The drnf1 Gene from the Drought-Adapted Cyanobacterium Nostoc flagelliforme Improved Salt Tolerance in Transgenic Synechocystis and Arabidopsis Plant
Abstract
:1. Introduction
2. Materials and Methods
2.1. Organisms and Culture Conditions
2.2. Salt Stress Treatment of Nostoc flagelliforme and drnf1 Transcription
2.3. Transgene in Synechocystis and Salt Stress Treatments
2.4. Assay for Photosynthetic and Respiration Rates
2.5. Exopolysaccharide Determination
2.6. Transcriptional Analysis
2.7. Na+ and K+ Content Determination
2.8. Transgene in Arabidopsis, Salt Stress Treatments, and Phenotypic Observation
3. Results
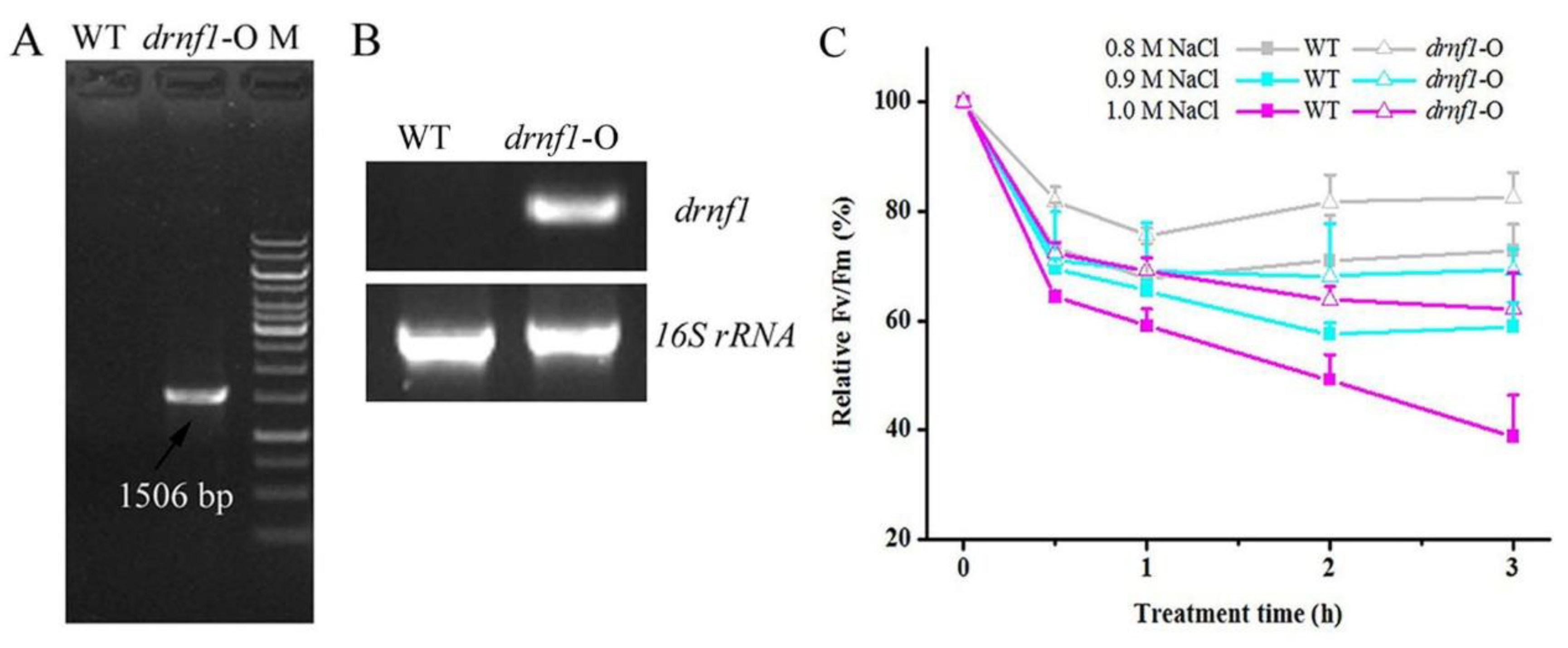
3.1. Transcriptional Induction of drnf1 by Salt Stress in Nostoc flagelliforme
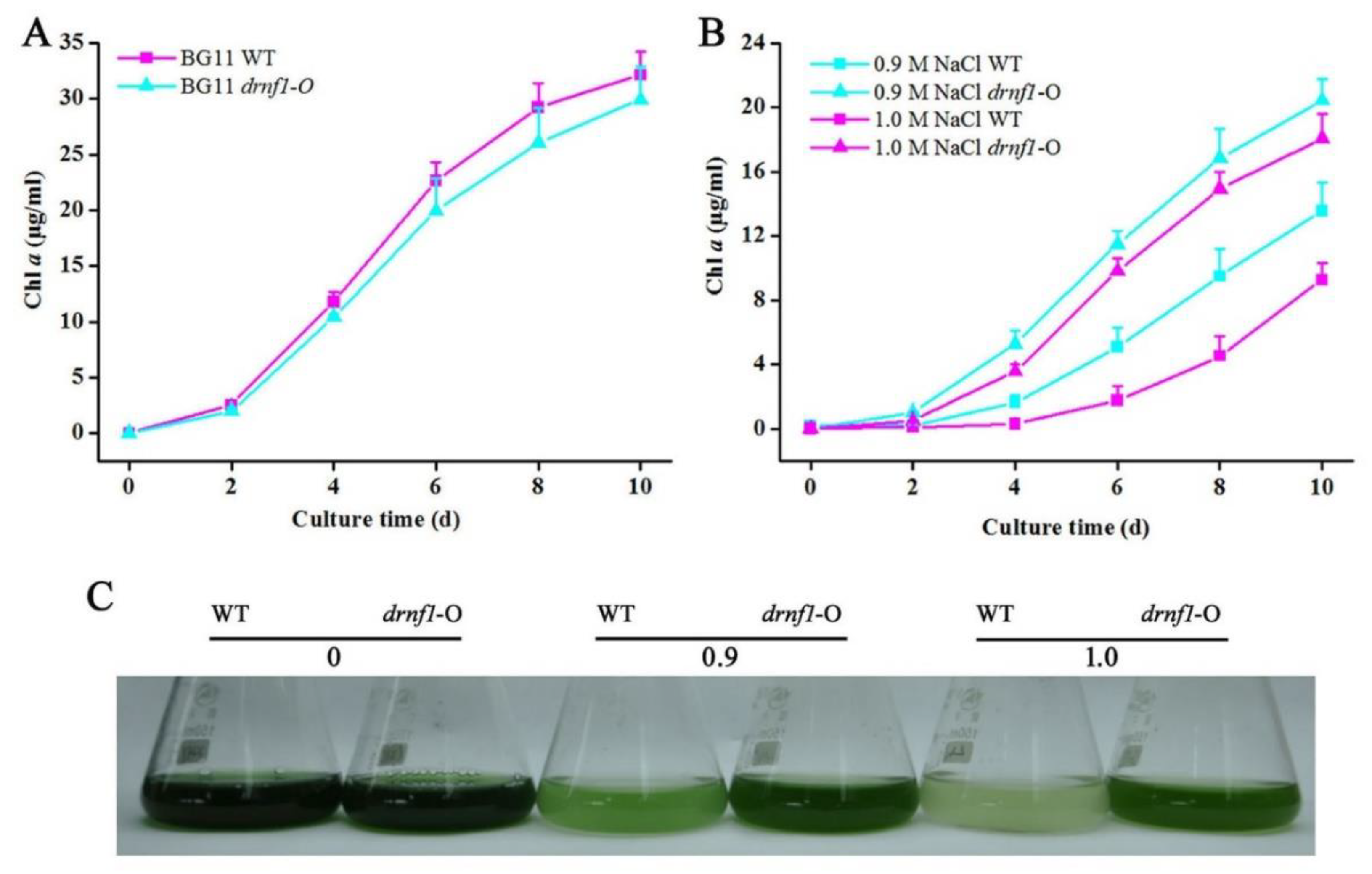
3.2. Increased Salt Tolerance in Transgenic drnf1 Synechocystis
3.3. Photosynthesis and Respiration of Transgenic Synechocystis upon Salt Stress
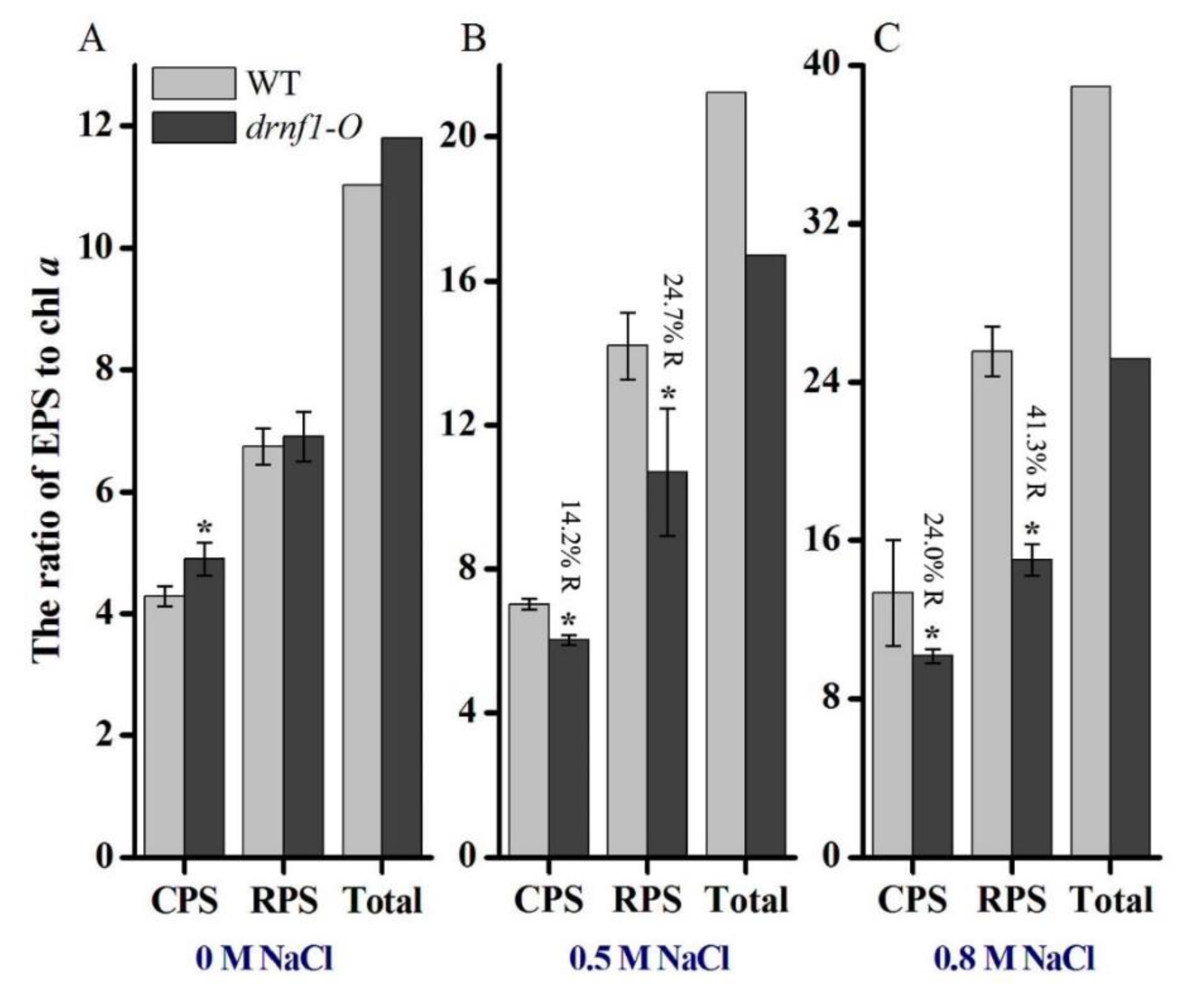
3.4. Exopolysaccharide Production in Transgenic Synechocystis
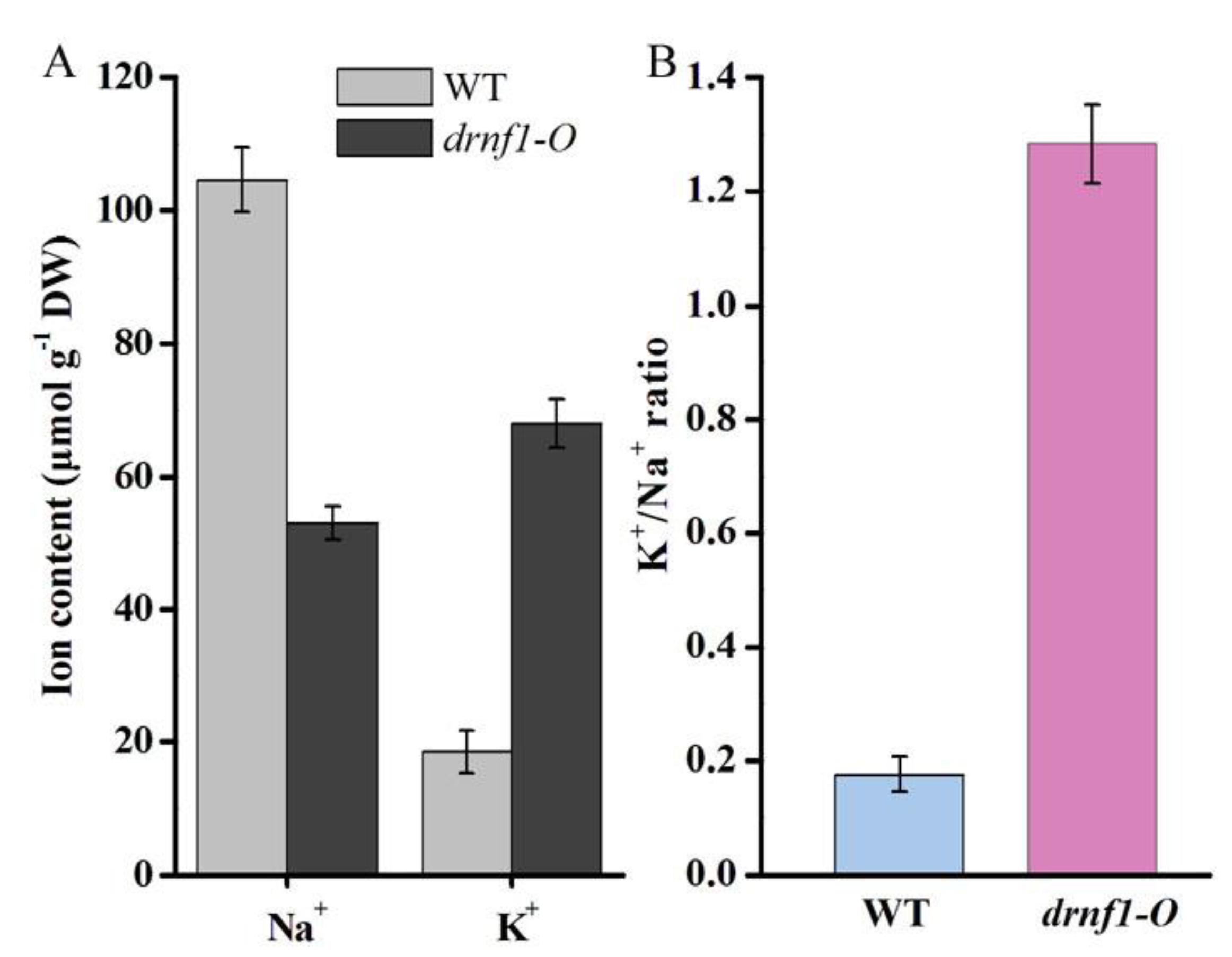
3.5. Transcription, K+ and Na+ Contents in Transgenic Synechocystis
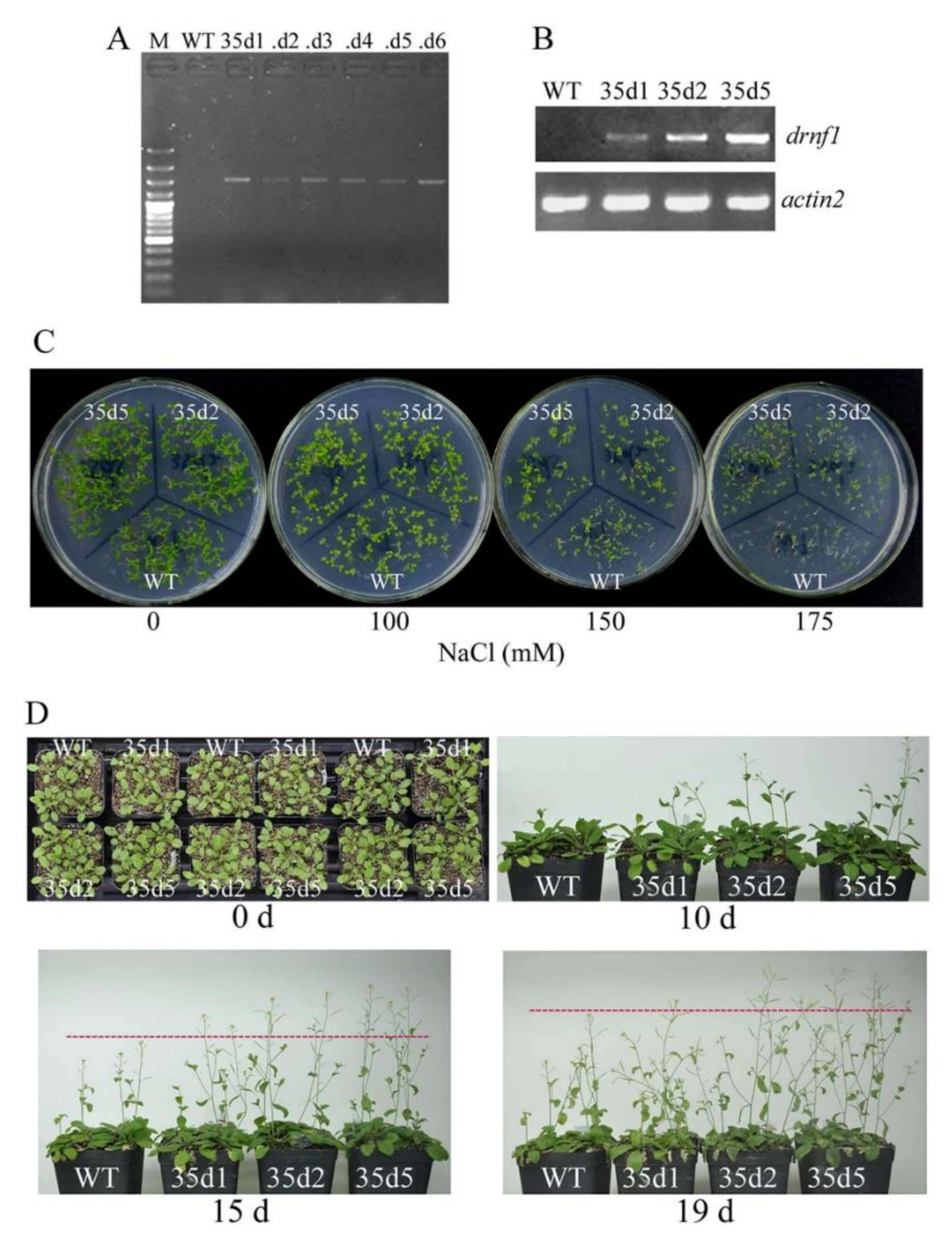
3.6. Increased Salt Tolerance in Transgenic drnf1 Arabidopsis
4. Discussion
Author Contributions
Funding
Conflicts of Interest
References
- Hu, H.; Xiong, L. Genetic engineering and breeding of drought-resistant crops. Annu. Rev. Plant Biol. 2014, 65, 715–741. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.Y.; Wang, H.L.; Shao, H.B.; Tang, X.L. Recent advances in utilizing transcription factors to improve plant abiotic stress tolerance by transgenic technology. Front. Plant Sci. 2016, 7, 67. [Google Scholar] [CrossRef] [PubMed]
- Nouri, M.Z.; Komatsu, S. Subcellular protein overexpression to develop abiotic stress tolerant plants. Front. Plant Sci. 2013, 4, 2. [Google Scholar] [CrossRef] [PubMed]
