Ochratoxigenic Black Species of Aspergilli in Grape Fruits of Northern Italy Identified by an Improved PCR-RFLP Procedure
Abstract
:1. Introduction
2. Materials and Methods
2.1. Collection of Grape Samples and Isolation of Aspergillus spp.
2.2. Culture Conditions and Extraction of DNA
2.3. PCR Conditions and DNA Digestions
2.4. Sequencing Analysis
2.5. Production of OTA by Aspergillus spp.
3. Results
3.1. ITS-RFLP of Aspergillus Isolates
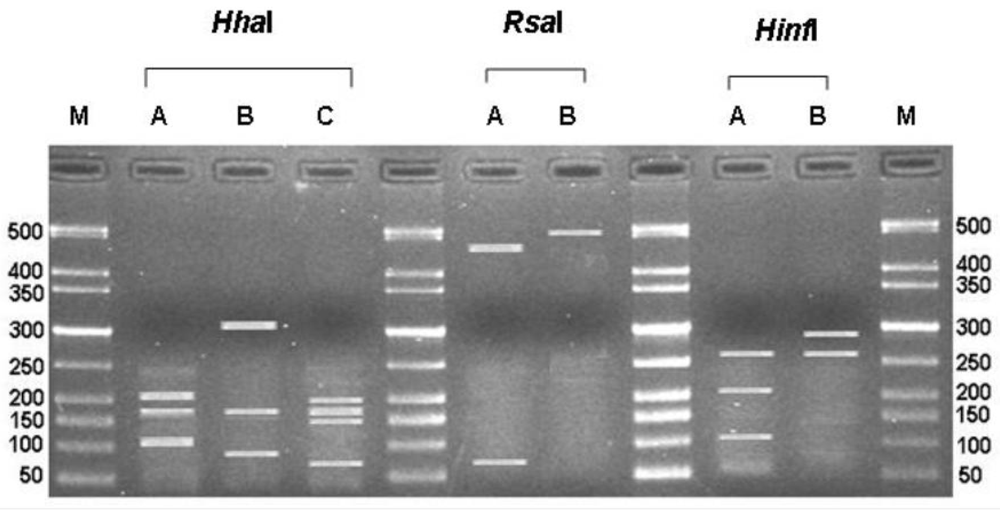
| Type | Species | No of isolates | Restriction patterns & size of the fragments (bp) | ||
|---|---|---|---|---|---|
| HhaI | RsaI | HinfI | |||
| N | A. niger | 153 | A | A | A |
| 207, 178, 114 | 480, 66 | 269, 202, 110 | |||
| C | A. carbonarius | 107 | B | A | B |
| 318, 178, 91 | 480, 66 | 294, 270 | |||
| T | A. tubingensis | 72 | A | B | A |
| 207, 178, 114 | 500 | 269, 202, 110 | |||
| A | A. japonicus | 20 | C | A | B |
| 185, 174, 156, 75 | 480, 66 | 294, 270 | |||
| J | A. aculeatus | 4 | C | A | A |
| 185, 174, 156, 75 | 480, 66 | 269, 202, 110 | |||
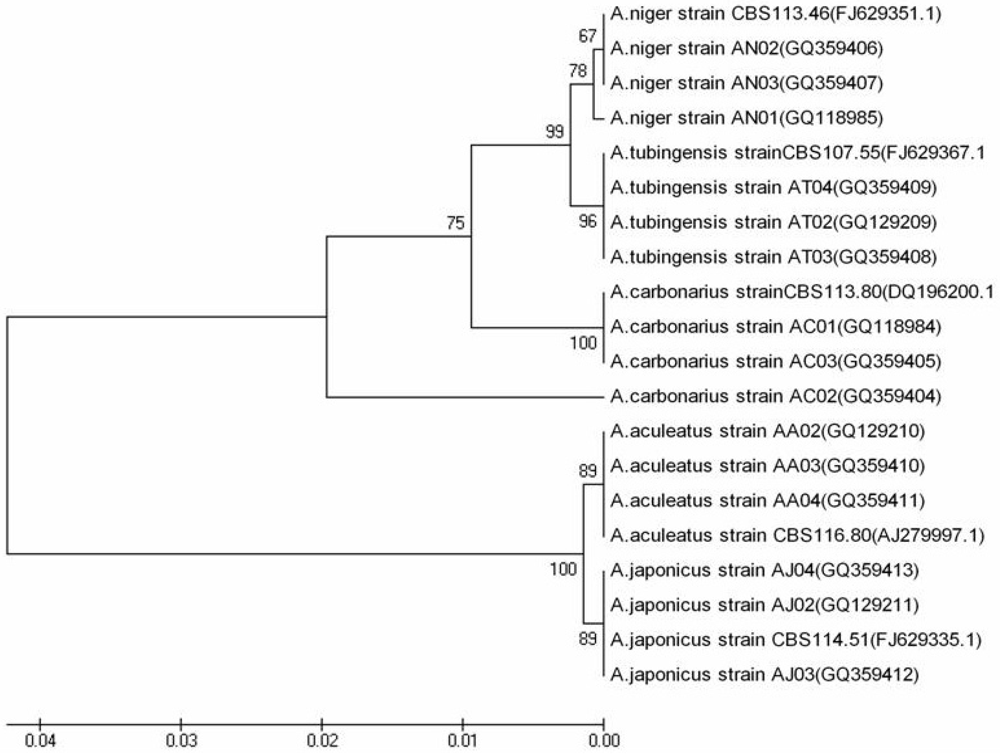

3.2. OTA Producing Ability of Aspergillus Isolates
| Species | Total No. of isolates and percentage | No of OTA producers (%) | OTA content (range µg mL−1) | ||
|---|---|---|---|---|---|
| 2006 | 2007 | 2006 | 2007 | ||
| A. niger | (57.4%) 112 | (25.5%) 41 | 0 | 0 | ND |
| A. carbonarius | (16.4%) 32 | (46.6%) 75 | 11 (34.4%) | 35 (46.7%) | 0.30 ± 0.10–3.0 ± 0.60 |
| A. tubingensis | (14.4%) 28 | (27.3%) 44 | 6 (21.4%) | 5 (11.4%) | 0.05 ± 0.01–0.8 ± 0.38 |
| A. japonicus | (9.7%) 19 | (6.2%) 1 | 5 (26.3%) | 1 (100.0%) | 0.05 ± 0.02–0.4 ± 0.09 |
