RNA Dependent RNA Polymerases: Insights from Structure, Function and Evolution
Abstract
1. Introduction
2. Structural Features
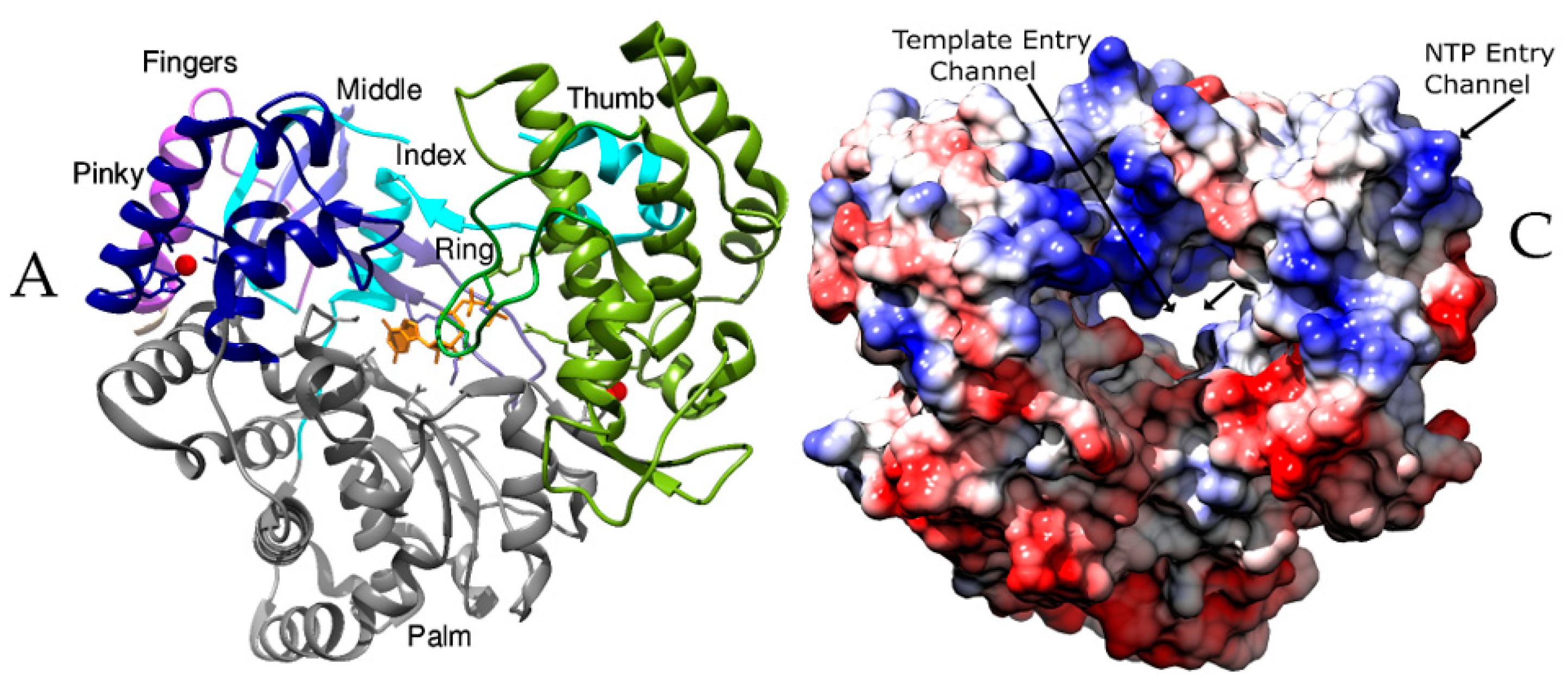
2.1. The Subdomains
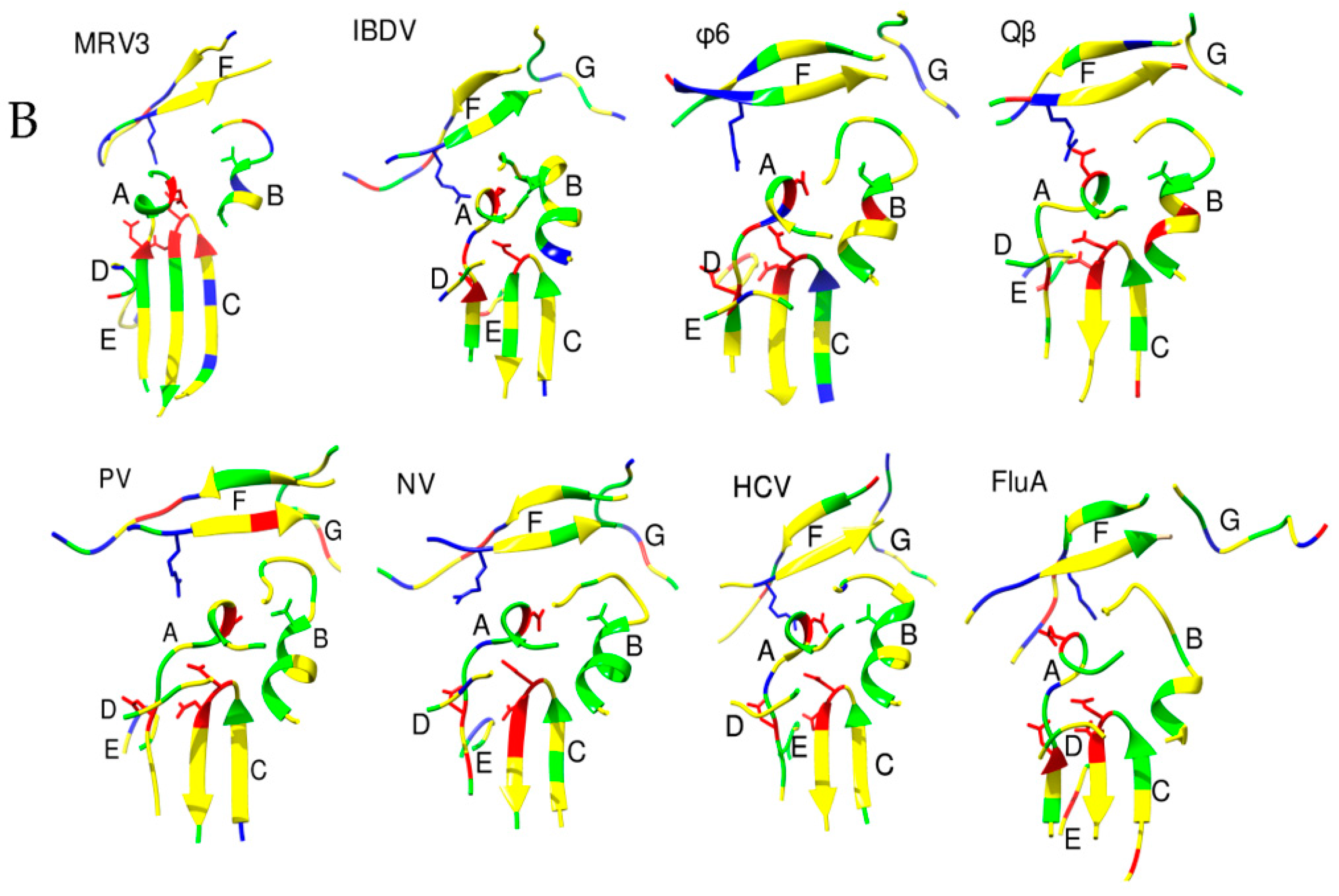
2.2. The Motifs
2.3. The Channels
2.4. Additional Structural Elements
3. Structure-Based Phylogeny of RdRps
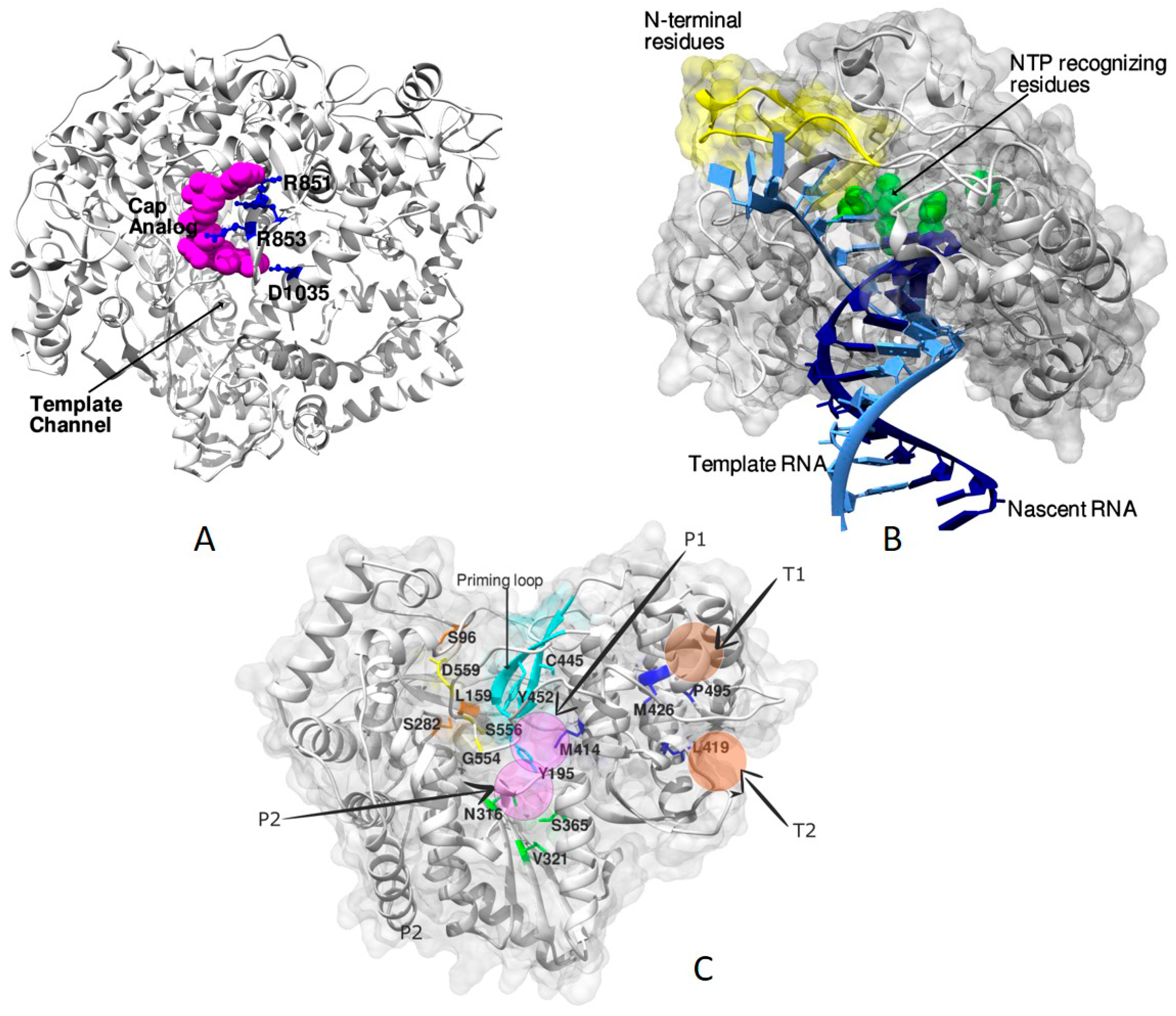
4. Analysis of RdRp Complexes
4.1. RdRp Complexes in Reoviridae
4.2. RdRp Complexes in Bacteriophages
4.3. RdRp Complexes of Caliciviridae
4.4. RdRp Complexes of Picornaviridae
4.5. RdRp Complexes of Flaviviridae
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Elena, S.F.; Sanjuan, R. Adaptive value of high mutation rates of RNA viruses: Separating causes from consequences. J. Virol. 2005, 79, 11555–11558. [Google Scholar] [CrossRef] [PubMed]
- Crotty, S.; Cameron, C.E.; Andino, R. RNA virus error catastrophe: Direct molecular test by using ribavirin. Proc. Natl. Acad. Sci. USA 2001, 98, 6895–6900. [Google Scholar] [CrossRef] [PubMed]
- Lai, M.M. Cellular factors in the transcription and replication of viral RNA genomes: A parallel to DNA-dependent RNA transcription. Virology 1998, 244, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Bruenn, J.A. A structural and primary sequence comparison of the viral RNA-dependent RNA polymerases. Nucleic Acids Res. 2003, 31, 1821–1829. [Google Scholar] [CrossRef] [PubMed]
- Ng, K.K.S.; Arnold, J.J.; Cameron, C.E. Structure-function relationships among RNA-dependent RNA polymerases. Curr. Top. Microbiol. Immunol. 2008, 320, 137–156. [Google Scholar] [PubMed]
- Te Velthuis, A.J.W. Common and unique features of viral RNA-dependent polymerases. Cell. Mol. Life Sci. 2014, 71, 4403–4420. [Google Scholar] [CrossRef] [PubMed]
- Ferrer-Orta, C.; Arias, A.; Escarmís, C.; Verdaguer, N. A comparison of viral RNA-dependent RNA polymerases. Curr. Opin. Struct. Biol. 2006, 16, 27–34. [Google Scholar] [CrossRef] [PubMed]
