More and More Coronaviruses: Human Coronavirus HKU1
Abstract
:1. Introduction
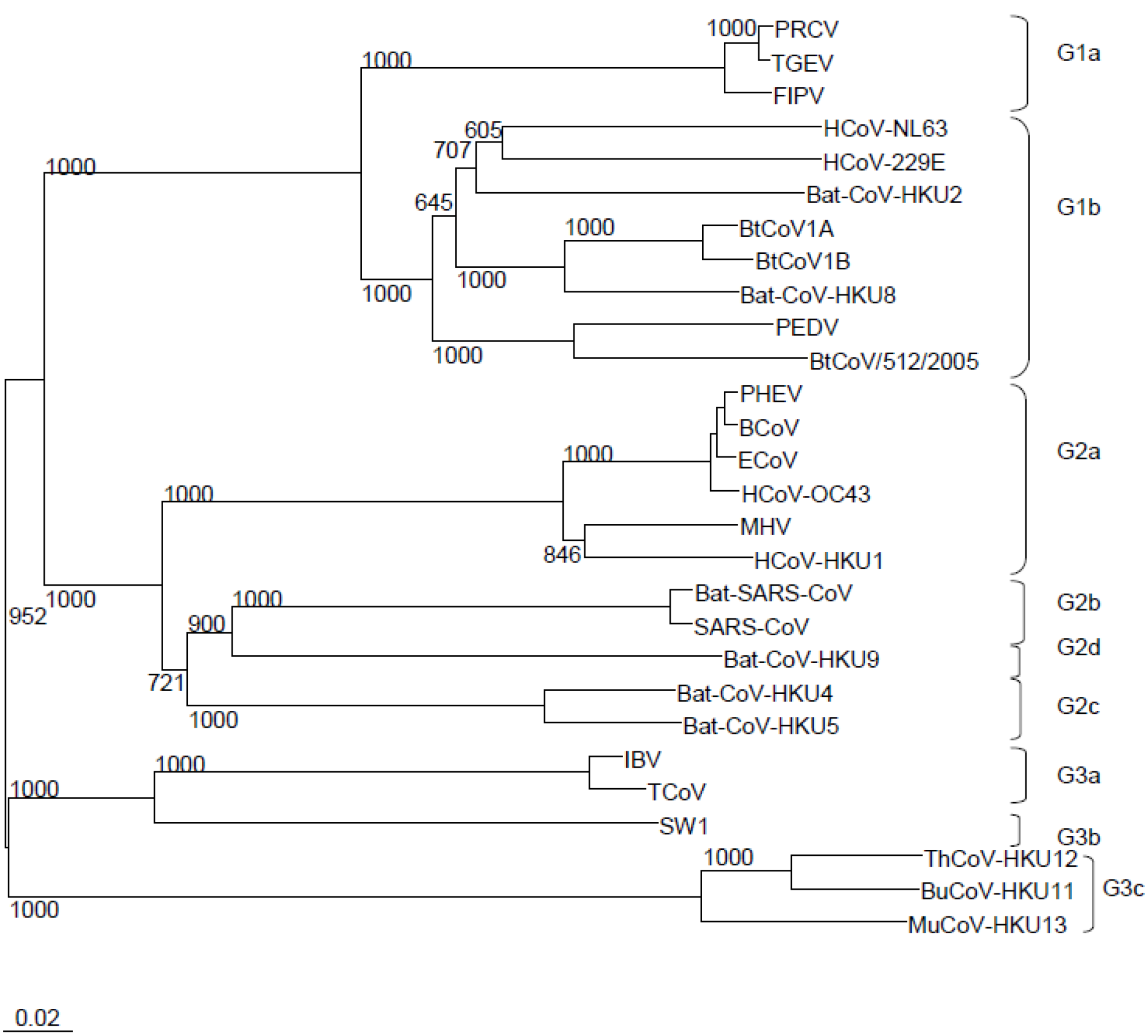
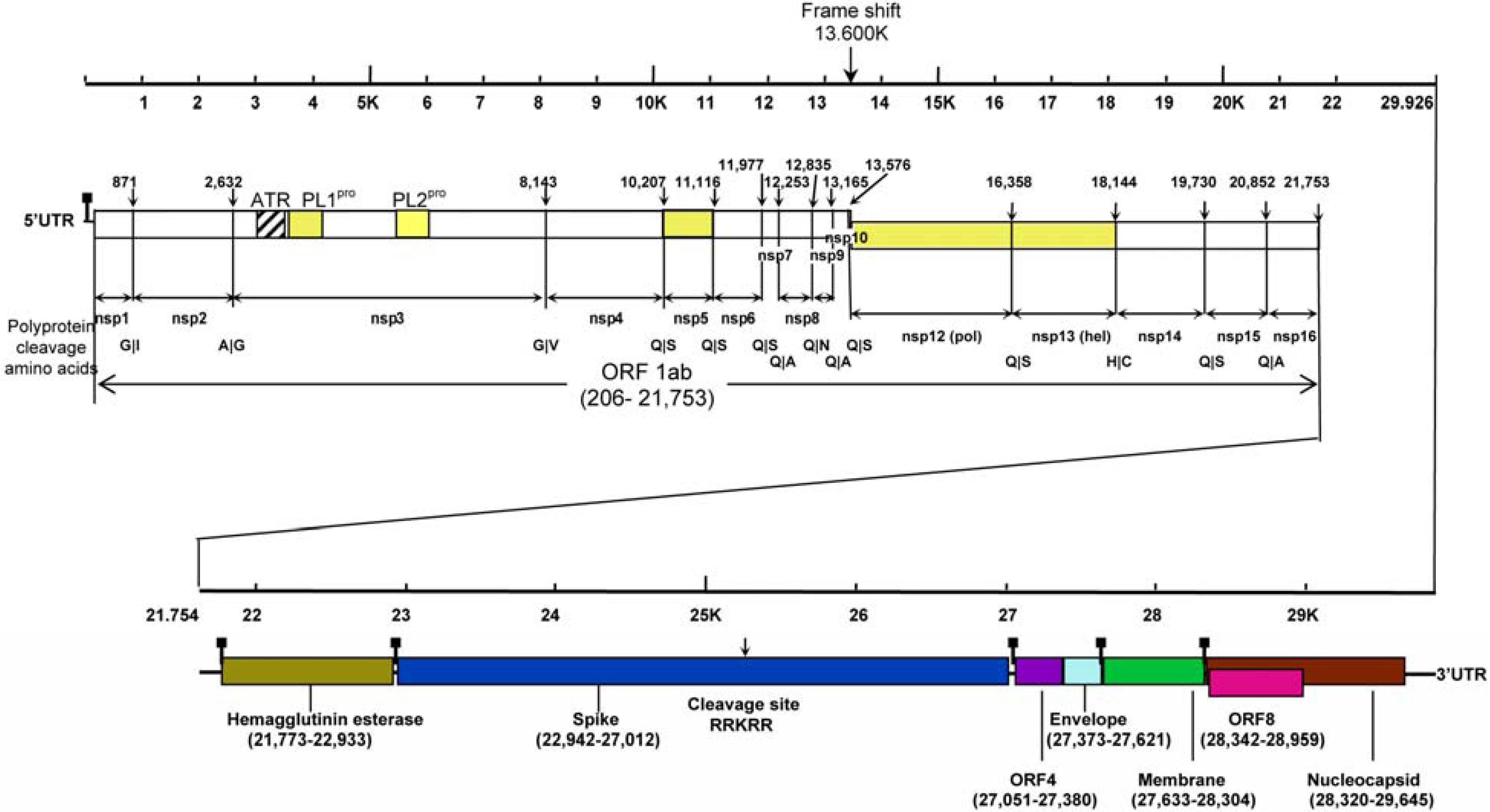
2 .Classification and Virology
3. Epidemiology
4. Clinical diseases
5. Laboratory Diagnosis
6. Concluding Remarks
Acknowledgments
References and Notes
- Bosis, S.; Esposito, S.; Niesters, H.G.; Zuccotti, G.V.; Marseglia, G.; Lanari, M.; Zuin, G.; Pelucchi, C.; Osterhaus, A.D.; Principi, N. Role of respiratory pathogens in infants hospitalized for a first episode of wheezing and their impact on recurrences. Clin. Microbiol. Infect. 2008, 14, 677–684. [Google Scholar] [CrossRef] [PubMed]
- Brian, D.A.; Baric, R.S. Coronavirus genome structure and replication. Curr. Top. Microbiol. Immunol. 2005, 287, 1–30. [Google Scholar] [PubMed]
- Canducci, F.; Debiaggi, M.; Sampaolo, M.; Marinozzi, M.C.; Berre, S.; Terulla, C.; Gargantini, G.; Cambieri, P.; Romero, E.; Clementi, M. Two-year prospective study of single infections and co-infections by respiratory syncytial virus and viruses identified recently in infants with acute respiratory disease. J. Med. Virol. 2008, 80, 716–723. [Google Scholar] [CrossRef] [PubMed]
- Chan, C.M.; Lau, S.K.; Woo, P.C.; Tse, H.; Zheng, B.J.; Chen, L.; Huang, J.D.; Yuen, K.Y. Identification of major histocompatibility complex class I C molecule as an attachment factor that facilitates coronavirus HKU1 spike-mediated infection. J. Virol. 2009, 83, 1026–1035. [Google Scholar] [CrossRef] [PubMed]
- Chan, C.M.; Tse, H.; Wong, S.S.; Woo, P.C.; Lau, S.K.; Chen, L.; Zheng, B.J.; Huang, J.D.; Yuen, K.Y. Examination of seroprevalence of coronavirus HKU1 infection with S protein-based ELISA and neutralization assay against viral spike pseudotyped virus. J. Clin. Virol. 2009, 45, 54–60. [Google Scholar] [CrossRef] [PubMed]
- Chan, C.M.; Woo, P.C.; Lau, S.K.; Tse, H.; Chen, H.L.; Li, F.; Zheng, B.J.; Chen, L.; Huang, J.D.; Yuen, K.Y. Spike protein, S, of human coronavirus HKU1: role in viral life cycle and application in antibody detection. Exp. Biol. Med. (Maywood) 2008, 233, 1527–1536. [Google Scholar] [CrossRef] [PubMed]
- Chu, D.K.; Peiris, J.S.; Chen, H.; Guan, Y.; Poon, L.L. Genomic characterizations of bat coronaviruses (1A, 1B and HKU8) and evidence for co-infections in Miniopterus bats. J. Gen. Virol. 2008, 89, 1282–1287. [Google Scholar] [CrossRef] [PubMed]
