Serine Protease Autotransporters of Enterobacteriaceae (SPATEs): Biogenesis and Function
Abstract
:1. Introduction
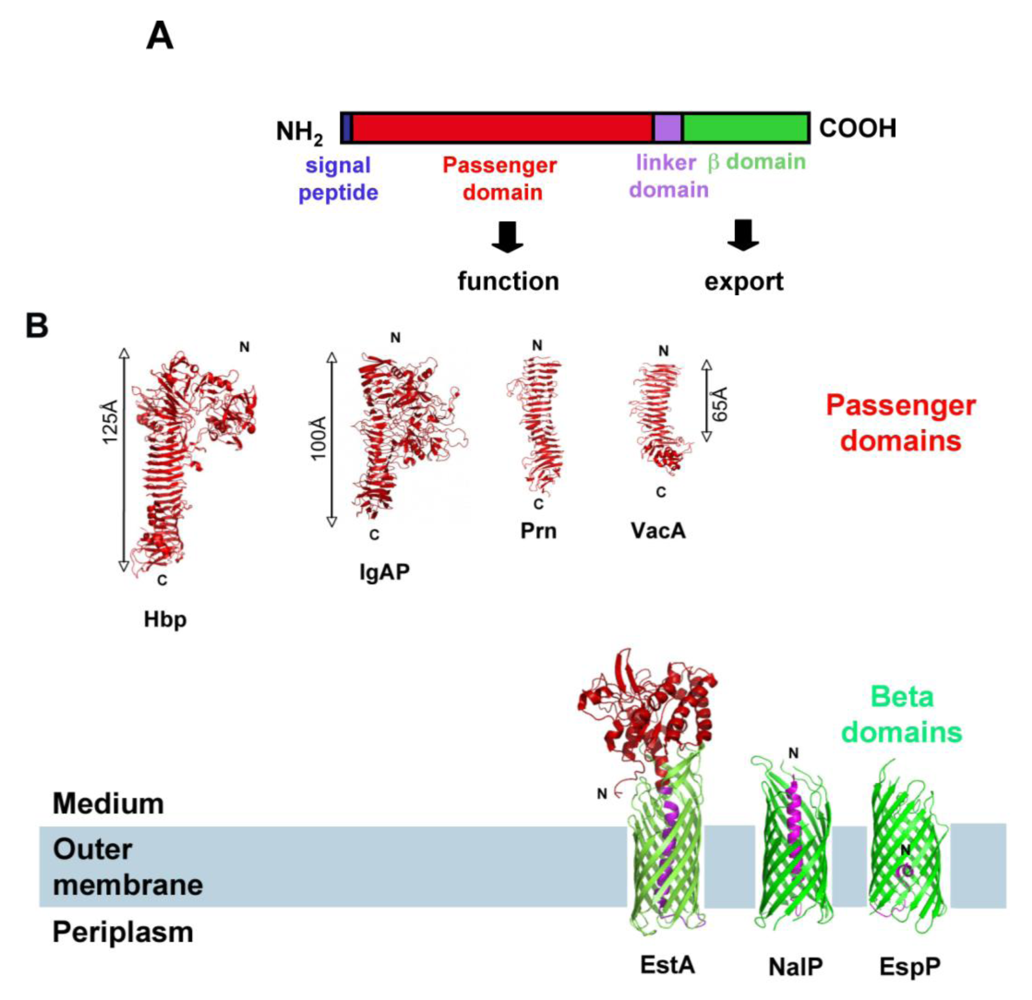
1.1. The Autotransporter Pathway
1.2. The SPATE Subfamily of Autotransporters
2. SPATEs Genes: Location and Evolution
3. Regulation of Expression
4. Biogenesis
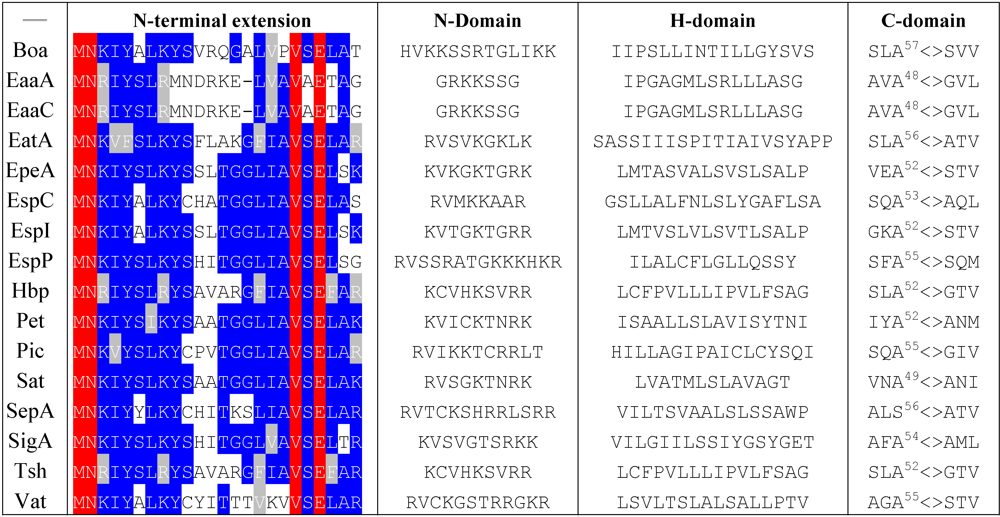
4.1. Export through the Inner Membrane
 |
4.2. Transport through the Periplasm
4.3. Transport through the Outer Membrane
4.4. Cleavage
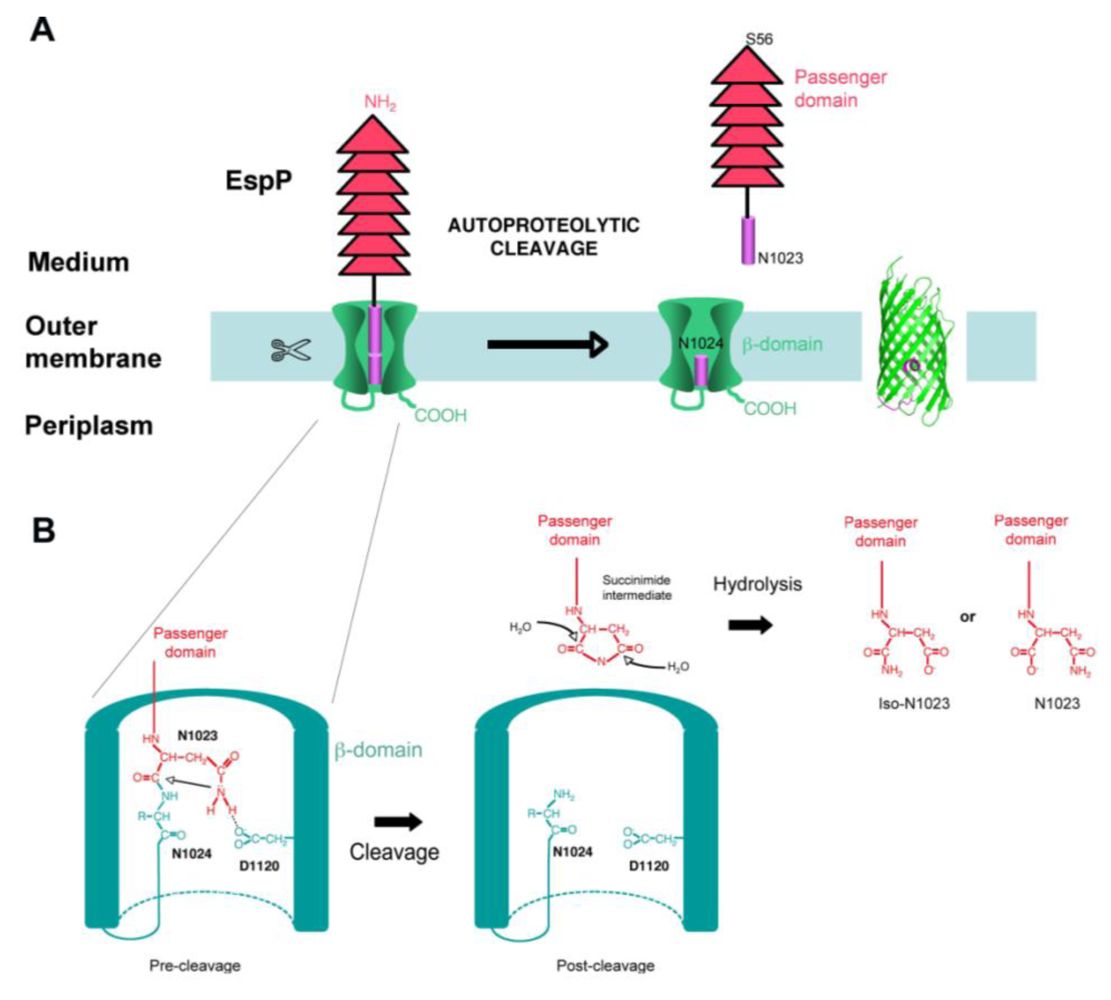
5. Function
5.1. Proteolytic Activity
| Cleaved | Not cleaved | |
|---|---|---|
| EatA | AAPM-pNA, AAPL-pNA [25] | |
| EpeA | pepsin A, gelatin, mucin [16] | |
| EspC | fodrin [71], hemoglobin [72], pepsin, factor V, spectrin (fodrin) [20,40] | Human IgA1 [71], mucin [40], lysozyme [73] |
| EspI | pepsin A, apolipoprotein A1 [26] | IgA1, hemoglobin, bovine serum albumin, α2-macroglobulin, haptoglobin, thrombin, collagene type 3, trypsin, high density lipoprotein, low density lipoprotein, very low density lipoprotein, trypsin, transferrin, lactoferrin, pepsinogen, gelatin, casein [26] |
| EspP | pepsin A, human coagulation factor V [14], casein [15], apolipoprotein A1 [26], AAPL-pNA [40] | human IgA1, bovine serum albumin, α2-macroglobulin, transferrin, lactoferrin, pepsinogen [14], mucin, spectrin (fodrin) [40] |
| Hbp | hemoglobin [5] | albumin, human lactoferrin, human immunoglobulin A1 [5] |
| Pet | casein, gelatin [17], pepsin, human coagulation factor V, spectrin [19,40] | actin [75], mucin [19,40] |
| Pic | gelatin, ovomucin, bovine mucin, murine mucin [18], human spectrin (fodrin), pepsin A, human coagulation factor V [19,40], mucin [19] | casein, IgA, IgM, IgG, hog gastric mucin [18] ovine spectrin [40] |
| Sat | casein [22], factor V, spectrin [40] | IgA1 [22], hemoglobin, mucin, pepsin [40] |
| SepA | FLF-pNA, VPF-pNA, AAPF-pNA, AAPM-pNA [70] | IgA1, gelatin [28], angiotensin-I, egg lysozyme [70], fibronectin, mucin, pepsin, factor V, spectrin (fodrin) [40] |
| SigA | casein [29], fodrin [45] | |
| Tsh | mucin, factor V [40] | human and chicken IgA, casein, pepsin A [40,42], spectrin [40] |
| Vat | casein [35] |
5.2. Role in Pathogenesis
6. Conclusions
Acknowledgments
References
- Henderson, I.R.; Nataro, J.P. Virulence functions of autotransporter proteins. Infect. Immun. 2001, 69, 1231–1243. [Google Scholar]
- Henderson, I.R.; Navarro-Garcia, F.; Nataro, J.P. The great escape: structure and function of the autotransporter proteins. Trends Microbiol. 1998, 6, 370–378. [Google Scholar] [PubMed]
- Dautin, N.; Bernstein, H.D. Protein secretion in gram-negative bacteria via the autotransporter pathway. Annu. Rev. Microbiol. 2007, 61, 89–11. [Google Scholar]
- Emsley, P.; Charles, I.G.; Fairweather, N.F.; Isaacs, N.W. Structure of Bordetella pertussis virulence factor P.69 pertactin. Nature 1996, 381, 90–92. [Google Scholar] [CrossRef] [PubMed]
- Otto, B.R.; Sijbrandi, R.; Luirink, J.; Oudega, B.; Heddle, J.G.; Mizutani, K.; Park, S.Y.; Tame, J.R. Crystal structure of hemoglobin protease, a heme binding autotransporter protein from pathogenic Escherichia coli. J. Biol. Chem. 2005, 280, 17339–17345. [Google Scholar] [PubMed]
