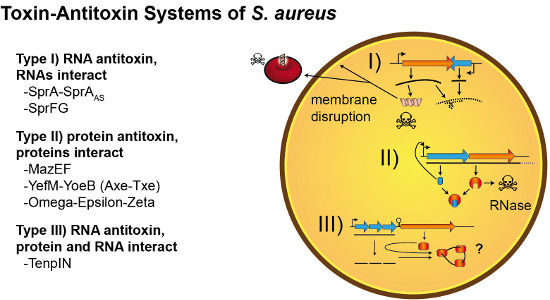
Toxin-Antitoxin Systems of Staphylococcus aureus
Abstract
:1. Introduction
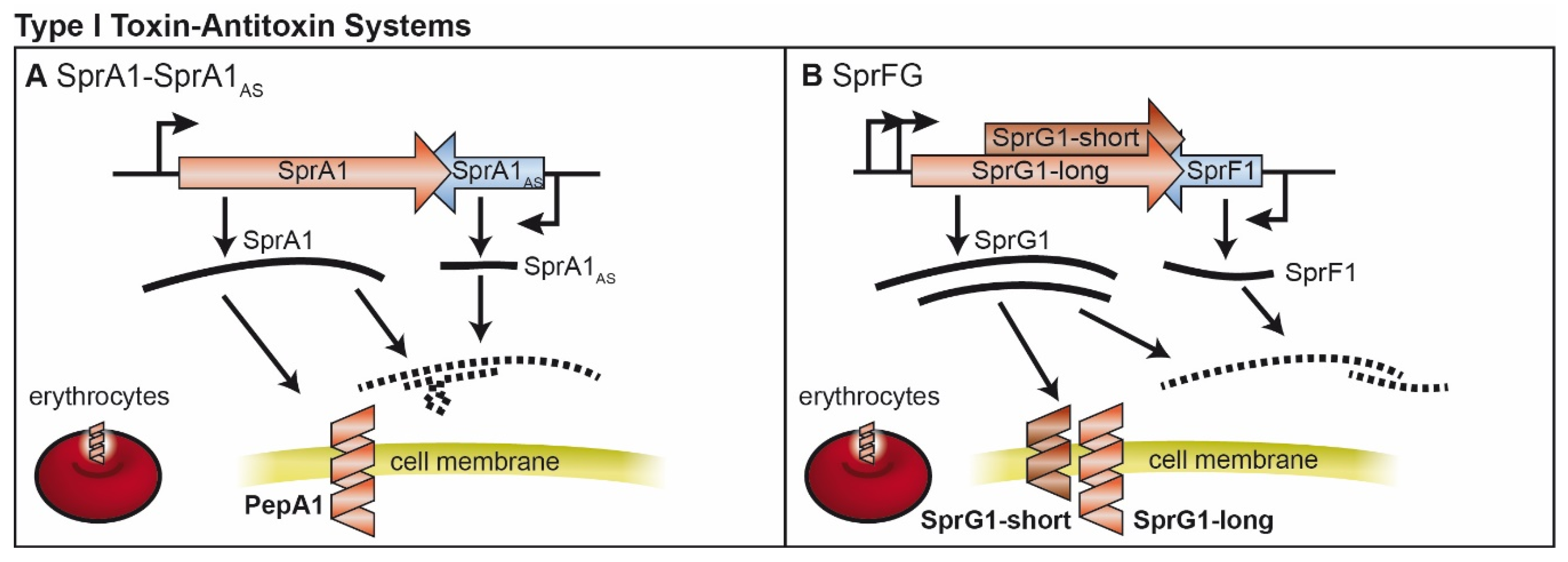
2. Type I TA Systems
2.1. SprA1/SprA1AS and Further Fst-Like Systems
2.2. SprF/SprG
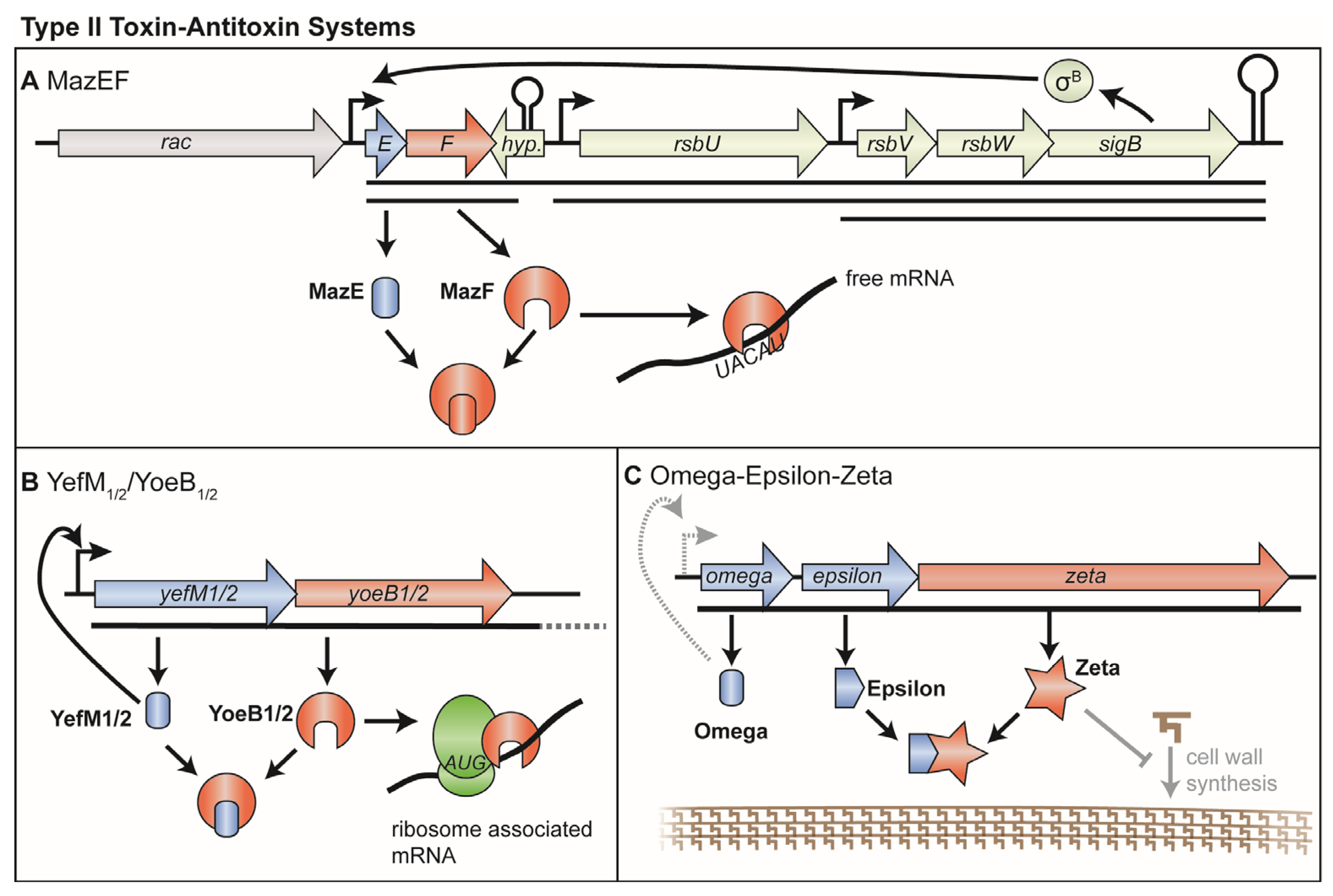
3. Type II TA Systems
3.1. MazEF and PemIK
3.2. YefM-YoeB (Axe-Txe)
3.3. Omega/Epsilon/Zeta
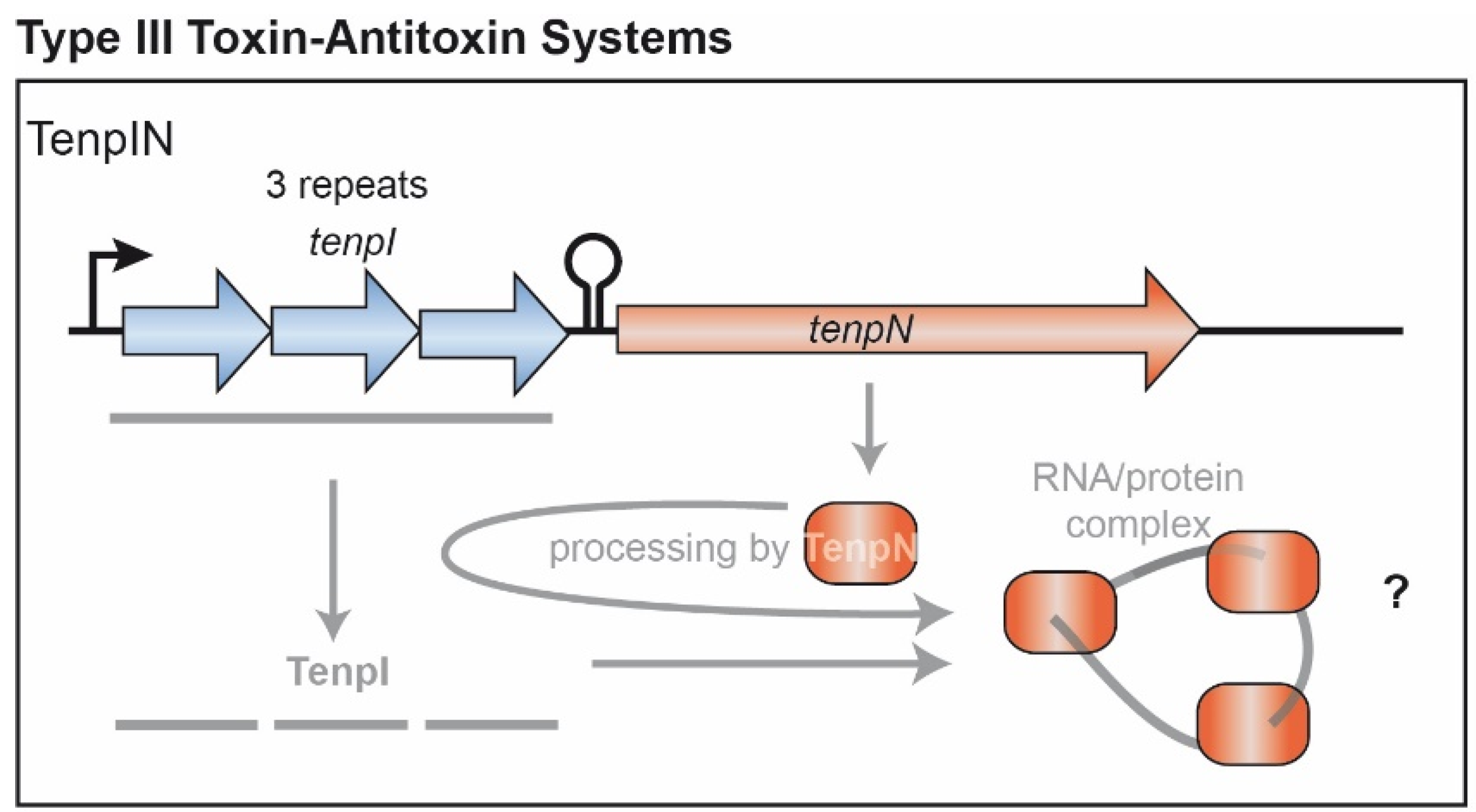
4. Type III TA Candidate Systems
5. Final Remarks
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| TA | toxin-antitoxin |
References
- DeLeo, F.R.; Otto, M.; Kreiswirth, B.N.; Chambers, H.F. Community-associated meticillin-resistant Staphylococcus aureus. Lancet 2010, 375, 1557–1568. [Google Scholar] [CrossRef]
- Lowy, F.D. Staphylococcus aureus infections. N. Engl. J. Med. 1998, 339, 520–532. [Google Scholar] [CrossRef] [PubMed]
- Otto, M. Staphylococcus aureus toxins. Curr. Opin. Microbiol. 2014, 17, 32–37. [Google Scholar] [CrossRef] [PubMed]
- Henkel, J.S.; Baldwin, M.R.; Barbieri, J.T. Toxins from bacteria. EXS 2010, 100, 1–29. [Google Scholar] [PubMed]
- Mruk, I.; Kobayashi, I. To be or not to be: Regulation of restriction-modification systems and other toxin-antitoxin systems. Nucleic Acids Res. 2014, 42, 70–86. [Google Scholar] [CrossRef] [PubMed]
- Schuster, C.F.; Bertram, R. Toxin-antitoxin systems are ubiquitous and versatile modulators of prokaryotic cell fate. FEMS Microbiol. Lett. 2013, 340, 73–85. [Google Scholar] [CrossRef] [PubMed]
- Unterholzner, S.J.; Poppenberger, B.; Rozhon, W. Toxin-antitoxin systems: Biology, identification, and application. Mob. Genet. Elem. 2013, 3. [Google Scholar] [CrossRef] [PubMed]
- Jaffé, A.; Ogura, T.; Hiraga, S. Effects of the ccd function of the F plasmid on bacterial growth. J. Bacteriol. 1985, 163, 841–849. [Google Scholar] [PubMed]
- Ogura, T.; Hiraga, S. Mini-F plasmid genes that couple host cell division to plasmid proliferation. Proc. Natl. Acad. Sci. USA 1983, 80, 4784–4788. [Google Scholar] [CrossRef] [PubMed]
- Gerdes, K.; Bech, F.W.; Jørgensen, S.T.; Løbner-Olesen, A.; Rasmussen, P.B.; Atlung, T.; Boe, L.; Karlstrom, O.; Molin, S.; von Meyenburg, K. Mechanism of postsegregational killing by the hok gene product of the parB system of plasmid R1 and its homology with the relF gene product of the E. coli relB operon. EMBO J. 1986, 5, 2023–2029. [Google Scholar] [PubMed]
- Wen, J.; Fozo, E.M. sRNA antitoxins: More than one way to repress a toxin. Toxins (Basel) 2014, 6, 2310–2335. [Google Scholar] [CrossRef] [PubMed]
- Brantl, S.; Jahn, N. sRNAs in bacterial type I and type III toxin-antitoxin systems. FEMS Microbiol. Rev. 2015, 39, 413–427. [Google Scholar] [CrossRef] [PubMed]
- Fozo, E.M.; Hemm, M.R.; Storz, G. Small toxic proteins and the antisense RNAs that repress them. Microbiol. Mol. Biol. Rev. 2008, 72, 579–589. [Google Scholar] [CrossRef] [PubMed]