- Chung, J.Y.; Han, T.H.; Kim, S.W.; Kim, C.K.; Hwang, E.S. Detection of viruses identified recently in children with acute wheezing. J. Med. Virol. 2007, 79, 1238–1243. [Google Scholar] [CrossRef] [PubMed]
- Dare, R.K.; Fry, A.M.; Chittaganpitch, M.; Sawanpanyalert, P.; Olsen, S.J.; Erdman, D.D. Human coronavirus infections in rural Thailand: a comprehensive study using real-time reverse-transcription polymerase chain reaction assays. J. Infect. Dis. 2007, 196, 1321–1328. [Google Scholar] [CrossRef] [PubMed]
- de Souza Luna, L.K.; Heiser, V.; Regamey, N.; Panning, M.; Drexler, J.F.; Mulangu, S.; Poon, L.; Baumgarte, S.; Haijema, B.J.; Kaiser, L.; Drosten, C. Generic detection of coronaviruses and differentiation at the prototype strain level by reverse transcription-PCR and nonfluorescent low-density microarray. J. Clin. Microbiol. 2007, 45, 1049–1052. [Google Scholar] [CrossRef] [PubMed]
- Esper, F.; Weibel, C.; Ferguson, D.; Landry, M.L.; Kahn, J.S. Coronavirus HKU1 infection in the United States. Emerg. Infect. Dis. 2006, 12, 775–779. [Google Scholar] [PubMed]
- Esposito, S.; Bosis, S.; Niesters, H.G.; Tremolati, E.; Begliatti, E.; Rognoni, A.; Tagliabue, C.; Principi, N.; Osterhaus, A.D. Impact of human coronavirus infections in otherwise healthy children who attended an emergency department. J. Med. Virol. 2006, 78, 1609–1615. [Google Scholar] [CrossRef] [PubMed]
- Gagneur, A.; Dirson, E.; Audebert, S.; Vallet, S.; Legrand-Quillien, M.C.; Laurent, Y.; Collet, M.; Sizun, J.; Oger, E.; Payan, C. Materno-fetal transmission of human coronaviruses: a prospective pilot study. Eur. J. Clin. Microbiol. Infect. Dis. 2008, 27, 863–866. [Google Scholar] [CrossRef] [PubMed]
- Garbino, J.; Crespo, S.; Aubert, J.D.; Rochat, T.; Ninet, B.; Deffernez, C.; Wunderli, W.; Pache, J.C.; Soccal, P.M.; Kaiser, L. A prospective hospital-based study of the clinical impact of non-severe acute respiratory syndrome (Non-SARS)-related human coronavirus infection. Clin. Infect. Dis. 2006, 43, 1009–1015. [Google Scholar] [CrossRef] [PubMed]
- Gerna, G.; Percivalle, E.; Sarasini, A.; Campanini, G.; Piralla, A.; Rovida, F.; Genini, E.; Marchi, A.; Baldanti, F. Human respiratory coronavirus HKU1 versus other coronavirus infections in Italian hospitalised patients. J. Clin. Virol. 2007, 38, 244–250. [Google Scholar] [CrossRef] [PubMed]