- Mönttinen, H.A.M.; Ravantti, J.J.; Stuart, D.I.; Poranen, M.M. Automated structural comparisons clarify the phylogeny of the right-hand-shaped polymerases. Mol. Biol. Evol. 2014, 31, 2741–2752. [Google Scholar] [CrossRef] [PubMed]
- Hyodo, K.; Kaido, M.; Okuno, T. Host and viral RNA-binding proteins involved in membrane targeting, replication and intercellular movement of plant RNA virus genomes. Front. Plant Sci. 2014, 5, 321. [Google Scholar] [CrossRef] [PubMed]
- Nagy, P.D.; Pogany, J.; Xu, K. Cell-free and cell-based approaches to explore the roles of host membranes and lipids in the formation of viral replication compartment induced by tombusviruses. Viruses 2016, 8, 68. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.-P.; Chen, I.-H.; Tsai, C.-H. Host Factors in the Infection Cycle of Bamboo mosaic virus. Front. Microbiol. 2017, 8, 437. [Google Scholar] [CrossRef] [PubMed]
- Miller, S.; Krijnse-Locker, J. Modification of intracellular membrane structures for virus replication. Nat. Rev. Microbiol. 2008, 6, 363–374. [Google Scholar] [CrossRef] [PubMed]
- Romero-Brey, I.; Bartenschlager, R. Membranous replication factories induced by plus-strand RNA viruses. Viruses 2014, 6, 2826–2857. [Google Scholar] [CrossRef] [PubMed]
- Sigrist, C.J.A.; Cerutti, L.; De Castro, E.; Langendijk-Genevaux, P.S.; Bulliard, V.; Bairoch, A.; Hulo, N. PROSITE, a protein domain database for functional characterization and annotation. Nucleic Acids Res. 2009, 38, D161–D166. [Google Scholar] [CrossRef] [PubMed]
- Berman, J.; Westbrook, Z.; Feng, G.; Gilliland, T.N.; Bhat, H.; Weissig, I.N.; Shindyalov, P.E.B. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera—A Visualization System for Exploratory Research and Analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef] [PubMed]
- DeLano, W.L. The PyMOL Molecular Graphics System, version 1.8; Schrödinger LLC: New York, NY, USA, 2014.
