Arenavirus Budding: A Common Pathway with Mechanistic Differences
Abstract
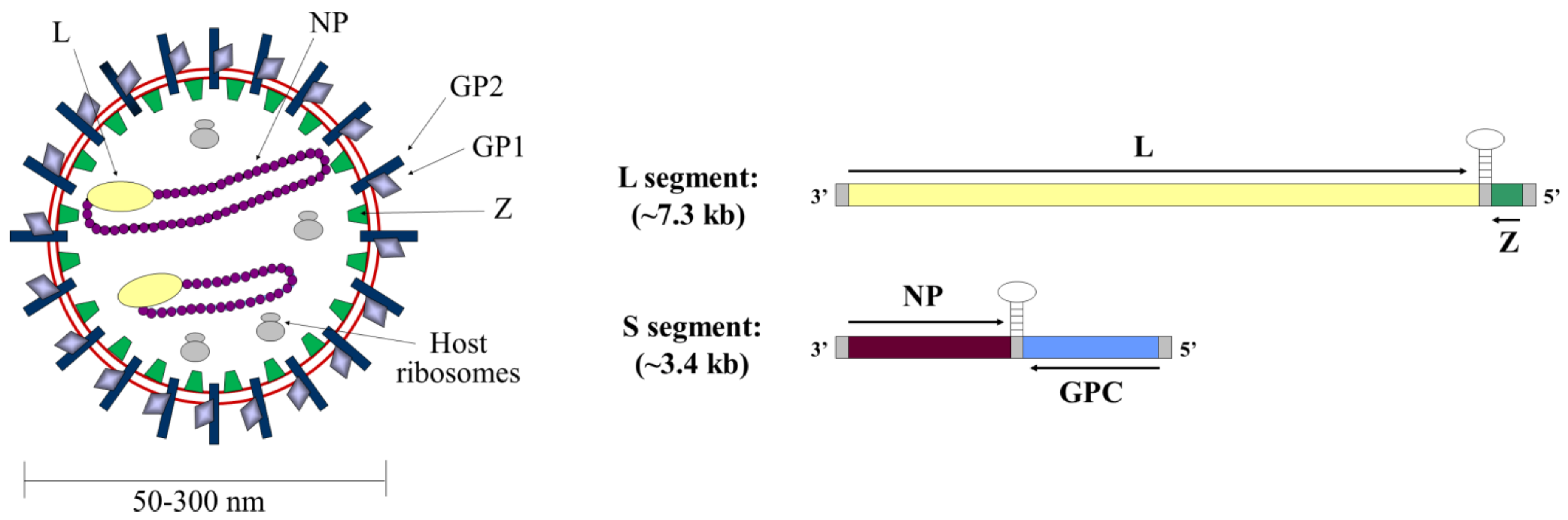
:1. Introduction
| Virus | Distribution | Reservoir | Human Disease | |
|---|---|---|---|---|
| Old World Arenviruses | Dandenong virus* | Yugoslavia (?) Australia (?) | Unknown | Febrile illness with encephalopathy (transplant-related) |
| Gbagroube virus* | Côte d'Ivoire | Mus (Nannomys) setulosus | None known | |
| Ippy virus | Central African Republic | Arvicanthus spp. | None known | |
| Lassa virus | Western Africa | Mastomys natalensis | Febrile illness, hemorrhagic fever in severe cases | |
| Lymphocytic Choriomeningitis virus | Worldwide | Mus musculus | Febrile illness, aseptic meningitis in severe cases | |
| Lujo virus | Zambia | Unknown | Hemorrhagic fever | |
| Luna virus* | Zambia | Mastomys natalensis | None known | |
| Kodoko virus* | Guinea | Mus (Nannomys) minutoides | None known | |
| Menekre virus* | Côte d'Ivoire | Hylomyscus spp. | None known | |
| Merino Walk virus* | South Africa | Myotomis unisulcatus | None known | |
| Mobala virus | Central African Republic | Praomys jacksoni | None known | |
| Mopeia virus | Mozambique | Mastomys natalensis | None known | |
| Morogoro virus* | Tanzania | Mastomys spp. | None known | |
| New World Arenaviruses | Allpahuayo virus | Peru | Oecomys spp. | None known |
| Amapari virus | Brazil | Oryzomys gaeldi Neacomys guianae | None known | |
| Bear Canyon virus | USA | Peromyscus californicus | None known | |
| Big Brushy Tank virus* | USA | Neotoma albigula | None known | |
| Catarina virus* | USA | Neotoma micropus | None known | |
| Chapare virus | Bolivia | Unknown | Hemorrhagic fever | |
| Cupixi virus | Brazil | Oryzomys spp. | None known | |
| Flexal virus | Brazil | Oryzomys spp. | Febrile illness(Lab-acquired) | |
| Guanarito virus | Venezuela | Zygodontomys brevicauda | Hemorrhagic fever | |
| Junín virus | Argentina | Calomys musculinus | Hemorrhagic fever | |
| Latino virus | Bolivia | Calomys callosus | None known | |
| Machupo virus | Bolivia | Calomys callosus | Hemorrhagic fever | |
| Oliveros virus | Argentina | Bolomys spp. | None known | |
| Paraná virus | Paraguay | Oryzomys buccinatus | None known | |
| Pichinde virus | Columbia | Oryzomys albigularis | None known | |
| Pinhal virus | Brazil | Calomys tener | None known | |
| Pirital virus | Venezuela | Sigmodon alstoni | None known | |
| Real de Catorce virus* | Mexico | Neotoma leucodon | None known | |
| Sabiá virus | Brazil | Unknown | Hemorrhagic fever | |
| Skinner Tank virus* | USA | Neotoma mexicana | None known | |
| Tacaribe virus | Trinidad | Artibeus spp. (bat) | Possible febrile illness (Lab-acquired) | |
| Tamiami virus | USA | Sigmodon hispidus | None known | |
| Tonto Creek virus | USA | Neotoma albigula | None known | |
| Whitewater Arroyo virus | USA | Neotoma albigula | Possible hemorrhagic fever |
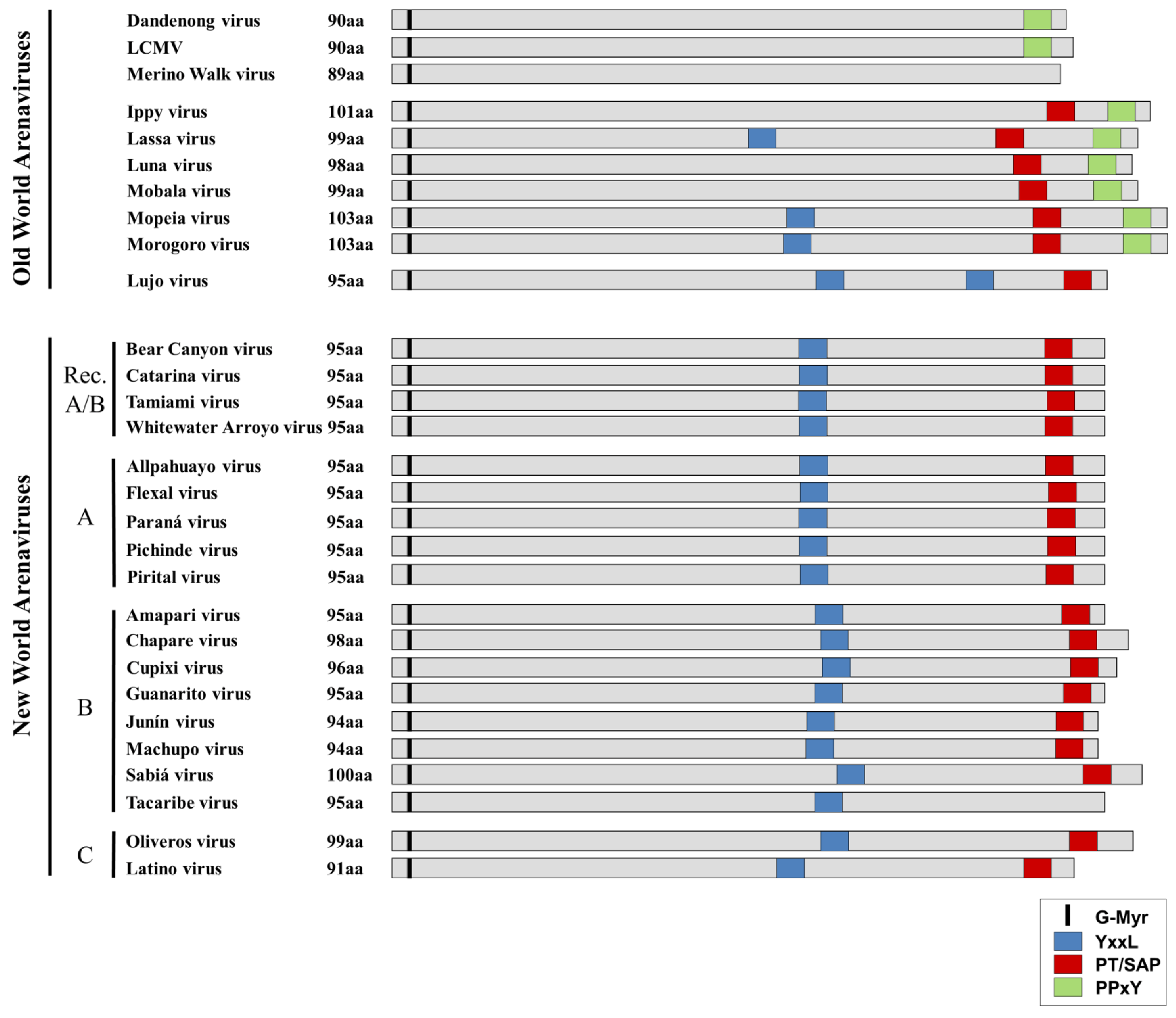
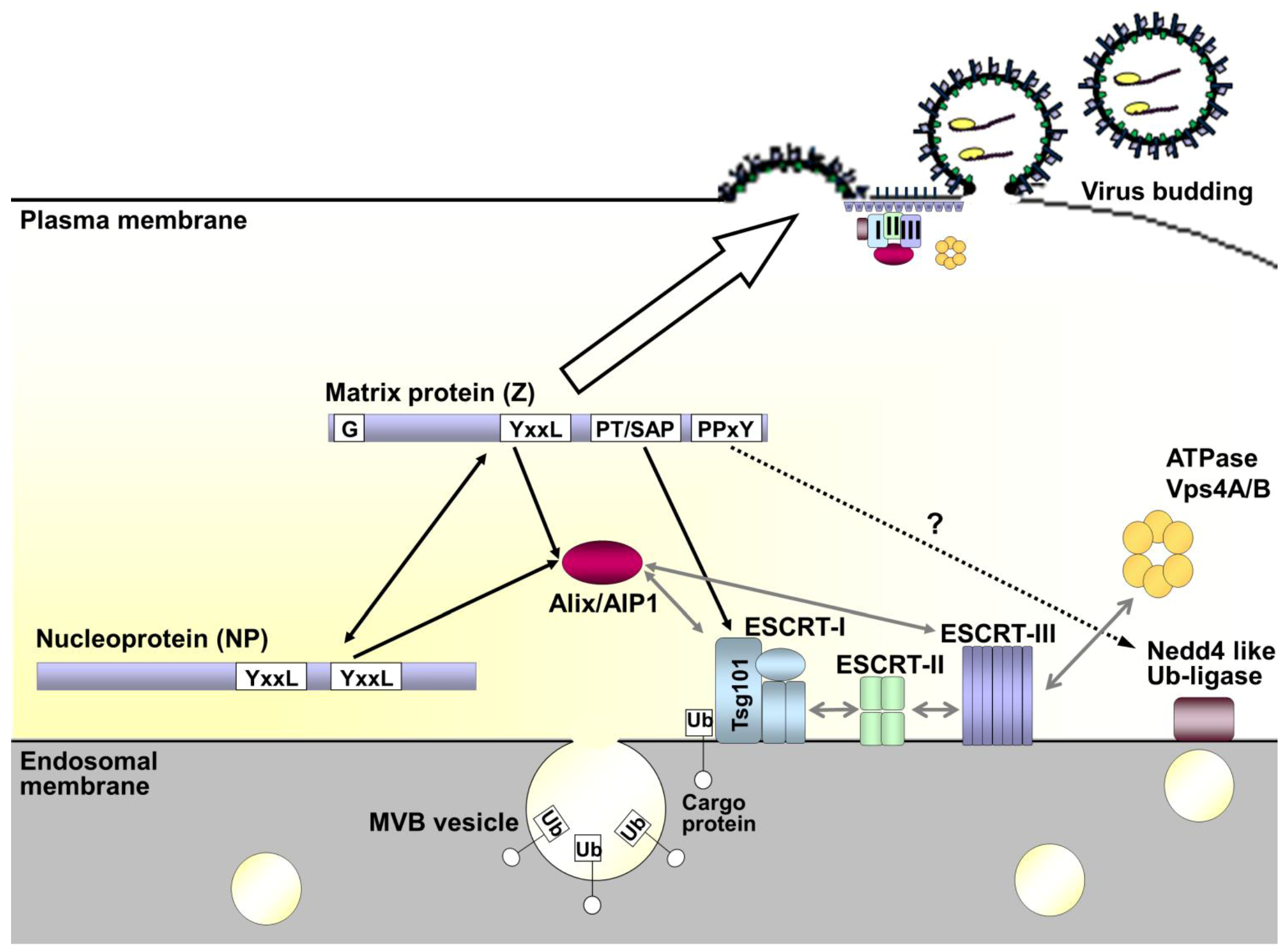
3. Virus Budding and Host Cell Sorting
| Motif | Virus families using this motif (interacting proteins) | References |
|---|---|---|
| YxxL | Arenaviridae (NP, Z) Retroviridae (Gag) Paramyxoviridae (C, M) | [60,61,62,63,64] |
