1. Introduction
The influenza A virus (IAV) pandemic of 2009 involving the H1N1 strain has raised international concern over the potential for outbreaks of increased severity and highlighted the need for improved global surveillance and vaccination strategies [
1,
2]. The most effective way to prevent IAV infection is through vaccination. The proteins of IAV are used for vaccination. However, these vaccines need to be updated annually because IAVs are constantly undergoing change. It is therefore of substantial interest to develop a vaccine that targets conserved viral proteins (e.g., M2e, NP) and could be used to protect against unpredicted antigenic variation in both epidemic and pandemic outbreaks. These conserved viral protein vaccines do not usually target the surface molecules of IAV. This has resulted in a substantial drop in the efficacy of vaccination. It is important to develop a vaccine that targets the surface molecule of IAV.
Hemagglutinin (HA) is the most abundant protein on the viral coat and is highly immunogenic. Antibody responses against the viral surface protein HA are the major determinants of protection against IAV [
3,
4]. However, variations in HA arise rapidly due to antigenic shift and drift, allowing the virus to evade the immunity conferred by seasonal vaccines [
5]. HA is synthesized as the precursor HA0, which is cleaved into the HA1 and HA2 subunits. Whereas the membrane distal domain (HA1) can be highly variable, the stalk region containing the core fusion machinery constituted primarily by the HA2 subunit is relatively conserved [
6,
7]. Therefore, the development of an IAV vaccine promoting enhanced immune responses to these conserved IAV domains should provide more effective protection against this rapidly evolving virus. Antibodies targeting the HA2 subunit can provide broad protection against both seasonal and pandemic IAV infections [
7]. However, HA2 is masked by the membrane-distal portion of HA, a bulky and highly immunogenic globular head domain [
8]. Generation of broadly neutralizing antibodies (NAbs) is difficult because the use of the conserved fusion peptide of HA alone as a vaccine is only weakly immunogenic and does not confer adequate protection against IAV [
8]. Therefore, strategies to boost immune responses targeting HA2 are needed.
