The Role of Innate Immunity in Conditioning Mosquito Susceptibility to West Nile Virus
Abstract
:1. Introduction
2. Small RNA Regulatory Pathways
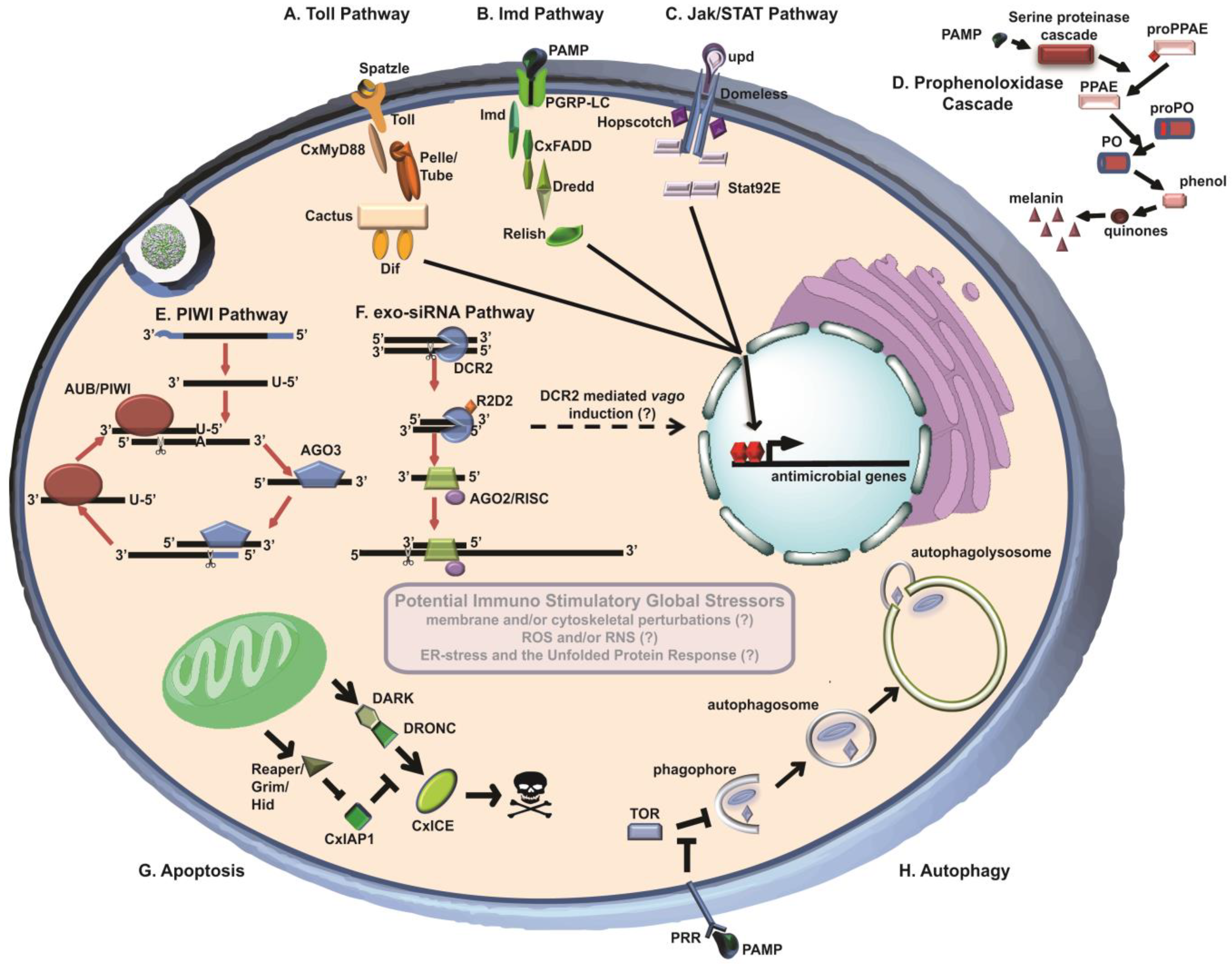
2.1. Exo-siRNA Pathway
2.2. Vago
2.3. PIWI-Interacting RNA Pathway
2.4. WNV sfRNA as a Viral Suppressor of RNAi (VSR)
3. Immune Signaling Cascades
3.1. Toll Pathway
3.2. Immune Deficiency (Imd) Pathway
3.3. Jak/STAT Pathway
3.4. Phenoloxidase (PO) Cascade
4. Cellular Processes
4.1. Autophagy
4.2. Apoptosis
5. Conclusions
Acknowledgements
Conflicts of Interest
References
- Kramer, L.D.; Ebel, G.D. Dynamics of flavivirus infection in mosquitoes. Adv. Virus Res. 2003, 60, 187–232. [Google Scholar] [CrossRef]
- Kilpatrick, A.M.; Randolph, S.E. Drivers, dynamics, and control of emerging vector-borne zoonotic diseases. Lancet 2012, 380, 1946–1955. [Google Scholar] [CrossRef]
- Kilpatrick, A.M.; Fonseca, D.M.; Ebel, G.D.; Reddy, M.R.; Kramer, L.D. Spatial and temporal variation in vector competence of Culex pipiens and Cx. restuans mosquitoes for West Nile virus. Am. J. Trop. Med. Hyg. 2010, 83, 607–613. [Google Scholar] [CrossRef]
- Kilpatrick, A.M.; Meola, M.A.; Moudy, R.M.; Kramer, L.D. Temperature, viral genetics, and the transmission of West Nile virus by Culex pipiens mosquitoes. PLoS Pathog. 2008, 4, e1000092. [Google Scholar] [CrossRef]
- Samuel, M.A.; Diamond, M.S. Alpha/beta interferon protects against lethal West Nile virus infection by restricting cellular tropism and enhancing neuronal survival. J. Virol. 2005, 79, 13350–13361. [Google Scholar] [CrossRef]
- Daffis, S.; Samuel, M.A.; Suthar, M.S.; Gale, M., Jr.; Diamond, M.S. Toll-like receptor 3 has a protective role against West Nile virus infection. J. Virol. 2008, 82, 10349–10358. [Google Scholar] [CrossRef]
- Ben-Nathan, D.; Huitinga, I.; Lustig, S.; van Rooijen, N.; Kobiler, D. West Nile virus neuroinvasion and encephalitis induced by macrophage depletion in mice. Arch. Virol. 1996, 141, 459–469. [Google Scholar] [CrossRef]
- Durrant, D.M.; Robinette, M.L.; Klein, R.S. IL-1R1 is required for dendritic cell-mediated T cell reactivation within the CNS during West Nile virus encephalitis. J. Exp. Med. 2013, 210, 503–516. [Google Scholar] [CrossRef]
- Ma, D.Y.; Suthar, M.S.; Kasahara, S.; Gale, M., Jr.; Clark, E.A. CD22 is required for protection against West Nile virus Infection. J. Virol. 2013, 87, 3361–3375. [Google Scholar] [CrossRef]
- Diamond, M.S.; Shrestha, B.; Marri, A.; Mahan, D.; Engle, M. B cells and antibody play critical roles in the immediate defense of disseminated infection by West Nile encephalitis virus. J. Virol. 2003, 77, 2578–2586. [Google Scholar] [CrossRef]
- Mehlhop, E.; Diamond, M.S. Protective immune responses against West Nile virus are primed by distinct complement activation pathways. J. Exp. Med. 2006, 203, 1371–1381. [Google Scholar] [CrossRef]
- Fuchs, A.; Pinto, A.K.; Schwaeble, W.J.; Diamond, M.S. The lectin pathway of complement activation contributes to protection from West Nile virus infection. Virology 2011, 412, 101–109. [Google Scholar] [CrossRef]
- Mehlhop, E.; Fuchs, A.; Engle, M.; Diamond, M.S. Complement modulates pathogenesis and antibody-dependent neutralization of West Nile virus infection through a C5-independent mechanism. Virology 2009, 393, 11–15. [Google Scholar] [CrossRef]
- Bosio, C.F.; Fulton, R.E.; Salasek, M.L.; Beaty, B.J.; Black, W.C.T. Quantitative trait loci that control vector competence for dengue-2 virus in the mosquito Aedes aegypti. Genetics 2000, 156, 687–698. [Google Scholar]
- Bennett, K.E.; Flick, D.; Fleming, K.H.; Jochim, R.; Beaty, B.J.; Black, W.C.T. Quantitative trait loci that control dengue-2 virus dissemination in the mosquito Aedes aegypti. Genetics 2005, 170, 185–194. [Google Scholar] [CrossRef]
- Richards, S.L.; Anderson, S.L.; Lord, C.C.; Smartt, C.T.; Tabachnick, W.J. Relationships between infection, dissemination, and transmission of West Nile virus RNA in Culex pipiens quinquefasciatus (Diptera: Culicidae). J. Med. Entomol. 2012, 49, 132–142. [Google Scholar] [CrossRef]
- Lindbo, J.A.; Silva-Rosales, L.; Proebsting, W.M.; Dougherty, W.G. Induction of a highly specific antiviral state in transgenic plants: implications for regulation of gene expression and virus resistance. Plant Cell 1993, 5, 1749–1759. [Google Scholar]
- Ratcliff, F.; Harrison, B.D.; Baulcombe, D.C. A similarity between viral defense and gene silencing in plants. Science 1997, 276, 1558–1560. [Google Scholar] [CrossRef]
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 1998, 391, 806–811. [Google Scholar] [CrossRef]
- Kennerdell, J.R.; Carthew, R.W. Use of dsRNA-mediated genetic interference to demonstrate that frizzled and frizzled 2 act in the wingless pathway. Cell 1998, 95, 1017–1026. [Google Scholar] [CrossRef]
- Liu, Q.; Rand, T.A.; Kalidas, S.; Du, F.; Kim, H.E.; Smith, D.P.; Wang, X. R2D2, a bridge between the initiation and effector steps of the Drosophila RNAi pathway. Science 2003, 301, 1921–1925. [Google Scholar] [CrossRef]