- Cook, G.M.; Robson, J.R.; Frampton, R.A.; McKenzie, J.; Przybilski, R.; Fineran, P.C.; Arcus, V.L. Ribonucleases in bacterial toxin-antitoxin systems. Biochim. Biophys. Acta 2013, 1829, 523–531. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Pogliano, J.; Helinski, D.R.; Konieczny, I. ParE toxin encoded by the broad-host-range plasmid RK2 is an inhibitor of Escherichia coli gyrase. Mol. Microbiol. 2002, 44, 971–979. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Inouye, M. RatA (YfjG), an Escherichia coli toxin, inhibits 70S ribosome association to block translation initiation. Mol. Microbiol. 2011, 79, 1418–1429. [Google Scholar] [CrossRef] [PubMed]
- Mutschler, H.; Gebhardt, M.; Shoeman, R.L.; Meinhart, A. A novel mechanism of programmed cell death in bacteria by toxin-antitoxin systems corrupts peptidoglycan synthesis. PLoS Biol. 2011, 9. [Google Scholar] [CrossRef] [PubMed]
- Aakre, C.D.; Phung, T.N.; Huang, D.; Laub, M.T. A bacterial toxin inhibits DNA replication elongation through a direct interaction with the beta sliding clamp. Mol. Cell 2013, 52, 617–628. [Google Scholar] [CrossRef] [PubMed]
- Masuda, H.; Tan, Q.; Awano, N.; Wu, K.P.; Inouye, M. YeeU enhances the bundling of cytoskeletal polymers of MreB and FtsZ, antagonizing the CbtA (YeeV) toxicity in Escherichia coli. Mol. Microbiol. 2012, 84, 979–989. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Lord, D.M.; Cheng, H.Y.; Osbourne, D.O.; Hong, S.H.; Sanchez-Torres, V.; Quiroga, C.; Zheng, K.; Herrmann, T.; Peti, W.; et al. A new type V toxin-antitoxin system where mRNA for toxin GhoT is cleaved by antitoxin GhoS. Nat. Chem. Biol. 2012, 8, 855–861. [Google Scholar] [CrossRef] [PubMed]
- Blower, T.R.; Short, F.L.; Rao, F.; Mizuguchi, K.; Pei, X.Y.; Fineran, P.C.; Luisi, B.F.; Salmond, G.P. Identification and classification of bacterial Type III toxin-antitoxin systems encoded in chromosomal and plasmid genomes. Nucleic Acids Res. 2012, 40, 6158–6173. [Google Scholar] [CrossRef] [PubMed]
- Pandey, D.P.; Gerdes, K. Toxin-antitoxin loci are highly abundant in free-living but lost from host-associated prokaryotes. Nucleic Acids Res. 2005, 33, 966–976. [Google Scholar] [CrossRef] [PubMed]
- Van Melderen, L. Toxin-antitoxin systems: Why so many, what for? Curr. Opin. Microbiol. 2010, 13, 781–785. [Google Scholar] [CrossRef] [PubMed]
- Fozo, E.M.; Makarova, K.S.; Shabalina, S.A.; Yutin, N.; Koonin, E.V.; Storz, G. Abundance of type I toxin-antitoxin systems in bacteria: Searches for new candidates and discovery of novel families. Nucleic Acids Res. 2010, 38, 3743–3759. [Google Scholar] [CrossRef] [PubMed]
- Sevin, E.W.; Barloy-Hubler, F. RASTA-Bacteria: A web-based tool for identifying toxin-antitoxin loci in prokaryotes. Genome Biol. 2007, 8. [Google Scholar] [CrossRef] [PubMed]
