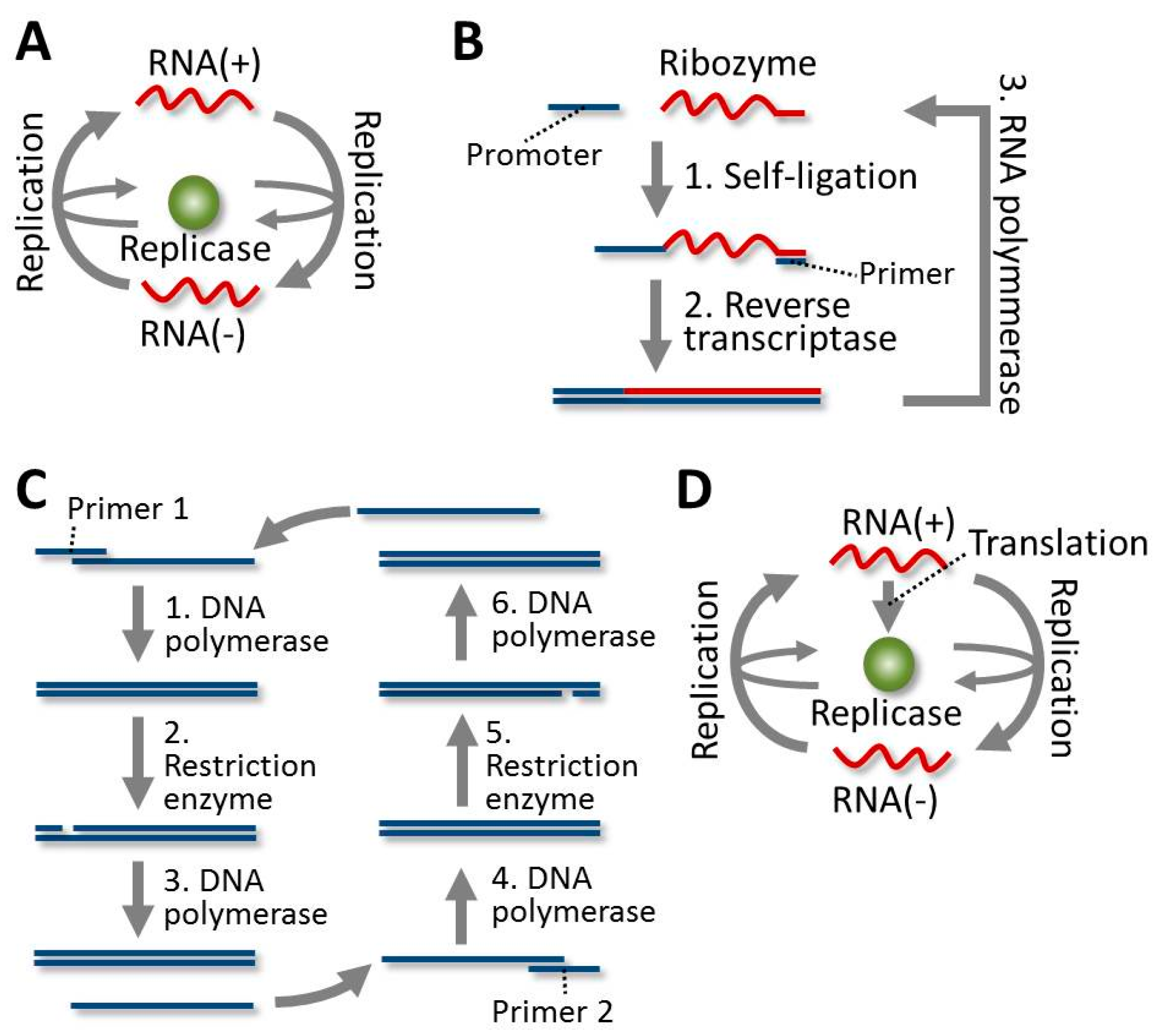
4.1. Evolution of the RNA in Spiegelman’s RNA System
4.2. Evolution of the RNA in the 3SR System
4.3. Evolution of the DNA in the SDA System
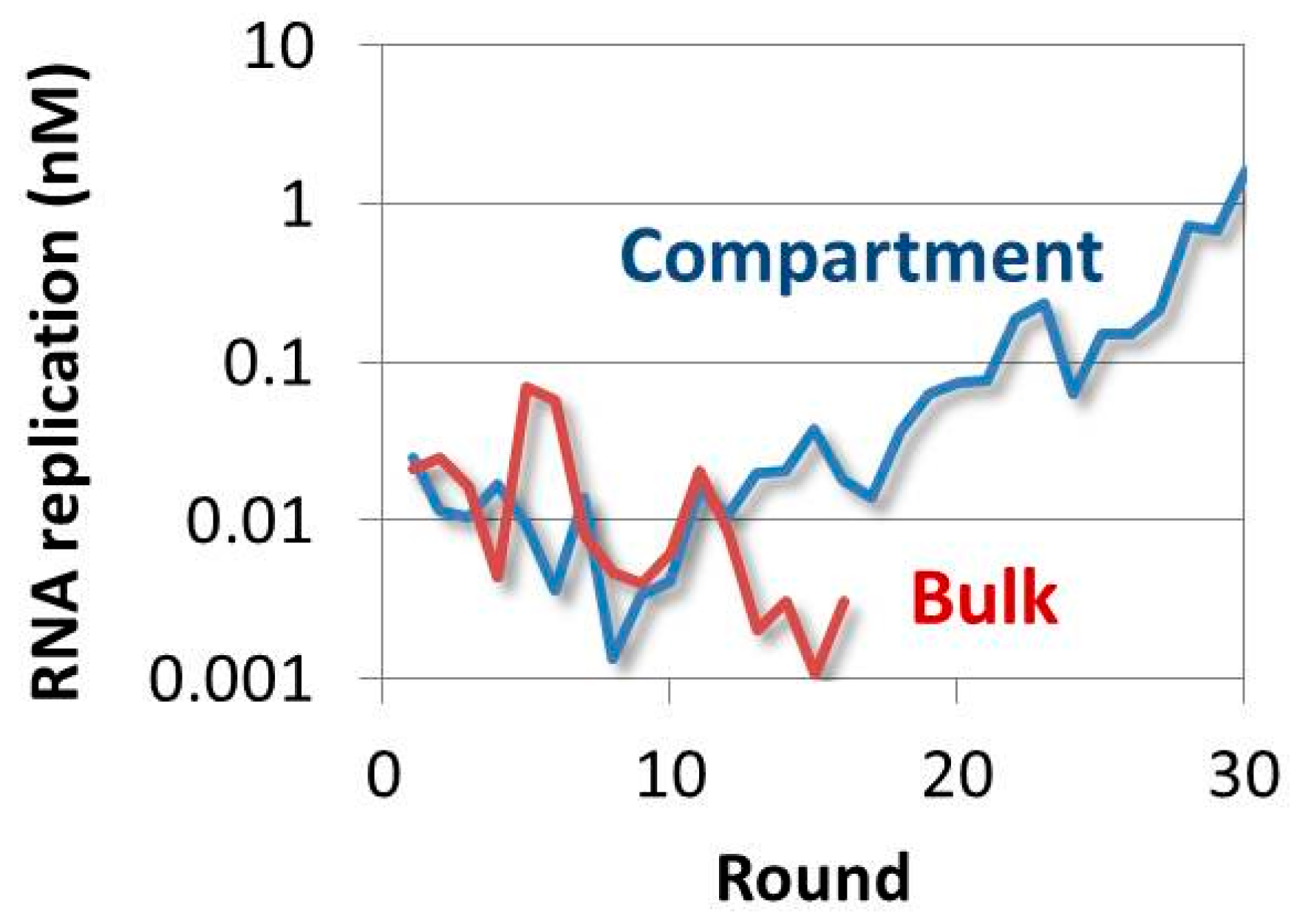
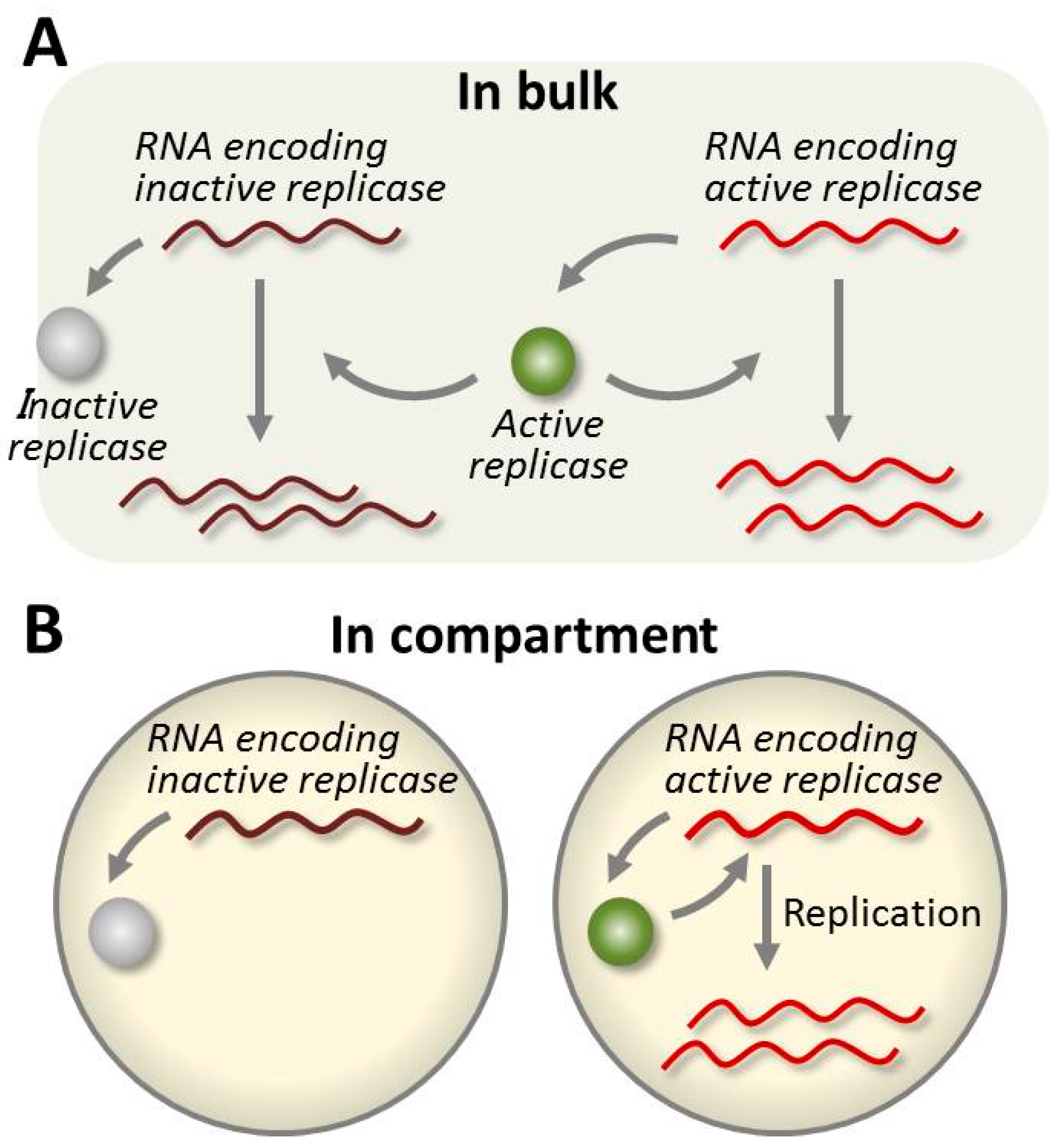
4.4. Evolution of the RNA in the TcRR System and the Requirement of a Compartment
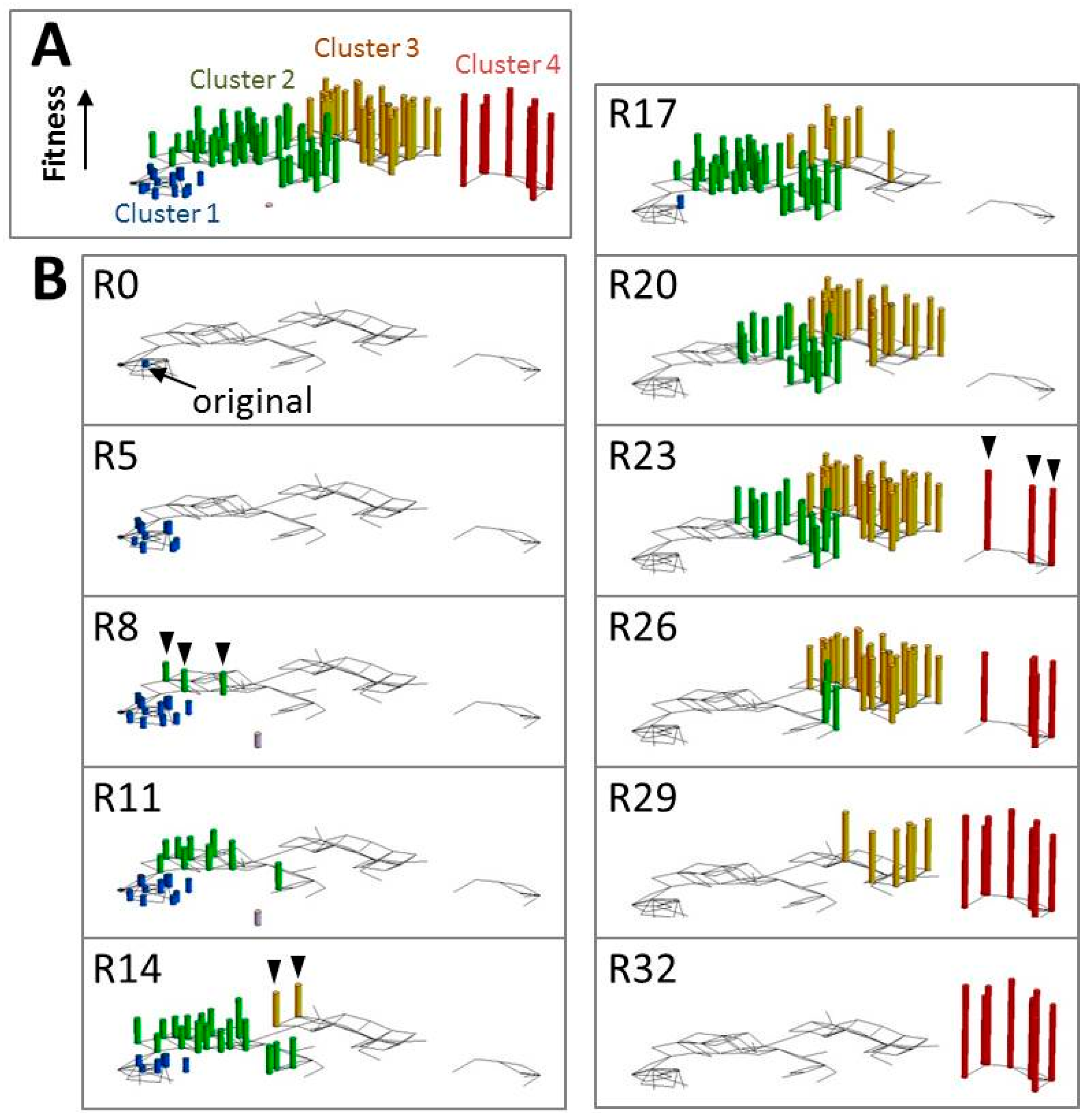
5. Analysis of the Evolutionary Process Using an in Vitro Self-Replicating System
6. Lessons from the Constructive Approach
7. Future Directions
Acknowledgments
Conflicts of Interest
References
- Gesteland, T.R.C.R.; Atkins, J.F. RNA World: The Nature of Modern RNA Suggests a Prebiotic RNA World; Cold Spring Harbor Laboratory Press: New York, NY, USA, 2005. [Google Scholar]
- Ruiz-Mirazo, K.; Briones, C.; de la Escosura, A. Prebiotic systems chemistry: New perspectives for the origins of life. Chem. Rev. 2014, 114, 285–366. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P.; Szostak, J.W. Isolation of New Ribozymes from a Large Pool of Random Sequences. Science 1993, 261, 1411–1418. [Google Scholar] [CrossRef] [PubMed]
- Keefe, A.D.; Szostak, J.W. Functional proteins from a random-sequence library. Nature 2001, 410, 715–718. [Google Scholar] [CrossRef] [PubMed]