The Bacille Calmette-Guérin (BCG) vaccine against
Mycobacterium tuberculosis (
M. tuberculosis) has demonstrated marked immunomodulatory effects and enhanced antigen-specific antibody production when used in combination with influenza vaccines [
9,
10,
11]. As a major secreted antigen of M. tuberculosis and an immunodominant antigen of BCG,
M. tuberculosis secreted antigen Ag85A (Ag85A) is known to increase T helper type 1 (Th1) cytokine responses [
12], such as interferon (IFN)‑γ, which are important for cell-mediated immunity against IAV [
13].
Based on the hypothesis that the Ag85A may serve as an effective immune adjuvant for HA2, we constructed a novel influenza vaccine vector expressing the conserved region of the HA protein linked to Ag85A. The efficacy of this vaccine vector in preventing mortality and morbidity in mice was evaluated after challenge with IAV, and the cytokine profiles of the sera and cultured splenocytes were determined.
3. Discussion
While antibodies against HA are the major determinants of host protection against IAV infection, HA can also induce high frequencies of effector and memory CD4 T cells [
1]. The variability of IAV, particularly that of the HA1 subunit, is one of the major difficulties in developing prophylactic vaccines. Therefore, the development of an IAV vaccine that can induce immune responses to conserved domains, such as the HA2 subunit, would be highly advantageous.
In this study, we developed a DNA vaccine vector encoding the Ag85A-sHA2 fusion protein with the aim of enhancing the protection against IAV that is afforded by the HA2 immunogen alone. Following the confirmation of successful expression in mammalian cells, the novel vaccine vector and corresponding controls were used to immunize mice via the intramuscular route before the challenge with the PR8 virus. Interestingly, while the sera from mice vaccinated with the pAg85A-sHA2 fusion vaccine vector exhibited apparently low functional antibody titers in the HI test, they were relatively highly effective in the neutralization assay. Furthermore, the viral loads of the mice vaccinated with the fusion construct (pAg85A-sHA2) were the lowest after challenge compared with those of the other groups given DNA vaccine vectors expressing either