- Černý, J.; Bolfíková, B.Č.; Valdés, J.J.; Grubhoffer, L.; Růžek, D. Evolution of tertiary structure of viral RNA dependent polymerases. PLoS ONE 2014, 9, e96070. [Google Scholar] [CrossRef] [PubMed]
- Ferrer-Orta, C.; Ferrero, D.; Verdaguer, N. RNA-dependent RNA polymerases of picornaviruses: From the structure to regulatory mechanisms. Viruses 2015, 7, 4438–4460. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Chung, M.S.; Kim, K.H. Structure and function of caliciviral RNA polymerases. Viruses 2017, 9, 329. [Google Scholar] [CrossRef] [PubMed]
- Pan, J.; Vakharia, V.N.; Tao, Y.J. The structure of a birnavirus polymerase reveals a distinct active site topology. Proc. Natl. Acad. Sci. USA 2007, 104, 7385–7390. [Google Scholar] [CrossRef] [PubMed]
- Graham, S.C.; Sarin, L.P.; Bahar, M.W.; Myers, R.A.; Stuart, D.I.; Bamford, D.H.; Grimes, J.M. The N-Terminus of the RNA polymerase from infectious pancreatic necrosis virus is the determinant of genome attachment. PLoS Pathog. 2011, 7. [Google Scholar] [CrossRef] [PubMed]
- Gong, P.; Peersen, O.B. Structural basis for active site closure by the poliovirus RNA-dependent RNA polymerase. Proc. Natl. Acad. Sci. USA 2010, 107, 22505–22510. [Google Scholar] [CrossRef] [PubMed]
- Lu, G.; Gong, P. A structural view of the RNA-dependent RNA polymerases from the Flavivirus genus. Virus Res. 2016, 234, 34–43. [Google Scholar] [CrossRef] [PubMed]
- Choi, K.H.; Rossmann, M.G. RNA-dependent RNA polymerases from Flaviviridae. Curr. Opin. Struct. Biol. 2009, 19, 746–751. [Google Scholar] [CrossRef] [PubMed]
- Te Velthuis, A.J.W.; Robb, N.C.; Kapanidis, A.N.; Fodor, E. The role of the priming loop in RNA synthesis. Nat. Microbiol. 2016, 1. [Google Scholar] [CrossRef] [PubMed]
- Te Velthuis, A.J.W.; Oymans, J. Initiation, elongation and realignment during influenza virus mRNA synthesis. J. Virol. 2017, 92, e01775-17. [Google Scholar] [CrossRef] [PubMed]
- Kidmose, R.T.; Vasiliev, N.N.; Chetverin, A.B.; Andersen, G.R.; Knudsen, C.R. Structure of the Qβ replicase, an RNA-dependent RNA polymerase consisting of viral and host proteins. Proc. Natl. Acad. Sci. USA 2010, 107, 10884–10889. [Google Scholar] [CrossRef] [PubMed]
- Tao, Y.; Farsetta, D.L.; Nibert, M.L.; Harrison, S.C. RNA synthesis in a cage—Structural studies of reovirus polymerase λ3. Cell 2002, 111, 733–745. [Google Scholar] [CrossRef]
- Wu, J.; Liu, W.; Gong, P.; Gong, P. A structural overview of RNA-dependent RNA polymerases from the Flaviviridae family. Int. J. Mol. Sci. 2015, 16, 12943–12957. [Google Scholar] [CrossRef] [PubMed]
- Lu, G.; Gong, P. Crystal Structure of the Full-length Japanese encephalitis virus NS5 reveals a conserved methyltransferase-polymerase interface. PLoS Pathog. 2013, 9. [Google Scholar] [CrossRef] [PubMed]
- Surana, P.; Satchidanandam, V.; Nair, D.T. RNA-dependent RNA polymerase of Japanese encephalitis virus binds the initiator nucleotide GTP to form a mechanistically important pre-initiation state. Nucleic Acids Res. 2014, 42, 2758–2773. [Google Scholar] [CrossRef] [PubMed]
- Choi, K.H.; Gallei, A.; Becher, P.; Rossmann, M.G. The structure of bovine viral diarrhea virus RNA-dependent RNA polymerase and its amino-terminal domain. Structure 2006, 14, 1107–1113. [Google Scholar] [CrossRef] [PubMed]
- Choi, K.H.; Groarke, J.M.; Young, D.C.; Kuhn, R.J.; Smith, J.L.; Pevear, D.C.; Rossmann, M.G. The structure of the RNA-dependent RNA polymerase from bovine viral diarrhea virus establishes the role of GTP in de novo initiation. Proc. Natl. Acad. Sci. USA 2004, 101, 4425–4430. [Google Scholar] [CrossRef] [PubMed]
- Ago, H.; Adachi, T.; Yoshida, A.; Yamamoto, M.; Habuka, N.; Yatsunami, K.; Miyano, M. Crystal structure of the RNA-dependent RNA polymerase of hepatitis C virus. Structure 1999, 7, 1417–1426. [Google Scholar] [CrossRef]
- Lesburg, C.A.; Cable, M.B.; Ferrari, E.; Hong, Z.; Mannarino, A.F.; Weber, P.C. Crystal structure of the RNA-dependent RNA polymerase from hepatitis C virus reveals a fully encircled active site. Nat. Struct. Biol. 1999, 6, 937–943. [Google Scholar] [CrossRef] [PubMed]
- O’Farrell, D.; Trowbridge, R.; Rowlands, D.; Jäger, J. Substrate complexes of hepatitis C virus RNA polymerase (HC-J4): Structural evidence for nucleotide import and De-novo initiation. J. Mol. Biol. 2003, 326, 1025–1035. [Google Scholar] [CrossRef]
- Mastrangelo, E.; Pezzullo, M.; Tarantino, D.; Petazzi, R.; Germani, F.; Kramer, D.; Robel, I.; Rohayem, J.; Bolognesi, M.; Milani, M. Structure-based inhibition of norovirus RNA-dependent RNA polymerases. J. Mol. Biol. 2012, 419, 198–210. [Google Scholar] [CrossRef] [PubMed]
- Croci, R.; Pezzullo, M.; Tarantino, D.; Milani, M.; Tsay, S.C.; Sureshbabu, R.; Tsai, Y.J.; Mastrangelo, E.; Rohayem, J.; Bolognesi, M.; et al. STructural bases of norovirus RNA dependent RNA polymerase inhibition by novel suramin-related compounds. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Fullerton, S.W.B.; Blaschke, M.; Coutard, B.; Gebhardt, J.; Gorbalenya, A.; Canard, B.; Tucker, P.A.; Rohayem, J. Structural and functional characterization of sapovirus RNA-dependent RNA polymerase. J. Virol. 2007, 81, 1858–1871. [Google Scholar] [CrossRef] [PubMed]