- Yamauchi, A.; Nakashima, T.; Tokuriki, N.; Hosokawa, M.; Nogami, H.; Arioka, S.; Urabe, I.; Yomo, T. Evolvability of random polypeptides through functional selection within a small library. Protein Eng. 2002, 15, 619–626. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, Y.; Aita, T.; Toyota, H.; Husimi, Y.; Urabe, I.; Yomo, T. PLoS ONE 2006, 1, e96. [PubMed]
- Toyota, H.; Hosokawa, M.; Urabe, I.; Yomo, T. Emergence of Polyproline II-Like Structure at Early Stages of Experimental Evolution from Random Polypeptides. Mol. Biol. Evol. 2008, 25, 1113–1119. [Google Scholar] [CrossRef] [PubMed]
- Dix, D.E. What Is Life? Prerequisites for a Definition. Yale J. Biol. Med. 2002, 75, 313–321. [Google Scholar] [PubMed]
- Ruiz-Mirazo, K.; Pereto, J.; Moreno, A. Defining life or bringing biology to life. Origins Life Evol. B 2010, 40, 203–213. [Google Scholar] [CrossRef] [PubMed]
- Kaneko, K. Life: An Introduction to Complex Systems Biology; Springer: Berlin, Germany, 2006. [Google Scholar]
- Ichihashi, N.; Matsuura, T.; Kita, H.; Sunami, T.; Suzuki, H.; Yomo, T. Cold Spring Harb. Perspect. Biol. 2010, 2, a004945.
- Oberholzer, T.; Wick, R.; Luisi, P.L.; Biebricher, C.K. Enzymatic RNA replication in self-reproducing vesicles: an approach to a minimal cell. Biochem. Biophys. Res. Commun. 1995, 207, 250–257. [Google Scholar] [CrossRef] [PubMed]
- Luisi, P.L.; Ferri, F.; Stano, P. Approaches to semi-synthetic minimal cells: a review. Naturwissenschaften 2006, 93, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Stano, P.; Carrara, P.; Kuruma, Y.; de Souza, T.P.; Luisi, P.L. Compartmentalized reactions as a case of soft-matter biotechnology: synthesis of proteins and nucleic acids inside lipid vesicles. J. Mater. Chem. 2011, 21, 18887–18902. [Google Scholar] [CrossRef]