- Gomaa, M.H.; Barta, J.R.; Ojkic, D.; Yoo, D. Complete genomic sequence of turkey coronavirus. Virus Res. 2008, 135, 237–246. [Google Scholar] [CrossRef] [PubMed]
- Hamre, D.; Procknow, J.J. A new virus isolated from the human respiratory tract. Proc. Soc. Exp. Biol. Med. 1966, 121, 190–193. [Google Scholar] [PubMed]
- Herrewegh, A.A.; Smeenk, I.; Horzinek, M.C.; Rottier, P.J.; de Groot, R.J. Feline coronavirus type II strains 79-1683 and 79-1146 originate from a double recombination between feline coronavirus type I and canine coronavirus. J. Virol. 1998, 72, 4508–4514. [Google Scholar] [PubMed]
- Huang, Y.; Lau, S.K.; Woo, P.C.; Yuen, K.Y. CoVDB: a comprehensive database for comparative analysis of coronavirus genes and genomes. Nucleic Acids Res. 2008, 36, D504–D511. [Google Scholar] [CrossRef] [PubMed]
- Kaplan, N.M.; Dove, W.; Abd-Eldayem, S.A.; Abu-Zeid, A.F.; Shamoon, H.E.; Hart, C. Molecular epidemiology and disease severity of respiratory syncytial virus in relation to other potential pathogens in children hospitalized with acute respiratory infection in Jordan. J. Med. Virol. 2008, 80, 168–174. [Google Scholar] [CrossRef] [PubMed]
- Kuypers, J.; Martin, E.T.; Heugel, J.; Wright, N.; Morrow, R.; Englund, J. A. Clinical disease in children associated with newly described coronavirus subtypes. Pediatrics 2007, 119, e70–e76. [Google Scholar] [CrossRef] [PubMed]
- Lai, M.M.; Cavanagh, D. The molecular biology of coronaviruses. Adv. Virus Res. 1997, 48, 1–100. [Google Scholar] [PubMed]
- Lau, S.K.; Woo, P.C.; Li, K.S.; Huang, Y.; Tsoi, H.W.; Wong, B.H.; Wong, S.S.; Leung, S.Y.; Chan, K.H.; Yuen, K.Y. Severe acute respiratory syndrome coronavirus-like virus in Chinese horseshoe bats. Proc. Natl. Acad. Sci. USA 2005, 102, 14040–14045. [Google Scholar] [CrossRef]
- Lau, S.K.; Woo, P.C.; Li, K. S.; Huang, Y.; Wang, M.; Lam, C.S.; Xu, H.; Guo, R.; Chan, K.H.; Zheng, B.J.; Yuen, K.Y. Complete genome sequence of bat coronavirus HKU2 from Chinese horseshoe bats revealed a much smaller spike gene with a different evolutionary lineage from the rest of the genome. Virology 2007, 367, 428–439. [Google Scholar] [CrossRef] [PubMed]