- Ng, K.K.S.; Cherney, M.M.; Vázquez, A.L.; Machín, Á.; Martín Alonso, J.M.; Parra, F.; James, M.N.G. Crystal structures of active and inactive conformations of a caliciviral RNA-dependent RNA polymerase. J. Biol. Chem. 2002, 277, 1381–1387. [Google Scholar] [CrossRef] [PubMed]
- Butcher, S.J.; Grimes, J.M.; Makeyev, E.V.; Bamford, D.H.; Stuart, D.I. A mechanism for initiating RNA-dependent RNA polymerization. Nature 2001, 410, 235–240. [Google Scholar] [CrossRef] [PubMed]
- Huiskonen, J.T.; de Haas, F.; Bubeck, D.; Bamford, D.H.; Fuller, S.D.; Butcher, S.J. Structure of the Bacteriophage φ{symbol}6 Nucleocapsid Suggests a Mechanism for Sequential RNA Packaging. Structure 2006, 14, 1039–1048. [Google Scholar] [CrossRef] [PubMed]
- Ren, Z.; Franklin, M.C.; Ghose, R. Structure of the RNA-directed RNA Polymerase from the cystovirus φ12. Proteins Struct. Funct. Bioinform. 2013, 81, 1479–1484. [Google Scholar] [CrossRef] [PubMed]
- Garriga, D.; Navarro, A.; Querol-Audí, J.; Abaitua, F.; Rodríguez, J.F.; Verdaguer, N. Activation mechanism of a noncanonical RNA-dependent RNA polymerase. Proc. Natl. Acad. Sci. USA 2007, 104, 20540–20545. [Google Scholar] [CrossRef] [PubMed]
- Collier, A.M.; Lyytinen, O.L.; Guo, Y.R.; Toh, Y.; Poranen, M.M.; Tao, Y.J. Initiation of RNA Polymerization and Polymerase Encapsidation by a Small dsRNA Virus. PLoS Pathog. 2016, 12. [Google Scholar] [CrossRef] [PubMed]
- McDonald, S.; Block, A.; Beaucourt, S.; Moratorio, G.; Vignuzzi, M.; Peersen, O.B. Design of a genetically stable high fidelity coxsackievirus B3 polymerase that attenuates virus growth in vivo. J. Biol. Chem. 2016, 291, 13999–14011. [Google Scholar] [CrossRef] [PubMed]
- Jacome, R.; Becerra, A.; De Leon, S.P.; Lazcano, A. Structural analysis of monomeric RNA-dependent polymerases: Evolutionary and therapeutic implications. PLoS ONE 2015, 10. [Google Scholar] [CrossRef] [PubMed]
- Lukarska, M.; Fournier, G.; Pflug, A.; Resa-Infante, P.; Reich, S.; Naffakh, N.; Cusack, S. Structural basis of an essential interaction between influenza polymerase and Pol II CTD. Nature 2017, 541, 117–121. [Google Scholar] [CrossRef] [PubMed]
- Reich, S.; Guilligay, D.; Pflug, A.; Malet, H.; Berger, I.; Crepin, T.; Hart, D.; Lunardi, T.; Nanao, M.; Ruigrok, R.W.H.; et al. Structural insight into cap-snatching and RNA synthesis by influenza polymerase. Nature 2014, 516, 361–366. [Google Scholar] [CrossRef] [PubMed]
- Hengrung, N.; El Omari, K.; Serna Martin, I.; Vreede, F.T.; Cusack, S.; Rambo, R.P.; Vonrhein, C.; Bricogne, G.; Stuart, D.I.; Grimes, J.M.; et al. Crystal structure of the RNA-dependent RNA polymerase from influenza C virus. Nature 2015, 527, 114–117. [Google Scholar] [CrossRef] [PubMed]
- Garriga, D.; Ferrer-Orta, C.; Querol-Audí, J.; Oliva, B.; Verdaguer, N. Role of motif B loop in allosteric regulation of RNA-dependent RNA polymerization activity. J. Mol. Biol. 2013, 425, 2279–2287. [Google Scholar] [CrossRef] [PubMed]
- Sankar, S.; Porter, A.G. Point mutations which drastically affect the polymerization activity of encephalomyocarditis virus RNA-dependent RNA polymerase correspond to the active site of Escherichia coli DNA polymerase I. J. Biol. Chem. 1992, 267, 10168–10176. [Google Scholar] [PubMed]
- Vazquez, A.L.; Alonso, J.M.M.; Parra, F. Mutation analysis of the GDD sequence motif of a calicivirus RNA-dependent RNA polymerase. J. Virol. 2000, 74, 3888–3891. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Xiao, M.; Chen, J.; Zhang, W.; Luo, J.; Bao, K.; Nie, M.; Li, B. Mutational analysis of the GDD sequence motif of classical swine fever virus RNA-dependent RNA polymerases. Virus Genes 2007, 34, 63–65. [Google Scholar] [CrossRef] [PubMed]
- Stevaert, A.; Naesens, L. The influenza virus polymerase complex: an update on its structure, functions, and significance for antiviral drug design. Med. Res. Rev. 2016, 36, 1127–1173. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Smidansky, E.D.; Maksimchuk, K.R.; Lum, D.; Welch, J.L.; Arnold, J.J.; Cameron, C.E.; Boehr, D.D. Motif D of viral RNA-dependent RNA polymerases determines efficiency and fidelity of nucleotide addition. Structure 2012, 20, 1519–1527. [Google Scholar] [CrossRef] [PubMed]
- Lang, D.M.; Zemla, A.T.; Ecale Zhou, C.L. Highly similar structural frames link the template tunnel and NTP entry tunnel to the exterior surface in RNA-dependent RNA polymerases. Nucleic Acids Res. 2013, 41, 1464–1482. [Google Scholar] [CrossRef] [PubMed]
- Castro, C.; Smidansky, E.D.; Arnold, J.J.; Maksimchuk, K.R.; Moustafa, I.; Uchida, A.; Götte, M.; Konigsberg, W.; Cameron, C.E. Nucleic acid polymerases use a general acid for nucleotidyl transfer. Nat. Struct. Mol. Biol. 2009, 16, 212–218. [Google Scholar] [CrossRef] [PubMed]
- Campagnola, G.; McDonald, S.; Beaucourt, S.; Vignuzzi, M.; Peersen, O.B. Structure-function relationships underlying the replication fidelity of viral RNA-dependent RNA polymerases. J. Virol. 2015, 89, 275–286. [Google Scholar] [CrossRef] [PubMed]
- Jacobo-Molina, A.; Ding, J.; Nanni, R.G.; Clark, A.D.; Lu, X.; Tantillo, C.; Williams, R.L.; Kamer, G.; Ferris, A.L.; Clark, P. Crystal structure of human immunodeficiency virus type 1 reverse transcriptase complexed with double-stranded DNA at 3.0 A resolution shows bent DNA. Proc. Natl. Acad. Sci. USA 1993, 90, 6320–6324. [Google Scholar] [CrossRef] [PubMed]