| A. aculeatus | (2.1%) 4 | (0.0%) 0 | 0 | 0 | ND |
| Total | 195 | 161 | 22 (11.3%) | 41 (25.5%) | - |
4. Discussion
Acknowledgements
Conflict of Interest
References
- Battilani, P.; Giorni, P.; Bertuzzi, T.; Formenti, S.; Pietri, A. Black aspergilli and ochratoxin A in grapes in Italy. Int. J. Food Microbiol. 2006, 111, S53–S60. [Google Scholar]
- Serra, R.; Mendonca, C.; Venàncio, A. Ochratoxin A occurrence and formation in Portuguese wine grapes at various stages of maturation. Int. J. Food Microbiol. 2006, 111, S35–S39. [Google Scholar]
- Spadaro, D.; Lorè, A.; Garibaldi, A.; Gullino, M.L. Occurrence of ochratoxin A before bottling in DOC and DOCG wines produced in Piedmont (northern Italy). Food Control 2010, 21, 1294–1297. [Google Scholar]
- Castegnaro, M.; Mohr, U.; Pfohl-Leszkowicz, A.; Esteve, J.; Steinmann, J.; Tillmann, J.; Michelson, T.; Bartsch, J. Sex- and strain-specific induction of renal tumors by ochratoxin A in rats correlated with DNA adduction. Int. J. Cancer 1998, 77, 70–75. [Google Scholar]
- Radic, B.; Fuchs, R.; Peraica, M.; Lucis, A. Ochratoxin A in human sera in the area with endemic nephropathy in Croatia. Toxicol. Lett. 1997, 91, 105–109. [Google Scholar]
- Wafa, E.W.; Yahya, R.S.; Sobh, M.A.; Eraky, I.; El-Baz, M.; El-Gayar, H.A.M.; Betbeder, A.M.; Creppy, E.E. Human ochratoxicosis and nephropathy in Egypt: A preliminary study. Hum. Exp. Toxicol. 1998, 17, 124–129. [Google Scholar]
- International Agency for Research on Cancer (IARC), Some Naturally Occurring Substances: Food Items and Constituents. Heterocyclic Aromatic Amines and Mycotoxins; World Health Organization: Lyon, France, 1993; 56.