- Kajava, A.V.; Steven, A.C. The turn of the screw: variation of the abundant β-solenoid motif in passenger domains of type V secretory proteins. J. Struct. Biol. 2006, 155, 306–315. [Google Scholar]
- Gangwer, K.A.; Mushrush, D.J.; Stauff, D.L.; Spiller, B.; McClain, M.S.; Cover, T.L.; Borden Lacy, D. Crystal structure of the Helicobacter pylori vacuolating toxin p55 domain. Proc. Natl. Acad. Sci. USA 2007, 104, 16293–16298. [Google Scholar]
- Johnson, T.A.; Qiu, J.; Plaut, A.G.; Holyoak, T. Active-site gating regulates substrate selectivity in a chymotrypsin-like serine protease: the structure of Haemophilus influenzae immunoglobulin A1 protease. J. Mol. Biol. 2009, 389, 559–574. [Google Scholar]
- Van den Berg, B. Crystal structure of a full-length autotransporter. J. Mol. Biol. 2010, 396, 627–633. [Google Scholar]
- Oomen, C.J.; van Ulsen, P.; van Gelder, P.; Feijen, M.; Tommassen, J.; Gros, P. Structure of the translocator domain of a bacterial autotransporter. EMBO J. 2004, 23, 1257–1266. [Google Scholar]
- Barnard, T.J.; Dautin, N.; Lukacik, P.; Bernstein, H.D.; Buchanan, S. Autotransporter structure reveals intra-barrel cleavage followed by conformational changes. Nat. Struct. Mol. Biol. 2007, 14, 1214–1220. [Google Scholar]
- De, E.; Saint, N.; Glinel, K.; Meli, A.C.; Levy, D.; Jacob-Dubuisson, F. Influence of the passenger domain of a model autotransporter on the properties of its translocator domain. Mol. Membr. Biol. 2008, 25, 192–202. [Google Scholar]
- Ieva, R.; Skillman, K.; Bernstein, H.D. Incorporation of a polypeptide segment into the beta-domain pore during the assembly of a bacterial autotransporter. Mol. Microbiol. 2008, 67, 188–201. [Google Scholar]
- Brunder, W.; Schmidt, H.; Karch, H. EspP, a novel extracellular serine protease of enterohaemorrhagic Escherichia coli O157:H7 cleaves human coagulation factor V. Mol Microbiol. 1997, 24, 767–778. [Google Scholar] [PubMed]
- Djafari, S.; Ebel, F.; Deibel, C.; Kramer, S.; Hudel, M.; Chakraborty, T. Characterization of an exported protease from Shiga toxin-producing Escherichia coli. Mol. Microbiol. 1997, 25, 771–784. [Google Scholar]
- Leyton, D.L.; Sloan, J.; Hill, R.E.; Doughty, S.; Hartland, E.L. Transfer region of pO113 from enterohemorrhagic Escherichia coli: similarity with R64 and identification of a novel plasmid-encoded autotransporter, EpeA. Infect. Immun. 2003, 71, 6307–6319. [Google Scholar]
- Eslava, C.E.; Navarro-García, F.; Czeczulin, J.R.; Henderson, I.R.; Cravioto, A.; Nataro, J.P. Pet, an autotransporter protein enterotoxin from enteroaggregative Escherichia coli. Infect. Immun. 1998, 66, 3155–3163. [Google Scholar]
- Henderson, I.R.; Czeczulin, J.; Eslava, C.; Noriega, F.; Nataro, J.P. Characterization of Pic, a secreted protease of Shigella flexneri and enteroaggregative Escherichia coli. Infect. Immun. 1999, 67, 5587–5596. [Google Scholar]
- Parham, N.J.; Srinivasan, U.; Desvaux, M.; Foxman, B.; Marrs, C.F.; Henderson, I.R. PicU, a second serine protease autotransporter of uropathogenic Escherichia coli. FEMS Microbiol. Lett. 2004, 230, 73–83. [Google Scholar]