- Shao, Y.; Harrison, E.M.; Bi, D.; Tai, C.; He, X.; Ou, H.Y.; Rajakumar, K.; Deng, Z. TADB: A web-based resource for Type 2 toxin-antitoxin loci in bacteria and archaea. Nucleic Acids Res. 2011, 39, D606–D611. [Google Scholar] [CrossRef] [PubMed]
- Gerdes, K.; Christensen, S.K.; Lobner-Olesen, A. Prokaryotic toxin-antitoxin stress response loci. Nat. Rev. Microbiol. 2005, 3, 371–382. [Google Scholar] [CrossRef] [PubMed]
- Gerdes, K.; Maisonneuve, E. Bacterial persistence and toxin-antitoxin loci. Annu. Rev. Microbiol. 2012, 66, 103–123. [Google Scholar] [CrossRef] [PubMed]
- Van Melderen, L.; Saavedra De Bast, M. Bacterial toxin-antitoxin systems: More than selfish entities? PLoS Genet. 2009, 5. [Google Scholar] [CrossRef] [PubMed]
- Fineran, P.C.; Blower, T.R.; Foulds, I.J.; Humphreys, D.P.; Lilley, K.S.; Salmond, G.P. The phage abortive infection system, ToxIN, functions as a protein-RNA toxin-antitoxin pair. Proc. Natl. Acad. Sci. USA 2009, 106, 894–899. [Google Scholar] [CrossRef] [PubMed]
- Koga, M.; Otsuka, Y.; Lemire, S.; Yonesaki, T. Escherichia coli rnlA and rnlB compose a novel toxin-antitoxin system. Genetics 2011, 187, 123–130. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, P.; Arora, G.; Singh, M.; Kidwai, S.; Narayan, O.P.; Singh, R. MazF ribonucleases promote Mycobacterium tuberculosis drug tolerance and virulence in guinea pigs. Nat. Commun. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Hazan, R.; Sat, B.; Engelberg-Kulka, H. Escherichia coli mazEF-mediated cell death is triggered by various stressful conditions. J. Bacteriol. 2004, 186, 3663–3669. [Google Scholar] [CrossRef] [PubMed]
- Yarmolinsky, M.B. Programmed cell death in bacterial populations. Science 1995, 267, 836–837. [Google Scholar] [CrossRef] [PubMed]
- Nolle, N.; Schuster, C.F.; Bertram, R. Two paralogous yefM-yoeB loci from Staphylococcus equorum encode functional toxin-antitoxin systems. Microbiology 2013, 159, 1575–1585. [Google Scholar] [CrossRef] [PubMed]
- Schuster, C.F.; Park, J.H.; Prax, M.; Herbig, A.; Nieselt, K.; Rosenstein, R.; Inouye, M.; Bertram, R. Characterization of a mazEF toxin-antitoxin homologue from Staphylococcus equorum. J. Bacteriol. 2013, 195, 115–125. [Google Scholar] [CrossRef] [PubMed]
- Weaver, K.E.; Jensen, K.D.; Colwell, A.; Sriram, S.I. Functional analysis of the Enterococcus faecalis plasmid pAD1-encoded stability determinant par. Mol. Microbiol. 1996, 20, 53–63. [Google Scholar] [CrossRef] [PubMed]
- Greenfield, T.J.; Ehli, E.; Kirshenmann, T.; Franch, T.; Gerdes, K.; Weaver, K.E. The antisense RNA of the par locus of pAD1 regulates the expression of a 33-amino-acid toxic peptide by an unusual mechanism. Mol. Microbiol. 2000, 37, 652–660. [Google Scholar] [CrossRef] [PubMed]