| PPxY | Arenaviridae (Z) Filoviridae (VP40) Retroviridae (Gag) Rhabdoviridae (M) | [52,65,66,67] |
| PT/SAP | Arenaviridae (Z) Filoviridae (NP, VP40) Retroviridae (Gag) Rhabdoviridae (M)* | [52,66,68,69,70] |
| θ PxV | Paramyxoviridae (M) | [59] |

6. The Budding Pathway as a Potential Target for Therapeutics
7. Conclusions
Acknowledgments
Conflict of Interest
References and Notes
- Salvato, M.S.; Clegg, J.C.S.; Buchmeier, M.J.; Charrel, R.N.; Gonzalez, J.P.; Lukashevich, I.S.; Peters, C.J.; Romanowski, V. Family arenaviridae. In: Virus taxonomy: Classification and nomenclature of viruses: Ninth report of the international committee on taxonomy of viruses. Academic Press, Elsevier 2012, 715–723. [Google Scholar]
- Charrel, R.N.; de Lamballerie, X.; Emonet, S. Phylogeny of the genus arenavirus. Curr. Opin. Microbiol. 2008, 11, 362–368. [Google Scholar] [CrossRef]
- Ishii, A.; Thomas, Y.; Moonga, L.; Nakamura, I.; Ohnuma, A.; Hang'ombe, B.; Takada, A.; Mweene, A.; Sawa, H. Novel arenavirus, zambia. Emerg Infect Dis 2011, 17, 1921–1924. [Google Scholar] [CrossRef]
- Inizan, C.C.; Cajimat, M.N.; Milazzo, M.L.; Barragan-Gomez, A.; Bradley, R.D.; Fulhorst, C.F. Genetic evidence for a tacaribe serocomplex virus, mexico. Emerg. Infect. Dis. 2010, 16, 1007–1010. [Google Scholar]
- Gunther, S.; Hoofd, G.; Charrel, R.; Roser, C.; Becker-Ziaja, B.; Lloyd, G.; Sabuni, C.; Verhagen, R.; van der Groen, G.; Kennis, J.; et al. Mopeia virus-related arenavirus in natal multimammate mice, morogoro, tanzania. Emerg. Infect. Dis. 2009, 15, 2008–2012. [Google Scholar]
- Palacios, G.; Savji, N.; Hui, J.; Travassos da Rosa, A.; Popov, V.; Briese, T.; Tesh, R.; Lipkin, W.I. Genomic and phylogenetic characterization of merino walk virus, a novel arenavirus isolated in south africa. J. Gen. Virol. 2010, 91, 1315–1324. [Google Scholar] [CrossRef]
- Briese, T.; Paweska, J.T.; McMullan, L.K.; Hutchison, S.K.; Street, C.; Palacios, G.; Khristova, M.L.; Weyer, J.; Swanepoel, R.; Egholm, M.; et al. Genetic detection and characterization of lujo virus, a new hemorrhagic fever-associated arenavirus from southern africa. PLoS Pathog. 2009, 5, e1000455. [Google Scholar]
- Delgado, S.; Erickson, B.R.; Agudo, R.; Blair, P.J.; Vallejo, E.; Albarino, C.G.; Vargas, J.; Comer, J.A.; Rollin, P.E.; Ksiazek, T.G.; et al. Chapare virus, a newly discovered arenavirus isolated from a fatal hemorrhagic fever case in bolivia. PLoS Pathog. 2008, 4, e1000047. [Google Scholar] [CrossRef]
- Lecompte, E.; ter Meulen, J.; Emonet, S.; Daffis, S.; Charrel, R.N. Genetic identification of kodoko virus, a novel arenavirus of the african pigmy mouse (mus nannomys minutoides) in west africa. Virology 2007, 364, 178–183. [Google Scholar] [CrossRef]
- Coulibaly-N'Golo, D.; Allali, B.; Kouassi, S.K.; Fichet-Calvet, E.; Becker-Ziaja, B.; Rieger, T.; Olschlager, S.; Dosso, H.; Denys, C.; Ter Meulen, J.; et al. Novel arenavirus sequences in hylomyscus sp. And mus (nannomys) setulosus from cote d'ivoire: Implications for evolution of arenaviruses in africa. PLoS One 2011, 6, e20893. [Google Scholar]
- Cajimat, M.N.; Milazzo, M.L.; Borchert, J.N.; Abbott, K.D.; Bradley, R.D.; Fulhorst, C.F. Diversity among tacaribe serocomplex viruses (family arenaviridae) naturally associated with the mexican woodrat (neotoma mexicana). Virus Res. 2008, 133, 211–217. [Google Scholar]