- Stein, M.; Kenny, B.; Stein, M.A.; Finlay, B.B. Characterization of EspC, a 110-kilodalton protein secreted by enteroaggregative Escherichia coli produces cellular damage associated with fodrin disruption. Infect. Immun. 1996, 68, 5920–5927. [Google Scholar]
- Otto, B.R.; Van Dooren, S.J.M.; Nuijens, J.H.; Luirink, J.; Oudega, B. Characterization of a hemoglobin protease secreted by the pathogenic Escherichia coli strain EB1. J. Exp. Med. 1998, 188, 1091–1103. [Google Scholar]
- Guyer, D.M.; Henderson, I.R.; Nataro, J.P.; Mobley, H.L. Identification of Sat, an autotransporter toxin produced by uropathogenic Escherichia coli. Mol. Microbiol. 2000, 38, 53–66. [Google Scholar]
- Provence, D.L.; Curtis, R.I. Isolation and characterization of a gene involved in hemagglutination by an avian pathogenic Escherichia coli strain. Infect. Immun. 1994, 62, 1369–1380. [Google Scholar]
- Salvadori, M.R.; Yano, T.; Carvalho, H.E.; Parreira, V.R.; Gyles, C.L. Vacuolating cytotoxin produced by avian pathogenic Escherichia coli. Avian Dis. 2001, 45, 43–51. [Google Scholar]
- Patel, S.K.; Dotson, J.; Allen, K.P.; Fleckenstein, J.M. Identification and molecular characterization of EatA, an autotransporter protein of enterotoxigenic Escherichia coli. Infect. Immun. 2004, 72, 1786–1794. [Google Scholar]
- Schmidt, H.; Zhang, W.L.; Hemmrich, U.; Jelacic, S.; Brunder, W.; Tarr, P.I.; Dobrindt, U.; Hacker, J.; Karch, H. Identification and characterization of a novel genomic island integrated at selC in locus of enterocyte effacement-negative, Shiga toxin-producing Escherichia coli. Infect. Immun. 2001, 69, 6863–6873. [Google Scholar]
- Sandt, C.H.; Hill, C.H. Four different genes responsible for nonimmune immunoglobulin-binding activities within a single strain of Escherichia coli. Infect. Immun. 2000, 68, 2205–2214. [Google Scholar]
- Benjelloun-Touimi, Z.; Sansonetti, P.J.; Parsot, C. SepA, the major extracellular protein of Shigella flexneri: autonomous secretion and involvement in tissue invasion. Mol. Microbiol. 1995, 17, 123–135. [Google Scholar]
- Al-Hasani, K.; Henderson, I.R.; Sakellaris, H.; Rajakumar, K.; Grant, T.; Nataro, J.P.; Robins-Browne, R.; Adler, B. The sigA gene, which is borne on the she pathogenicity island of Shigella flexneri 2a encodes an exported cytopathic protease involved in intestinal fluid accumulation. Infect. Immun. 2000, 68, 2457–2463. [Google Scholar]
- Parham, N.J.; Pollard, S.J.; Desvaux, M.; Scott-Tucker, A.; Liu, C.; Fivian, A.; Henderson, I.R. Distribution of the serine protease autotransporters of the Enterobacteriaceae among extraintestinal clinical isolates of Escherichia coli. J. Clin. Microbiol. 2005, 43, 4076–4082. [Google Scholar]
- Yen, Y.T.; Kostakioti, M.; Henderson, I.R.; Stathopoulos, C. Common themes and variations in serine protease autotransporters. Trends Microbiol. 2008, 16, 370–379. [Google Scholar]
- Dozois, C.M.; Dho-Moulin, M.; Bree, A.; Fairbrother, J.M.; Desautels, C.; Curtiss, R., 3rd. Relationship between the Tsh autotransporter and pathogenicity of avian Escherichia coli and localization and analysis of the Tsh genetic region. Infect. Immun. 2000, 68, 4145–4154. [Google Scholar]