Ag85A or HA2 epitopes alone. The HI antibody titers were low in these groups for both the primary and booster responses, as might be expected based upon the absence of the HA1 antigen in the vaccine vector, which could induce antibodies against the receptor binding site or other epitopes involved in HI. This observation is consistent with a previous study describing NAbs to HA2 epitopes that could not completely prevent the hemagglutination of erythrocytes [
7]. It is speculated that such antibodies may prevent infection at a step other than that of the initial virus attachment to the target cells [
7], although the mechanism has not yet been fully explained. The epitopes of sHA2 exist in many homologous strains, as showed in
Table 2. As shown in
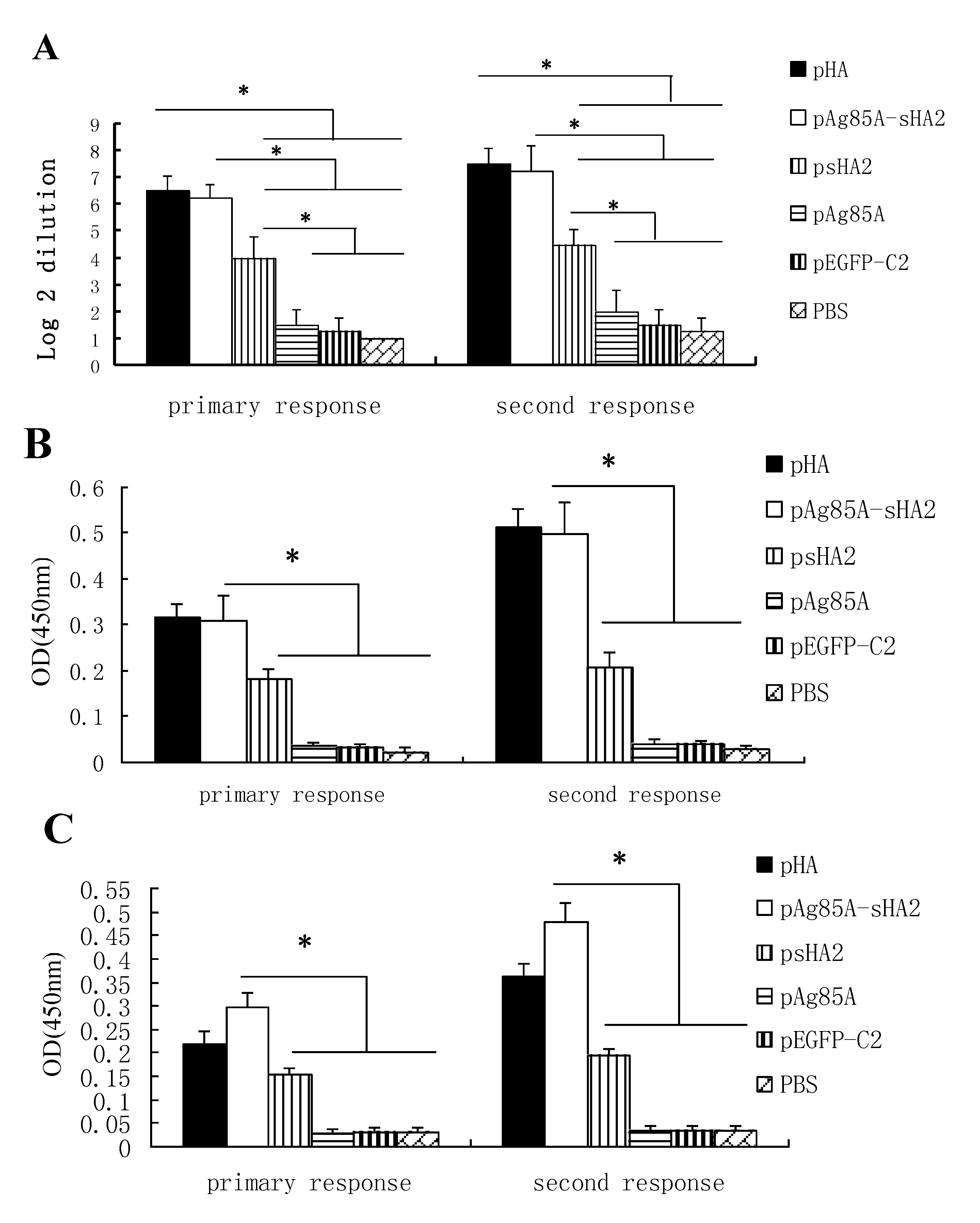
Figure 3C, the IgG antibodies induced by pAg85A-sHA2 could bind recombinant H5 HA proteins of the A/Anhui/1/2005 better, compared with those induced by pHA. Nevertheless, our results demonstrated that the humoral immune responses to HA2 epitopes were successfully induced by the pAg85A-sHA2 and psHA2 vaccine vectors and the IgG antibody induced by pAg85A-sHA2 or psHA2 could bind recombinant H5 HA proteins of the A/Anhui/1/2005, although further tests are warranted to determine whether these responses to HA2 epitopes may confer protection against diverse types of IAV.
The sHA2 construct did not include a signal peptide. The posttranslational processing of sHA2 did not involve its secretion into the extracellular space. We assumed that the high expression of the sHA2 ORF in our experiments, which was driven by the plasmid’s strong cytomegalovirus (CMV) promoter, led to the release of free sHA2, which was responsible for the induction of antibodies and cell damage.
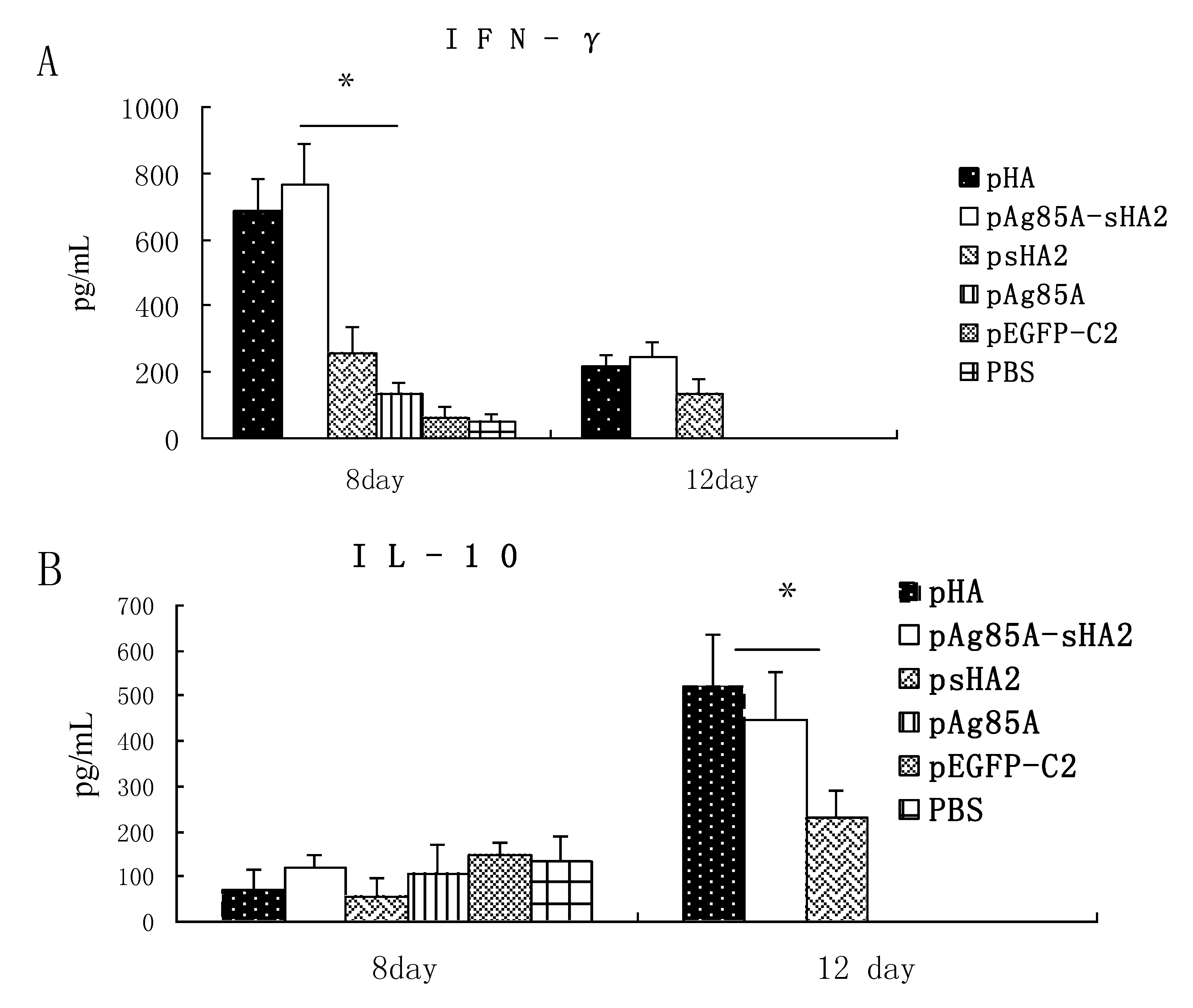
Our analysis of cytokine production showed that the sera IFN-γ levels in the psHA2 group were significantly higher than those in the pEGFP-C2 and PBS groups after PR8 virus challenge. IFN-γ is a representative Th1 cytokine associated with inhibition of IAV proliferation [
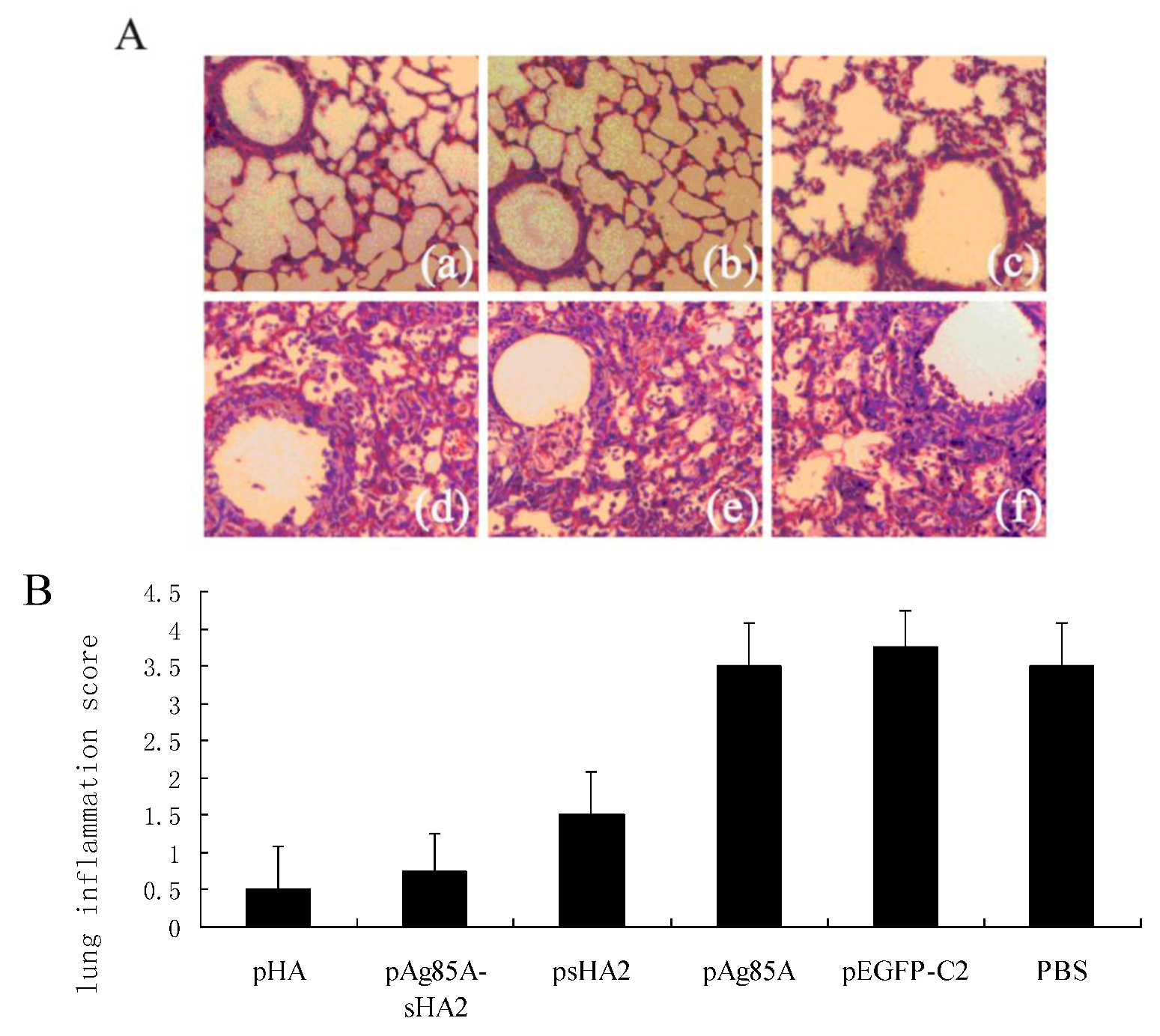
13]. The alveolar inflammatory cell numbers were reduced in the psHA2 group. These results confirmed that sHA2 alone could provide limited protection against the PR8 virus.