- Milazzo, M.L.; Cajimat, M.N.; Haynie, M.L.; Abbott, K.D.; Bradley, R.D.; Fulhorst, C.F. Diversity among tacaribe serocomplex viruses (family arenaviridae) naturally associated with the white-throated woodrat (neotoma albigula) in the southwestern united states. Vector Borne Zoonotic Dis. 2008, 8, 523–540. [Google Scholar] [CrossRef]
- Cajimat, M.N.; Milazzo, M.L.; Bradley, R.D.; Fulhorst, C.F. Catarina virus, an arenaviral species principally associated with neotoma micropus (southern plains woodrat) in texas. Am. J. Trop. Med. Hyg. 2007, 77, 732–736. [Google Scholar]
- Palacios, G.; Druce, J.; Du, L.; Tran, T.; Birch, C.; Briese, T.; Conlan, S.; Quan, P.L.; Hui, J.; Marshall, J.; et al. A new arenavirus in a cluster of fatal transplant-associated diseases. N. Engl. J. Med. 2008, 358, 991–998. [Google Scholar] [CrossRef]
- Gunther, S.; Lenz, O. Lassa virus. Crit. Rev. Clin. Lab. Sci. 2004, 41, 339–390. [Google Scholar] [CrossRef]
- McCormick, J.B.; Webb, P.A.; Krebs, J.W.; Johnson, K.M.; Smith, E.S. A prospective study of the epidemiology and ecology of lassa fever. J. Infect. Dis. 1987, 155, 437–444. [Google Scholar]
- Fischer, S.A.; Graham, M.B.; Kuehnert, M.J.; Kotton, C.N.; Srinivasan, A.; Marty, F.M.; Comer, J.A.; Guarner, J.; Paddock, C.D.; DeMeo, D.L.; et al. Transmission of lymphocytic choriomeningitis virus by organ transplantation. N. Engl. J. Med. 2006, 354, 2235–2249. [Google Scholar]
- Charrel, R.N.; de Lamballerie, X. Arenaviruses other than lassa virus. Antiviral Res. 2003, 57, 89–100. [Google Scholar] [CrossRef]
- Charrel, R.N.; de Lamballerie, X.; Fulhorst, C.F. The whitewater arroyo virus: Natural evidence for genetic recombination among tacaribe serocomplex viruses (family arenaviridae). Virology 2001, 283, 161–166. [Google Scholar]
- Radoshitzky, S.R.; Abraham, J.; Spiropoulou, C.F.; Kuhn, J.H.; Nguyen, D.; Li, W.; Nagel, J.; Schmidt, P.J.; Nunberg, J.H.; Andrews, N.C.; et al. Transferrin receptor 1 is a cellular receptor for new world haemorrhagic fever arenaviruses. Nature 2007, 446, 92–96. [Google Scholar]
- Helguera, G.; Jemielity, S.; Abraham, J.; Cordo, S.M.; Martinez, M.G.; Rodriguez, J.A.; Bregni, C.; Wang, J.J.; Farzan, M.; Penichet, M.L.; et al. An antibody recognizing the apical domain of human transferrin receptor 1 efficiently inhibits the entry of all new world hemorrhagic fever arenaviruses. J. Virol. 2012, 86, 4024–4028. [Google Scholar]
- Abraham, J.; Kwong, J.A.; Albarino, C.G.; Lu, J.G.; Radoshitzky, S.R.; Salazar-Bravo, J.; Farzan, M.; Spiropoulou, C.F.; Choe, H. Host-species transferrin receptor 1 orthologs are cellular receptors for nonpathogenic new world clade b arenaviruses. PLoS Pathog. 2009, 5, e1000358. [Google Scholar]
- Flanagan, M.L.; Oldenburg, J.; Reignier, T.; Holt, N.; Hamilton, G.A.; Martin, V.K.; Cannon, P.M. New world clade b arenaviruses can use transferrin receptor 1 (tfr1)-dependent and -independent entry pathways, and glycoproteins from human pathogenic strains are associated with the use of tfr1. J. Virol. 2008, 82, 938–948. [Google Scholar]
- Spiropoulou, C.F.; Kunz, S.; Rollin, P.E.; Campbell, K.P.; Oldstone, M.B. New world arenavirus clade c, but not clade a and b viruses, utilizes alpha-dystroglycan as its major receptor. J. Virol. 2002, 76, 5140–5146. [Google Scholar]