- Mellies, J.L.; Navarro-Garcia, F.; Okeke, I.; Frederickson, J.; Nataro, J.P.; Kaper, J.B. espC pathogenicity island of enteropathogenic Escherichia coli encodes an enterotoxin. Infect. Immun. 2001, 69, 315–324. [Google Scholar]
- Al-Hasani, K.; Rajakumar, K.; Bulach, D.; Robins-Browne, R.; Adler, B.; Sakellaris, H. Genetic organization of the she pathogenicity island in Shigella flexneri 2a. Microb. Pathog. 2001, 30, 1–8. [Google Scholar]
- Parreira, V.R.; Gyles, C.L. A novel pathogenicity island integrated adjacent to the thrW tRNA gene of avian pathogenic Escherichia coli encodes a vacuolating autotransporter toxin. Infect. Immun. 2003, 71, 5087–5096. [Google Scholar]
- Davis, J.; Smith, A.L.; Hughes, W.R.; Golomb, M. Evolution of an autotransporter: Domain shuffling and lateral transfer from pathogenic Haemophilus to Neisseria. J. Bacteriol. 2001, 183, 4626–4635. [Google Scholar]
- Yen, M.R.; Peabody, C.R.; Partovi, S.M.; Zhai, Y.; Tseng, Y.-H.; Saier, M.H., Jr. Protein-translocating outer membrane porins of gram-negative bacteria. Biochim. Biophys. Acta 2002, 1562, 6–31. [Google Scholar]
- Loveless, B.J.; Saier, M.H., Jr. A novel family of channel-forming, autotransporting, bacterial virulence factors. Mol. Membr. Biol. 1997, 14, 113–123. [Google Scholar]
- Brockmeyer, J.; Bielaszewska, M.; Fruth, A.; Bonn, M.L.; Mellmann, A.; Humpf, H.A.; Karch, H. Subtypes of the plasmid-encoded serine protease EspP in Shiga toxin-producing Escherichia coli: distribution, secretion, and proteolytic activity. Appl. Environ. Microbiol. 2007, 73, 6351–6359. [Google Scholar] [CrossRef] [PubMed]
- Dutta, P.R.; Cappello, R.; Navarro-Garcia, F.; Nataro, J.P. Functional comparison of serine protease autotransporters of Enterobactericeae. Infect. Immun. 2002, 70, 7105–7113. [Google Scholar]
- Kenny, B.; Finlay, B.B. Protein secretion by enteropathogenic Escherichia coli is essential for transducing signals to epithelial cells. Proc. Natl. Acad. Sci. USA 1995, 92, 7991–7995. [Google Scholar]
- Stathopoulos, C.; Provence, D.L.; Curtiss, R., III. Characterization of the avian pathogenic Escherichia coli hemagglutinin Tsh, a member of the immunoglobulin A protease-type family of autotransporters. Infect. Immun. 1999, 67, 772–781. [Google Scholar]
- Jarvis, K.G.; Giron, J.A.; Jerse, A.E.; McDaniel, T.K.; Donnenberg, M.S.; Kaper, J.B. Enteropathogenic Escherichia coli contains a putative type III secretion system necessary for the export of proteins involved in attaching and effacing lesion formation. Proc. Natl. Acad. Sci. USA 1995, 92, 7996–8000. [Google Scholar]
- Bellini, E.M.; Elias, W.P.; Gomes, T.A.; Tanaka, T.L.; Taddei, C.R.; Huerta, E.; Navarro-Garcia, F.; Martinez, M.B. Antibody response against plasmid-encoded toxin (Pet) and the protein involved in intestinal colonization (Pic) in children with diarrhea produced by enteroaggregative Escherichia coli. FEMS Immunol. Med. Microbial. 2005, 43, 259–264. [Google Scholar]
- Al-Hasani, K.; Navarro-Garcia, F.; Huerta, J.; Sakellaris, H.; Adler, B. The immunogenic SigA enterotoxin of Shigella flexneri 2a binds to HEp-2 cells and induces fodrin redistribution in intoxicated cells. PLoS ONE 2010, 4, e8223. [Google Scholar]