- Mishra, A.; Tanna, B. Halophytes: Potential resources for salt stress tolerance genes and promoters. Front. Plant Sci. 2017, 8, 829. [Google Scholar] [CrossRef] [PubMed]
- Giarola, V.; Hou, Q.; Bartels, D. Angiosperm plant desiccation tolerance: Hints from transcriptomics and genome sequencing. Trends Plant Sci. 2017, 22, 705–717. [Google Scholar] [CrossRef] [PubMed]
- Ye, S.F.; Gao, X. Excavating abiotic stress-related gene resources of terrestrial macroscopic cyanobacteria for crop genetic engineering: Dawn and challenge. Bioengineered 2015, 6, 313–315. [Google Scholar] [CrossRef] [PubMed]
- Dodds, W.K.; Gudder, D.A.; Mollenhauer, D. The ecology of Nostoc. J. Phycol. 1995, 31, 2–18. [Google Scholar] [CrossRef]
- Potts, M. Desiccation tolerance: A simple process? Trends Microbiol. 2001, 9, 553–559. [Google Scholar] [CrossRef]
- Raanan, H.; Oren, N.; Treves, H.; Keren, N.; Ohad, I.; Berkowicz, S.M.; Hagemann, M.; Koch, M.; Shotland, Y.; Kaplan, A. Towards clarifying what distinguishes cyanobacteria able to resurrect after desiccation from those that cannot: The photosynthetic aspect. Biochim. Biophys. Acta 2016, 1857, 715–722. [Google Scholar] [CrossRef] [PubMed]
- Hagemann, M. Molecular biology of cyanobacterial salt acclimation. FEMS Microbiol. Rev. 2010, 35, 87–123. [Google Scholar] [CrossRef] [PubMed]
- Gao, K. Chinese studies on the edible blue-green alga, Nostoc flagelliforme: A review. J. Appl. Phycol. 1998, 10, 37–49. [Google Scholar] [CrossRef]
- Liang, W.Y.; Jiao, G.F.; Zhou, Y.W.; You, X.R.; Zhang, Y.P.; Chen, W. Differential expression and gene cloning of peroxiredoxin from Nostoc flagelliforme subjected to dehydration and rehydration. Chin. J. Appl. Environ. Biol. 2011, 17, 666–672. [Google Scholar]