- Mills, D.R.; Peterson, R.L.; Spiegelman, S. An extracellular Darwinian experiment with a self-duplicating nucleic acid molecule. Proc. Natl. Acad. Sci. USA 1967, 58, 217–224. [Google Scholar] [CrossRef] [PubMed]
- Guatelli, J.C.; Whitfield, K.M.; Kwoh, D.Y.; Barringer, K.J.; Richman, D.D.; Gingeras, T.R. Isothermal, in vitro amplification of nucleic acids by a multienzyme reaction modeled after retroviral replication. Proc. Natl. Acad. Sci. USA 1990, 87, 1874–1878. [Google Scholar] [CrossRef] [PubMed]
- Breaker, R.R.; Joyce, G.F. Emergence of a replicating species from an in vitro RNA evolution reaction. Proc. Natl. Acad. Sci. USA 1994, 91, 6093–6097. [Google Scholar] [CrossRef] [PubMed]
- Walker, G.T.; Little, M.C.; Nadeau, J.G.; Shank, D.D. Isothermal in vitro amplification of DNA by a restriction enzyme/DNA polymerase system. Proc. Natl. Acad. Sci. USA 1992, 89, 392–396. [Google Scholar] [CrossRef] [PubMed]
- Kita, H.; Matsuura, T.; Sunami, T.; Hosoda, K.; Ichihashi, N.; Tsukada, K.; Urabe, I.; Yomo, T. Replication of Genetic Information with Self-Encoded Replicase in Liposomes. Chembiochem 2008, 9, 2403–2410. [Google Scholar] [CrossRef] [PubMed]