- Elliott, S.J.; Sperandio, V.; Giron, J.A.; Shin, S.; Mellies, J.L.; Wainwright, L.; Hutcheson, S.W.; McDaniel, T.K.; Kaper, J.P. The locus of enterocyte effacement (LEE)-encoded regulator controls expression of both LEE- and non-LEE-encoded virulence factors in enteropathogenic and enterohemorrhagic Escherichia coli. Infect. Immun. 2000, 68, 6115–6126. [Google Scholar]
- Szabadi, R.L.; Peterson, J.H.; Skillman, K.M.; Bernstein, H.D. An unusual signal peptide facilitates late steps in the biogenesis of a bacterial autotransporter. Proc. Natl. Acad. Sci. USA 2005, 102, 221–226. [Google Scholar]
- Peterson, J.H.; Szabadi, R.L.; Bernstein, H.D. An unusual signal peptide extension inhibits the binding of bacterial presecretory proteins to the signal recognition particle, trigger factor, and the SecYEG complex. J. Biol. Chem. 2006, 281, 9038–9048. [Google Scholar]
- Desvaux, M.; Scott-Tucker, A.; Turner, S.M.; Cooper, L.M.; Huber, D.; Nataro, J.P.; Henderson, I.R. A conserved extended signal peptide region directs posttranslational protein translocation via a novel mechanism. Microbiology 2007, 153, 59–70. [Google Scholar] [CrossRef] [PubMed]
- Jong, W.S.; Luirink, J. The conserved extension of the Hbp autotransporter signal peptide does not determine targeting pathway specificity. Biochem. Biophys. Res. Commun. 2008, 368, 522–527. [Google Scholar]
- Sijbrandi, R.; Urbanus, M.L.; ten Hagen-Jongman, C.M.; Bernstein, H.D.; Oudega, B.; Otto, B.R.; Luirink, J. Signal recognition particle (SRP)-mediated targeting and sec-dependent translocation of an extracellular Escherichia coli protein. J. Biol. Chem. 2003, 278, 4654–4659. [Google Scholar]
- Junker, M.; Schuster, C.C.; McDonnell, A.V.; Sorg, K.A.; Finn, M.C.; Berger, B.; Clark, P.L. Pertactin beta-helix folding mechanism suggests common themes for the secretion and folding of autotransporter proteins. Proc. Natl. Acad. Sci. USA 2006, 103, 4918–4923. [Google Scholar]
- Renn, J.; Clark, P.L. A conserved stable core structure in the passenger domain β-helix of autotransporter virulence proteins. Biopolymers 2008, 89, 420–427. [Google Scholar]
- Sauri, A.; Soprova, Z.; Wickstrom, D.; de Gier, J.W.; van der Schors, R.C.; Smit, A.B.; Jong, W.S.; Luirink, J. The Bam (Omp85) complex is involved in secretion of the autotransporter haemoglobin protease. Microbiology 2009, 155, 3982–3291. [Google Scholar]
- Ieva, R.; Bernstein, H.D. Interaction of an autotransporter passenger domain with BamA during its translocation across the bacterial outer membrane. Proc. Natl. Acad. Sci. USA 2009, 106, 19120–19125. [Google Scholar]
- Ruiz-Perez, F.; Henderson, I.R.; Leyton, D.L.; Rossiter, A.E.; Zhang, Y.; Nataro, J.P. Roles of periplasmic chaperone proteins in the biogenesis of serine protease autotransporters of Enterobacteriaceae. J. Bacteriol. 2009, 191, 6571–6583. [Google Scholar]
- Pohlner, J.; Halter, R.; Beyreuther, K.; Meyer, T.F. Gene structure and extracellular secretion of Neisseria gonorrhoeae IgA protease. Nature 1987, 325, 458–462. [Google Scholar] [CrossRef] [PubMed]
- Bernstein, H.D. Are bacterial “autotransporters” really transporters? Trends Microbiol. 2007, 15, 441–447. [Google Scholar]