- Beaume, M.; Hernandez, D.; Farinelli, L.; Deluen, C.; Linder, P.; Gaspin, C.; Romby, P.; Schrenzel, J.; Francois, P. Cartography of methicillin-resistant S. aureus transcripts: Detection, orientation and temporal expression during growth phase and stress conditions. PLoS ONE 2010, 5. [Google Scholar] [CrossRef] [PubMed]
- Pichon, C.; Felden, B. Small RNA genes expressed from Staphylococcus aureus genomic and pathogenicity islands with specific expression among pathogenic strains. Proc. Natl. Acad. Sci. USA 2005, 102, 14249–14254. [Google Scholar] [CrossRef] [PubMed]
- Geissmann, T.; Chevalier, C.; Cros, M.J.; Boisset, S.; Fechter, P.; Noirot, C.; Schrenzel, J.; Francois, P.; Vandenesch, F.; Gaspin, C.; et al. A search for small noncoding RNAs in Staphylococcus aureus reveals a conserved sequence motif for regulation. Nucleic Acids Res. 2009, 37, 7239–7257. [Google Scholar] [CrossRef] [PubMed]
- Bohn, C.; Rigoulay, C.; Chabelskaya, S.; Sharma, C.M.; Marchais, A.; Skorski, P.; Borezée-Durant, E.; Barbet, R.; Jacquet, E.; Jacq, A.; et al. Experimental discovery of small RNAs in Staphylococcus aureus reveals a riboregulator of central metabolism. Nucleic Acids Res. 2010, 38, 6620–6636. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weaver, K.E.; Reddy, S.G.; Brinkman, C.L.; Patel, S.; Bayles, K.W.; Endres, J.L. Identification and characterization of a family of toxin-antitoxin systems related to the Enterococcus faecalis plasmid pAD1 par addiction module. Microbiology 2009, 155, 2930–2940. [Google Scholar] [CrossRef] [PubMed]
- Kwong, S.M.; Jensen, S.O.; Firth, N. Prevalence of Fst-like toxin-antitoxin systems. Microbiology 2010, 156, 975–977; discussion 977. [Google Scholar] [CrossRef] [PubMed]
- Jensen, S.O.; Apisiridej, S.; Kwong, S.M.; Yang, Y.H.; Skurray, R.A.; Firth, N. Analysis of the prototypical Staphylococcus aureus multiresistance plasmid pSK1. Plasmid 2010, 64, 135–142. [Google Scholar] [CrossRef] [PubMed]
- Sayed, N.; Jousselin, A.; Felden, B. A cis-antisense RNA acts in trans in Staphylococcus aureus to control translation of a human cytolytic peptide. Nat. Struct. Mol. Biol. 2012, 19, 105–112. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sayed, N.; Nonin-Lecomte, S.; Réty, S.; Felden, B. Functional and structural insights of a Staphylococcus aureus apoptotic-like membrane peptide from a toxin-antitoxin Module. J. Biol. Chem. 2012, 287, 43454–43463. [Google Scholar] [CrossRef] [PubMed]
- Silvaggi, J.M.; Perkins, J.B.; Losick, R. Small untranslated RNA antitoxin in Bacillus subtilis. J. Bacteriol. 2005, 187, 6641–6650. [Google Scholar] [CrossRef] [PubMed]
- Pinel-Marie, M.L.; Brielle, R.; Felden, B. Dual toxic-peptide-coding Staphylococcus aureus RNA under antisense regulation targets host cells and bacterial rivals unequally. Cell Rep. 2014, 7, 424–435. [Google Scholar] [CrossRef] [Green Version]