- Yang, Z.; Ebright, Y.W.; Yu, B.; Chen, X. HEN1 recognizes 21–24 nt small RNA duplexes and deposits a methyl group onto the 2' OH of the 3' terminal nucleotide. Nucleic Acids Res. 2006, 34, 667–675. [Google Scholar] [CrossRef]
- Horwich, M.D.; Li, C.; Matranga, C.; Vagin, V.; Farley, G.; Wang, P.; Zamore, P.D. The Drosophila RNA methyltransferase, DmHen1, modifies germline piRNAs and single-stranded siRNAs in RISC. Curr. Biol. 2007, 17, 1265–1272. [Google Scholar] [CrossRef]
- Rand, T.A.; Ginalski, K.; Grishin, N.V.; Wang, X. Biochemical identification of Argonaute 2 as the sole protein required for RNA-induced silencing complex activity. Proc. Natl. Acad. Sci. USA 2004, 101, 14385–14389. [Google Scholar] [CrossRef]
- Pusch, O.; Boden, D.; Silbermann, R.; Lee, F.; Tucker, L.; Ramratnam, B. Nucleotide sequence homology requirements of HIV-1-specific short hairpin RNA. Nucleic Acids Res. 2003, 31, 6444–6449. [Google Scholar] [CrossRef]
- Westerhout, E.M.; Ooms, M.; Vink, M.; Das, A.T.; Berkhout, B. HIV-1 can escape from RNA interference by evolving an alternative structure in its RNA genome. Nucleic Acids Res. 2005, 33, 796–804. [Google Scholar] [CrossRef]
- Aliyari, R.; Wu, Q.; Li, H.W.; Wang, X.H.; Li, F.; Green, L.D.; Han, C.S.; Li, W.X.; Ding, S.W. Mechanism of induction and suppression of antiviral immunity directed by virus-derived small RNAs in Drosophila. Cell Host Microbe 2008, 4, 387–397. [Google Scholar] [CrossRef]
- Galiana-Arnoux, D.; Dostert, C.; Schneemann, A.; Hoffmann, J.A.; Imler, J.L. Essential function in vivo for Dicer-2 in host defense against RNA viruses in Drosophila. Nat. Immunol. 2006, 7, 590–597. [Google Scholar]
- Van Rij, R.P.; Saleh, M.C.; Berry, B.; Foo, C.; Houk, A.; Antoniewski, C.; Andino, R. The RNA silencing endonuclease Argonaute 2 mediates specific antiviral immunity in Drosophila melanogaster. Genes Dev. 2006, 20, 2985–2995. [Google Scholar] [CrossRef]
- Keene, K.M.; Foy, B.D.; Sanchez-Vargas, I.; Beaty, B.J.; Blair, C.D.; Olson, K.E. RNA interference acts as a natural antiviral response to O'nyong-nyong virus (Alphavirus; Togaviridae) infection of Anopheles gambiae. Proc. Natl. Acad. Sci. USA 2004, 101, 17240–17245. [Google Scholar] [CrossRef]
- Myles, K.M.; Wiley, M.R.; Morazzani, E.M.; Adelman, Z.N. Alphavirus-derived small RNAs modulate pathogenesis in disease vector mosquitoes. Proc. Natl. Acad. Sci. USA 2008, 105, 19938–19943. [Google Scholar] [CrossRef]
- Sanchez-Vargas, I.; Scott, J.C.; Poole-Smith, B.K.; Franz, A.W.; Barbosa-Solomieu, V.; Wilusz, J.; Olson, K.E.; Blair, C.D. Dengue virus type 2 infections of Aedes aegypti are modulated by the mosquito’s RNA interference pathway. PLoS Pathog. 2009, 5, e1000299. [Google Scholar] [CrossRef]
- Chotkowski, H.L.; Ciota, A.T.; Jia, Y.; Puig-Basagoiti, F.; Kramer, L.D.; Shi, P.Y.; Glaser, R.L. West Nile virus infection of Drosophila melanogaster induces a protective RNAi response. Virology 2008, 377, 197–206. [Google Scholar] [CrossRef]
- Brackney, D.E.; Scott, J.C.; Sagawa, F.; Woodward, J.E.; Miller, N.A.; Schilkey, F.D.; Mudge, J.; Wilusz, J.; Olson, K.E.; Blair, C.D.; et al. C6/36 Aedes albopictus cells have a dysfunctional antiviral RNA interference response. PLoS Negl. Trop. Dis. 2010, 4, e856. [Google Scholar] [CrossRef]
- Morazzani, E.M.; Wiley, M.R.; Murreddu, M.G.; Adelman, Z.N.; Myles, K.M. Production of virus-derived ping-pong-dependent piRNA-like small RNAs in the Mosquito Soma. PLoS Pathog. 2012, 8, e1002470. [Google Scholar] [CrossRef]
- Brackney, D.E.; Beane, J.E.; Ebel, G.D. RNAi targeting of West Nile virus in mosquito midguts promotes virus diversification. PLoS Pathog. 2009, 5, e1000502. [Google Scholar] [CrossRef]
- Konishi, M.; Wu, C.H.; Kaito, M.; Hayashi, K.; Watanabe, S.; Adachi, Y.; Wu, G.Y. siRNA-Resistance in treated HCV replicon cells is correlated with the development of specific HCV mutations. J. Viral. Hepat. 2006, 13, 756–761. [Google Scholar] [CrossRef]
- Martínez, F.; Lafforgue, G.; Morelli, M.J.; González-Candelas, F.; Chua, N.-H.; Daròs, J.-A.; Elena, S.F. Ultradeep sequencing analysis of population dynamics of virus escape mutants in RNAi-mediated resistant plants. Mol. Biol. Evol. 2012, 29, 3297–3307. [Google Scholar] [CrossRef]
- Gitlin, L.; Stone, J.K.; Andino, R. Poliovirus escape from RNA interference: Short interfering RNA-target recognition and implications for therapeutic approaches. J. Virol. 2005, 79, 1027–1035. [Google Scholar] [CrossRef]
- Jerzak, G.; Bernard, K.A.; Kramer, L.D.; Ebel, G.D. Genetic variation in West Nile virus from naturally infected mosquitoes and birds suggests quasispecies structure and strong purifying selection. J. Gen. Virol. 2005, 86, 2175–2183. [Google Scholar] [CrossRef]
- Coffey, L.L.; Beeharry, Y.; Borderia, A.V.; Blanc, H.; Vignuzzi, M. Arbovirus high fidelity variant loses fitness in mosquitoes and mice. Proc. Natl. Acad. Sci. USA 2011, 108, 16038–16043. [Google Scholar] [CrossRef]
- Voinnet, O. Non-cell autonomous RNA silencing. FEBS Lett. 2005, 579, 5858–5871. [Google Scholar] [CrossRef]
- Feinberg, E.H.; Hunter, C.P. Transport of dsRNA into cells by the transmembrane protein SID-1. Science 2003, 301, 1545–1547. [Google Scholar]