- Wu, S.X.; He, L.; Shen, R.R.; Zhang, X.; Wang, Q.X. Molecular cloning of maltooligosyltrehalose trehalohydrolase gene from Nostoc flagelliforme and trehalose-related response to stresses. J. Microbiol. Biotechnol. 2011, 21, 830–837. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.H.; Liu, K.; Ai, Y.F.; Jiang, H.B.; Gao, X.; Qiu, B.S. Differential display analysis of cDNA fragments potentially involved in Nostoc flagelliforme response to osmotic stress. J. Appl. Phycol. 2012, 24, 1487–1494. [Google Scholar] [CrossRef]
- Liang, W.Y.; Wang, L.X.; Zhang, Y.P.; Lei, X.T.; Yang, J.; You, X.R.; Cheng, C.L.; Zhou, Y.W.; Chen, W. Comparative proteomic and physiological analysis of diurnal changes in Nostoc flagelliforme. J. Appl. Phycol. 2013, 25, 1709–1721. [Google Scholar] [CrossRef]
- Ai, Y.F.; Yang, Y.W.; Gao, X.; Qiu, B.S. Heterologous expression of three stress-responsive genes from Nostoc flagelliforme confers tolerance to abiotic stresses in Escherichia coli. J. Appl. Phycol. 2014, 26, 123–129. [Google Scholar] [CrossRef]
- Gao, X.; Liu, K.; Qiu, B.S. An investigation on the genetic background of Nostoc flagelliforme by similarity analysis of its partial genomic DNA and phylogenetic comparison of deduced related species. Acta Physiol. Plant. 2011, 33, 1301–1318. [Google Scholar] [CrossRef]
- Pécsi, I.; Szabó, J.E.; Adams, S.D.; Simon, I.; Sellers, J.R.; Vértessy, B.G.; Tóth, J. Nucleotide pyrophosphatase employs a P-loop-like motif to enhance catalytic power and NDP/NTP discrimination. Proc. Natl. Acad. Sci. USA 2011, 108, 14437–14442. [Google Scholar] [CrossRef] [PubMed]
- Arya, P.; Acharya, V. Plant STAND P-loop NTPases: A current perspective of genome distribution, evolution, and function. Mol. Genet. Genom. 2018, 293, 17–31. [Google Scholar] [CrossRef] [PubMed]
- Nachin, L.; Loiseau, L.; Expert, D.; Barras, F. SufC: An unorthodox cytoplasmic ABC/ATPase required for [Fe-S] biogenesis under oxidative stress. EMBO J. 2003, 22, 427–437. [Google Scholar] [CrossRef] [PubMed]