- Lau, S.K.; Woo, P.C.; Yip, C.C.; Tse, H.; Tsoi, H.W.; Cheng, V.C.; Lee, P.; Tang, B.S.; Cheung, C.H.; Lee, R.A.; So, L.Y.; Lau, Y.L.; Chan, K.H.; Yuen, K.Y. Coronavirus HKU1 and other coronavirus infections in Hong Kong. J. Clin. Microbiol. 2006, 44, 2063–2071. [Google Scholar] [CrossRef] [PubMed]
- Lehmann, C.; Wolf, H.; Xu, J.; Zhao, Q.; Shao, Y.; Motz, M.; Lindner, P. A line immunoassay utilizing recombinant nucleocapsid proteins for detection of antibodies to human coronaviruses. Diagn. Microbiol. Infect. Dis. 2008, 61, 40–88. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Shi, Z.; Yu, M.; Ren, W.; Smith, C.; Epstein, J.H.; Wang, H.; Crameri, G.; Hu, Z.; Zhang, H.; Zhang, J.; McEachern, J.; Field, H.; Daszak, P.; Eaton, B.T.; Zhang, S.; Wang, L. F. Bats are natural reservoirs of SARS-like coronaviruses. Science 2005, 310, 676–679. [Google Scholar] [CrossRef] [PubMed]
- Mihindukulasuriya, K.A.; Wu, G.; St Leger, J.; Nordhausen, R.W; Wang, D. Identification of a novel coronavirus from a beluga whale by using a panviral microarray. J. Virol. 2008, 82, 5084–5088. [Google Scholar] [CrossRef] [PubMed]
- Pierangeli, A.; Gentile, M.; Di Marco, P.; Pagnotti, P.; Scagnolari, C.; Trombetti, S.; Lo Russo, L.; Tromba, V.; Moretti, C.; Midulla, F.; Antonelli, G. Detection and typing by molecular techniques of respiratory viruses in children hospitalized for acute respiratory infection in Rome, Italy. J. Med. Virol. 2007, 79, 463–468. [Google Scholar] [CrossRef] [PubMed]
- Regamey, N.; Kaiser, L.; Roiha, H.L.; Deffernez, C.; Kuehni, C.E.; Latzin, P.; Aebi, C.; Frey, U. Viral etiology of acute respiratory infections with cough in infancy: a community-based birth cohort study. Pediatr. Infect. Dis. J. 2008, 27, 100–105. [Google Scholar] [PubMed]
- Severance, E.G.; Bossis, I.; Dickerson, F.B.; Stallings, C.R.; Origoni, A.E.; Sullens, A.; Yolken, R .H.; Viscidi, R.P. Development of a nucleocapsid-based human coronavirus immunoassay and estimates of individuals exposed to coronavirus in a U.S. metropolitan population. Clin. Vaccine Immunol. 2008, 15, 1805–1810. [Google Scholar] [CrossRef] [PubMed]