- Cao, W.; Henry, M.D.; Borrow, P.; Yamada, H.; Elder, J.H.; Ravkov, E.V.; Nichol, S.T.; Compans, R.W.; Campbell, K.P.; Oldstone, M.B. Identification of alpha-dystroglycan as a receptor for lymphocytic choriomeningitis virus and lassa fever virus. Science 1998, 282, 2079–2081. [Google Scholar]
- Albarino, C.G.; Bergeron, E.; Erickson, B.R.; Khristova, M.L.; Rollin, P.E.; Nichol, S.T. Efficient reverse genetics generation of infectious junin viruses differing in glycoprotein processing. J. Virol. 2009, 83, 5606–5614. [Google Scholar]
- Hass, M.; Golnitz, U.; Muller, S.; Becker-Ziaja, B.; Gunther, S. Replicon system for lassa virus. J. Virol. 2004, 78, 13793–13803. [Google Scholar] [CrossRef]
- Lopez, N.; Jacamo, R.; Franze-Fernandez, M.T. Transcription and rna replication of tacaribe virus genome and antigenome analogs require n and l proteins: Z protein is an inhibitor of these processes. J. Virol. 2001, 75, 12241–12251. [Google Scholar]
- Young, P.R.; Howard, C.R. Fine structure analysis of pichinde virus nucleocapsids. J. Gen. Virol. 1983, 64 (Pt 4), 833–842. [Google Scholar]
- Lee, K.J.; Novella, I.S.; Teng, M.N.; Oldstone, M.B.; de La Torre, J.C. Np and l proteins of lymphocytic choriomeningitis virus (lcmv) are sufficient for efficient transcription and replication of lcmv genomic rna analogs. J. Virol. 2000, 74, 3470–3477. [Google Scholar] [CrossRef]
- Qi, X.; Lan, S.; Wang, W.; Schelde, L.M.; Dong, H.; Wallat, G.D.; Ly, H.; Liang, Y.; Dong, C. Cap binding and immune evasion revealed by lassa nucleoprotein structure. Nature 2010, 468, 779–783. [Google Scholar] [CrossRef]
- Martinez-Sobrido, L.; Giannakas, P.; Cubitt, B.; Garcia-Sastre, A.; de la Torre, J.C. Differential inhibition of type i interferon induction by arenavirus nucleoproteins. J. Virol. 2007, 81, 12696–12703. [Google Scholar]
- Agnihothram, S.S.; York, J.; Nunberg, J.H. Role of the stable signal peptide and cytoplasmic domain of g2 in regulating intracellular transport of the junin virus envelope glycoprotein complex. J. Virol. 2006, 80, 5189–5198. [Google Scholar] [CrossRef]
- Eichler, R.; Lenz, O.; Strecker, T.; Eickmann, M.; Klenk, H.D.; Garten, W. Identification of lassa virus glycoprotein signal peptide as a trans-acting maturation factor. EMBO Rep. 2003, 4, 1084–1088. [Google Scholar]
- Eichler, R.; Lenz, O.; Strecker, T.; Garten, W. Signal peptide of lassa virus glycoprotein gp-c exhibits an unusual length. FEBS Lett. 2003, 538, 203–206. [Google Scholar]
- Beyer, W.R.; Popplau, D.; Garten, W.; von Laer, D.; Lenz, O. Endoproteolytic processing of the lymphocytic choriomeningitis virus glycoprotein by the subtilase ski-1/s1p. J. Virol. 2003, 77, 2866–2872. [Google Scholar]
- Kunz, S.; Edelmann, K.H.; de la Torre, J.C.; Gorney, R.; Oldstone, M.B. Mechanisms for lymphocytic choriomeningitis virus glycoprotein cleavage, transport, and incorporation into virions. Virology 2003, 314, 168–178. [Google Scholar]
- Lenz, O.; ter Meulen, J.; Klenk, H.D.; Seidah, N.G.; Garten, W. The lassa virus glycoprotein precursor gp-c is proteolytically processed by subtilase ski-1/s1p. Proc. Natl. Acad. Sci. U.S.A 2001, 98, 12701–12705. [Google Scholar]