- Ichihashi, N.; Matsuura, T.; Kita, H.; Hosoda, K.; Sunami, T.; Tsukada, K.; Yomo, T. Importance of Translation–Replication Balance for Efficient Replication by the Self-Encoded Replicase. Chembiochem 2008, 9, 3023–3028. [Google Scholar] [CrossRef] [PubMed]
- Lincoln, T.A.; Joyce, G.F. Self-sustained replication of an RNA enzyme. Science 2009, 323, 1229–1232. [Google Scholar] [CrossRef] [PubMed]
- Vaidya, N.; Manapat, M.L.; Chen, I.A.; Xulvi-Brunet, R.; Hayden, E.J.; Lehman, N. Spontaneous network formation among cooperative RNA replicators. Nature 2012, 491, 72–77. [Google Scholar] [CrossRef] [PubMed]
- Zaher, H.S.; Unrau, P.J. Selection of an improved RNA polymerase ribozyme with superior extension and fidelity. RNA 2007, 13, 1017–1026. [Google Scholar] [CrossRef] [PubMed]
- Attwater, J.; Wochner, A.; Holliger, P. In-ice evolution of RNA polymerase ribozyme activity. Nat. Chem. 2013, 5, 1011–1018. [Google Scholar] [CrossRef] [PubMed]
- Higgs, P.G.; Lehman, N. The RNA World: molecular cooperation at the origins of life. Nat. Rev. Genet. 2015, 16, 7–17. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.H.; Granja, J.R.; Martinez, J.A.; Severin, K.; Ghadri, M.R. A self-replicating peptide. Nature 1996, 382, 525–528. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.H.; Severin, K.; Yokobayashi, Y.; Ghadiri, M.R. Emergence of symbiosis in peptide self-replication through a hypercyclic network. Nature 1997, 390, 591–594. [Google Scholar] [PubMed]