- Winston, W.M.; Molodowitch, C.; Hunter, C.P. Systemic RNAi in C. elegans requires the putative transmembrane protein SID-1. Science 2002, 295, 2456–2459. [Google Scholar] [CrossRef]
- Saleh, M.C.; van Rij, R.P.; Hekele, A.; Gillis, A.; Foley, E.; O’Farrell, P.H.; Andino, R. The endocytic pathway mediates cell entry of dsRNA to induce RNAi silencing. Nat. Cell Biol. 2006, 8, 793–802. [Google Scholar] [CrossRef]
- Saleh, M.C.; Tassetto, M.; van Rij, R.P.; Goic, B.; Gausson, V.; Berry, B.; Jacquier, C.; Antoniewski, C.; Andino, R. Antiviral immunity in Drosophila requires systemic RNA interference spread. Nature 2009, 458, 346–350. [Google Scholar] [CrossRef]
- Attarzadeh-Yazdi, G.; Fragkoudis, R.; Chi, Y.; Siu, R.W.; Ulper, L.; Barry, G.; Rodriguez-Andres, J.; Nash, A.A.; Bouloy, M.; Merits, A.; et al. Cell-to-cell spread of the RNA interference response suppresses Semliki Forest virus (SFV) infection of mosquito cell cultures and cannot be antagonized by SFV. J. Virol. 2009, 83, 5735–5748. [Google Scholar] [CrossRef]
- Sijen, T.; Fleenor, J.; Simmer, F.; Thijssen, K.L.; Parrish, S.; Timmons, L.; Plasterk, R.H.; Fire, A. On the role of RNA amplification in dsRNA-triggered gene silencing. Cell 2001, 107, 465–476. [Google Scholar]
- Pak, J.; Fire, A. Distinct populations of primary and secondary effectors during RNAi in C. elegans. Science 2007, 315, 241–244. [Google Scholar] [CrossRef]
- Sijen, T.; Steiner, F.A.; Thijssen, K.L.; Plasterk, R.H. Secondary siRNAs result from unprimed RNA synthesis and form a distinct class. Science 2007, 315, 244–247. [Google Scholar] [CrossRef]
- Goic, B.; Vodovar, N.; Mondotte, J.A.; Monot, C.; Frangeul, L.; Blanc, H.; Gausson, V.; Vera-Otarola, J.; Cristofari, G.; Saleh, M.C. RNA-mediated interference and reverse transcription control the persistence of RNA viruses in the insect model Drosophila. Nat. Immunol. 2013, 14, 396–403. [Google Scholar] [CrossRef]
- Rechavi, O.; Minevich, G.; Hobert, O. Transgenerational inheritance of an acquired small RNA-based antiviral response in C. elegans. Cell 2011, 147, 1248–1256. [Google Scholar] [CrossRef]
- Deddouche, S.; Matt, N.; Budd, A.; Mueller, S.; Kemp, C.; Galiana-Arnoux, D.; Dostert, C.; Antoniewski, C.; Hoffmann, J.A.; Imler, J.L. The DExD/H-box helicase Dicer-2 mediates the induction of antiviral activity in drosophila. Nat. Immunol. 2008, 9, 1425–1432. [Google Scholar] [CrossRef]
- Paradkar, P.N.; Trinidad, L.; Voysey, R.; Duchemin, J.B.; Walker, P.J. Secreted Vago restricts West Nile virus infection in Culex mosquito cells by activating the Jak-STAT pathway. Proc. Natl. Acad. Sci. USA 2012, 109, 18915–18920. [Google Scholar] [CrossRef]
- Lin, H.; Spradling, A.C. A novel group of pumilio mutations affects the asymmetric division of germline stem cells in the Drosophila ovary. Development 1997, 124, 2463–2476. [Google Scholar]
- Aravin, A.; Gaidatzis, D.; Pfeffer, S.; Lagos-Quintana, M.; Landgraf, P.; Iovino, N.; Morris, P.; Brownstein, M.J.; Kuramochi-Miyagawa, S.; Nakano, T.; et al. A novel class of small RNAs bind to MILI protein in mouse testes. Nature 2006, 442, 203–207. [Google Scholar]
- Girard, A.; Sachidanandam, R.; Hannon, G.J.; Carmell, M.A. A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 2006, 442, 199–202. [Google Scholar]
- Grivna, S.T.; Beyret, E.; Wang, Z.; Lin, H. A novel class of small RNAs in mouse spermatogenic cells. Genes Dev. 2006, 20, 1709–1714. [Google Scholar] [CrossRef]
- Saito, K.; Nishida, K.M.; Mori, T.; Kawamura, Y.; Miyoshi, K.; Nagami, T.; Siomi, H.; Siomi, M.C. Specific association of Piwi with rasiRNAs derived from retrotransposon and heterochromatic regions in the Drosophila genome. Genes Dev. 2006, 20, 2214–2222. [Google Scholar] [CrossRef]
- Watanabe, T.; Takeda, A.; Tsukiyama, T.; Mise, K.; Okuno, T.; Sasaki, H.; Minami, N.; Imai, H. Identification and characterization of two novel classes of small RNAs in the mouse germline: retrotransposon-derived siRNAs in oocytes and germline small RNAs in testes. Genes Dev. 2006, 20, 1732–1743. [Google Scholar] [CrossRef]
- Saito, K.; Sakaguchi, Y.; Suzuki, T.; Siomi, H.; Siomi, M.C. Pimet, the Drosophila homolog of HEN1, mediates 2'-O-methylation of Piwi-interacting RNAs at their 3' ends. Genes Dev. 2007, 21, 1603–1608. [Google Scholar] [CrossRef]
- Lee, E.J.; Banerjee, S.; Zhou, H.; Jammalamadaka, A.; Arcila, M.; Manjunath, B.S.; Kosik, K.S. Identification of piRNAs in the central nervous system. RNA 2011, 17, 1090–1099. [Google Scholar] [CrossRef]
- Yan, Z.; Hu, H.Y.; Jiang, X.; Maierhofer, V.; Neb, E.; He, L.; Hu, Y.; Hu, H.; Li, N.; Chen, W.; et al. Widespread expression of piRNA-like molecules in somatic tissues. Nucleic Acids Res. 2011, 39, 6596–6607. [Google Scholar] [CrossRef]
- Nishida, K.M.; Saito, K.; Mori, T.; Kawamura, Y.; Nagami-Okada, T.; Inagaki, S.; Siomi, H.; Siomi, M.C. Gene silencing mechanisms mediated by Aubergine piRNA complexes in Drosophila male gonad. RNA 2007, 13, 1911–1922. [Google Scholar] [CrossRef]
- Brennecke, J.; Aravin, A.A.; Stark, A.; Dus, M.; Kellis, M.; Sachidanandam, R.; Hannon, G.J. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 2007, 128, 1089–1103. [Google Scholar] [CrossRef]