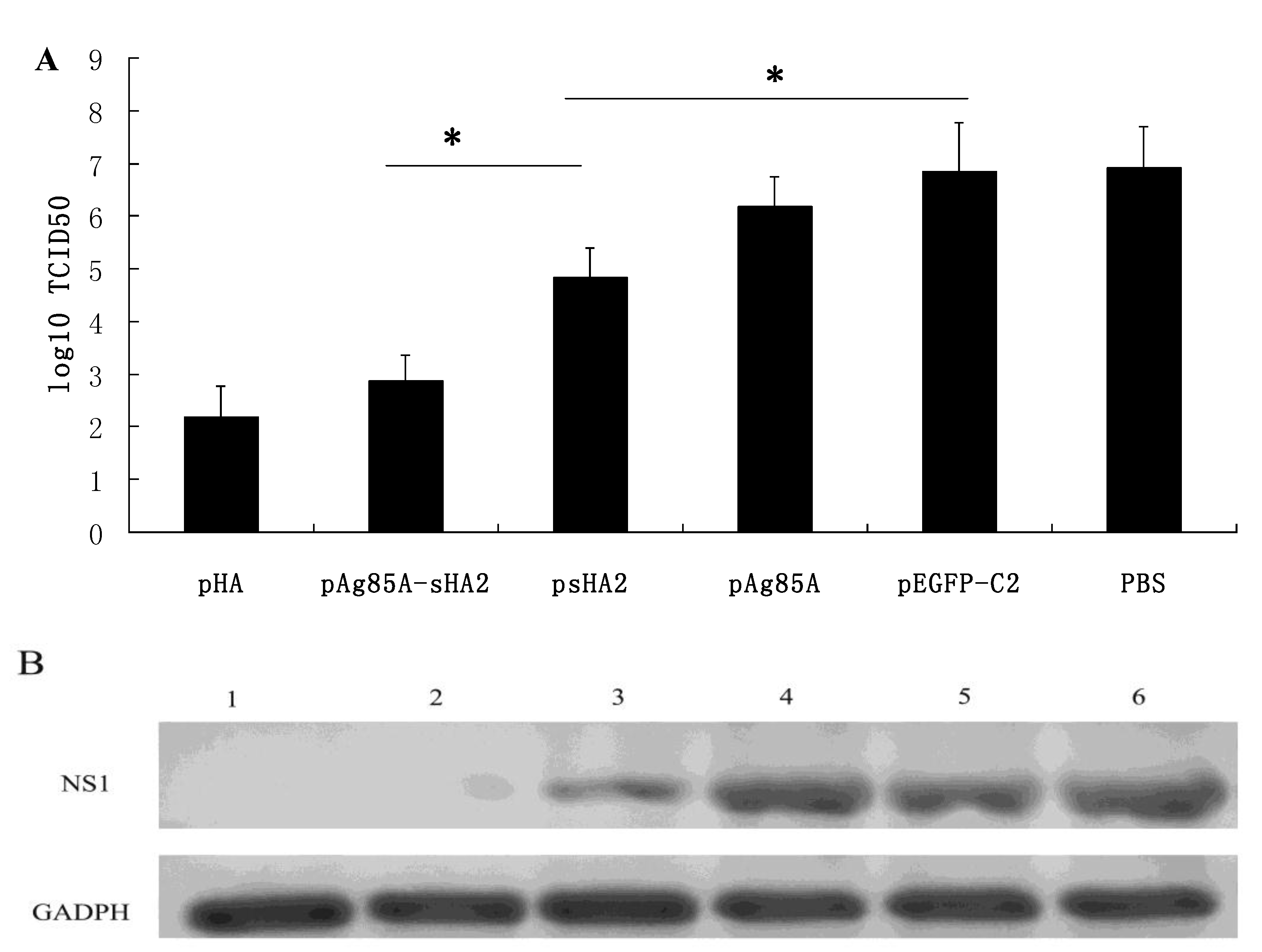
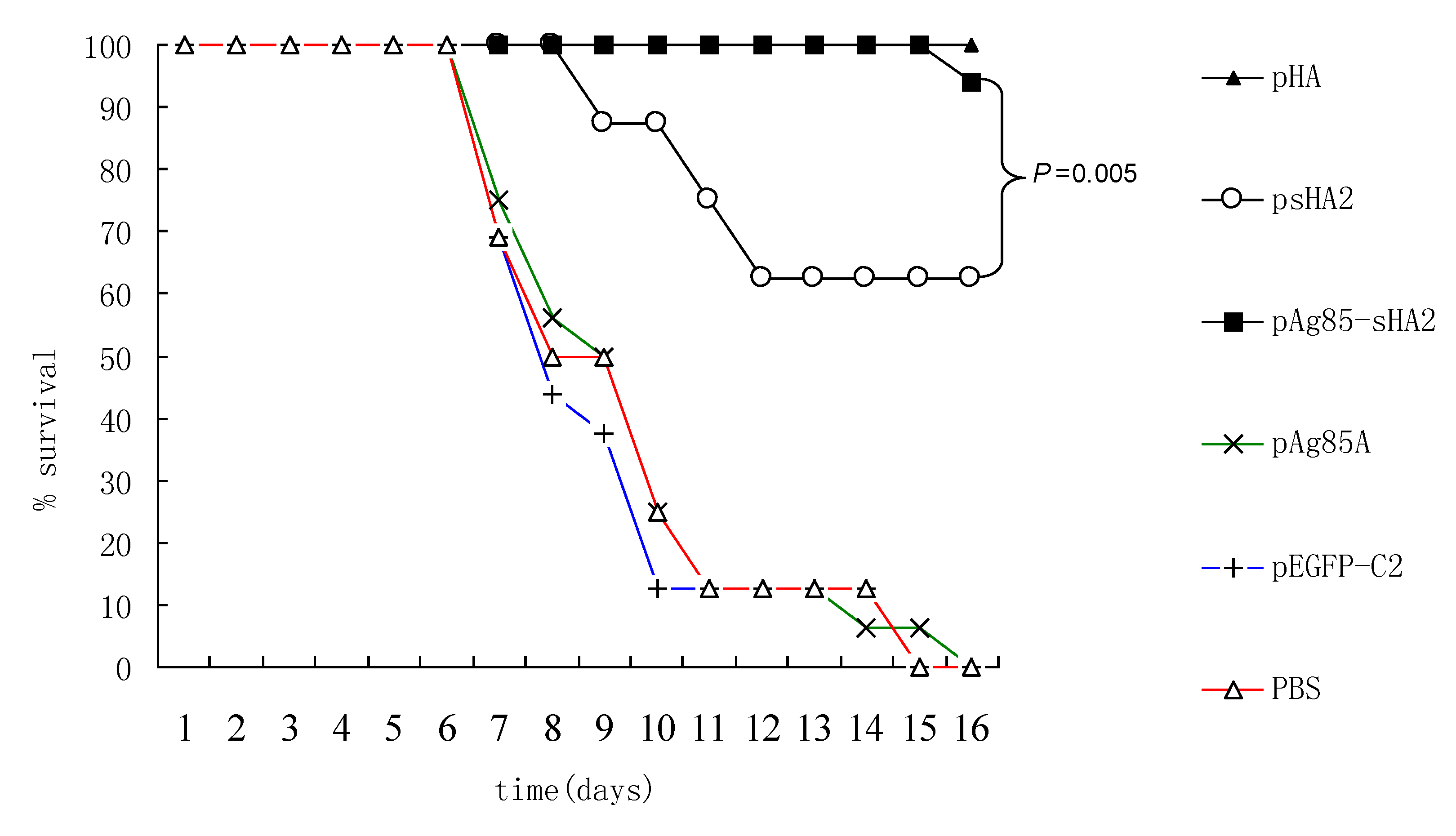
Importantly, vaccination with the pAg85A-sHA2 fusion construct produced significantly higher sera IFN-γ levels and resulted in higher survival rates than vaccination with pHA2. The IAV titer from the pAg85A-sHA2 group was lower compared to those from the pEGFP-C2, psHA2 and pAg85A groups. The alveolar inflammatory cell numbers were reduced in the psHA2, pAg85A-sHA2 and pHA groups. The increased survival rate after challenge in the pAg85A-sHA2 group was likely due to Ag85A-mediated immune activation, which enhanced the protective effects of sHA2 against PR8 virus challenge, thereby decreasing the viral load. As IAV can suppress the functions of immune cells [
20,
21], the elimination of IAV in our experiments may have thus resulted in the enhancement of these functions. Finally, the decreased viral loads relieved pneumonia in mice, as determined by the histopathological analysis.