- Juhnke, H.; Charizanis, C.; Latifi, F.; Krems, B.; Entian, K.D. The essential protein Fap7 is involved in the oxidative stress response of Saccharomyces cerevisiae. Mol. Microbiol. 2000, 35, 936–948. [Google Scholar] [CrossRef] [PubMed]
- Jou, Y.; Chiang, C.P.; Jauh, G.Y.; Yen, H.E. Functional characterization of rice plant SKD1, an AAA-type ATPase associated with the endoplasmic reticulum-Golgi network, and its role in adaptation to salt stress. Plant Physiol. 2006, 141, 135–146. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.W.; Vierling, E. Mutants of Arabidopsis thaliana defective in the acquisition of tolerance to high temperature stress. Proc. Natl. Acad. Sci. USA 2000, 97, 4392–4397. [Google Scholar] [CrossRef] [PubMed]
- Cheung, M.Y.; Li, X.; Miao, R.; Fong, Y.H.; Li, K.P.; Yung, Y.L.; Yua, M.H.; Wonga, K.B.; Chenc, Z.; Lam, H.M. ATP binding by the P-loop NTPase OsYchF1 (an unconventional G protein) contributes to biotic but not abiotic stress responses. Proc. Natl. Acad. Sci. USA 2016, 113, 2648–2653. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, X.M.; Bi, Y.H.; Chen, L.; Hu, S.; Hu, Z.Y. Responses of photosynthetic activity in the drought-tolerant cyanobacterium, Nostoc flagelliforme to rehydration at different temperature. J. Arid Environ. 2008, 72, 370–377. [Google Scholar] [CrossRef]
- Liu, Y.H.; Yu, L.; Ke, W.T.; Gao, X.; Qiu, B.S. Photosynthetic recovery of Nostoc flagelliforme (Cyanophyceae) upon rehydration after 2 years and 8 years dry storage. Phycologia 2010, 49, 429–437. [Google Scholar] [CrossRef]
- Gao, K.S.; Qiu, B.S.; Xia, J.R.; Yu, A.J.; Li, Y.G. Effect of wind speed on loss of water from Nostoc flagelliforme colonies. J. Appl. Phycol. 1998, 10, 55–58. [Google Scholar] [CrossRef]
- Ye, C.P.; Gao, K.S. Photosynthetic response to salt of aquatic-living colonies of the terrestrial cyanobacterium Nostoc flagelliforme. J. Appl. Phycol. 2004, 16, 477–481. [Google Scholar] [CrossRef]