- Gunawardane, L.S.; Saito, K.; Nishida, K.M.; Miyoshi, K.; Kawamura, Y.; Nagami, T.; Siomi, H.; Siomi, M.C. A slicer-mediated mechanism for repeat-associated siRNA 5' end formation in Drosophila. Science 2007, 315, 1587–1590. [Google Scholar] [CrossRef]
- Schnettler, E.; Donald, C.L.; Human, S.; Watson, M.; Siu, R.W.; McFarlane, M.; Fazakerley, J.K.; Kohl, A.; Fragkoudis, R. Knockdown of piRNA pathway proteins results in enhanced Semliki Forest virus production in mosquito cells. J. Gen. Virol. 2013, 94, 1680–1689. [Google Scholar] [CrossRef]
- Scott, J.C.; Brackney, D.E.; Campbell, C.L.; Bondu-Hawkins, V.; Hjelle, B.; Ebel, G.D.; Olson, K.E.; Blair, C.D. Comparison of dengue virus type 2-specific small RNAs from RNA interference-competent and -incompetent mosquito cells. PLoS Negl. Trop. Dis. 2010, 4, e848. [Google Scholar] [CrossRef]
- Wu, Q.; Luo, Y.; Lu, R.; Lau, N.; Lai, E.C.; Li, W.X.; Ding, S.W. Virus discovery by deep sequencing and assembly of virus-derived small silencing RNAs. Proc. Natl. Acad. Sci. USA 2010, 107, 1606–1611. [Google Scholar] [CrossRef]
- Hess, A.M.; Prasad, A.N.; Ptitsyn, A.; Ebel, G.D.; Olson, K.E.; Barbacioru, C.; Monighetti, C.; Campbell, C.L. Small RNA profiling of Dengue virus-mosquito interactions implicates the PIWI RNA pathway in anti-viral defense. BMC Microbiol. 2011, 11. [Google Scholar] [CrossRef]
- Vodovar, N.; Bronkhorst, A.W.; van Cleef, K.W.; Miesen, P.; Blanc, H.; van Rij, R.P.; Saleh, M.C. Arbovirus-derived piRNAs exhibit a ping-pong signature in Mosquito cells. PLoS One 2012, 7, e30861. [Google Scholar]
- Leger, P.; Lara, E.; Jagla, B.; Sismeiro, O.; Mansuroglu, Z.; Coppee, J.Y.; Bonnefoy, E.; Bouloy, M. Dicer-2 and piwi mediated RNA interference in Rift Valley Fever Virus infected Mosquito cells. J. Virol. 2012, 87. [Google Scholar] [CrossRef]
- Schnettler, E.; Ratinier, M.; Watson, M.; Shaw, A.E.; McFarlane, M.; Varela, M.; Elliott, R.M.; Palmarini, M.; Kohl, A. RNA interference targets arbovirus replication in Culicoides cells. J. Virol. 2012, 87, 2441–2454. [Google Scholar]
- Li, H.; Li, W.X.; Ding, S.W. Induction and suppression of RNA silencing by an animal virus. Science 2002, 296, 1319–1321. [Google Scholar] [CrossRef]
- Blakqori, G.; Delhaye, S.; Habjan, M.; Blair, C.D.; Sanchez-Vargas, I.; Olson, K.E.; Attarzadeh-Yazdi, G.; Fragkoudis, R.; Kohl, A.; Kalinke, U.; et al. La Crosse bunyavirus nonstructural protein NSs serves to suppress the type I interferon system of mammalian hosts. J. Virol. 2007, 81, 4991–4999. [Google Scholar] [CrossRef]
- Soldan, S.S.; Plassmeyer, M.L.; Matukonis, M.K.; Gonzalez-Scarano, F. La Crosse virus nonstructural protein NSs counteracts the effects of short interfering RNA. J. Virol. 2005, 79, 234–244. [Google Scholar] [CrossRef]
- Kakumani, P.K.; Ponia, S.S.; S, R.K.; Sood, V.; Chinnappan, M.; Banerjea, A.C.; Medigeshi, G.R.; Malhotra, P.; Mukherjee, S.K.; Bhatnagar, R.K. Role of RNA interference (RNAi) in dengue virus replication and identification of NS4B as an RNAi suppressor. J. Virol. 2013, 87, 8870–8883. [Google Scholar] [CrossRef]
- Pijlman, G.P.; Funk, A.; Kondratieva, N.; Leung, J.; Torres, S.; van der Aa, L.; Liu, W.J.; Palmenberg, A.C.; Shi, P.Y.; Hall, R.A.; et al. A highly structured, nuclease-resistant, noncoding RNA produced by flaviviruses is required for pathogenicity. Cell Host Microbe 2008, 4, 579–591. [Google Scholar] [CrossRef]
- Funk, A.; Truong, K.; Nagasaki, T.; Torres, S.; Floden, N.; Balmori Melian, E.; Edmonds, J.; Dong, H.; Shi, P.Y.; Khromykh, A.A. RNA structures required for production of subgenomic flavivirus RNA. J. Virol. 2010, 84, 11407–11417. [Google Scholar] [CrossRef]
- Silva, P.A.; Pereira, C.F.; Dalebout, T.J.; Spaan, W.J.; Bredenbeek, P.J. An RNA pseudoknot is required for production of yellow fever virus subgenomic RNA by the host nuclease XRN1. J. Virol. 2010, 84, 11395–11406. [Google Scholar]
- Schnettler, E.; Sterken, M.G.; Leung, J.Y.; Metz, S.W.; Geertsema, C.; Goldbach, R.W.; Vlak, J.M.; Kohl, A.; Khromykh, A.A.; Pijlman, G.P. Noncoding flavivirus RNA displays RNA interference suppressor activity in insect and Mammalian cells. J. Virol. 2012, 86, 13486–13500. [Google Scholar] [CrossRef]
- Ding, S.W.; Voinnet, O. Antiviral immunity directed by small RNAs. Cell 2007, 130, 413–426. [Google Scholar] [CrossRef]
- Moon, S.L.; Anderson, J.R.; Kumagai, Y.; Wilusz, C.J.; Akira, S.; Khromykh, A.A.; Wilusz, J. A noncoding RNA produced by arthropod-borne flaviviruses inhibits the cellular exoribonuclease XRN1 and alters host mRNA stability. RNA 2012, 18, 2029–2040. [Google Scholar] [CrossRef] [Green Version]
- Christensen, B.M.; Li, J.; Chen, C.C.; Nappi, A.J. Melanization immune responses in mosquito vectors. Trends Parasitol. 2005, 21, 192–199. [Google Scholar] [CrossRef]
- Lemaitre, B.; Hoffmann, J. The host defense of Drosophila melanogaster. Annu. Rev. Immunol. 2007, 25, 697–743. [Google Scholar] [CrossRef]