- York, J.; Romanowski, V.; Lu, M.; Nunberg, J.H. The signal peptide of the junin arenavirus envelope glycoprotein is myristoylated and forms an essential subunit of the mature g1-g2 complex. J. Virol. 2004, 78, 10783–10792. [Google Scholar]
- Jacamo, R.; Lopez, N.; Wilda, M.; Franze-Fernandez, M.T. Tacaribe virus z protein interacts with the l polymerase protein to inhibit viral rna synthesis. J. Virol. 2003, 77, 10383–10393. [Google Scholar]
- Wilda, M.; Lopez, N.; Casabona, J.C.; Franze-Fernandez, M.T. Mapping of the tacaribe arenavirus z-protein binding sites on the l protein identified both amino acids within the putative polymerase domain and a region at the n terminus of l that are critically involved in binding. J. Virol. 2008, 82, 11454–11460. [Google Scholar]
- Loureiro, M.E.; Wilda, M.; Levingston Macleod, J.M.; D'Antuono, A.; Foscaldi, S.; Marino Buslje, C.; Lopez, N. Molecular determinants of arenavirus z protein homo-oligomerization and l polymerase binding. J. Virol. 2011, 85, 12304–12314. [Google Scholar]
- Cornu, T.I.; de la Torre, J.C. Ring finger z protein of lymphocytic choriomeningitis virus (lcmv) inhibits transcription and rna replication of an lcmv s-segment minigenome. J. Virol. 2001, 75, 9415–9426. [Google Scholar]
- Borden, K.L.; Campbell Dwyer, E.J.; Salvato, M.S. An arenavirus ring (zinc-binding) protein binds the oncoprotein promyelocyte leukemia protein (pml) and relocates pml nuclear bodies to the cytoplasm. J. Virol. 1998, 72, 758–766. [Google Scholar]
- Kentsis, A.; Dwyer, E.C.; Perez, J.M.; Sharma, M.; Chen, A.; Pan, Z.Q.; Borden, K.L. The ring domains of the promyelocytic leukemia protein pml and the arenaviral protein z repress translation by directly inhibiting translation initiation factor eif4e. J. Mol. Biol. 2001, 312, 609–623. [Google Scholar]
- Borden, K.L.; Campbelldwyer, E.J.; Carlile, G.W.; Djavani, M.; Salvato, M.S. Two ring finger proteins, the oncoprotein pml and the arenavirus z protein, colocalize with the nuclear fraction of the ribosomal p proteins. J. Virol. 1998, 72, 3819–3826. [Google Scholar]
- Campbell Dwyer, E.J.; Lai, H.; MacDonald, R.C.; Salvato, M.S.; Borden, K.L. The lymphocytic choriomeningitis virus ring protein z associates with eukaryotic initiation factor 4e and selectively represses translation in a ring-dependent manner. J. Virol. 2000, 74, 3293–3300. [Google Scholar]
- Fan, L.; Briese, T.; Lipkin, W.I. Z proteins of new world arenaviruses bind rig-i and interfere with type i interferon induction. J. Virol. 2010, 84, 1785–1791. [Google Scholar]
- Casabona, J.C.; Levingston Macleod, J.M.; Loureiro, M.E.; Gomez, G.A.; Lopez, N. The ring domain and the l79 residue of z protein are involved in both the rescue of nucleocapsids and the incorporation of glycoproteins into infectious chimeric arenavirus-like particles. J. Virol. 2009, 83, 7029–7039. [Google Scholar]
- Eichler, R.; Strecker, T.; Kolesnikova, L.; ter Meulen, J.; Weissenhorn, W.; Becker, S.; Klenk, H.D.; Garten, W.; Lenz, O. Characterization of the lassa virus matrix protein z: Electron microscopic study of virus-like particles and interaction with the nucleoprotein (np). Virus Res. 2004, 100, 249–255. [Google Scholar]