- Williams, J.J.; Halvorsen, E.M.; Dwyer, E.M.; DiFazio, R.M.; Hergenrother, P.J. Toxin-antitoxin (TA) systems are prevalent and transcribed in clinical isolates of Pseudomonas aeruginosa and methicillin-resistant Staphylococcus aureus. FEMS Microbiol. Lett. 2011, 322, 41–50. [Google Scholar] [CrossRef] [PubMed]
- Aizenman, E.; Engelberg-Kulka, H.; Glaser, G. An Escherichia coli chromosomal “addiction module” regulated by guanosine-3′,5′ -bispyrophosphate: A model for programmed bacterial cell death. Proc. Natl. Acad. Sci. USA 1996, 93, 6059–6063. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; de Lencastre, H.; Tomasz, A. Sigma-B, a putative operon encoding alternate sigma factor of Staphylococcus aureus RNA polymerase: Molecular cloning and DNA sequencing. J. Bacteriol. 1996, 178, 6036–6042. [Google Scholar] [PubMed]
- Kullik, I.; Jenni, R.; Berger-Bächi, B. Sequence of the putative alanine racemase operon in Staphylococcus aureus: Insertional interruption of this operon reduces D-alanine substitution of lipoteichoic acid and autolysis. Gene 1998, 219, 9–17. [Google Scholar] [CrossRef]
- Mittenhuber, G. Occurrence of mazEF-like antitoxin/toxin systems in bacteria. J. Mol. Microbiol. Biotechnol. 1999, 1, 295–302. [Google Scholar] [PubMed]
- Schuster, C.F.; Mechler, L.; Nolle, N.; Krismer, B.; Zelder, M.E.; Götz, F.; Bertram, R. The mazEF toxin-antitoxin system alters the β-Lactam susceptibility of Staphylococcus aureus. PLoS ONE 2015, 10. [Google Scholar] [CrossRef] [PubMed]
- Gertz, S.; Engelmann, S.; Schmid, R.; Ohlsen, K.; Hacker, J.; Hecker, M. Regulation of sigmaB-dependent transcription of sigB and asp23 in two different Staphylococcus aureus strains. Mol. Gen. Genet. 1999, 261, 558–566. [Google Scholar] [CrossRef] [PubMed]
- Donegan, N.P.; Cheung, A.L. Regulation of the mazEF toxin-antitoxin module in Staphylococcus aureus and its impact on sigB expression. J. Bacteriol. 2009, 191, 2795–2805. [Google Scholar] [CrossRef] [PubMed]
- Senn, M.M.; Giachino, P.; Homerova, D.; Steinhuber, A.; Strassner, J.; Kormanec, J.; Flückiger, U.; Berger-Bächi, B.; Bischoff, M. Molecular analysis and organization of the σB operon in Staphylococcus aureus. J. Bacteriol. 2005, 187, 8006–8019. [Google Scholar] [CrossRef] [PubMed]
- De Hoon, M.J.; Makita, Y.; Nakai, K.; Miyano, S. Prediction of transcriptional terminators in Bacillus subtilis and related species. PLoS Comput. Biol. 2005, 1. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Donegan, N.P.; Thompson, E.T.; Fu, Z.; Cheung, A.L. Proteolytic regulation of toxin-antitoxin systems by ClpPC in Staphylococcus aureus. J. Bacteriol. 2010, 192, 1416–1422. [Google Scholar] [CrossRef] [PubMed]