- Otteneder, H.; Majerus, P. Occurrence of ochratoxin A (OTA) in wines: influence of the type of wine and its geographical origin. Food Addit. Contam. 2000, 17, 793–798. [Google Scholar]
- Zimmerli, B.; Dick, R. Ochratoxin A in table wine and grape-juice: Occurrence and risk assessment. Food Addit. Contam. 1996, 13, 655–668. [Google Scholar]
- Leong, S.L.; Hocking, A.D.; Scott, E.S. Aspergillus species producing ochratoxin A: Isolation from vineyard soils and infection of Semillon bunches in Australia. J. Appl. Microbiol. 2007, 102, 124–133. [Google Scholar] [CrossRef]
- Magnoli, C.; Astoreca, A.; Ponsone, M.L.; Fernández-Juri, M.G.; Barberis, C.; Dalcero, A.M. Ochratoxin A and Aspergillus section Nigri in peanut seeds at different months of storage in Córdoba, Argentina. Int. J. Microbiol. 2007, 119, 213–218. [Google Scholar] [CrossRef]
- Spadaro, D.; Patharajan, S.; Karthikeyan, M.; Lorè, A.; Garibaldi, A.; Gullino, M.L. Specific PCR primers for the detection of isolates of Aspergillus carbonarius producing ochratoxin A on grapevine. Ann. Microbiol. 2011, 61, 267–272. [Google Scholar] [CrossRef]
- Cabañes, F.J.; Accensi, F.; Bragulat, M.R.; Abarca, M.L.; Castellá, G.; Minguez, S.; Pons, A. What is the source of ochratoxin A in wine? Int. J. Food Microbiol. 2002, 79, 213–215. [Google Scholar] [CrossRef]
- Perrone, G.; Mule, G.; Susca, A.; Battilani, P.; Pietri, A.; Logrieco, A. Ochratoxin A production and amplified fragment length polymorphism analysis of Aspergillus carbonarius, Aspergillus tubingensis, and Aspergillus niger strains isolated from grapes in Italy. Appl. Environ. Microbiol. 2006, 72, 680–685. [Google Scholar] [CrossRef]
- Horie, Y. Productivity of ochratoxin A of Aspergillus carbonarius in Aspergillus section Nigri. Nippon Kingakukai Kaiho 1995, 36, 73–76. [Google Scholar]
- Abarca, M.L.; Bragulat, M.R.; Castellá, G.; Cabañes, F.J. Ochratoxin A production by strains of Aspergillus niger var. niger. Appl. Environ. Microbiol. 1994, 60, 2650–2652. [Google Scholar]
- Kusters van Someren, M.A.; Samson, R.A.; Visser, J. The use of RFLP analysis in classification of the black Aspergilli: Reinterpretation of Aspergillus niger aggregate. Curr. Genet. 1991, 19, 21–26. [Google Scholar] [CrossRef]
- Battilani, P.; Giorni, P.; Pietri, A. Epidemiology of toxin-producing fungi and ochratoxin A occurrence in grape. Eur. J. Plant Pathol. 2003, 109, 715–722. [Google Scholar]
- Abarca, M.L.; Accensi, F.; Bragulat, M.R.; Cabañes, F.J. Current importance of ochratoxin A-producing Aspergillus species. J. Food Prot. 2001, 64, 903–906. [Google Scholar]
- Lopez de Cerain, A.; Gonzàlez-Peñas, E.; Jiménez, A.M.; Bello, J. Contribution to the study of ochratoxin A in Spanish wines. Food Addit. Contam. 2002, 19, 1058–1064. [Google Scholar]
- Battilani, P.; Pietri, A.; Bertuzzi, T.; Languasco, L.; Giorni, P.; Kozakiewicz, Z. Occurrence of ochratoxin A-producing fungi in grapes grown in Italy. J. Food Prot. 2003, 66, 633–636. [Google Scholar]
- Bellí, N.; Mitchell, D.; Marín, S.; Alegre, I.; Ramos, A.J.; Magan, N.; Sanchis, V. Ochratoxin A-producing fungi in Spanish wine grapes and their relationship with meteorological conditions. Eur. J. Plant Pathol. 2005, 113, 233–239. [Google Scholar]
- Gullino, M.L.; Lorè, A.; Muthusamy, K.; Spadaro, D.; Garibaldi, A. Monitoring of ochratoxin producing Aspergillus carbonarius in grapevine using molecular markers. Phytopathology 2008, 98, S64. [Google Scholar]
- Serra, R.; Braga, A.; Venàncio, A. Mycotoxin-producing and other fungi isolated from grapes for wine production, with particular emphasis on ochratoxin A. Res. Microbiol. 2005, 156, 515–521. [Google Scholar]