- Perez, M.; Craven, R.C.; de la Torre, J.C. The small ring finger protein z drives arenavirus budding: Implications for antiviral strategies. Proc. Natl. Acad. Sci. U.S.A. 2003, 100, 12978–12983. [Google Scholar]
- Strecker, T.; Eichler, R.; Meulen, J.; Weissenhorn, W.; Dieter Klenk, H.; Garten, W.; Lenz, O. Lassa virus z protein is a matrix protein and sufficient for the release of virus-like particles. J. Virol. 2003, 77, 10700–10705. [Google Scholar]
- Urata, S.; Yasuda, J.; de la Torre, J.C. The z protein of the new world arenavirus tacaribe virus has bona fide budding activity that does not depend on known late domain motifs. J. Virol. 2009, 83, 12651–12655. [Google Scholar]
- Hoenen, T.; Kolesnikova, L.; Becker, S. Recent advances in filovirus- and arenavirus-like particles. Future Virology 2007, 2, 193–203. [Google Scholar]
- Chen, B.J.; Lamb, R.A. Mechanisms for enveloped virus budding: Can some viruses do without an escrt? Virology 2008, 372, 221–232. [Google Scholar] [CrossRef]
- Bieniasz, P.D. Late budding domains and host proteins in enveloped virus release. Virology 2006, 344, 55–63. [Google Scholar] [CrossRef]
- Hurley, J.H.; Hanson, P.I. Membrane budding and scission by the escrt machinery: It's all in the neck. Nat. Rev. Mol. Cell. Biol. 2010, 11, 556–566. [Google Scholar]
- Freed, E.O. Viral late domains. J. Virol. 2002, 76, 4679–4687. [Google Scholar]
- Schmitt, A.P.; Leser, G.P.; Morita, E.; Sundquist, W.I.; Lamb, R.A. Evidence for a new viral late-domain core sequence, fpiv, necessary for budding of a paramyxovirus. J. Virol. 2005, 79, 2988–2997. [Google Scholar]
- Shtanko, O.; Watanabe, S.; Jasenosky, L.D.; Watanabe, T.; Kawaoka, Y. Alix/aip1 is required for np incorporation into mopeia virus z-induced virus-like particles. J. Virol. 2011, 85, 3631–3641. [Google Scholar]
- Strack, B.; Calistri, A.; Craig, S.; Popova, E.; Gottlinger, H.G. Aip1/alix is a binding partner for hiv-1 p6 and eiav p9 functioning in virus budding. Cell 2003, 114, 689–699. [Google Scholar] [CrossRef]
- Dilley, K.A.; Gregory, D.; Johnson, M.C.; Vogt, V.M. An lypsl late domain in the gag protein contributes to the efficient release and replication of rous sarcoma virus. J. Virol. 2010, 84, 6276–6287. [Google Scholar]
- Sakaguchi, T.; Kato, A.; Sugahara, F.; Shimazu, Y.; Inoue, M.; Kiyotani, K.; Nagai, Y.; Yoshida, T. Aip1/alix is a binding partner of sendai virus c protein and facilitates virus budding. J. Virol. 2005, 79, 8933–8941. [Google Scholar]
- Irie, T.; Shimazu, Y.; Yoshida, T.; Sakaguchi, T. The yldl sequence within sendai virus m protein is critical for budding of virus-like particles and interacts with alix/aip1 independently of c protein. J. Virol. 2007, 81, 2263–2273. [Google Scholar]
- Harty, R.N.; Paragas, J.; Sudol, M.; Palese, P. A proline-rich motif within the matrix protein of vesicular stomatitis virus and rabies virus interacts with ww domains of cellular proteins: Implications for viral budding. J. Virol. 1999, 73, 2921–2929. [Google Scholar]
- Timmins, J.; Schoehn, G.; Ricard-Blum, S.; Scianimanico, S.; Vernet, T.; Ruigrok, R.W.; Weissenhorn, W. Ebola virus matrix protein vp40 interaction with human cellular factors tsg101 and nedd4. J. Mol. Biol. 2003, 326, 493–502. [Google Scholar]