- Sloots, T.P.; McErlean, P.; Speicher, D.J.; Arden, K.E.; Nissen, M.D.; Mackay, I.M. Evidence of human coronavirus HKU1 and human bocavirus in Australian children. J. Clin. Virol. 2006, 35, 99–102. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.C.; Zhang, J.X.; Zhang, S.Y.; Wang, P.; Fan, X.H.; Li, L.F.; Li, G.; Dong, B.Q.; Liu, W.; Cheung, C.L.; Xu, K.M.; Song, W.J.; Vijaykrishna, D.; Poon, L.L.; Peiris, J.S.; Smith, G.J.; Chen, H.; Guan, Y. Prevalence and genetic diversity of coronaviruses in bats from China. J. Virol. 2006, 80, 7481–7490. [Google Scholar] [CrossRef] [PubMed]
- Tyrrell, D.A.; Bynoe, M.L. Cultivation of a Novel Type of Common-Cold Virus in Organ Cultures. Br. Med. J. 1965, 1, 1467–1470. [Google Scholar] [CrossRef] [PubMed]
- Vabret, A.; Dina, J.; Gouarin, S.; Petitjean, J.; Corbet, S.; Freymuth, F. Detection of the new human coronavirus HKU1: a report of 6 cases. Clin. Infect. Dis. 2006, 42, 634–639. [Google Scholar] [CrossRef] [PubMed]
- Vabret, A.; Dina, J.; Gouarin, S.; Petitjean, J.; Tripey, V.; Brouard, J.; Freymuth, F. Human (non-severe acute respiratory syndrome) coronavirus infections in hospitalised children in France. J Paediatr. Child Health 2008, 44, 176–181. [Google Scholar] [CrossRef] [PubMed]
- van der Hoek, L.; Pyrc, K.; Jebbink, M.F.; Vermeulen-Oost, W.; Berkhout, R.J.; Wolthers, K.C.; Wertheim-van Dillen, P. M.; Kaandorp, J.; Spaargaren, J.; Berkhout, B. Identification of a new human coronavirus. Nat. Med. 2004, 10, 368–373. [Google Scholar] [CrossRef] [PubMed]
- Woo, P.C.; Huang, Y.; Lau, S.K.; Tsoi, H.W.; Yuen, K.Y. In silico analysis of ORF1ab in coronavirus HKU1 genome reveals a unique putative cleavage site of coronavirus HKU1 3C-like protease. Microbiol. Immunol. 2005, 49, 899–908. [Google Scholar] [PubMed]
- Woo, P.C.; Lau, S.K.; Chu, C.M.; Chan, K.H.; Tsoi, H.W.; Huang, Y.; Wong, B.H.; Poon, R.W.; Cai, J.J.; Luk, W.K.; Poon, L.L.; Wong, S.S.; Guan, Y.; Peiris, J.S.; Yuen, K.Y. Characterization and complete genome sequence of a novel coronavirus, coronavirus HKU1, from patients with pneumonia. J. Virol. 2005, 79, 884–895. [Google Scholar] [CrossRef] [PubMed]
- Woo, P.C.; Lau, S.K.; Lam, C.S.; Lai, K.K.; Huang, Y.; Lee, P.; Luk, G.S.; Dyrting, K.C.; Chan, K.H.; Yuen, K.Y. Comparative analysis of complete genome sequences of three avian coronaviruses reveals a novel group 3c coronavirus. J. Virol. 2009, 83, 908–917. [Google Scholar] [CrossRef] [PubMed]