- Geiser, D.M.; Klich, M.A.; Frisvad, J.C.; Peterson, S.W.; Varga, J.; Samson, R.A. The current status of species recognition and identification in Aspergillus. Stud. Mycol. 2007, 59, 1–10. [Google Scholar] [CrossRef]
- Samson, R.A.; Noomin, P.; Meijer, M.; Houbraken, J.; Frisvad, J.C.; Varga, J. Diagnostic tools to identify black aspergilli. Stud. Mycol. 2007, 59, 129–145. [Google Scholar]
- Perrone, G.; Susca, A.; Cozzi, K.; Ehrlich, J.; Varga, J.; Frisvad, J.C.; Meijer, M.; Noomin, P.; Mahakarnchanakul, W.; Samson, R.A. Biodiversity of Aspergillus species in some important agricultural products. Stud. Mycol. 2007, 59, 53–66. [Google Scholar] [CrossRef]
- Ponsone, M.L.; Combina, M.; Dalcero, A.; Chulze, S.N. Ochratoxin A and ochratoxigenic Aspergillus species in Argentinean wine grapes cultivated under organic and non-organic systems. Int. J. Food Microbiol. 2007, 114, 131–135. [Google Scholar] [CrossRef]
- Samson, R.A.; Houbraken, J.A.M.P.; Kuijpers, A.F.A.; Frank, J.M.; Frisvad, J.C. New ochratoxin A or sclerotium producing species in Aspergillus section Nigri. Stud. Mycol. 2004, 50, 45–61. [Google Scholar]
- Mégnégneau, B.; Debets, F.; Hoekstra, R.F. Genetic variability and relatedness in the complex group of black aspergilli based on random amplification of polymorphic DNA. Curr. Genet. 1993, 23, 323–329. [Google Scholar]
- Varga, J.; Kevei, F.; Fekete, C.; Coenen, A.; Kozakiewicz, Z.; Croft, J.H. Restriction fragment length polymorphism in the mitochondrial DNAs of Aspergillus niger aggregate. Mycol. Res. 1993, 97, 1207–1212. [Google Scholar] [CrossRef]
- Varga, J.; Kevei, F.; Vriesema, A.; Debets, F.; Kozakiewicz, Z.; Croft, J.H. Mitochondrial DNA restriction fragment length polymorphism in field isolates of the Aspergillus niger aggregate. Can. J. Microbiol. 1994, 40, 612–621. [Google Scholar] [CrossRef]
- Accensi, F.; Cano, J.; Figuera, L.; Abarca, M.L.; Cabañes, F.J. New PCR method to differentiate species in the Aspergillus niger aggregate. FEMS Microbiol. Lett. 1999, 180, 191–196. [Google Scholar] [CrossRef]
- Martínez-Culebras, P.V.; Ramón, D. An ITS-RFLP method to identify black Aspergillus isolates responsible for OTA contamination in grapes and wine. Int. J. Food Microbiol. 2007, 13, 147–153. [Google Scholar]
- Klich, M.A. Identification of Common Aspergillus Species; Centraalbureau voor Schimmelcultures: Utrecht, The Netherlands, 2002. [Google Scholar]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and Direct Sequencing of Fungal Ribosomal RNA Genes for Phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Shinsky, J.J., White, T.J., Eds.; Academic Press: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar]
- Hanahan, D. Studies on transformation of Escherichia coli with plasmids. J. Mol. Biol. 1983, 166, 557–580. [Google Scholar] [CrossRef]
- Hills, D.M.; Bull, J.J. An empirical test of bootstrapping as a method for assessing confidence in phylogenetic analysis. Syst. Biol. 1993, 42, 182–192. [Google Scholar]
- Kumar, S.; Tamura, K.; Jakobsen, I.B.; Nei, M. MEGA2: Molecular evolutionary genetic analysis software. Bioinformatics 2001, 17, 1244–1245. [Google Scholar]
- Parenicová, L.; Skouboe, P.; Frisvad, J.; Samson, R.; Rossen, L.; Hoor-Suykerbyuk, M.T.; Visser, J. Combined molecular and biochemical approach identifies Aspergillus japonicus and Aspergillus aculeatus as two species. Appl. Environ. Microbiol. 2001, 67, 521–527. [Google Scholar] [CrossRef]
- Martínez-Culebras, V.; Sempere, C.A.; Hervás, S.M.; Elizaquivel, P.; Aznar, R.; Ramón, D. Molecular characterization of the black Aspergillus isolates responsible for ochratoxin A contamination in grapes and wine in relation to taxonomy of Aspergillus section Nigri. Int. J. Food Microbiol. 2009, 132, 33–41. [Google Scholar] [CrossRef]