- Anderson, K.V.; Bokla, L.; Nusslein-Volhard, C. Establishment of dorsal-ventral polarity in the Drosophila embryo: The induction of polarity by the Toll gene product. Cell 1985, 42, 791–798. [Google Scholar] [CrossRef]
- Lemaitre, B.; Meister, M.; Govind, S.; Georgel, P.; Steward, R.; Reichhart, J.M.; Hoffmann, J.A. Functional analysis and regulation of nuclear import of dorsal during the immune response in Drosophila. EMBO J. 1995, 14, 536–545. [Google Scholar]
- Lemaitre, B.; Reichhart, J.M.; Hoffmann, J.A. Drosophila host defense: Differential induction of antimicrobial peptide genes after infection by various classes of microorganisms. Proc. Natl. Acad. Sci. USA 1997, 94, 14614–14619. [Google Scholar] [CrossRef]
- Zambon, R.A.; Nandakumar, M.; Vakharia, V.N.; Wu, L.P. The Toll pathway is important for an antiviral response in Drosophila. Proc. Natl. Acad. Sci. USA 2005, 102, 7257–7262. [Google Scholar] [CrossRef]
- Xi, Z.; Ramirez, J.L.; Dimopoulos, G. The Aedes aegypti toll pathway controls dengue virus infection. PLoS Pathog. 2008, 4, e1000098. [Google Scholar] [CrossRef]
- Ramirez, J.L.; Dimopoulos, G. The Toll immune signaling pathway control conserved anti-dengue defenses across diverse Ae. aegypti strains and against multiple dengue virus serotypes. Dev. Comp. Immunol. 2010, 34, 625–629. [Google Scholar] [CrossRef]
- Luplertlop, N.; Surasombatpattana, P.; Patramool, S.; Dumas, E.; Wasinpiyamongkol, L.; Saune, L.; Hamel, R.; Bernard, E.; Sereno, D.; Thomas, F.; et al. Induction of a peptide with activity against a broad spectrum of pathogens in the Aedes aegypti salivary gland, following Infection with Dengue Virus. PLoS Pathog. 2011, 7, e1001252. [Google Scholar] [CrossRef]
- Sanders, H.R.; Foy, B.D.; Evans, A.M.; Ross, L.S.; Beaty, B.J.; Olson, K.E.; Gill, S.S. Sindbis virus induces transport processes and alters expression of innate immunity pathway genes in the midgut of the disease vector, Aedes aegypti. Insect Biochem. Mol. Biol. 2005, 35, 1293–1307. [Google Scholar] [CrossRef]
- Colpitts, T.M.; Cox, J.; Vanlandingham, D.L.; Feitosa, F.M.; Cheng, G.; Kurscheid, S.; Wang, P.; Krishnan, M.N.; Higgs, S.; Fikrig, E. Alterations in the Aedes aegypti transcriptome during infection with West Nile, dengue and yellow fever viruses. PLoS Pathog. 2011, 7, e1002189. [Google Scholar] [CrossRef]
- Fragkoudis, R.; Chi, Y.; Siu, R.W.; Barry, G.; Attarzadeh-Yazdi, G.; Merits, A.; Nash, A.A.; Fazakerley, J.K.; Kohl, A. Semliki Forest virus strongly reduces mosquito host defence signaling. Insect Mol. Biol. 2008, 17, 647–656. [Google Scholar] [CrossRef]
- Bartholomay, L.C.; Waterhouse, R.M.; Mayhew, G.F.; Campbell, C.L.; Michel, K.; Zou, Z.; Ramirez, J.L.; Das, S.; Alvarez, K.; Arensburger, P.; et al. Pathogenomics of Culex quinquefasciatus and meta-analysis of infection responses to diverse pathogens. Science 2010, 330, 88–90. [Google Scholar] [CrossRef]
- Corbo, J.C.; Levine, M. Characterization of an immunodeficiency mutant in Drosophila. Mech. Dev. 1996, 55, 211–220. [Google Scholar] [CrossRef]
- Levashina, E.A.; Ohresser, S.; Lemaitre, B.; Imler, J.L. Two distinct pathways can control expression of the gene encoding the Drosophila antimicrobial peptide metchnikowin. J. Mol. Biol. 1998, 278, 515–527. [Google Scholar] [CrossRef]
- Costa, A.; Jan, E.; Sarnow, P.; Schneider, D. The Imd pathway is involved in antiviral immune responses in Drosophila. PLoS One 2009, 4, e7436. [Google Scholar] [CrossRef]
- De Gregorio, E.; Spellman, P.T.; Tzou, P.; Rubin, G.M.; Lemaitre, B. The Toll and Imd pathways are the major regulators of the immune response in Drosophila. EMBO J. 2002, 21, 2568–2579. [Google Scholar] [CrossRef]
- Avadhanula, V.; Weasner, B.P.; Hardy, G.G.; Kumar, J.P.; Hardy, R.W. A novel system for the launch of alphavirus RNA synthesis reveals a role for the Imd pathway in arthropod antiviral response. PLoS Pathog. 2009, 5, e1000582. [Google Scholar] [CrossRef]
- Huang, Z.; Kingsolver, M.B.; Avadhanula, V.; Hardy, R.W. An antiviral role for antimicrobial peptides during the arthropod response to alphavirus replication. J. Virol. 2013, 87, 4272–4280. [Google Scholar] [CrossRef]
- Tsai, C.W.; McGraw, E.A.; Ammar, E.D.; Dietzgen, R.G.; Hogenhout, S.A. Drosophila melanogaster mounts a unique immune response to the Rhabdovirus sigma virus. Appl. Environ. Microbiol. 2008, 74, 3251–3256. [Google Scholar] [CrossRef]
- Dostert, C.; Jouanguy, E.; Irving, P.; Troxler, L.; Galiana-Arnoux, D.; Hetru, C.; Hoffmann, J.A.; Imler, J.L. The Jak-STAT signaling pathway is required but not sufficient for the antiviral response of drosophila. Nat. Immunol. 2005, 6, 946–953. [Google Scholar]
- Waldock, J.; Olson, K.E.; Christophides, G.K. Anopheles gambiae antiviral immune response to systemic O’nyong-nyong infection. PLoS Negl. Trop. Dis. 2012, 6, e1565. [Google Scholar] [CrossRef]
- Yan, R.; Small, S.; Desplan, C.; Dearolf, C.R.; Darnell, J.E., Jr. Identification of a Stat gene that functions in Drosophila development. Cell 1996, 84, 421–430. [Google Scholar] [CrossRef]
- Brown, S.; Hu, N.; Hombria, J.C. Identification of the first invertebrate interleukin JAK/STAT receptor, the Drosophila gene domeless. Curr. Biol. 2001, 11, 1700–1705. [Google Scholar] [CrossRef]