- Gao, X.; Ren, F.; Lu, Y.T. The Arabidopsis mutant stg1 identifies a function for TBP-associated factor 10 in plant osmotic stress adaptation. Plant Cell Physiol. 2006, 47, 1285–1294. [Google Scholar] [CrossRef] [PubMed]
- Elhai, J.; Wolk, C.P. A versatile class of positive-selection vectors based on the nonviability of palindrome-containing plasmids that allows cloning into long polylinkers. Gene 1988, 68, 119–138. [Google Scholar] [CrossRef]
- Jiang, H.B.; Kong, R.Q.; Xu, X.D. The N-acetylmuramic acid 6-phosphate etherase gene promotes growth and cell differentiation of cyanobacteria under light-limiting conditions. J. Bacteriol. 2010, 192, 2239–2245. [Google Scholar] [CrossRef] [PubMed]
- Williams, J.G. Construction of specific mutations in photosystem II photosynthetic reaction center by genetic engineering methods in Synechocystis 6803. Methods Enzymol. 1988, 167, 766–778. [Google Scholar]
- Gao, X.; Yang, Y.W.; Ai, Y.F.; Luo, H.Y.; Qiu, B.S. Quality evaluation of the edible blue-green alga Nostoc flagelliforme using a chlorophyll fluorescence parameter and several biochemical markers. Food Chem. 2014, 143, 307–312. [Google Scholar] [CrossRef] [PubMed]
- Dai, G.Z.; Qiu, B.S.; Forchhammer, K. Ammonium tolerance in the cyanobacterium Synechocystis sp. strain PCC 6803 and the role of the psbA multigene family. Plant Cell Environ. 2014, 37, 840–851. [Google Scholar] [CrossRef] [PubMed]
- Kehr, J.C.; Dittmann, E. Biosynthesis and function of extracellular glycans in cyanobacteria. Life 2015, 5, 164–180. [Google Scholar] [CrossRef] [PubMed]
- DuBois, M.; Gilles, K.A.; Hamilton, J.K.; Rebers, P.T.; Smith, F. Colorimetric method for determination of sugars and related substances. Anal. Chem. 1956, 28, 350–356. [Google Scholar] [CrossRef]
- Hagemann, M.; Schoor, A.; Jeanjean, R.; Zuther, E.; Joset, F. The stpA gene from Synechocystis sp. strain PCC 6803 encodes the glucosylglycerol-phosphate phosphatase involved in cyanobacterial osmotic response to salt shock. J. Bacteriol. 1997, 179, 1727–1733. [Google Scholar] [CrossRef] [PubMed]