- Wick, R.; Walde, P.; Luisi, P.L. Light microscopic investigations of the autocatalytic self-reproduction of giant vesicles. J. Am. Chem. Soc. 1995, 117, 1435–1436. [Google Scholar] [CrossRef]
- Takakura, K.; Toyota, T.; Sugawara, T. A Novel System of Self-Reproducing Giant Vesicles. J. Am. Chem. Soc. 2003, 125, 8134–8140. [Google Scholar] [CrossRef] [PubMed]
- Hardya, M.D.; Yanga, J.; Selimkhanovb, J.; Colea, C.M.; Tsimringb, L.S.; Devaraj, N.K. Self-reproducing catalyst drives repeated phospholipid synthesis and membrane growth. Proc. Natl. Acad. Sci. USA 2015, 112, 8187–8192. [Google Scholar] [CrossRef] [PubMed]
- Kurihara, K.; Tamura, M.; Shohda, K.; Toyota, T.; Suzuki, K.; Sugawara, T. Self-reproduction of supramolecular giant vesicles combined with the amplification of encapsulated DNA. Nat. Chem. 2011, 3, 775–781. [Google Scholar] [CrossRef] [PubMed]
- Zielinski, W.S.; Orgel, L.E. Autocatalytic synthesis of a tetranucleotide analogue. Nature 1987, 327, 346–347. [Google Scholar] [CrossRef] [PubMed]
- Tjivikua, T.; Ballester, P.; Rebek, J. Convergent functional groups. VIII. Flexible model receptors for adenine derivatives. J. Am. Chem. Soc. 1990, 112, 8408–8414. [Google Scholar] [CrossRef]
- Li, T.; Nicolaou, K.C. Chemical self-replication of palindromic duplex DNA. Nature 1994, 369, 218–221. [Google Scholar] [CrossRef] [PubMed]