- Veiga, E.; Sugawara, E.; Nikaido, H.; de Lorenzo, V.; Fernandez, L.A. Export of autotransported proteins proceeds through an oligomeric ring shaped by C-terminal domains. EMBO J. 2002, 21, 2122–2131. [Google Scholar]
- Voulhoux, R.; Bos, M.P.; Geurtsen, J.; Mols, M.; Tommassen, J. Role of a highly conserved bacterial protein in outer membrane protein assembly. Science 2003, 299, 262–265. [Google Scholar] [CrossRef] [PubMed]
- Hritonenko, V.; Kostakioti, M.; Stathopoulos, C. Quaternary structure of a SPATE autotransporter protein. Mol. Membr. Biol. 2006, 23, 466–474. [Google Scholar]
- Skillman, K.M.; Barnard, T.J.; Peterson, J.H.; Ghirlando, R.; Bernstein, H.D. Efficient secretion of a folded protein domain by a monomeric bacterial autotransporter. Mol. Microbiol. 2005, 58, 954–958. [Google Scholar]
- Jong, W.S.; ten Hagen-Jongman, C.M.; den Blaauwen, T.; Slotboom, D.J.; Tame, J.R.; Wickstrom, D.; de Gier, J.W.; Otto, B.R.; Luirink, J. Limited tolerance towards folded elements during secretion of the autotransporter Hbp. Mol. Microbiol. 2007, 63, 1524–1536. [Google Scholar]
- Dautin, N.; Barnard, T.J.; Anderson, D.E.; Bernstein, H.D. Cleavage of a bacterial autotransporter by an evolutionarily convergent autocatalytic mechanism. EMBO J. 2007, 26, 1942–1952. [Google Scholar]
- Schneemann, A.; Zhong, W.; Gallagher, T.M.; Rueckert, R.R. Maturation cleavage required for infectivity of a nodavirus. J. Virol. 1992, 66, 6728–6734. [Google Scholar]
- Kostakioti, M.; Stathopoulos, C. Role of the alpha-helical linker of the C-terminal translocator in the biogenesis of the serine protease subfamily of autotransporters. Infect. Immun. 2006, 74, 4961–4969. [Google Scholar]
- Boisen, N.; Ruiz-Perez, F.; Scheutz, F.; Krogfelt, K.A.; Nataro, J.P. High prevalence of serine protease autotransporter cytotoxins among strains of enteroaggregative Escherichia coli. Am. J. Trop. Med. Hyg. 2009, 80, 294–301. [Google Scholar]
- Brockmeyer, J.; Spelten, S.; Kucziua, T.; Bielaszewska, M.; Karch, H. Structure and function relationship of the autotransport and proteolytic activity of EspP from Shiga toxin-producing Escherichia coli. PLos ONE 2009, 4, e6100. [Google Scholar]
- Nishimura, K.; Tajima, N.; Yoon, Y.-H.; Park, S.-Y.; Tame, J.R.H. Autotransporter passenger proteins: virulence factors with common structural themes. J. Mol. Med. 2010. [Ahead of print].. [Google Scholar]
- Benjelloun-Touimi, Z.; Si Tahar, M.; Montecucco, C.; Sansonetti, P.J.; Parsot, C. SepA, the 110kDa protein secreted by Shigella flexneri: two-domain structure and proteolytic activity. Microbiology 1998, 144, 1815–1822. [Google Scholar] [CrossRef] [PubMed]
- Navarro-Garcia, F.; Canizalez-Roman, A.; Bao Quan, S.; Nataro, J.P.; Azamar, Y. The serine protease motif of EspC from enteropathogenic Escherichia coli produces epithelial damage by a mechanism different from that of Pet toxin from enteroaggregative E. coli. Infect. Immun. 2004, 72, 3609–3621. [Google Scholar] [CrossRef] [PubMed]
- Drago-Serrano, M.E.; Gavilanes Parra, S.; Manjarrez-Hernandez, A.H. EspC, an autotransporter protein secreted by enteropathogenic Escherichia coli (EPEC), displays protease activity on human hemoglobin. FEMS Microbiol. Lett. 2006, 265, 35–40. [Google Scholar]