In addition to analyzing the IFN-γ responses above, we also investigated the IL-4 levels following vaccination with the various constructs. IL-4 is a cytokine of the Th2 response [
22]. Sera IL-4 levels were significantly higher in the pAg85A-sHA2 group or the pHA vaccinated group compared to those in the psHA2 group (
P < 0.05) on Day 12 after infection with the PR8 virus. These results demonstrated that pHA or pAg85A-sHA2 vaccine could increase the Th2 cytokines response.
HA2 is masked by the membrane-distal portion of HA, which is a bulky and highly immunogenic globular head domain [
8]. The HA2 subunit is relatively conserved [
6,
7]. Experiments are ongoing to determine whether the responses to HA2 epitopes may confer protection against diverse types of IAV. The preliminary results, which will be reported in a future paper, have shown that the antibodies against sHA2 epitopes can bind H5 HA.
4. Materials and Methods
4.1. Design of Epitopes
The HA (ABO21709.1) and Ag85A (AY207395.1) sequences were downloaded from the National Center for Biotechnology Information (NCBI). The B-cell and T-cell epitopes for HA2 and Ag85A were predicted bioinformatically using the network servers BCEPRED [
23] and SYFPEITHI [
24], respectively. With the NCBI Blast algorithm [
25], conserved epitopes were clearly identified in HA2 between amino acids (aa) 350 and 498 in the N-terminal region of HA Subtypes 1, 5 and 6 (
Table 2). The predicted results were similar to those that were reported by Sui
et al. [
7]. The short HA2 (sHA2) sequence (from aa 350 to 498) was used to construct the novel vaccine vector. The immunodominant epitopes of Ag85A were identified within aa 1 to 250 of the N-terminal sequence (data not shown), which was consistent with previous reports [
14,
15].
Table 2.
Predicted B and T cell epitopes in sHA2.
Table 2.
Predicted B and T cell epitopes in sHA2.
| Epitope | Region | Amino acid sequence |
|---|
| B epitope | H1HA368–375 | YHHQNEQG |
| B epitope | H1HA379–386 | YAADQKSTQ |
| B epitope | H1HA391–397, H5HA347–353 | GITNKVN |
| B epitope | H1HA453–460, H5HA455–462 | DFHDSNVK |
| B epitope | H1HA491–498, H5HA493–500 | ECMESVRN |
| T epitope | H1HA373–387 | EQGSGYAADQKSTQN |
| T epitope | H1HA439–453 | NAELLVLLENERTLD |
| T epitope | H1HA434–448 | DIWTYNAELLVLLEN |
4.2. Construction of Recombinant Eukaryotic Expression Plasmid Vaccine Vectors
The pEGFP/Ag85A (pAg85A) plasmid was constructed by first cloning the Ag85A sequence (aa 1 to 250 of the N-terminus) [
14,
15,
16] from BCG, and pEGFP/sHA2 (psHA2) was constructed by cloning the sHA2 sequence (aa 350–498 of the HA N-terminus) from the IAV strain A/PR/8/34 (H1N1) as described previously [
17]. These sequences contain the immunodominant epitopes of Ag85A and sHA2. The hydrophilicity of the proximal 250 amino acids of Ag85A is similar to that of the HA N‑terminal amino acids 290–321 [
16]. Ag85A and sHA2 were fused using a 5 glycine linker as described previously [
8] and the resulting fusion was designated as Ag85A-sHA2. The pEGFP/Ag85A-sHA2 (p Ag85A-sHA2) expression vector was constructed by inserting Ag85A-sHA2 into pEGFP-C2 (Clontech, Houston, TX, USA). The pEGFP/HA (pHA) expression vector was constructed by inserting HA (Reference Sequence: NC_002017.1) into pEGFP-C2. The successful insertion of all fragments was confirmed by restriction endonuclease digestion. Sequencing was performed using an ABI Prism 377XL DNA sequencer. Large scale extraction and purification of pEGFP-C2, pHA, pAg85A-sHA2, pAg85A and psHA2 were performed using the EndoFree Plasmid Maxi Kit (Qiagen, Düsseldorf, Germany).