- Sievers, D.; von Kiedrowski, G. Self-replication of complementary nucleotide-based oligomers. Nature 1994, 369, 221–224. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.M. The Problems of Biology; Oxford University Press: Oxford, UK, 1986. [Google Scholar]
- Oliphant, A.R.; Brandl, C.J.; Struhl, K. Defining the sequence specificity of DNA-binding proteins by selecting binding sites from random-sequence oligonucleotides: analysis of yeast GCN4 protein. Mol. Cell. Biol. 1989, 9, 2944–2949. [Google Scholar] [CrossRef] [PubMed]
- Joyce, G.F. Amplification, mutation and selection of catalytic RNA. Gene 1989, 82, 83–87. [Google Scholar] [CrossRef]
- Tuerk, C.; Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 1990, 249, 505–510. [Google Scholar] [CrossRef] [PubMed]
- Ellington, A.D.; Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 1990, 346, 818–822. [Google Scholar] [CrossRef] [PubMed]
- Chetverin, A.B.; Chetverina, H.V.; Demidenko, A.A.; Ugarov, V.I. Nonhomologous RNA Recombination in a Cell-Free System: Evidence for a Transesterification Mechanism Guided by Secondary Structure. Cell 1997, 88, 503–513. [Google Scholar] [CrossRef]
- Kramer, F.R.; Mills, D.R.; Cole, P.E.; Nishihara, T.; Spiegelman, S. Evolution in vitro: Sequence and phenotype of a mutant RNA resistant to ethidium bromide. J. Mol. Biol. 1974, 89, 719–736. [Google Scholar] [CrossRef]
- Wright, M.C.; Joyce, G.F. Continuous in Vitro Evolution of Catalytic Function. Science 1997, 276, 614–617. [Google Scholar] [CrossRef] [PubMed]