- Elanskaya, I.V.; Karandashova, I.V.; Bogachev, A.V.; Hagemann, M. Functional analysis of the Na+/H+ antiporter encoding genes of the cyanobacterium Synechocystis PCC 6803. Biochemistry 2002, 67, 432–440. [Google Scholar] [PubMed]
- Marin, K.; Kanesaki, Y.; Los, D.A.; Murata, N.; Suzuki, I.; Hagemann, M. Gene expression profiling reflects physiological processes in salt acclimation of Synechocystis sp. strain PCC 6803. Plant Physiol. 2004, 136, 3290–3300. [Google Scholar] [CrossRef] [PubMed]
- Nikkinen, H.L.; Hakkila, K.; Gunnelius, L.; Huokko, T.; Pollari, M.; Tyystjärvi, T. The SigB σ factor regulates multiple salt acclimation responses of the cyanobacterium Synechocystis sp. PCC 6803. Plant Physiol. 2012, 158, 514–523. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhang, X.; Shi, M.; Gao, L.; Niu, X.; Te, R.; Chen, L.; Zhang, W. Metabolomic analysis of the salt-sensitive mutants reveals changes in amino acid and fatty acid composition important to long-term salt stress in Synechocystis sp. PCC 6803. Funct. Integr. Genom. 2014, 14, 431–440. [Google Scholar] [CrossRef] [PubMed]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Juneau, P.; Qiu, B. Effects of iron on the growth and minimal fluorescence yield of three marine Synechococcus strains (Cyanophyceae). Phycol. Res. 2012, 60, 61–69. [Google Scholar] [CrossRef]
- Marin, K.; Stirnberg, M.; Eisenhut, M.; Krämer, R.; Hagemann, M. Osmotic stress in Synechocystis sp. PCC 6803: Low tolerance towards nonionic osmotic stress results from lacking activation of glucosylglycerol accumulation. Microbiology 2006, 152, 2023–2030. [Google Scholar] [CrossRef] [PubMed]
- Allakhverdiev, S.I.; Murata, N. Salt stress inhibits photosystems II and I in cyanobacteria. Photosynth. Res. 2008, 98, 529–539. [Google Scholar] [CrossRef] [PubMed]