- Gerlach, P.; Malet, H.; Cusack, S.; Reguera, J. Structural insights into bunyavirus replication and its regulation by the vRNA promoter. Cell 2015, 161, 1267–1279. [Google Scholar] [CrossRef] [PubMed]
- Reguera, J.; Gerlach, P.; Cusack, S. Towards a structural understanding of RNA synthesis by negative strand RNA viral polymerases. Curr. Opin. Struct. Biol. 2016, 36, 75–84. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; McDonald, S.M.; Tortorici, M.A.; Tao, Y.J.; Vasquez-Del Carpio, R.; Nibert, M.L.; Patton, J.T.; Harrison, S.C. Mechanism for coordinated RNA packaging and genome replication by rotavirus polymerase VP1. Structure 2008, 16, 1678–1688. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhou, N.; Chen, W.; Zhu, B.; Wang, X.; Xu, B.; Wang, J.; Liu, H.; Cheng, L. Near-Atomic Resolution Structure Determination of a Cypovirus Capsid and Polymerase Complex Using Cryo-EM at 200 kV. J. Mol. Biol. 2017, 429, 79–87. [Google Scholar] [CrossRef] [PubMed]
- Chu, C.; Fan, S.; Li, C.; Macken, C.; Kim, J.H.; Hatta, M.; Neumann, G.; Kawaoka, Y. Functional analysis of conserved motifs in influenza virus PB1 protein. PLoS ONE 2012, 7. [Google Scholar] [CrossRef] [PubMed]
- Stubbs, T.M.; Te Velthuis, A.J. The RNA-dependent RNA polymerase of the influenza A virus. Futur. Virol. 2014, 9, 863–876. [Google Scholar] [CrossRef] [PubMed]
- Van der Linden, L.; Vives-Adrian, L.; Selisko, B.; Ferrer-Orta, C.; Liu, X.; Lanke, K.; Ulferts, R.; De Palma, A.M.; Tanchis, F.; Goris, N.; et al. The RNA Template channel of the RNA-dependent RNA polymerase as a target for development of antiviral therapy of multiple genera within a virus family. PLoS Pathog. 2015, 11, 1–23. [Google Scholar] [CrossRef] [PubMed]
- McDonald, S.M.; Tao, Y.J.; Patton, J.T. The ins and outs of four-tunneled Reoviridae RNA-dependent RNA polymerases. Curr. Opin. Struct. Biol. 2009, 19, 775–782. [Google Scholar] [CrossRef] [PubMed]
- Gruez, A.; Selisko, B.; Roberts, M.; Bricogne, G.; Bussetta, C.; Jabafi, I.; Coutard, B.; De Palma, A.M.; Neyts, J.; Canard, B. The crystal structure of coxsackievirus B3 RNA-dependent RNA polymerase in complex with its protein primer VPg confirms the existence of a second VPg binding site on Picornaviridae polymerases. J. Virol. 2008, 82, 9577–9590. [Google Scholar] [CrossRef] [PubMed]
- Ogden, K.M.; Ramanathan, H.N.; Patton, J.T. Mutational analysis of residues involved in nucleotide and divalent cation stabilization in the rotavirus RNA-dependent RNA polymerase catalytic pocket. Virology 2012, 431, 12–20. [Google Scholar] [CrossRef] [PubMed]
- Gytz, H.; Mohr, D.; Seweryn, P.; Yoshimura, Y.; Kutlubaeva, Z.; Dolman, F.; Chelchessa, B.; Chetverin, A.B.; Mulder, F.A.A.; Brodersen, D.E.; et al. Structural basis for RNA-genome recognition during bacteriophage Qβ replication. Nucleic Acids Res. 2015, 43, 10893–10906. [Google Scholar] [CrossRef] [PubMed]
- Yap, T.L.; Xu, T.; Chen, Y.-L.; Malet, H.; Egloff, M.-P.; Canard, B.; Vasudevan, S.G.; Lescar, J. Crystal structure of the dengue virus RNA-dependent RNA polymerase catalytic domain at 1.85-Angstrom resolution. J. Virol. 2007, 81, 4753–4765. [Google Scholar] [CrossRef] [PubMed]
- Noble, C.G.; Lim, S.P.; Chen, Y.-L.; Liew, C.W.; Yap, L.; Lescar, J.; Shi, P.-Y. Conformational flexibility of the dengue virus RNA-dependent RNA polymerase revealed by a complex with an inhibitor. J. Virol. 2013, 87, 5291–5295. [Google Scholar] [CrossRef] [PubMed]
- Noble, C.G.; Chen, Y.L.; Dong, H.; Gu, F.; Lim, S.P.; Schul, W.; Wang, Q.Y.; Shi, P.Y. Strategies for development of dengue virus inhibitors. Antivir. Res. 2010, 85, 450–462. [Google Scholar] [CrossRef] [PubMed]
- El Sahili, A.; Lescar, J. Dengue Virus Non-Structural Protein 5. Viruses 2017, 9, 91. [Google Scholar] [CrossRef] [PubMed]
- Upadhyay, A.K.; Cyr, M.; Longenecker, K.; Tripathi, R.; Sun, C.; Kempf, D.J. Crystal structure of full-length Zika virus NS5 protein reveals a conformation similar to Japanese encephalitis virus NS5. Acta Crystallogr. Sect. F Struct. Biol. Commun. 2017, 73, 116–122. [Google Scholar] [CrossRef] [PubMed]
- Godoy, A.S.; Lima, G.M.A.; Oliveira, K.I.Z.; Torres, N.U.; Maluf, F.V.; Guido, R.V.C.; Oliva, G. Crystal structure of Zika virus NS5 RNA-dependent RNA polymerase. Nat. Commun. 2017, 9, 156–170. [Google Scholar] [CrossRef] [PubMed]
- Saiz, J.-C.; Martín-Acebes, M.A. Zika virus: A race in search for antivirals. Antimicrob. Agents Chemother. 2017, 61, e00411-17. [Google Scholar] [CrossRef] [PubMed]
- Malet, H.; Egloff, M.-P.; Selisko, B.; Butcher, R.E.; Wright, P.J.; Roberts, M.; Gruez, A.; Sulzenbacher, G.; Vonrhein, C.; Bricogne, G.; et al. Crystal structure of the RNA polymerase domain of the West Nile virus non-structural protein 5. J. Biol. Chem. 2007, 282, 10678–10689. [Google Scholar] [CrossRef] [PubMed]
- De Clercq, E. Antivirals: Past, present and future. Biochem. Pharmacol. 2013, 85, 727–744. [Google Scholar] [CrossRef] [PubMed]