- Zorzini, V.; Buts, L.; Sleutel, M.; Garcia-Pino, A.; Talavera, A.; Haesaerts, S.; De Greve, H.; Cheung, A.; van Nuland, N.A.; Loris, R. Structural and biophysical characterization of Staphylococcus aureus SaMazF shows conservation of functional dynamics. Nucleic Acids Res. 2014, 42, 6709–6725. [Google Scholar] [CrossRef] [PubMed]
- Zorzini, V.; Haesaerts, S.; Cheung, A.; Loris, R.; van Nuland, N.A. 1H, 13C, and 15N backbone and side-chain chemical shift assignment of the staphylococcal MazF mRNA interferase. Biomol. NMR Assign. 2011, 5, 157–160. [Google Scholar] [CrossRef] [PubMed]
- Zorzini, V.; Haesaerts, S.; Donegan, N.P.; Fu, Z.; Cheung, A.L.; van Nuland, N.A.; Loris, R. Crystallization of the Staphylococcus aureus MazF mRNA interferase. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 2011, 67, 386–389. [Google Scholar] [CrossRef] [PubMed]
- Simanshu, D.K.; Yamaguchi, Y.; Park, J.H.; Inouye, M.; Patel, D.J. Structural basis of mRNA recognition and cleavage by toxin MazF and its regulation by antitoxin MazE in Bacillus subtilis. Mol. Cell 2013, 52, 447–458. [Google Scholar] [CrossRef] [PubMed]
- Fu, Z.; Donegan, N.P.; Memmi, G.; Cheung, A.L. Characterization of MazFSa, an endoribonuclease from Staphylococcus aureus. J. Bacteriol. 2007, 189, 8871–8879. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Inoue, K.; Yoshizumi, S.; Kobayashi, H.; Zhang, Y.; Ouyang, M.; Kato, F.; Sugai, M.; Inouye, M. Staphylococcus aureus MazF specifically cleaves a pentad sequence, UACAU, which is unusually abundant in the mRNA for pathogenic adhesive factor SraP. J. Bacteriol. 2009, 191, 3248–3255. [Google Scholar] [CrossRef] [PubMed]
- Van Rensburg, J.J.; Hergenrother, P.J. Detection of endogenous MazF enzymatic activity in Staphylococcus aureus. Anal. Biochem. 2013, 443, 81–87. [Google Scholar] [CrossRef] [PubMed]
- Fu, Z.; Tamber, S.; Memmi, G.; Donegan, N.P.; Cheung, A.L. Overexpression of MazFSa in Staphylococcus aureus induces bacteriostasis by selectively targeting mRNAs for cleavage. J. Bacteriol. 2009, 191, 2051–2059. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Corvaglia, A.R.; Leo, S.; Cheung, A.; Francois, P. Characterization of RNA Helicase CshA and Its Role in Protecting mRNAs and Small RNAs of Staphylococcus aureus Strain Newman. Infect. Immun. 2016, 84, 833–844. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, D.; Natarajan, J. Network analysis of S. aureus response to ramoplanin reveals modules for virulence factors and resistance mechanisms and characteristic novel genes. Gene 2015, 574, 149–162. [Google Scholar] [CrossRef] [PubMed]
- Bukowski, M.; Lyzen, R.; Helbin, W.M.; Bonar, E.; Szalewska-Palasz, A.; Wegrzyn, G.; Dubin, G.; Dubin, A.; Wladyka, B. A regulatory role for Staphylococcus aureus toxin-antitoxin system PemIKSa. Nat. Commun. 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Pomerantsev, A.P.; Golovliov, I.R.; Ohara, Y.; Mokrievich, A.N.; Obuchi, M.; Norqvist, A.; Kuoppa, K.; Pavlov, V.M. Genetic organization of the Francisella plasmid pFNL10. Plasmid 2001, 46, 210–222. [Google Scholar] [CrossRef] [PubMed]