- Binari, R.; Perrimon, N. Stripe-specific regulation of pair-rule genes by hopscotch, a putative Jak family tyrosine kinase in Drosophila. Genes Dev. 1994, 8, 300–312. [Google Scholar] [CrossRef]
- Kemp, C.; Mueller, S.; Goto, A.; Barbier, V.; Paro, S.; Bonnay, F.; Dostert, C.; Troxler, L.; Hetru, C.; Meignin, C.; et al. Broad RNA interference-mediated antiviral immunity and virus-specific inducible responses in Drosophila. J. Immunol. 2013, 190, 650–658. [Google Scholar] [CrossRef]
- Souza-Neto, J.A.; Sim, S.; Dimopoulos, G. An evolutionary conserved function of the JAK-STAT pathway in anti-dengue defense. Proc. Natl. Acad. Sci. USA 2009, 106, 17841–17846. [Google Scholar] [CrossRef]
- Popham, H.J.; Shelby, K.S.; Brandt, S.L.; Coudron, T.A. Potent virucidal activity in larval Heliothis virescens plasma against Helicoverpa zea single capsid nucleopolyhedrovirus. J. Gen. Virol. 2004, 85, 2255–2261. [Google Scholar] [CrossRef]
- Shelby, K.S.; Popham, H.J. Plasma phenoloxidase of the larval tobacco budworm, Heliothis virescens, is virucidal. J. Insect Sci. 2006, 6, 1–12. [Google Scholar] [CrossRef]
- Zhao, P.; Lu, Z.; Strand, M.R.; Jiang, H. Antiviral, anti-parasitic, and cytotoxic effects of 5,6-dihydroxyindole (DHI), a reactive compound generated by phenoloxidase during insect immune response. Insect Biochem. Mol. Biol. 2011, 41, 645–652. [Google Scholar] [CrossRef]
- Tamang, D.; Tseng, S.M.; Huang, C.Y.; Tsao, I.Y.; Chou, S.Z.; Higgs, S.; Christensen, B.M.; Chen, C.C. The use of a double subgenomic Sindbis virus expression system to study mosquito gene function: Effects of antisense nucleotide number and duration of viral infection on gene silencing efficiency. Insect Mol. Biol. 2004, 13, 595–602. [Google Scholar] [CrossRef]
- Rodriguez-Andres, J.; Rani, S.; Varjak, M.; Chase-Topping, M.E.; Beck, M.H.; Ferguson, M.C.; Schnettler, E.; Fragkoudis, R.; Barry, G.; et al. Phenoloxidase activity acts as a mosquito innate immune response against infection with Semliki Forest virus. PLoS Pathog. 2012, 8, e1002977. [Google Scholar] [CrossRef]
- Gutierrez, M.G.; Master, S.S.; Singh, S.B.; Taylor, G.A.; Colombo, M.I.; Deretic, V. Autophagy is a defense mechanism inhibiting BCG and Mycobacterium tuberculosis survival in infected macrophages. Cell 2004, 119, 753–766. [Google Scholar] [CrossRef]
- Ling, Y.M.; Shaw, M.H.; Ayala, C.; Coppens, I.; Taylor, G.A.; Ferguson, D.J.; Yap, G.S. Vacuolar and plasma membrane stripping and autophagic elimination of Toxoplasma gondii in primed effector macrophages. J. Exp. Med. 2006, 203, 2063–2071. [Google Scholar] [CrossRef]
- Kudchodkar, S.B.; Levine, B. Viruses and autophagy. Rev. Med. Virol. 2009, 19, 359–378. [Google Scholar] [CrossRef]
- Elmore, S. Apoptosis: A review of programmed cell death. Toxicol. Pathol. 2007, 35, 495–516. [Google Scholar] [CrossRef]
- Best, S.M. Viral subversion of apoptotic enzymes: Escape from death row. Annu. Rev. Microbiol. 2008, 62, 171–192. [Google Scholar] [CrossRef]
- Levine, B.; Mizushima, N.; Virgin, H.W. Autophagy in immunity and inflammation. Nature 2011, 469, 323–335. [Google Scholar] [CrossRef]
- Tanida, I. Autophagy basics. Microbiol. Immunol. 2011, 55, 1–11. [Google Scholar] [CrossRef]
- Shelly, S.; Lukinova, N.; Bambina, S.; Berman, A.; Cherry, S. Autophagy is an essential component of Drosophila immunity against vesicular stomatitis virus. Immunity 2009, 30, 588–598. [Google Scholar] [CrossRef]
- Nakamoto, M.; Moy, R.H.; Xu, J.; Bambina, S.; Yasunaga, A.; Shelly, S.S.; Gold, B.; Cherry, S. Virus recognition by Toll-7 activates antiviral autophagy in Drosophila. Immunity 2012, 36, 658–667. [Google Scholar] [CrossRef]
- Kerr, J.F.; Wyllie, A.H.; Currie, A.R. Apoptosis: A basic biological phenomenon with wide-ranging implications in tissue kinetics. Br. J. Cancer 1972, 26, 239–257. [Google Scholar] [CrossRef]
- Fulda, S.; Debatin, K.M. Extrinsic versus intrinsic apoptosis pathways in anticancer chemotherapy. Oncogene 2006, 25, 4798–4811. [Google Scholar] [CrossRef]
- Courtiade, J.; Pauchet, Y.; Vogel, H.; Heckel, D.G. A comprehensive characterization of the caspase gene family in insects from the order Lepidoptera. BMC Genomics 2011, 12, 357. [Google Scholar] [CrossRef]
- Settles, E.W.; Friesen, P.D. Flock house virus induces apoptosis by depletion of Drosophila inhibitor-of-apoptosis protein DIAP1. J. Virol. 2008, 82, 1378–1388. [Google Scholar] [CrossRef]
- Vandergaast, R.; Schultz, K.L.; Cerio, R.J.; Friesen, P.D. Active depletion of host cell inhibitor-of-apoptosis proteins triggers apoptosis upon baculovirus DNA replication. J. Virol. 2011, 85, 8348–8358. [Google Scholar] [CrossRef]
- Hershberger, P.A.; Dickson, J.A.; Friesen, P.D. Site-specific mutagenesis of the 35-kilodalton protein gene encoded by Autographa californica nuclear polyhedrosis virus: Cell line-specific effects on virus replication. J. Virol. 1992, 66, 5525–5533. [Google Scholar]
- Clem, R.J.; Miller, L.K. Apoptosis reduces both the in vitro replication and the in vivo infectivity of a baculovirus. J. Virol. 1993, 67, 3730–3738. [Google Scholar]