- Gemma, S.; Brogi, S.; Novellino, E.; Campiani, G.; Maga, G.; Brindisi, M.; Butini, S. HCV-targeted antivirals: Current status and future challenges. Curr. Pharm. Des. 2014, 20, 3445–3464. [Google Scholar] [CrossRef] [PubMed]
- Laurila, M.R.L.; Makeyev, E.V.; Bamford, D.H. Bacteriophage φ6 RNA-dependent RNA polymerase. Molecular details of initiating nucleic acid synthesis without primer. J. Biol. Chem. 2002, 277, 17117–17124. [Google Scholar] [CrossRef] [PubMed]
- Hansen, J.L.; Long, A.M.; Schultz, S.C. Structure of the RNA-dependent RNA polymerase of poliovirus. Structure 1997, 5, 1109–1122. [Google Scholar] [CrossRef]
- Gohara, D.W.; Arnold, J.J.; Cameron, C.E. Poliovirus RNA-dependent RNA polymerase (3Dpol): Kinetic, thermodynamic, and structural analysis of ribonucleotide selection. Biochemistry 2004, 43, 5149–5158. [Google Scholar] [CrossRef] [PubMed]
- Love, R.A.; Maegley, K.A.; Yu, X.; Ferre, R.A.; Lingardo, L.K.; Diehl, W.; Parge, H.E.; Dragovich, P.S.; Fuhrman, S.A. The crystal structure of the RNA-dependent RNA polymerase from human rhinovirus: A dual function target for common cold antiviral therapy. Structure 2004, 12, 1533–1544. [Google Scholar] [CrossRef] [PubMed]
- Ferrer-Orta, C.; Arias, A.; Perez-Luque, R.; Escarmís, C.; Domingo, E.; Verdaguer, N. Structure of foot-and-mouth disease virus RNA-dependent RNA polymerase and its complex with a template-primer RNA. J. Biol. Chem. 2004, 279, 47212–47221. [Google Scholar] [CrossRef] [PubMed]
- Vives-Adrian, L.; Lujan, C.; Oliva, B.; van der Linden, L.; Selisko, B.; Coutard, B.; Canard, B.; van Kuppeveld, F.J.M.; Ferrer-Orta, C.; Verdaguer, N. The crystal structure of a cardiovirus RNA-dependent RNA polymerase reveals an unusual conformation of the polymerase active site. J. Virol. 2014, 88, 5595–5607. [Google Scholar] [CrossRef] [PubMed]
- Shu, B.; Gong, P. Structural basis of viral RNA-dependent RNA polymerase catalysis and translocation. Proc. Natl. Acad. Sci. USA 2016, 113, E4005–E4014. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Sheng, J.; Fokine, A.; Meng, G.; Shin, W.H.; Long, F.; Kuhn, R.J.; Kihara, D.; Rossmann, M.G. Structure and inhibition of EV-D68, a virus that causes respiratory illness in children. Science 2015, 347, 71–74. [Google Scholar] [CrossRef] [PubMed]
- Kranzusch, P.J.; Schenk, A.D.; Rahmeh, A.A.; Radoshitzky, S.R.; Bavari, S.; Walz, T.; Whelan, S.P.J. Assembly of a functional Machupo virus polymerase complex. Proc. Natl. Acad. Sci. USA 2010, 107, 20069–20074. [Google Scholar] [CrossRef] [PubMed]
- Liang, B.; Li, Z.; Jenni, S.; Rahmeh, A.A.; Morin, B.M.; Grant, T.; Grigorieff, N.; Harrison, S.C.; Whelan, S.P.J. Structure of the L protein of vesicular stomatitis virus from electron cryomicroscopy. Cell 2015, 162, 314–327. [Google Scholar] [CrossRef] [PubMed]
- Trask, S.D.; McDonald, S.M.; Patton, J.T. Structural insights into the coupling of virion assembly and rotavirus replication. Nat. Rev. Microbiol. 2012, 10, 165–177. [Google Scholar] [CrossRef] [PubMed]
- Takeshita, D.; Yamashita, S.; Tomita, K. Molecular insights into replication initiation by Qβ replicase using ribosomal protein S1. Nucleic Acids Res. 2014, 42, 10809–10822. [Google Scholar] [CrossRef] [PubMed]
- Takeshita, D.; Tomita, K. Molecular basis for RNA polymerization by Qβ replicase. Nat. Struct. Mol. Biol. 2012, 19, 229–237. [Google Scholar] [CrossRef] [PubMed]
- Padmanabhan, R.; Takhampunya, R.; Teramoto, T.; Choi, K.H. Flavivirus RNA synthesis in vitro. Methods 2015, 91, 20–34. [Google Scholar] [CrossRef] [PubMed]
- Klema, V.J.; Ye, M.; Hindupur, A.; Teramoto, T.; Gottipati, K.; Padmanabhan, R.; Choi, K.H. Dengue virus nonstructural protein 5 (NS5) assembles into a dimer with a unique methyltransferase and polymerase interface. PLoS Pathog. 2016, 12. [Google Scholar] [CrossRef] [PubMed]
- Green, T.J. Structure of the vesicular stomatitis virus nucleoprotein-RNA complex. Science 2006, 313, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Baltimore, D.; Huang, A.S.; Stampfer, M. ribonucleic acid synthesis of vesicular stomatitis virus. II. An RNA polymerase in the virion. Proc. Natl. Acad. Sci. USA 1970, 66, 572–576. [Google Scholar] [CrossRef] [PubMed]
- Koonin, E.V.; Dolja, V.V. Expanding networks of RNA virus evolution. BMC Biol. 2012, 10, 54. [Google Scholar] [CrossRef] [PubMed]
- Krupovič, M.; Bamford, D.H. Does the evolution of viral polymerases reflect the origin and evolution of viruses? Nat. Rev. Microbiol. 2009, 7, 250. [Google Scholar] [CrossRef] [PubMed]
- Koonin, E.V.; Dolja, V.V.; Krupovic, M. Origins and evolution of viruses of eukaryotes: The ultimate modularity. Virology 2015, 479–480, 2–25. [Google Scholar] [CrossRef] [PubMed]
- Zemla, A.; Geisbrecht, B.; Smith, J.; Lam, M.; Kirkpatrick, B.; Wagner, M.; Slezak, T.; Zhou, C.E. STRALCP—Structure alignment-based clustering of proteins. Nucleic Acids Res. 2007, 35. [Google Scholar] [CrossRef] [PubMed]
- Papageorgiou, L.; Loukatou, S.; Sofia, K.; Maroulis, D.; Vlachakis, D. An updated evolutionary study of Flaviviridae NS3 helicase and NS5 RNA-dependent RNA polymerase reveals novel invariable motifs as potential pharmacological targets. Mol. BioSyst. 2016, 12, 2080–2093. [Google Scholar] [CrossRef] [PubMed]