4.3. Cell Transfection Assay
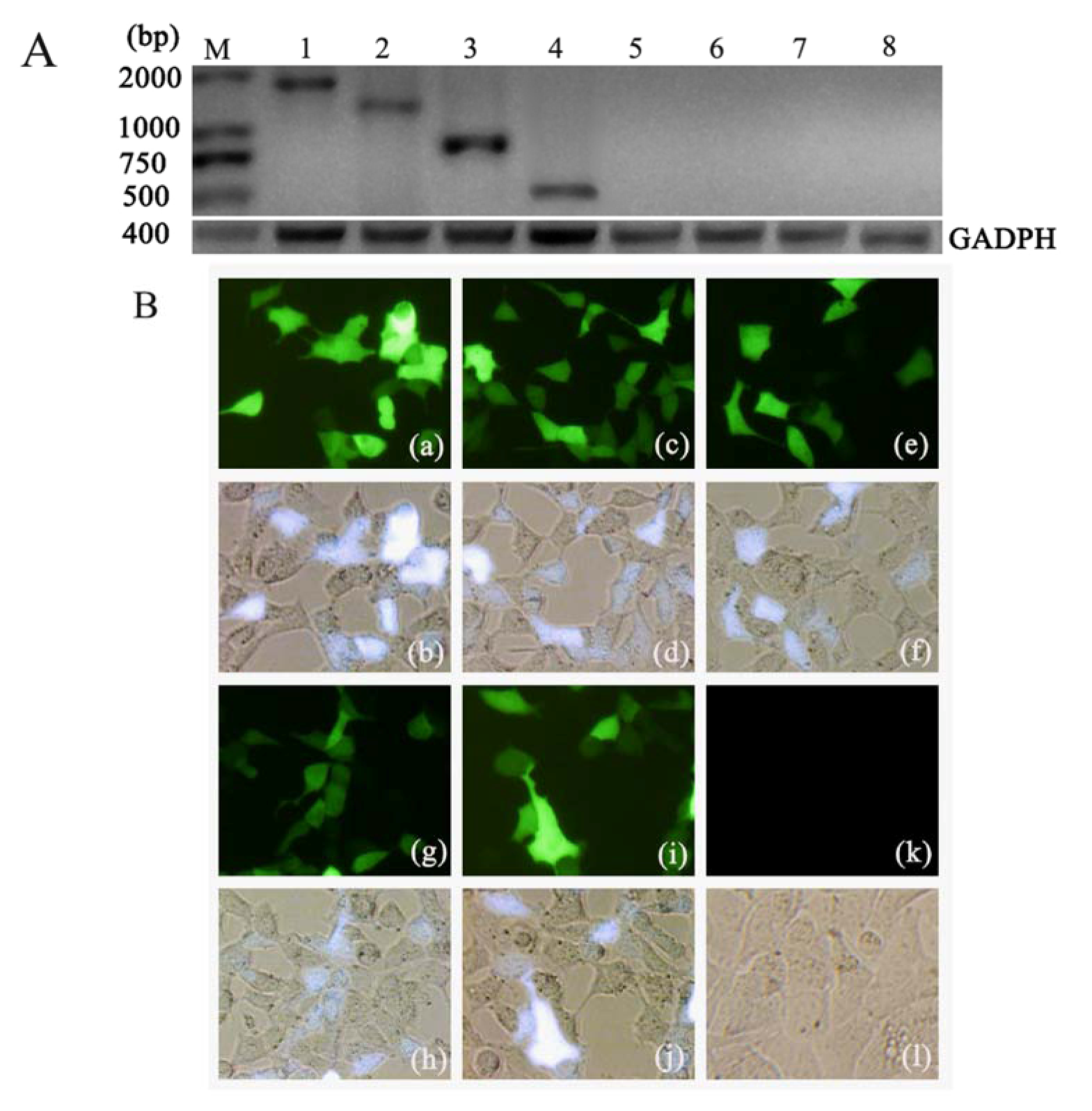
Using the LipofectamineTM 2000 transfection reagent (Invitrogen, Carlsbad, CA, USA), 0.8 μg of each of the purified eukaryotic expression plasmids pHA, pAg85A-sHA2, pAg85A, pEGFP-C2 and psHA2 were transfected into 1 × 105 HEK293 cells (CRL-1573™) obtained from the American Type Culture Collection (ATCC) according to the manufacturer’s instructions. Each plasmid was transfected in triplicate wells, and PBS was applied as a mock transfection control. Plasmid expression in all sets of transfected cells was assessed by fluorescence microscopy at 48 h after transfection.
Total RNA from each set of transfected cells was isolated by using 1 mL TRIpure Reagent according to the manufacturer’s instructions (Biotake, Beijing, China). Contaminating genomic DNA was then eliminated from the RNA preparations by using RNase-free DNase I (Biotake, Beijing, China). Reverse transcription (RT) was performed with the ReverAidTM H Minus First Strand cDNA Synthesis Kit (Fermentas, Pittsburg, PA, USA). cDNA samples were amplified by PCR using 2× TransTaq High Fidelity (HiFi) PCR SuperMix (Transgene, Beijing, China) in a 50 μL reaction mixture according to the manufacturer’s instructions. PCR primer pairs for mouse GAPDH were obtained from the RevertAidTM H Minus First Strand cDNA Synthesis Kit (400 bp). The following PCR primer pairs were used: pAg85A-sHA2, 5'-CCCTCGAGCATGCAGCTTGTTGACAG-3' and
5'-CGCGGATCCTCATTTGGGGTAATCATAA-3';
pAg85A, 5'-CCCTCGAGCATGCAGCTTGTTGACAG-3' and
5'-CGGGATCCTTAGCCGTTGCCGCAGTACAC-3';
psHA2, 5'-CCGCTCGAGTATTGAAGGGGGATGGA-3' and
5'-CGCGGATCCTCATTTGGGGTAATCATAA-3'; and
pHA, 5'-CATAAGCTTAACCAAAATGAAGGCAACC-3' and
5'-CGCGGATCCTCATATTTCTGAAATTCTAAT-3'. Control PCRs using glyceraldehyde 3-phosphate dehydrogenase (GAPDH) primers were performed on cDNA from the pHA, pAg85A-sHA2, pAg85A, pEGFP-C2 and psHA2 groups. Test PCRs for the pEGFP-C2 group were performed with the pHA primers, psHA2 primers, pAg85A primers and pAg85A-sHA2 primers. Test PCRs for the pHA, pAg85A-sHA2, pAg85A and psHA2 groups were performed with their respective primers. All PCRs were performed simultaneously under the same conditions as follows: five minutes at 95 °C, followed by 30 cycles consisting of 30 s at 95 °C, 30 s at 50 °C and 2 min at 72 °C. Following the amplification, 10 μL of each PCR product were subjected to electrophoresis on a 2% agarose gel and visualized by ethidium bromide staining. PCR amplification and sequencing were performed using an ABI Prism 377XL DNA Sequencer.
4.4. DNA Vaccination and Challenge Infection with the PR8 Virus
A total of 120 female BALB/c mice, 4–6 weeks old, were purchased from the animal care center of Sichuan University and divided into six groups, with 20 mice in each group. They were bred under pathogen-free conditions. Each group of mice was injected separately with one of the following: pAg85A-sHA2, pAg85A, psHA2, pEGFP-C2, PBS or pHA. The mice were immunized by intramuscular injection in the quadriceps of both legs (50 μg plasmid in 50 μL PBS per leg) twice with a 3-week interval between injections. The pHA group was the positive control group.