- Grady, R.; Hayes, F. Axe-Txe, a broad-spectrum proteic toxin-antitoxin system specified by a multidrug-resistant, clinical isolate of Enterococcus faecium. Mol. Microbiol. 2003, 47, 1419–1432. [Google Scholar] [CrossRef] [PubMed]
- Yoshizumi, S.; Zhang, Y.; Yamaguchi, Y.; Chen, L.; Kreiswirth, B.N.; Inouye, M. Staphylococcus aureus YoeB homologues inhibit translation initiation. J. Bacteriol. 2009, 191, 5868–5872. [Google Scholar] [CrossRef] [PubMed]
- Kasari, V.; Mets, T.; Tenson, T.; Kaldalu, N. Transcriptional cross-activation between toxin-antitoxin systems of Escherichia coli. BMC Microbiol. 2013, 13. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Inouye, M. The inhibitory mechanism of protein synthesis by YoeB, an Escherichia coli toxin. J. Biol. Chem. 2009, 284, 6627–6638. [Google Scholar] [CrossRef] [PubMed]
- Larson, A.S.; Hergenrother, P.J. Light activation of Staphylococcus aureus toxin YoeBSa1 reveals guanosine-specific endoribonuclease activity. Biochemistry 2014, 53, 188–201. [Google Scholar] [CrossRef] [PubMed]
- Locke, J.B.; Rahawi, S.; Lamarre, J.; Mankin, A.S.; Shaw, K.J. Genetic environment and stability of cfr in methicillin-resistant Staphylococcus aureus CM05. Antimicrob. Agents Chemother. 2012, 56, 332–340. [Google Scholar] [CrossRef] [PubMed]
- Zielenkiewicz, U.; Ceglowski, P. The toxin-antitoxin system of the streptococcal plasmid pSM19035. J. Bacteriol. 2005, 187, 6094–6105. [Google Scholar] [CrossRef] [PubMed]
- Blower, T.R.; Fineran, P.C.; Johnson, M.J.; Toth, I.K.; Humphreys, D.P.; Salmond, G.P. Mutagenesis and functional characterization of the RNA and protein components of the toxIN abortive infection and toxin-antitoxin locus of Erwinia. J. Bacteriol. 2009, 191, 6029–6039. [Google Scholar] [CrossRef] [PubMed]
- Bélanger, M.; Moineau, S. Mutational Analysis of the Antitoxin in the Lactococcal Type III Toxin-Antitoxin System AbiQ. Appl. Environ. Microbiol. 2015, 81, 3848–3855. [Google Scholar] [CrossRef] [PubMed]
- Short, F.L.; Monson, R.E.; Salmond, G.P. A Type III protein-RNA toxin-antitoxin system from Bacillus thuringiensis promotes plasmid retention during spore development. RNA Biol. 2015, 12, 933–937. [Google Scholar] [CrossRef] [PubMed]
- Conlon, B.P. Staphylococcus aureus chronic and relapsing infections: Evidence of a role for persister cells: An investigation of persister cells, their formation and their role in S. aureus disease. Bioessays 2014, 36, 991–996. [Google Scholar] [CrossRef] [PubMed]
- Prax, M.; Bertram, R. Metabolic aspects of bacterial persisters. Front. Cell. Infect. Microbiol. 2014, 4. [Google Scholar] [CrossRef] [PubMed]