- Vlachakis, D.; Koumandou, V.L.; Kossida, S. A holistic evolutionary and structural study of Flaviviridae provides insights into the function and inhibition of HCV helicase. PeerJ 2013, 1, e74. [Google Scholar] [CrossRef] [PubMed]
- Ferrero, D.S.; Buxaderas, M.; Rodríguez, J.F.; Verdaguer, N. The structure of the RNA-dependent RNA polymerase of a permutotetravirus suggests a link between primer-dependent and primer-independent polymerases. PLoS Pathog. 2015, 11. [Google Scholar] [CrossRef] [PubMed]
- Shaik, M.M.; Bhattacharjee, N.; Feliks, M.; Ng, K.K.; Field, M.J. Norovirus RNA-dependent RNA polymerase: A computational study of metal-binding preferences. Proteins Struct. Funct. Bioinform. 2017, 85, 1435–1445. [Google Scholar] [CrossRef] [PubMed]
- Wright, S.; Poranen, M.M.; Bamford, D.H.; Stuart, D.I.; Grimes, J.M. Noncatalytic ions direct the RNA-dependent RNA polymerase of bacterial double-Stranded RNA virus 6 from de novo initiation to elongation. J. Virol. 2012, 86, 2837–2849. [Google Scholar] [CrossRef] [PubMed]
- Guglielmi, K.M.; McDonald, S.M.; Patton, J.T. Mechanism of intraparticle synthesis of the rotavirus double-stranded RNA genome. J. Biol. Chem. 2010, 285, 18123–18128. [Google Scholar] [CrossRef] [PubMed]
- Makeyev, E.V.; Grimes, J.M. RNA-dependent RNA polymerases of dsRNA bacteriophages. Virus Res. 2004, 101, 45–55. [Google Scholar] [CrossRef] [PubMed]
- Dreher, T.W. Role of tRNA-like structures in controlling plant virus replication. Virus Res. 2009, 139, 217–229. [Google Scholar] [CrossRef] [PubMed]
- Zamyatkin, D.F.; Parra, F.; Machín, Á.; Grochulski, P.; Ng, K.K.S. Binding of 2-Amino-2′-Deoxycytidine-5′-Triphosphate to Norovirus Polymerase Induces Rearrangement of the Active Site. J. Mol. Biol. 2009, 390, 10–16. [Google Scholar] [CrossRef] [PubMed]
- Zamyatkin, D.; Rao, C.; Hoffarth, E.; Jurca, G.; Rho, H.; Parra, F.; Grochulski, P.; Ng, K.K.S. Structure of a backtracked state reveals conformational changes similar to the state following nucleotide incorporation in human norovirus polymerase. Acta Crystallogr. Sect. D Biol. Crystallogr. 2014, 70, 3099–3109. [Google Scholar] [CrossRef] [PubMed]
- Zamyatkin, D.F.; Parra, F.; Alonso, J.M.M.; Harki, D.A.; Peterson, B.R.; Grochulski, P.; Ng, K.K.-S. Structural insights into mechanisms of catalysis and inhibition in Norwalk virus polymerase. J. Biol. Chem. 2008, 283, 7705–7712. [Google Scholar] [CrossRef] [PubMed]
- Gong, P.; Kortus, M.G.; Nix, J.C.; Davis, R.E.; Peersen, O.B. Structures of Coxsackievirus, Rhinovirus, and Poliovirus Polymerase Elongation Complexes Solved by Engineering RNA Mediated Crystal Contacts. PLoS ONE 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Thompson, A.A.; Peersen, O.B. Structural basis for proteolysis-dependent activation of the poliovirus RNA-dependent RNA polymerase. EMBO J. 2004, 23, 3462–3471. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Wang, C.; Li, Q.; Wang, Z.; Xie, W. Crystal structure and thermostability characterization of enterovirus D68 3D pol. J. Virol. 2017, 91, e00876-17. [Google Scholar] [CrossRef] [PubMed]
- Ferrer-Orta, C.; Arias, A.; Agudo, R.; Pérez-Luque, R.; Escarmís, C.; Domingo, E.; Verdaguer, N. The structure of a protein primer-polymerase complex in the initiation of genome replication. EMBO J. 2006, 25, 880–888. [Google Scholar] [CrossRef] [PubMed]
- De Clercq, E.; Li, G. Approved antiviral drugs over the past 50 years. Clin. Microbiol. Rev. 2016, 29, 695–747. [Google Scholar] [CrossRef] [PubMed]
- Eltahla, A.A.; Luciani, F.; White, P.A.; Lloyd, A.R.; Bull, R.A. Inhibitors of the hepatitis C virus polymerase; mode of action and resistance. Viruses 2015, 7, 5206–5224. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.P.; Noble, C.G.; Shi, P.-Y. The dengue virus NS5 protein as a target for drug discovery. Antivir. Res. 2015, 119, 57–67. [Google Scholar] [CrossRef] [PubMed]
- Sofia, M.J.; Chang, W.; Furman, P.A.; Mosley, R.T.; Ross, B.S. Nucleoside, nucleotide, and non-nucleoside inhibitors of hepatitis C virus NS5B RNA-dependent RNA-polymerase. J. Med. Chem. 2012, 55, 2481–2531. [Google Scholar] [CrossRef] [PubMed]
- Appleby, T.C.; Perry, J.K.; Murakami, E.; Barauskas, O.; Feng, J.; Cho, A.; Fox, D.; Wetmore, D.R.; Mcgrath, M.E.; Ray, A.S.; et al. Structural basis for RNA replication by the hepatitis C virus polymerase. Science 2015, 347, 771–775. [Google Scholar] [CrossRef] [PubMed]
- Haudecoeur, R.; Peuchmaur, M.; Ahmed-Belkacem, A.; Pawlotsky, J.-M.; Boumendjel, A. Structure-activity relationships in the development of allosteric hepatitis C virus RNA-dependent RNA polymerase inhibitors: Ten years of research. Med. Res. Rev. 2013, 33, 934–984. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Johnson, K.A. Thumb site 2 inhibitors of hepatitis C viral RNA-dependent RNA polymerase allosterically block the transition from initiation to elongation. J. Biol. Chem. 2016, 291, 10067–10077. [Google Scholar] [CrossRef] [PubMed]
- Yokokawa, F.; Nilar, S.; Noble, C.G.; Lim, S.P.; Rao, R.; Tania, S.; Wang, G.; Lee, G.; Hunziker, J.; Karuna, R.; et al. Discovery of potent non-nucleoside inhibitors of dengue viral RNA-dependent RNA polymerase from a fragment hit using structure-based drug design. J. Med. Chem. 2016, 59, 3935–3952. [Google Scholar] [CrossRef] [PubMed]