- Walter, N.G.; Strunk, G. Strand displacement amplification as an in vitro model for rolling-circle replication: deletion formation and evolution during serial transfer. Proc. Natl. Acad. Sci. USA 1994, 91, 7937–7941. [Google Scholar] [CrossRef] [PubMed]
- Ichihashi, N.; Usui, K.; Kazuta, Y.; Sunami, T.; Matsuura, T.; Yomo, T. Darwinian evolution in a translation-coupled RNA replication system within a cell-like compartment. Nat. Commun. 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Mizuuchi, R.; Ichihashi, N.; Usui, K.; Kazuta, Y.; Yomo, T. Adaptive Evolution of an Artificial RNA Genome to a Reduced Ribosome Environment. ACS Synth. Biol. 2014, 4, 292–298. [Google Scholar] [CrossRef] [PubMed]
- Bansho, Y.; Ichihashi, N.; Kazuta, Y.; Matsuura, T.; Suzuki, H.; Yomo, T. Importance of Parasite RNA Species Repression for Prolonged Translation-Coupled RNA Self-Replication. Chem. Biol. 2012, 19, 478–487. [Google Scholar] [CrossRef] [PubMed]
- Ichihashi, N.; Yomo, T. Positive roles of compartmentalization in internal reactions. Importance of Parasite RNA Species Repression for Prolonged Translation-Coupled RNA Self-Replication. Curr. Opin. Chem. Biol. 2014, 22, 12–17. [Google Scholar] [CrossRef] [PubMed]
- Wright, S. The roles of mutation, inbreeding, crossbreeding, and selection in evolution. In Proceedings of the Sixth International Congress on Genetics, Ithaca, NY, USA, 1932; Volume 1, pp. 355–366.