- Salinger, N.; Kokona, B.; Fairman, R.; Okeke, I.N. The plasmid-encoded regulator activates factors conferring lysozyme resistance on enteopathogenic Escherichia coli strains. Appl. Environ. Microb. 2009, 75, 275–280. [Google Scholar]
- Navarro-Garcia, F.; Sears, C.; Eslava, C.; Cravioto, A.; Nataro, J.P. Cytoskeletal effect of Pet, the serine protease of enteroaggregative Escherichia coli. Infect. Immun. 1999, 67, 2184–2192. [Google Scholar]
- Vidal, J.E.; Navarro-Garcia, F. Efficient translocation of EspC into epithelial cells depends on enteropathogenic Escherichia coli and host cell contact. Infect. Immun. 2006, 74, 2293–2303. [Google Scholar]
- Vidal, J.E.; Navarro-Garcia, F. EspC translocation into epithelial cells by enteropathogenic Escherichia coli requires a concerted participation of type V and type III secretions systems. Cell Microbiol. 2008, 10, 1975–1986. [Google Scholar]
- Deane, J.E.; Abrusci, P.; Johnson, S. Timing is everything: the regulation of type III secretion. Cell. Mol. Life Sci. 2010, 67, 1065–1075. [Google Scholar]
- Blocker, A.J.; Deane, J.E.; Veenendaal, A.K.; Roversi, P.; Hodgkinson, J.L.; Johnson, S.; Lea, S. What’s the point of the type III secretion needle? Proc. Natl. Acad. Sci. USA 2008, 105, 6507–6513. [Google Scholar]
- Leiman, P.G.; Basler, M.; Ramagopal, U.A.; Bonanno, J.B.; Sauder, J.M.; Pukatzki, S.; Burley, S.K.; Almo, S.C.; Mekalanos, J.J. Type VI secretion apparatus and phage tail-associated protein complexes share a common evolutionary origin. Proc. Natl. Acad. Sci. USA 2009, 106, 4154–4159. [Google Scholar]
- Szabo, I.; Brutsche, S.; Tombola, F.; Moschioni, M.; Satin, B.; Telford, J.L.; Rappuoli, R.; Montecucco, C.; Papini, E.; Zoratti, M. Formation of anion-selective channels in the cell plasma membrane by the toxin VacA of Helicobacter pylori is required for its biological activity. EMBO J. 1999, 18, 5517–5527. [Google Scholar]
- Van Diemen, P.M.; Dziza, F.; Stevens, P.M.; Wallis, T.M. Identification of enterohemorrhagic Escherichia coli O26: H-genes required for intestinal colonization in calves. Infect. Immun. 2005, 73, 1735–1743. [Google Scholar]
- Dziva, F.; Mahajan, A.; Cameron, P.; Currie, C.; McKendrick, I.J.; Wallis, T.S.; Smith, D.G.; Stevens, M.P. EspP, a type V-secreted protease of enterohaemorrhagic Escherichia coli O157:H7, influences intestinal colonization of calves and adherence to bovine primary intestinal epithelial cells. FEMS Microbiol. Lett. 2007, 271, 258–264. [Google Scholar]
- Puttamreddy, S.; Cornick, N.A.; Minion, C. Genome-wide transposon mutagenesis reveals a role for pO157 genes in biofilm development in Escherichia coli O157:H7 EDL933. Infect. Immun. 2010. [Epub ahead of print].. [Google Scholar]
- Tzipori, S.; Karch, H.; Wachsmuth, K.I.; Robins-Browne, R.M.; O’Brien, A.D.; Lior, H.; Cohen, M.L.; Smithers, J.; Levine, M.M. Role of the 60-megadalton plasmid and Shiga-like toxins in the pathogenesis of infection caused by enterohemorrhagic Escherichia coli O157:H7 in gnotobiotic piglets. Infect. Immun. 1987, 55, 3117–3125. [Google Scholar]