- Kikonyogo, A.; Bouamr, F.; Vana, M.L.; Xiang, Y.; Aiyar, A.; Carter, C.; Leis, J. Proteins related to the nedd4 family of ubiquitin protein ligases interact with the l domain of rous sarcoma virus and are required for gag budding from cells. Proc. Natl. Acad. Sci. U.S.A. 2001, 98, 11199–11204. [Google Scholar]
- Garrus, J.E.; von Schwedler, U.K.; Pornillos, O.W.; Morham, S.G.; Zavitz, K.H.; Wang, H.E.; Wettstein, D.A.; Stray, K.M.; Cote, M.; Rich, R.L.; et al. Tsg101 and the vacuolar protein sorting pathway are essential for hiv-1 budding. Cell 2001, 107, 55–65. [Google Scholar]
- Dolnik, O.; Kolesnikova, L.; Stevermann, L.; Becker, S. Tsg101 is recruited by a late domain of the nucleocapsid protein to support budding of marburg virus-like particles. J. Virol. 2010, 84, 7847–7856. [Google Scholar]
- Irie, T.; Licata, J.M.; McGettigan, J.P.; Schnell, M.J.; Harty, R.N. Budding of ppxy-containing rhabdoviruses is not dependent on host proteins tgs101 and vps4a. J. Virol. 2004, 78, 2657–2665. [Google Scholar]
- Urata, S.; Noda, T.; Kawaoka, Y.; Yokosawa, H.; Yasuda, J. Cellular factors required for lassa virus budding. J. Virol. 2006, 80, 4191–4195. [Google Scholar]
- Pornillos, O.; Alam, S.L.; Rich, R.L.; Myszka, D.G.; Davis, D.R.; Sundquist, W.I. Structure and functional interactions of the tsg101 uev domain. EMBO J. 2002, 21, 2397–2406. [Google Scholar]
- Harty, R.N.; Brown, M.E.; Wang, G.; Huibregtse, J.; Hayes, F.P. A ppxy motif within the vp40 protein of ebola virus interacts physically and functionally with a ubiquitin ligase: Implications for filovirus budding. Proc. Natl. Acad. Sci. U.S.A. 2000, 97, 13871–13876. [Google Scholar]
- Yasuda, J.; Hunter, E. A proline-rich motif (pppy) in the gag polyprotein of mason-pfizer monkey virus plays a maturation-independent role in virion release. J. Virol. 1998, 72, 4095–4103. [Google Scholar]
- Carpp, L.N.; Galler, R.; Bonaldo, M.C. Interaction between the yellow fever virus nonstructural protein ns3 and the host protein alix contributes to the release of infectious particles. Microbes Infect. 2010, 13, 85–95. [Google Scholar]
- Fisher, R.D.; Chung, H.Y.; Zhai, Q.; Robinson, H.; Sundquist, W.I.; Hill, C.P. Structural and biochemical studies of alix/aip1 and its role in retrovirus budding. Cell 2007, 128, 841–852. [Google Scholar]
- Perez, M.; Greenwald, D.L.; de la Torre, J.C. Myristoylation of the ring finger z protein is essential for arenavirus budding. J. Virol. 2004, 78, 11443–11448. [Google Scholar]
- Strecker, T.; Maisa, A.; Daffis, S.; Eichler, R.; Lenz, O.; Garten, W. The role of myristoylation in the membrane association of the lassa virus matrix protein z. Virol. J. 2006, 3, 93. [Google Scholar]
- Shtanko, O.; Imai, M.; Goto, H.; Lukashevich, I.S.; Neumann, G.; Watanabe, T.; Kawaoka, Y. A role for the c terminus of mopeia virus nucleoprotein in its incorporation into z protein-induced virus-like particles. J. Virol. 2010, 84, 5415–5422. [Google Scholar]
- Groseth, A.; Wolff, S.; Strecker, T.; Hoenen, T.; Becker, S. Efficient budding of the tacaribe virus matrix protein z requires the nucleoprotein. J. Virol. 2010, 84, 3603–3611. [Google Scholar]
- Schmitt, A.P.; Leser, G.P.; Waning, D.L.; Lamb, R.A. Requirements for budding of paramyxovirus simian virus 5 virus-like particles. J. Virol. 2002, 76, 3952–3964. [Google Scholar]