- Proctor, R.A.; Kriegeskorte, A.; Kahl, B.C.; Becker, K.; Löffler, B.; Peters, G. Staphylococcus aureus Small Colony Variants (SCVs): A road map for the metabolic pathways involved in persistent infections. Front. Cell. Infect. Microbiol. 2014, 4. [Google Scholar] [CrossRef]
- Maisonneuve, E.; Shakespeare, L.J.; Jørgensen, M.G.; Gerdes, K. Bacterial persistence by RNA endonucleases. Proc. Natl. Acad. Sci. USA 2011, 108, 13206–13211. [Google Scholar] [CrossRef] [PubMed]
- Conlon, B.P.; Rowe, S.E.; Gandt, A.B.; Nuxoll, A.S.; Donegan, N.P.; Zalis, E.A.; Clair, G.; Adkins, J.N.; Cheung, A.L.; Lewis, K. Persister formation in Staphylococcus aureus is associated with ATP depletion. Nat. Microbiol. 2016, 1. [Google Scholar] [CrossRef]
- Chan, W.T.; Balsa, D.; Espinosa, M. One cannot rule them all: Are bacterial toxins-antitoxins druggable? FEMS Microbiol. Rev. 2015, 39, 522–540. [Google Scholar] [CrossRef] [PubMed]
- Lioy, V.S.; Rey, O.; Balsa, D.; Pellicer, T.; Alonso, J.C. A toxin-antitoxin module as a target for antimicrobial development. Plasmid 2010, 63, 31–39. [Google Scholar] [CrossRef] [PubMed]
- Hayes, F.; Kedzierska, B. Regulating toxin-antitoxin expression: Controlled detonation of intracellular molecular timebombs. Toxins (Basel) 2014, 6, 337–358. [Google Scholar] [CrossRef] [PubMed]
- Williams, J.J.; Hergenrother, P.J. Artificial activation of toxin-antitoxin systems as an antibacterial strategy. Trends Microbiol. 2012, 20, 291–298. [Google Scholar] [CrossRef] [PubMed]
- Bacconi, M.; Haag, A.F.; Torre, A.; Castagnetti, A.; Chiarot, E.; Delany, I.; Bensi, G. A stable luciferase reporter plasmid for in vivo imaging in murine models of Staphylococcus aureus infections. Appl. Microbiol. Biotechnol. 2016, 100, 3197–3206. [Google Scholar] [CrossRef] [PubMed]
- Solecki, O.; Mosbah, A.; Baudy Floc’h, M.; Felden, B. Converting a Staphylococcus aureus toxin into effective cyclic pseudopeptide antibiotics. Chem. Biol. 2015, 22, 329–335. [Google Scholar] [CrossRef] [PubMed]
- Perros, M. Infectious disease. A sustainable model for antibiotics. Science 2015, 347, 1062–1064. [Google Scholar] [CrossRef] [PubMed]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Schuster, C.F.; Bertram, R. Toxin-Antitoxin Systems of Staphylococcus aureus. Toxins 2016, 8, 140. https://doi.org/10.3390/toxins8050140
Schuster CF, Bertram R. Toxin-Antitoxin Systems of Staphylococcus aureus. Toxins. 2016; 8(5):140. https://doi.org/10.3390/toxins8050140
Chicago/Turabian StyleSchuster, Christopher F., and Ralph Bertram. 2016. "Toxin-Antitoxin Systems of Staphylococcus aureus" Toxins 8, no. 5: 140. https://doi.org/10.3390/toxins8050140
APA StyleSchuster, C. F., & Bertram, R. (2016). Toxin-Antitoxin Systems of Staphylococcus aureus. Toxins, 8(5), 140. https://doi.org/10.3390/toxins8050140