| S. No. | Category | PROSITE ID | Total Reviewed Sequences/Exemplar UniProt Accession No. | No. of Unique structures/Exemplar Protein Data Bank (PDB) IDs |
|---|---|---|---|---|
| 1 | Bacteriophages (Group III and IV) | PS50522 | 07 | 03 |
| Leviviridae: | ||||
| Bacteriophage Qβ (Qβ) | P14647 | 3mmp (2.5 Å) | ||
| Cystoviridae: | ||||
| Pseudomonas phage φ6 (φ6) | P11124 | 1hhs (2.0 Å) | ||
| Pseudomonas phage φ12 (φ12) | Q94M06 | 4gzk (1.69 Å) | ||
| 2 | Reoviridae (Group III) | PS50523 | 45 | 03 |
| Mammalian orthoreovirus 3 (MRV3) | P0CK31 | 1n35 (2.5 Å) | ||
| Simian rotavirus SA11 (SiRV) | O37061 | 2r7r (2.6 Å) | ||
| Bombyx mori Cytoplasmic polyhedrosis virus (BmCPV) | A0A0S1LIW6 | 5h0r (3.9 Å) | ||
| 3 | Birnaviridae (Group III) | PS50524 | 09 | 02 |
| Infectious bursal disease virus (IBDV) | Q9Q6Q5 | 2pus (2.4 Å) | ||
| Infectious pancreatic necrosis virus (IPNV) | P22173 | 2yi9 (2.2 Å) | ||
| 4 | Picobirnaviridae (Group III) | 01 | 01 | |
| Picobirnaviridae | ||||
| Human picobirnavirus (hPBV) | Q50LE4 | 5i61 (2.4 Å) | ||
| 5 | Group IV viruses | PS50507 | 470 | 18 |
| Permutetraviridae: | ||||
| Thosea asigna virus (TAV) | Q6A562 | 4xhi (2.15 Å) | ||
| Picornaviridae: | ||||
| Coxsackievirus B3 (CVB3) | Q5UEA2 | 4zpc (1.59 Å) | ||
| Human rhinovirus 16 (HRV) | Q82122 | 1xr7 (2.3 Å) | ||
| Poliovirus type 1 (PV) | P03300 | 1ra6 (2.0 Å) | ||
| Foot-and-mouth disease virus (FMDV) | Q9QCE4 | 1u09 (1.91 Å) | ||
| Encephalomyocarditis virus 1 (EMCV) | P12296 | 4nyz (2.15 Å) | ||
| Enterovirus A71 (EV71) | E5RPG2 | 5f8n (2.48 Å) | ||
| Enterovirus D68 (EVD68) | F1T146 | 5xe0 (2.3 Å) | ||
| Caliciviridae: | ||||
| Murine Norovirus (mNoV) | Q80J95 | 3uqs (2.0 Å) | ||
| Human Norovirus (hNoV) | A0ZNP5 | 4nrt (2.02 Å) | ||
| Sapporo virus (SV) | Q69014 | 2uut (2.4 Å) | ||
| Rabbit hemorrhagic disease virus (RHDV) | P27410 | 1khw (2.7 Å) | ||
| Flaviviridae: | ||||
| Hepatitis C virus (HCV) | O92972 | 1nb4 (2.0 Å) | ||
| Bovine viral diarrhea virus (BVDV) | Q96662 | 2cjq (2.6 Å) | ||
| West nile virus (WNV) | P14335 | 2hcn (2.35 Å) | ||
| Dengue virus (DENV) | Q6YMS4 | 4hhj (1.79 Å) | ||
| Zika virus (ZIKV) | A0A109PRQ3 | 5wz3 (1.8 Å) | ||
| Japanese encephalitis virus (JEV) | P27395 | 4k6m (2.6 Å) | ||
| 6 | Group V viruses (segmented genome) | PS50525 | 142 | 04 |
| Orthomyxoviridae: | ||||
| Influenza A virus (FluA) | H6QM91 | 5m3h (2.5 Å) | ||
| Influenza B virus (FluB) | Q5V8Y6 | 4wrt (2.7 Å) | ||
| Influenza C virus (FluC) | Q9IMP4 | 5d98 (3.9 Å) | ||
| Bunyaviridae: | ||||
| La crosse virus (LACV) | A5HC98 | 5amq (3.0 Å) | ||
| 7 | Group V viruses (non-segmented genomes) | PS50526 | 81 | 01 |
| Rhabdoviridae: | ||||
| Vesicular stomatitis virus (VSV) | P03523 | 5a22 (3.8 Å) | ||
| Group | Family | Structures with Ligands | RdRps Complexed with nucleoside triphosphate (NTP) and Their Derivatives | PDB IDs of RdRp-NTP Complexes | RdRps Complexed with Inhibitors |
|---|---|---|---|---|---|
| dsRNA viruses | Reoviridae and Birnaviridae | 5 | 2 | 1MWH,1N35 | 0 |
| Bacteriophages | Cystoviridae and Leviviridae | 9 | 2 | 3AVX, 1UVK | 0 |
| (+) ssRNA Viruses | Caliciviridae | 6 | 4 | 3B5N, 3BS0, 3H5X, 3H5Y | 1 |
| (+) ssRNA Viruses | Flaviviridae | 133 | 5 | 2XI3, 1GX6, 1GX5, 4HDH, 4HDG | 120 |
| (+) ssRNA Viruses | Picornaviridae | 40 | 7 | 1RA7, 2ILZ, 2IM0, 2IM1, 2IM2, 3OLB, 5F8I | 0 |
| (−) ssRNA Viruses | Ortho-myxoviridae, Rhabdoviridae | 5 | 0 | NA | 0 |
| Binding Sites | PDB IDs | Total No. of Complexes |
|---|---|---|
| Palm Site Inhibitor Complexes | 3CDE, 3CWJ, 2YOJ, 3BR9, 3BSA, 3CO9, 3CVK, 3D28, 3D5M, 3E51, 3G86, 3GYN, 3H2L, 3H59, 3H5S, 3H5U, 3H98, 3HKW, 3HKY, 3LKH, 3SKA, 3SKE, 3SKH, 3TYQ, 3TYV, 3U4O, 3U4R, 3UPH, 3UPI, 4EAW, 4IH5, 4IH6, 4IH7, 4KAI, 4KB7, 4KBI, 4KE5, 4KHM, 4KHR, 4MIB, 4MK8, 4MK9, 4MKA, 4MKB, 4MZ4, 5PZK, 5PZL, 5PZN, 3FQK, 3FQL, 4JY0, 2GIQ, 2AWZ, 2AX0, 2AX1 | 55 |
| Thumb Site Inhibitor Complexes | 2D3U, 2D3Z, 2D41, 2HWH, 2HWI, 1NHU, 2DXS, 2O5D, 2WHO, 3CIZ, 3CJ0, 3CJ2, 3CJ3, 3CJ4, 3CJ5, 3FRZ, 3MF5, 3Q0Z, 4DRU, 4EO6, 4EO8, 4IZ0, 4J02, 4J04, 4J06, 4J08, 4J0A, 4JJS, 4JJU, 4JU3, 4JU4, 4JU6, 4JU7, 4JVQ, 4OBC, 4TLR, 2BRK, 2BRL, 2HAI, 2I1R, 2WRM, 3HVO, 2WCX, 2GIR | 44 |
| Primer Grip Inhibitor Complex | 2IJN, 5TWM,1YVF | 3 |
| At interfaces of Subdomains | 2GC8, 3GNW, 3QGF, 3QGH, 3QGI, 5TRI, 5TRK | 7 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Venkataraman, S.; Prasad, B.V.L.S.; Selvarajan, R. RNA Dependent RNA Polymerases: Insights from Structure, Function and Evolution. Viruses 2018, 10, 76. https://doi.org/10.3390/v10020076
Venkataraman S, Prasad BVLS, Selvarajan R. RNA Dependent RNA Polymerases: Insights from Structure, Function and Evolution. Viruses. 2018; 10(2):76. https://doi.org/10.3390/v10020076
Chicago/Turabian StyleVenkataraman, Sangita, Burra V. L. S. Prasad, and Ramasamy Selvarajan. 2018. "RNA Dependent RNA Polymerases: Insights from Structure, Function and Evolution" Viruses 10, no. 2: 76. https://doi.org/10.3390/v10020076
APA StyleVenkataraman, S., Prasad, B. V. L. S., & Selvarajan, R. (2018). RNA Dependent RNA Polymerases: Insights from Structure, Function and Evolution. Viruses, 10(2), 76. https://doi.org/10.3390/v10020076