- Li, M.; Schmitt, P.T.; Li, Z.; McCrory, T.S.; He, B.; Schmitt, A.P. Mumps virus matrix, fusion, and nucleocapsid proteins cooperate for efficient production of virus-like particles. J. Virol. 2009, 83, 7261–7272. [Google Scholar] [CrossRef]
- Bello, N.F.; Dussupt, V.; Sette, P.; Rudd, V.; Nagashima, K.; Bibollet-Ruche, F.; Chen, C.; Montelaro, R.C.; Hahn, B.H.; Bouamr, F. Budding of retroviruses utilizing divergent l domains requires nucleocapsid. J. Virol. 2012, 86, 4182–4193. [Google Scholar]
- Licata, J.M.; Johnson, R.F.; Han, Z.; Harty, R.N. Contribution of ebola virus glycoprotein, nucleoprotein, and vp24 to budding of vp40 virus-like particles. J. Virol. 2004, 78, 7344–7351. [Google Scholar] [CrossRef]
- Dussupt, V.; Sette, P.; Bello, N.F.; Javid, M.P.; Nagashima, K.; Bouamr, F. Basic residues in the nucleocapsid domain of gag are critical for late events of hiv-1 budding. J. Virol. 2011, 85, 2304–2315. [Google Scholar]
- Radoshitzky, S.R.; Dong, L.; Chi, X.; Clester, J.C.; Retterer, C.; Spurgers, K.; Kuhn, J.H.; Sandwick, S.; Ruthel, G.; Kota, K.; et al. Infectious lassa virus, but not filoviruses, is restricted by bst-2/tetherin. J. Virol. 2010 84, 10569–10580.
- Sakuma, T.; Noda, T.; Urata, S.; Kawaoka, Y.; Yasuda, J. Inhibition of lassa and marburg virus production by tetherin. J. Virol. 2009, 83, 2382–2385. [Google Scholar] [CrossRef]
- Neil, S.J.; Zang, T.; Bieniasz, P.D. Tetherin inhibits retrovirus release and is antagonized by hiv-1 vpu. Nature 2008, 451, 425–430. [Google Scholar]
- Van Damme, N.; Goff, D.; Katsura, C.; Jorgenson, R.L.; Mitchell, R.; Johnson, M.C.; Stephens, E.B.; Guatelli, J. The interferon-induced protein bst-2 restricts hiv-1 release and is downregulated from the cell surface by the viral vpu protein. Cell Host Microbe. 2008, 3, 245–252. [Google Scholar] [CrossRef]
- Franke, T.F. Intracellular signaling by akt: Bound to be specific. Sci Signal 2008, 1, pe29. [Google Scholar] [CrossRef]
- Saeed, M.F.; Kolokoltsov, A.A.; Freiberg, A.N.; Holbrook, M.R.; Davey, R.A. Phosphoinositide-3 kinase-akt pathway controls cellular entry of ebola virus. PLoS Pathog. 2008, 4, e1000141. [Google Scholar] [CrossRef]
- Sun, M.; Fuentes, S.M.; Timani, K.; Sun, D.; Murphy, C.; Lin, Y.; August, A.; Teng, M.N.; He, B. Akt plays a critical role in replication of nonsegmented negative-stranded rna viruses. J. Virol. 2008, 82, 105–114. [Google Scholar] [CrossRef]
- Linero, F.N.; Scolaro, L.A. Participation of the phosphatidylinositol 3-kinase/akt pathway in junin virus replication in vitro. Virus Res. 2009, 145, 166–170. [Google Scholar] [CrossRef]
- Urata, S.; Ngo, N.; de la Torre, J.C. The pi3k/akt pathway contributes to arenavirus budding. J. Virol. 2012, 86, 4578–4585. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Wolff, S.; Ebihara, H.; Groseth, A. Arenavirus Budding: A Common Pathway with Mechanistic Differences. Viruses 2013, 5, 528-549. https://doi.org/10.3390/v5020528
Wolff S, Ebihara H, Groseth A. Arenavirus Budding: A Common Pathway with Mechanistic Differences. Viruses. 2013; 5(2):528-549. https://doi.org/10.3390/v5020528
Chicago/Turabian StyleWolff, Svenja, Hideki Ebihara, and Allison Groseth. 2013. "Arenavirus Budding: A Common Pathway with Mechanistic Differences" Viruses 5, no. 2: 528-549. https://doi.org/10.3390/v5020528