Seven days after the booster immunization, the mice were challenged with IAV, as described previously [
17,
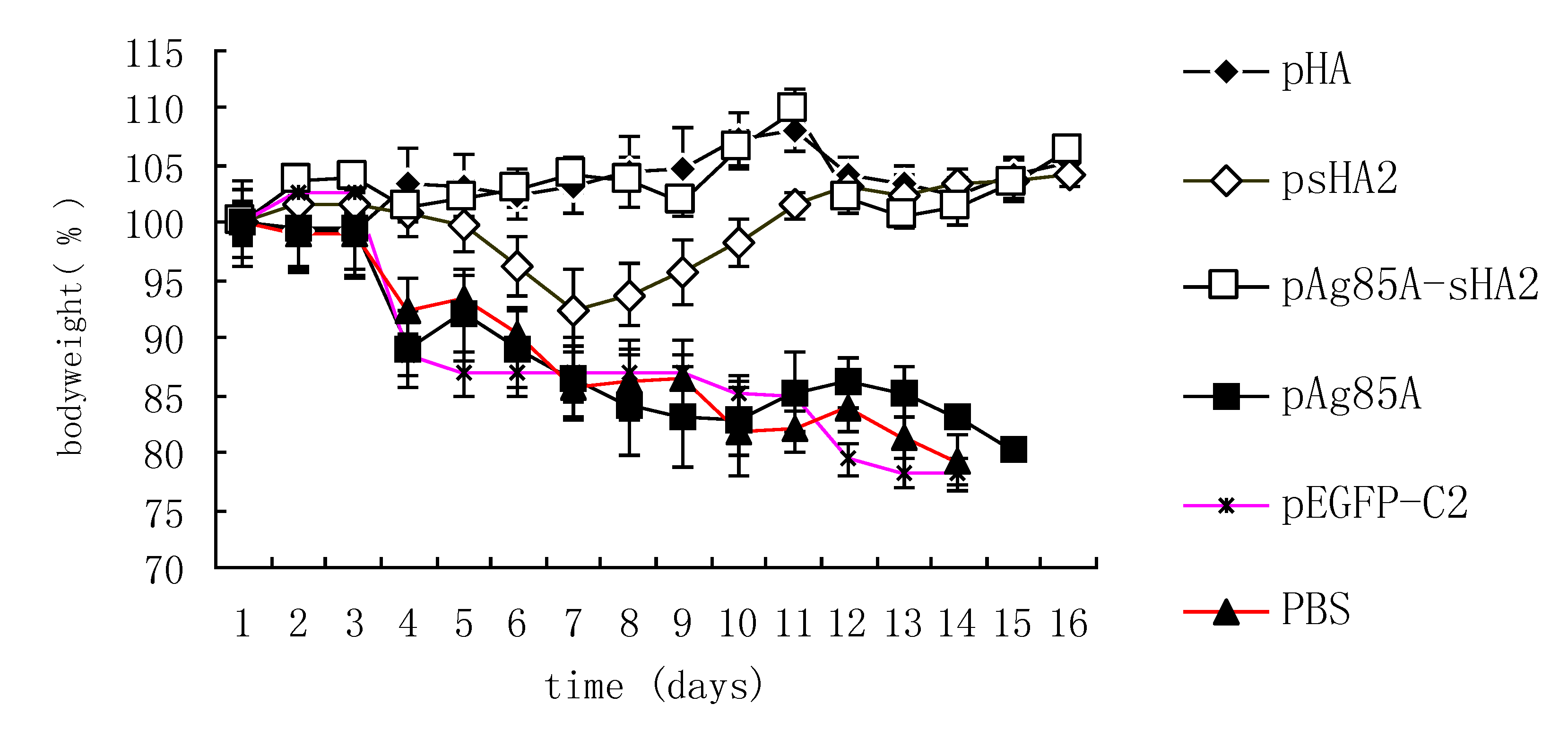
18]. In brief, the A/Puerto Rico/8/34 virus (H1N1, PR8 virus) was harvested from 10- to 11-day-old embryonated chicken eggs and titrated in a plaque assay. The mice were anesthetized by inhalation of isoflurane and challenged with IAV via intranasal inoculation of ten times of 50% mouse lethal doses (10 × LD50) of the PR8 virus diluted in PBS in a total volume of 50 μL. The survival rates and body weights were monitored daily, and mice losing more than 25% of their initial weights were sacrificed and scored as dead. All of the experiments were performed with institutional approval.
4.5. Specimen Collection
Specimens were collected as reported by Zhang
et al. [
17] and Tamura
et al. [
18]. In brief, the first immune serum samples were obtained from each mouse via tail bleeding on day 1 before the booster immunization. The second immune serum samples were obtained from four randomly chosen mice of each group by terminal cardiac bleeding under chloroform anesthesia with chloroform on Day 8 after the virus challenge. Splenocytes from the pAg85A-sHA2, psHA2, pAg85A, pEGFP-C2, PBS and pHA group were isolated and cultured to measure cytokine production. The trachea and lungs were collected and washed twice with 2 mL sterile PBS containing 0.1% BSA. The bronchoalveolar lavages were centrifuged to remove cellular debris, filtered through a 0.22 μM membrane and used for virus titration. Total RNA was isolated from 0.2 mg of left lung tissue from each group using 2 mL TRIpure Reagent according to the manufacturer’s instructions (Biotake, Beijing, China). Contaminating genomic DNA was then eliminated from the RNA preparations using RNase-free DNase I (Biotake). RT reactions were performed with RevertAid
TM H Minus First Strand cDNA Synthesis Kit (Fermentas, Pittsburg, PA, USA). Lung cDNA samples from all of the groups were stored at -80°C before use. A 0.6 mg left lung samples from each group was used for the histopathological analyses. The third serum samples for the cytokine measurements were obtained from each mouse via tail bleeding on Day 12 after the virus challenge. All of the sera and bronchoalveolar lavages samples were stored at −20 °C before use.