- Liu, B.; Behura, S.K.; Clem, R.J.; Schneemann, A.; Becnel, J.; Severson, D.W.; Zhou, L. P53-mediated rapid induction of apoptosis conveys resistance to viral infection in Drosophila melanogaster. PLoS Pathog. 2013, 9, e1003137. [Google Scholar] [CrossRef]
- Karpf, A.R.; Brown, D.T. Comparison of Sindbis virus-induced pathology in mosquito and vertebrate cell cultures. Virology 1998, 240, 193–201. [Google Scholar]
- Li, Y.G.; Siripanyaphinyo, U.; Tumkosit, U.; Noranate, N.; A-nueqoonpipat, A.; Tao, R.; Kurosu, T.; Ikuta, K.; Takeda, N.; Anantapreecha, S. Chikungunya virus induces a more moderate cytopathic effect in mosquito cells than in mammalian cells. Intervirology 2013, 56, 6–12. [Google Scholar] [CrossRef]
- Bowers, D.F.; Coleman, C.G.; Brown, D.T. Sindbis virus-associated pathology in Aedes albopictus (Diptera: Culicidae). J. Med. Entomol. 2003, 40, 698–705. [Google Scholar] [CrossRef]
- Girard, Y.A.; Schneider, B.S.; McGee, C.E.; Wen, J.; Han, V.C.; Popov, V.; Mason, P.W.; Higgs, S. Salivary gland morphology and virus transmission during long-term cytopathologic West Nile virus infection in Culex mosquitoes. Am. J. Trop. Med. Hyg. 2007, 76, 118–128. [Google Scholar]
- Mims, C.A.; Day, M.F.; Marshall, I.D. Cytopathic effect of Semliki Forest virus in the mosquito Aedes aegypti. Am. J. Trop. Med. Hyg. 1966, 15, 775–784. [Google Scholar]
- Vaidyanathan, R.; Scott, T.W. Apoptosis in mosquito midgut epithelia associated with West Nile virus infection. Apoptosis 2006, 11, 1643–1651. [Google Scholar] [CrossRef]
- Weaver, S.C.; Lorenz, L.H.; Scott, T.W. Pathologic changes in the midgut of Culex tarsalis following infection with Western equine encephalomyelitis virus. Am. J. Trop. Med. Hyg. 1992, 47, 691–701. [Google Scholar]
- Weaver, S.C.; Scott, T.W.; Lorenz, L.H.; Lerdthusnee, K.; Romoser, W.S. Togavirus-associated pathologic changes in the midgut of a natural mosquito vector. J. Virol. 1988, 62, 2083–2090. [Google Scholar]
- Kelly, E.M.; Moon, D.C.; Bowers, D.F. Apoptosis in mosquito salivary glands: Sindbis virus-associated and tissue homeostasis. J. Gen. Virol. 2012, 93, 2419–2424. [Google Scholar] [CrossRef]
- Girard, Y.A.; Popov, V.; Wen, J.; Han, V.; Higgs, S. Ultrastructural study of West Nile virus pathogenesis in Culex pipiens quinquefasciatus (Diptera: Culicidae). J. Med. Entomol. 2005, 42, 429–444. [Google Scholar] [CrossRef]
- Wang, H.; Blair, C.D.; Olson, K.E.; Clem, R.J. Effects of inducing or inhibiting apoptosis on Sindbis virus replication in mosquito cells. J. Gen. Virol. 2008, 89, 2651–2661. [Google Scholar] [CrossRef]
- Blitvich, B.J.; Blair, C.D.; Kempf, B.J.; Hughes, M.T.; Black, W.C.; Mackie, R.S.; Meredith, C.T.; Beaty, B.J.; Rayms-Keller, A. Developmental- and tissue-specific expression of an inhibitor of apoptosis protein 1 homologue from Aedes triseriatus mosquitoes. Insect Mol. Biol. 2002, 11, 431–442. [Google Scholar]
- Ocampo, C.B.; Caicedo, P.A.; Jaramillo, G.; Ursic Bedoya, R.; Baron, O.; Serrato, I.M.; Cooper, D.M.; Lowenberger, C. Differential expression of apoptosis related genes in selected strains of Aedes aegypti with different susceptibilities to dengue virus. PLoS One 2013, 8, e61187. [Google Scholar]
- Baron, O.L.; Ursic-Bedoya, R.J.; Lowenberger, C.A.; Ocampo, C.B. Differential gene expression from midguts of refractory and susceptible lines of the mosquito, Aedes aegypti, infected with Dengue-2 virus. J. Insect Sci. 2010, 10, 1–23. [Google Scholar]
- Wang, H.; Gort, T.; Boyle, D.L.; Clem, R.J. Effects of manipulating apoptosis on Sindbis virus infection of Aedes aegypti mosquitoes. J. Virol. 2012, 86, 6546–6554. [Google Scholar] [CrossRef]
- Espada-Murao, L.A.; Morita, K. Delayed cytosolic exposure of Japanese encephalitis virus double-stranded RNA impedes interferon activation and enhances viral dissemination in porcine cells. J. Virol. 2011, 85, 6736–6749. [Google Scholar] [CrossRef]
- Hastie, K.M.; Kimberlin, C.R.; Zandonatti, M.A.; MacRae, I.J.; Saphire, E.O. Structure of the Lassa virus nucleoprotein reveals a dsRNA-specific 3' to 5' exonuclease activity essential for immune suppression. Proc. Natl. Acad. Sci. USA 2011, 108, 2396–2401. [Google Scholar] [CrossRef]
- Noyce, R.S.; Taylor, K.; Ciechonska, M.; Collins, S.E.; Duncan, R.; Mossman, K.L. Membrane perturbation elicits an IRF3-dependent, interferon-independent antiviral response. J. Virol. 2011, 85, 10926–10931. [Google Scholar] [CrossRef]
- Noyce, R.S.; Collins, S.E.; Mossman, K.L. Identification of a novel pathway essential for the immediate-early, interferon-independent antiviral response to enveloped virions. J. Virol. 2006, 80, 226–235. [Google Scholar] [CrossRef]
- Collins, S.E.; Noyce, R.S.; Mossman, K.L. Innate cellular response to virus particle entry requires IRF3 but not virus replication. J. Virol. 2004, 78, 1706–1717. [Google Scholar] [CrossRef]
- Rosette, C.; Karin, M. Cytoskeletal control of gene expression: Depolymerization of microtubules activates NF-kappa B. J. Cell Biol. 1995, 128, 1111–1119. [Google Scholar] [CrossRef]