- Palencia, E.R.; Klich, M.A.; Glenn, A.E. Use of a rep-PCR system to predict species in the Aspergillus section Nigri. J. Microbiol. Methods 2009, 79, 1–7. [Google Scholar] [CrossRef]
- Battilani, P.; Pietri, A. Ochratoxin A in grapes and wine. Eur. J. Plant Pathol. 2002, 108, 639–643. [Google Scholar]
- Bejaoui, H.; Mathieu, F.; Taillandier, P.; Lebrihi, A. Black aspergilli and ochratoxin A production in French vineyards. Int. J. Food Microbiol. 2006, 111, S46–S52. [Google Scholar]
- Da Rocha Rosa, C.A.; Palacios, V.; Combina, M.; Fraga, M.E.; De Oliveira Rekson, A.; Magnoli, C.E.; Dalcero, A.M. Potential ochratoxin A producers from wine grapes in Argentina and Brazil. Food Addit. Contam. 2002, 19, 408–414. [Google Scholar]
- Bellí, N.; Bau, M.; Marín, S.; Abarca, M.L.; Ramos, A.J.; Bragulat, M.R. Mycobiota and ochratoxin A producing fungi from Spanish wine grapes. Int. J. Food Microbiol. 2006, 111, S40–S45. [Google Scholar]
- Leong, S.L.; Hocking, A.D.; Scott, E.S. Effect of temperature and water activity on growth and ochratoxin A production by Australian Aspergillus carbonarius and A. niger isolates on a simulated grape juice medium. Int. J. Food Microbiol. 2006, 110, 209–216. [Google Scholar] [CrossRef]
- Palacios-Cabrera, H.; Taniwaki, M.H.; Hashimoto, J.M.; Menezes, H.C. Growth of Aspergillus ochraceus, A. carbonarius and A. niger on culture media at different water activities and temperatures. Braz. J. Microbiol. 2005, 36, 24–28. [Google Scholar] [CrossRef]
- Spadaro, D.; Patharajan, S.; Lorè, A.; Gullino, M.L.; Garibaldi, A. Effect of pH, water activity and temperature on the growth and level of ochratoxin A produced by three strains of Aspergillus carbonarius isolated from Italian vineyards. Phytopathol. Mediterr. 2010, 49, 65–73. [Google Scholar]
- Bragulat, M.R.; Abarca, M.L.; Cabañes, F.J. An easy screening method for fungi producing ochratoxin A in pure culture. Int. J. Food Microbiol. 2001, 71, 139–144. [Google Scholar]
- Bau, M.; Bragulat, M.R.; Abarca, M.L.; Minguez, S.; Cabañes, F.J. Ochratoxin species from Spanish wine grapes. Int. J. Food Microbiol. 2005, 98, 125–130. [Google Scholar]
- Medina, A.; Mateo, R.; López-Ocaña, L.; Valle-Algarra, F.M.; Jimenez, M. Study of Spanish grape mycobiota and ochratoxin A production by isolates of Aspergillus tubingensis and other members of Aspergillus section Nigri. Appl. Environ. Microbiol. 2005, 71, 4696–4702. [Google Scholar] [CrossRef]
- Accensi, F.; Abarca, M.L.; Cano, J.; Figuera, L.; Cabañes, F.J. Distribution of ochratoxin A producing strains in the A. niger aggregate. Antonie Leeuwenhoek 2001, 79, 365–370. [Google Scholar] [CrossRef]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Spadaro, D.; Patharajan, S.; Lorè, A.; Garibaldi, A.; Gullino, M.L. Ochratoxigenic Black Species of Aspergilli in Grape Fruits of Northern Italy Identified by an Improved PCR-RFLP Procedure. Toxins 2012, 4, 42-54. https://doi.org/10.3390/toxins4020042
Spadaro D, Patharajan S, Lorè A, Garibaldi A, Gullino ML. Ochratoxigenic Black Species of Aspergilli in Grape Fruits of Northern Italy Identified by an Improved PCR-RFLP Procedure. Toxins. 2012; 4(2):42-54. https://doi.org/10.3390/toxins4020042
Chicago/Turabian StyleSpadaro, Davide, Subban Patharajan, Alessia Lorè, Angelo Garibaldi, and Maria Lodovica Gullino. 2012. "Ochratoxigenic Black Species of Aspergilli in Grape Fruits of Northern Italy Identified by an Improved PCR-RFLP Procedure" Toxins 4, no. 2: 42-54. https://doi.org/10.3390/toxins4020042
APA StyleSpadaro, D., Patharajan, S., Lorè, A., Garibaldi, A., & Gullino, M. L. (2012). Ochratoxigenic Black Species of Aspergilli in Grape Fruits of Northern Italy Identified by an Improved PCR-RFLP Procedure. Toxins, 4(2), 42-54. https://doi.org/10.3390/toxins4020042