4.6. HA Inhibition Assay
The HA inhibition (HI) assay was performed as reported by Zhang
et al. [
17]. In brief, the sera were collected at Day 1 before the booster immunization and at Day 8 after the virus challenge. Two-fold serial dilutions of the sera were prepared in PBS. Four HA units of virus (A/PR/8/34, H1N1) in 25 μL per well were added to a 96-well microtiter plate containing 25 μL of the diluted serum sample, mixed and incubated at 4 °C for 1 h. Chicken erythrocytes in 50 μL (0.5% v/v) were then added to each well and the plate was mixed and incubated at 4 °C for 1 h. The plate was further incubated at 4 °C for 30–45 min. The HI end-point titers were calculated as the reciprocals of the highest serum dilutions that completely inhibited hemagglutination.
4.7. Neutralization Assay
The neutralization assay was performed as previously described [
8]. In brief, the sera were collected at Day 1 before the booster immunization and at Day 8 after the virus challenge. Fifty microliters of each of the two-fold serial dilutions of treated sera were mixed with 50 μL of the 50% tissue culture-infective dose (TCID50) of the PR8 virus and incubated for 45 min at 37 °C. Madin-Darby canine kidney (MDCK) cells (ATCC, CCL-34) growing in a 24-well plate were washed with PBS and inoculated with the 100-μL virus-antibody mixtures. Following a 45 min incubation, Dulbecco's Modified Eagle's Medium
(DMEM) supplemented with 2 µg/mL trypsin (Boster Biotechnological, Wuhan, China) and 3% bovine serum albumin (Beijing Biosynthesis Biotechnology Co. LTD, Wuhan, China) were added to each well of the 24-well plate. Each dilution was performed in four replicate wells. The cytopathic effect (CPE) was assayed after incubation at 37 °C for 2 days. The titer from each sample, recorded as the dilution that reduced the CPE by 50%, was calculated by the Reed–Muench method.
4.8. Serological Assays by ELISA
Ninety-six-well plates (Costar) were coated with 0.1μg per well of purified recombinant HA protein in PBS. The recombinant HA proteins of the A/Anhui/1/2005 (H5N1, catalog number: 11048-V08H4) and A/PR/8/34 (H1N1, catalog number: 11684-V08H) viruses were obtained from Sino Biological Inc., China. 500-fold dilutions of antisera were incubated on the plates, bound antibody was detected with peroxidase-link Goat Anti-Mouse IgG antibody (Boster Biotechnological, Wuhan, China) according to the manufacturer’s instructions. The experiments were repeated three times, and the results from one experiment are shown.
4.9. Cytokine Measurements
Spleens were removed aseptically on Day 8 after the virus challenge and processed as follows. Cells were isolated, washed, adjusted to a concentration of 5 × 106 cells per mL, and grown in a 96-well flat-bottom plate (5 × 105 cells per well) in Roswell Park Memorial Institute medium (RPMI) 1640 medium, supplemented with HEPES (N-2-hydroxyethylpiperazine-N'-2-ethanesulfonic acid), glutamine and 10% heat-inactivated fetal calf serum (FCS). Cells were stimulated separately with 0.5 µg concanavalin A (ConA) or 1.25 µg recombinant HA of A/PR/8/34 (catalog number: 11684-V08H, Sino Biological Inc., Beijing, China) and incubated at 37 °C in a humidified CO2 incubator. After 72 h, supernatants were harvested, pooled and stored frozen at −20 °C until assayed. The experiments were repeated three times, and the results from one experiment were shown.
Sera were collected on Days 8 and 12 after infection with the PR8 virus. For the cytokine measurements, sera from each group were pooled and stored at −20 °C until assayed. IFN-γ and IL-4 from the sera and IFN-γ from the splenocyte culture supernatants were measured using specific enzyme-linked immunosorbent assays (ELISA; Boster Biotechnological, Wuhan, China) according to the manufacturer’s instructions.
4.10. IAV Titrations
IAV titering was performed as reported by Zhang
et al. [
17]. In brief, 10-fold serial dilutions of the sterile bronchoalveolar lavage fluid that were collected on Day 8 after the virus challenge were prepared and used to inoculate confluent monolayers of MDCK cells in 24-well plates. Each dilution was repeated in four wells. The CPE was assayed after incubation at 37 °C for 2 days. The virus titer of each specimen, recorded as the 50% tissue culture infection dose (TCID50), was calculated by the Reed–Muench method. The virus titer from each group was presented as the mean ± SD of all specimens in each group.
4.11. PCR-Amplification and Sequencing of PR8 Nucleic Acids Using RNA Extracted from Lungs of Infected Mice
Lung cDNA samples from each group were amplified by PCR using the 2× TransTaq High Fidelity (HiFi) PCR SuperMix (Transgene, Beijing, China) in a 50 μL reaction mixture according to the manufacturer’s instructions. PCR primer pairs for mouse GAPDH were obtained from the RevertAidTM H Minus First Strand cDNA Synthesis Kit. The following IAV NS1 (GenBank: V01104.1) PCR primer pairs were used: 5'- GCTTTCAGGTAGATTGCT -3' and 5'-CCATTCTCATTACTGCTTCT-3'; PCRs for the pHA, pAg85A-sHA2, PBS, pAg85A, pEGFP-C2 and psHA2 groups were performed with GAPDH primers and NS1 primers. All of PCRs were performed simultaneously under the same conditions as follows: Five minutes at 95 °C, followed by the 25 cycles consisting of 30 s at 95 °C, 30 s at 50 °C and 2 min at 72 °C. After amplification, 10 μL of each PCR product were subjected to electrophoresis on a 2% agarose gel and visualized by ethidium bromide staining. PCR amplification and sequencing were performed using an ABI Prism 377XL DNA sequencer.
4.12. Histopathological Analysis
Lungs harvested at the 8th day after the virus challenge were used for histological examination. The specimens were fixed in 10% formalin, and embedded in paraffin. Four micrometer sections were stained with H&E. All of the sections were examined for histopathological changes in a blind manner, as described previously [
19]. In brief, the entire lung surface was analyzed with respect to the following parameters: pleuritis, bronchitis, edema, interstitial inflammation, percentage of pneumonia and endothelialitis. The pathologist assigned scores to the tissue sections according to these parameters. For lung inflammation and damage, each parameter was graded on a scale of 0 to 4, with 0 meaning “absent” and 4 meaning “severe.” The lung inflammation score for each tissue section was expressed as the average of the scores for all parameters.
4.13. Statistical Analysis
Data are presented as means ± SD. Group-group comparisons were analyzed using t-tests. Survival rates were determined using the Kaplan-Meier method. P values < 0.05 were considered to be significant.