- de Visser, J.A.; Krug, J. Empirical fitness landscapes and the predictability of evolution. Nat. Rev. Genet. 2014, 15, 480–490. [Google Scholar] [CrossRef] [PubMed]
- Kun, Á.; Szathmáry, E. Fitness Landscapes of Functional RNAs. Life 2015, 5, 1497–1517. [Google Scholar] [CrossRef] [PubMed]
- Ichihashi, N.; Aita, T.; Motooka, D.; Nakamura, S.; Yomo, T. Periodic Pattern of Genetic and Fitness Diversity during Evolution of an Artificial Cell-Like System. Mol. Biol. Evol. 2015, 32, 3205–3214. [Google Scholar] [CrossRef] [PubMed]
- Gerrish, P.J.; Lenski, R.E. The fate of competing beneficial mutations in an asexual population. Genetica 1998, 127, 102–103. [Google Scholar]
- Darwin, C. On the Origin of Species by Means of Natural Selection; John Murray: London, UK, 1859. [Google Scholar]
- Walde, P.; Goto, A.; Monnard, P.A.; Wessicken, M.; Luisi, P.L. Oparin’s Reactions Revisited: Enzymic Synthesis of Poly(adenylic acid) in Micelles and Self-Reproducing Vesicles. J. Am. Chem. Soc. 1994, 116, 7541–7547. [Google Scholar] [CrossRef]
- Attwater, J.; Wochner, A.; Pinheiro, V.B.; Coulson, A.; Holliger, P. Ice as a protocellular medium for RNA replication. Nat. Commu. 2010, 1. [Google Scholar] [CrossRef] [PubMed]
- Monnard, P.-A.; Walde, P. Current Ideas about Prebiological Compartmentalization. Life 2015, 5, 1239–1263. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Usui, K.; Ichihashi, N.; Yomo, T. A design principle for a single-stranded RNA genome that replicates with less double-strand formation. Nucleic Acids Res. 2015, 43, 8033–8043. [Google Scholar] [CrossRef] [PubMed]
- Usui, K.; Ichihashi, N.; Kazuta, Y.; Matsuura, T.; Yomo, T. Kinetic model of double-stranded RNA formation during long RNA replication by Qβ replicase. FEBS Lett. 2013, 587, 2565–2571. [Google Scholar] [CrossRef] [PubMed]
- Fujiwara, K.; Katayama, T.; Nomura, S.M. Cooperative working of bacterial chromosome replication proteins generated by a reconstituted protein expression system. Nucleic Acids Res. 2013, 41, 7176–7183. [Google Scholar] [CrossRef] [PubMed]
- Sakatani, Y.; Ichihashi, N.; Kazuta, Y.; Yomo, T. A transcription and translation-coupled DNA replication system using rolling-circle replication. Sci. Rep. 2015, 5. [Google Scholar] [CrossRef] [PubMed]
- Zhu, T.F.; Szostak, J.W. Coupled Growth and Division of Model Protocell Membranes. J. Am. Chem. Soc. 2009, 131, 5705–5713. [Google Scholar] [CrossRef] [PubMed]
- Kuruma, Y.; Stano, P.; Ueda, T.; Luisi, P.L. A synthetic biology approach to the construction of membrane proteins in semi-synthetic minimal cells. Biochim. Biophys. Acta 2009, 1788, 567–574. [Google Scholar] [CrossRef] [PubMed]
- Terasawa, H.; Nishimura, K.; Suzuki, H.; Matsuura, T.; Yomo, T. Coupling of the fusion and budding of giant phospholipid vesicles containing macromolecules. Proc. Natl. Acad. Sci. USA 2012, 109, 5942–5947. [Google Scholar] [CrossRef] [PubMed]
- Murtas, G. Early self-reproduction, the emergence of division mechanisms in protocells. Mol. Biosyst. 2013, 9, 195–204. [Google Scholar] [CrossRef] [PubMed]
- Kobori, S.; Ichihashi, N.; Kazuta, Y.; Yomo, T. A controllable gene expression system in liposomes that includes a positive feedback loop. Mol. Biosyst. 2013, 9, 1282–1285. [Google Scholar] [CrossRef] [PubMed]
- Martini, L.; Mansy, S.S. Cell-like systems with riboswitch controlled gene expression. Chem. Commun. 2011, 47, 10734–10736. [Google Scholar] [CrossRef] [PubMed]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ichihashi, N.; Yomo, T. Constructive Approaches for Understanding the Origin of Self-Replication and Evolution. Life 2016, 6, 26. https://doi.org/10.3390/life6030026
Ichihashi N, Yomo T. Constructive Approaches for Understanding the Origin of Self-Replication and Evolution. Life. 2016; 6(3):26. https://doi.org/10.3390/life6030026
Chicago/Turabian StyleIchihashi, Norikazu, and Tetsuya Yomo. 2016. "Constructive Approaches for Understanding the Origin of Self-Replication and Evolution" Life 6, no. 3: 26. https://doi.org/10.3390/life6030026