- Klähn, S.; Hagemann, M. Compatible solute biosynthesis in cyanobacteria. Environ. Microbiol. 2011, 13, 551–562. [Google Scholar] [CrossRef] [PubMed]
- Pade, N.; Hagemann, M. Salt acclimation of cyanobacteria and their application in biotechnology. Life 2014, 5, 25–49. [Google Scholar] [CrossRef] [PubMed]
- Tabei, Y.; Okada, K.; Tsuzuki, M. Sll1330 controls the expression of glycolytic genes in Synechocystis sp. PCC 6803. Biochem. Biophys. Res. Commun. 2007, 355, 1045–1050. [Google Scholar] [CrossRef] [PubMed]
- Joseph, A.; Aikawa, S.; Sasaki, K.; Teramura, H.; Hasunuma, T.; Matsuda, F.; Osanaif, T.; Hiraig, M.Y.; Kondo, A. Rre37 stimulates accumulation of 2-oxoglutarate and glycogen under nitrogen starvation in Synechocystis sp. PCC 6803. FEBS Lett. 2014, 588, 466–471. [Google Scholar] [CrossRef] [PubMed]
- Hagemann, M.; Fulda, S.; Schubert, H. DNA, RNA, and protein synthesis in the cyanobacterium Synechocystis sp. PCC 6803 adapted to different salt concentrations. Curr. Microbiol. 1994, 28, 201–207. [Google Scholar] [CrossRef]
- Waditee, R.; Hibino, T.; Nakamura, T.; Incharoensakdi, A.; Takabe, T. Overexpression of a Na+/H+ antiporter confers salt tolerance on a freshwater cyanobacterium, making it capable of growth in sea water. Proc. Natl. Acad. Sci. USA 2002, 99, 4109–4114. [Google Scholar] [CrossRef] [PubMed]
- Szabados, L.; Savouré, A. Proline: A multifunctional amino acid. Trends Plant Sci. 2010, 15, 89–97. [Google Scholar] [CrossRef] [PubMed]
- Savakis, P.; Tan, X.; Qiao, C.; Song, K.; Lu, X.; Hellingwerf, K.J.; Branco dos Santos, F. Slr1670 from Synechocystis sp. PCC 6803 is required for the re-assimilation of the osmolyte glucosylglycerol. Front. Microbiol. 2016, 7, 1350. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.Z.; Wang, G.H.; Hong, S.; Liu, A.; Li, C.; Liu, Y.D. UV-B–induced oxidative damage and protective role of exopolysaccharides in desert cyanobacterium Microcoleus vaginatus. J. Integr. Plant Biol. 2009, 51, 194–200. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.B.; Wu, S.J.; Liu, D. Preparation of polysaccharides from cyanobacteria Nostoc commune and their antioxidant activities. Carbohydr. Polym. 2014, 99, 553–555. [Google Scholar] [CrossRef] [PubMed]
- Fry, I.V.; Huflejt, M.; Erber, W.W.A.; Peschek, G.A.; Packer, L. The role of respiration during adaptation of the freshwater cyanobacterium Synechococcus 6311 to salinity. Arch. Biochem. Biophys. 1986, 244, 686–691. [Google Scholar] [CrossRef]
- Deinlein, U.; Stephan, A.B.; Horie, T.; Luo, W.; Xu, G.; Schroeder, J.I. Plant salt-tolerance mechanisms. Trends Plant Sci. 2014, 19, 371–379. [Google Scholar] [CrossRef] [PubMed] [Green Version]


| Sample | Net Photosynthesis (µmol h−1 mg−1 Chl a) | Respiration (µmol h−1 mg−1 Chl a) |
|---|---|---|
| WT | 90.5 ± 11.1 | 38.6 ± 9.1 |
| drnf1-O | 105.6 ± 13.7 | 68.0 ± 12.5 * |
| WT + | 60.3 ± 10.3 | 28.1 ± 5.3 |
| drnf1-O + | 70.1 ± 10.5 | 53.0 ± 4.1 * |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cui, L.; Liu, Y.; Yang, Y.; Ye, S.; Luo, H.; Qiu, B.; Gao, X. The drnf1 Gene from the Drought-Adapted Cyanobacterium Nostoc flagelliforme Improved Salt Tolerance in Transgenic Synechocystis and Arabidopsis Plant. Genes 2018, 9, 441. https://doi.org/10.3390/genes9090441
Cui L, Liu Y, Yang Y, Ye S, Luo H, Qiu B, Gao X. The drnf1 Gene from the Drought-Adapted Cyanobacterium Nostoc flagelliforme Improved Salt Tolerance in Transgenic Synechocystis and Arabidopsis Plant. Genes. 2018; 9(9):441. https://doi.org/10.3390/genes9090441
Chicago/Turabian StyleCui, Lijuan, Yinghui Liu, Yiwen Yang, Shuifeng Ye, Hongyi Luo, Baosheng Qiu, and Xiang Gao. 2018. "The drnf1 Gene from the Drought-Adapted Cyanobacterium Nostoc flagelliforme Improved Salt Tolerance in Transgenic Synechocystis and Arabidopsis Plant" Genes 9, no. 9: 441. https://doi.org/10.3390/genes9090441