- Kustermans, G.; el Benna, J.; Piette, J.; Legrand-Poels, S. Perturbation of actin dynamics induces NF-kappaB activation in myelomonocytic cells through an NADPH oxidase-dependent pathway. Biochem. J. 2005, 387, 531–540. [Google Scholar] [CrossRef]
- Delorme-Axford, E.; Coyne, C.B. The actin cytoskeleton as a barrier to virus infection of polarized epithelial cells. Viruses 2011, 3, 2462–2477. [Google Scholar] [CrossRef]
- Hare, D.; Mossman, K.L. Novel paradigms of innate immune sensing of viral infections. Cytokine 2013, 63, 219–224. [Google Scholar] [CrossRef]
- Gonzalez-Dosal, R.; Horan, K.A.; Rahbek, S.H.; Ichijo, H.; Chen, Z.J.; Mieyal, J.J.; Hartmann, R.; Paludan, S.R. HSV infection induces production of ROS, which potentiate signaling from pattern recognition receptors: Role for S-glutathionylation of TRAF3 and 6. PLoS Pathog. 2011, 7, e1002250. [Google Scholar] [CrossRef]
- Indukuri, H.; Castro, S.M.; Liao, S.M.; Feeney, L.A.; Dorsch, M.; Coyle, A.J.; Garofalo, R.P.; Brasier, A.R.; Casola, A. Ikkepsilon regulates viral-induced interferon regulatory factor-3 activation via a redox-sensitive pathway. Virology 2006, 353, 155–165. [Google Scholar] [CrossRef]
- Soucy-Faulkner, A.; Mukawera, E.; Fink, K.; Martel, A.; Jouan, L.; Nzengue, Y.; Lamarre, D.; Vande Velde, C.; Grandvaux, N. Requirement of NOX2 and reactive oxygen species for efficient RIG-I-mediated antiviral response through regulation of MAVS expression. PLoS Pathog. 2010, 6, e1000930. [Google Scholar] [CrossRef]
- Chen, T.H.; Lo, Y.P.; Yang, C.F.; Chen, W.J. Additive protection by antioxidant and apoptosis-inhibiting effects on mosquito cells with dengue 2 virus infection. PLoS Negl. Trop. Dis. 2012, 6, e1613. [Google Scholar] [CrossRef]
- Pan, X.; Zhou, G.; Wu, J.; Bian, G.; Lu, P.; Raikhel, A.S.; Xi, Z. Wolbachia induces reactive oxygen species (ROS)-dependent activation of the Toll pathway to control dengue virus in the mosquito Aedes aegypti. Proc. Natl. Acad. Sci. USA 2012, 109, E23–E31. [Google Scholar]
- Glaser, R.L.; Meola, M.A. The native Wolbachia endosymbionts of Drosophila melanogaster and Culex quinquefasciatus increase host resistance to West Nile virus infection. PLoS One 2010, 5, e11977. [Google Scholar] [CrossRef]
- Schroder, M.; Kaufman, R.J. The mammalian unfolded protein response. Annu. Rev. Biochem. 2005, 74, 739–789. [Google Scholar] [CrossRef]
- Zeng, L.; Liu, Y.P.; Sha, H.; Chen, H.; Qi, L.; Smith, J.A. XBP-1 couples endoplasmic reticulum stress to augmented IFN-beta induction via a cis-acting enhancer in macrophages. J. Immunol. 2010, 185, 2324–2330. [Google Scholar] [CrossRef]
- Umareddy, I.; Pluquet, O.; Wang, Q.Y.; Vasudevan, S.G.; Chevet, E.; Gu, F. Dengue virus serotype infection specifies the activation of the unfolded protein response. Virol. J. 2007, 4. [Google Scholar] [CrossRef]
- Patramool, S.; Surasombatpattana, P.; Luplertlop, N.; Seveno, M.; Choumet, V.; Thomas, F.; Misse, D. Proteomic analysis of an Aedes albopictus cell line infected with Dengue serotypes 1 and 3 viruses. Parasit Vectors 2011, 4. [Google Scholar] [CrossRef]
- Pena, J.; Harris, E. Dengue virus modulates the unfolded protein response in a time-dependent manner. J. Biol. Chem. 2011, 286, 14226–14236. [Google Scholar] [CrossRef]
- Su, H.L.; Liao, C.L.; Lin, Y.L. Japanese encephalitis virus infection initiates endoplasmic reticulum stress and an unfolded protein response. J. Virol. 2002, 76, 4162–4171. [Google Scholar] [CrossRef]
- Rathore, A.P.; Ng, M.L.; Vasudevan, S.G. Differential unfolded protein response during Chikungunya and Sindbis virus infection: CHIKV nsP4 suppresses eIF2alpha phosphorylation. Virol. J. 2013, 10. [Google Scholar] [CrossRef]
- Nene, V.; Wortman, J.R.; Lawson, D.; Haas, B.; Kodira, C.; Tu, Z.J.; Loftus, B.; Xi, Z.; Megy, K.; Grabherr, M.; et al. Genome sequence of Aedes aegypti, a major arbovirus vector. Science 2007, 316, 1718–1723. [Google Scholar] [CrossRef]
- Holt, R.A.; Subramanian, G.M.; Halpern, A.; Sutton, G.G.; Charlab, R.; Nusskern, D.R.; Wincker, P.; Clark, A.G.; Ribeiro, J.M.; Wides, R.; et al. The genome sequence of the malaria mosquito Anopheles gambiae. Science 2002, 298, 129–149. [Google Scholar] [CrossRef]
- Arensburger, P.; Megy, K.; Waterhouse, R.M.; Abrudan, J.; Amedeo, P.; Antelo, B.; Bartholomay, L.; Bidwell, S.; Caler, E.; Camara, F.; et al. Sequencing of Culex quinquefasciatus establishes a platform for mosquito comparative genomics. Science 2010, 330, 86–88. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Prasad, A.N.; Brackney, D.E.; Ebel, G.D. The Role of Innate Immunity in Conditioning Mosquito Susceptibility to West Nile Virus. Viruses 2013, 5, 3142-3170. https://doi.org/10.3390/v5123142
Prasad AN, Brackney DE, Ebel GD. The Role of Innate Immunity in Conditioning Mosquito Susceptibility to West Nile Virus. Viruses. 2013; 5(12):3142-3170. https://doi.org/10.3390/v5123142
Chicago/Turabian StylePrasad, Abhishek N., Doug. E. Brackney, and Gregory D. Ebel. 2013. "The Role of Innate Immunity in Conditioning Mosquito Susceptibility to West Nile Virus" Viruses 5, no. 12: 3142-3170. https://doi.org/10.3390/v5123142