- Otto, B.R.; van Dooren, S.J.; Dozois, C.M.; Luirink, J.; Oudega, B. Escherichia coli hemoglobin protease autotransporter contributes to synergistic abscess formation and heme-dependent growth of Bacteroides fragilis. Infect. Immun. 2002, 70, 5–10. [Google Scholar] [CrossRef] [PubMed]
- Navarro-Garcia, F.; Eslava, C.; Villaseca, J.M.; Lopez-Revilla, R.; Czeczulin, J.R.; Srinivas, S.; Nataro, J.P.; Cravioto, A. In vitro effects of a high-molecular-weight heat-labile enterotoxin from enteroaggregative Escherichia coli. Infect. Immun. 1998, 66, 3149–3154. [Google Scholar] [PubMed]
- Navarro-Garcia, F.; Canizalez-Roman, J.E.; Vidal, M.I.; Salazar, M.I. Intoxication of epithelial cells by plasmid-encoded toxin requires clathrin-mediated endocytosis. Microbiology 2007, 153, 2828–2838. [Google Scholar]
- Navarro-Garcia, F.; Canizalez-Roman, A.; Burlingame, K.E.; Teter, K.; Vidal, J.E. Pet, a non-AB toxin, is retrograde transported and translocated into epithelial cells. Infect. Immun. 2007, 75, 2101. [Google Scholar]
- Heimer, S.R.; Rasko, D.A.; Lockatell, V.C.; Johnson, D.E.; Mobley, H.L. Autotransporter genes pic and tsh are associated with Escherichia coli strains that cause acute pyelonephritis and are expressed during urinary tract infection. Infect. Immun. 2004, 72, 593–597. [Google Scholar]
- Harrington, S.M.; Sheikh, J.; Henderson, I.R.; Ruiz-Perez, F.; Cohen, P.S.; Nataro, J.P. The Pic protease of enteroaggragative Escherichia coli promotes intestinal colonization and growth in the presence of mucin. Infect. Immun. 2009, 77, 2465–2473. [Google Scholar]
- Guyer, D.M.; Radulovic, S.; Jones, F.E.; Mobley, H.L. Sat, the secreted autotransporter toxin of uropathogenic Escherichia coli, is a vacuolating cytotoxin for bladder and kidney epithelial cells. Infect. Immun. 2002, 70, 4539–4546. [Google Scholar]
- Maroncle, N.M.; Sivick, K.E.; Brady, R.; Stokes, F.E.; Mobley, H.L. Protease activity, secretion, cell entry, cytotoxicity, and cellular targets of secreted autotransporter toxin of uropathogenic Escherichia coli. Infect. Immun. 2006, 74, 6124–6134. [Google Scholar]
- Guignot, J.; Chaplais, C.; Coconnier-Polter, M.H.; Servin, A.L. The secreted autotransporter toxin, Sat, functions as a virulence factor in Afa/Dr diffusely adhering Escherichia coli by promoting lesions in tight junction of polarized cells. Cell Microb. 2007, 9, 204–221. [Google Scholar] [CrossRef]
- Kostakioti, M.; Stathopoulos, C. Functional analysis of the Tsh autotransporter from an avian pathogenic Escherichia coli strain. Infect. Immun. 2004, 72, 5548–5554. [Google Scholar]
© 2010 by the authors; licensee MDPI, Basel, Switzerland This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Dautin, N. Serine Protease Autotransporters of Enterobacteriaceae (SPATEs): Biogenesis and Function. Toxins 2010, 2, 1179-1206. https://doi.org/10.3390/toxins2061179
Dautin N. Serine Protease Autotransporters of Enterobacteriaceae (SPATEs): Biogenesis and Function. Toxins. 2010; 2(6):1179-1206. https://doi.org/10.3390/toxins2061179
Chicago/Turabian StyleDautin, Nathalie. 2010. "Serine Protease Autotransporters of Enterobacteriaceae (SPATEs): Biogenesis and Function" Toxins 2, no. 6: 1179-1206. https://doi.org/10.3390/toxins2061179