- Woo, P.C.; Lau, S.K.; Li, K.S.; Poon, R.W.; Wong, B.H.; Tsoi, H.W.; Yip, B.C.; Huang, Y.; Chan, K.H.; Yuen, K.Y. Molecular diversity of coronaviruses in bats. Virology 2006, 351, 180–187. [Google Scholar] [CrossRef] [PubMed]
- Woo, P.C.; Lau, S.K.; Tsoi, H.W.; Huang, Y.; Poon, R.W.; Chu, C.M.; Lee, R.A.; Luk, W.K.; Wong, G.K.; Wong, B.H.; Cheng, V.C.; Tang, B.S.; Wu, A.K.; Yung, R.W.; Chen, H.; Guan, Y.; Chan, K.H.; Yuen, K.Y. Clinical and molecular epidemiological features of coronavirus HKU1-associated community-acquired pneumonia. J. Infect. Dis. 2005, 192, 1898–1907. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Woo, P.C.; Lau, S.K.; Yip, C.C.; Huang, Y.; Tsoi, H.W.; Chan, K.H.; Yuen, K.Y. Comparative analysis of 22 coronavirus HKU1 genomes reveals a novel genotype and evidence of natural recombination in coronavirus HKU1. J. Virol. 2006, 80, 7136–7145. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Woo, P.C.; Wang, M.; Lau, S.K.; Xu, H.; Poon, R.W.; Guo, R.; Wong, B.H.; Gao, K.; Tsoi, H.W.; Huang, Y.; Li, K.S.; Lam, C.S.; Chan, K.H.; Zheng, B.J.; Yuen, K.Y. Comparative analysis of twelve genomes of three novel group 2c and group 2d coronaviruses reveals unique group and subgroup features. J. Virol. 2007, 81, 1574–1585. [Google Scholar] [CrossRef] [PubMed]
- Woo, P.C.; Wong, B.H.; Huang, Y.; Lau, S.K.; Yuen, K.Y. Cytosine deamination and selection of CpG suppressed clones are the two major independent biological forces that shape codon usage bias in coronaviruses. Virology 2007, 369, 431–442. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Guy, J.S.; Snijder, E.J.; Denniston, D.A.; Timoney, P.J.; Balasuriya, U.B. Genomic characterization of equine coronavirus. Virology 2007, 369, 92–104. [Google Scholar] [CrossRef] [PubMed]
- Ziebuhr, J. Molecular biology of severe acute respiratory syndrome coronavirus. Curr. Opin. Microbiol. 2004, 7, 412–419. [Google Scholar] [CrossRef] [PubMed]
| Studies | Ref. | Place of study | No. of patients/specimens | Patient characteristics | Duration of study | No. (%) of patients/specimens positive for HCoV-HKU1 | No. (%) of HCoV-HKU1 positive patients with underlying diseases | No. (%) of patients/specimens positive for other human coronaviruses | Outcome | ||
|---|---|---|---|---|---|---|---|---|---|---|---|
| HCoV-OC43 | HCoV-229E | HCoV-NL63 | |||||||||
| 1 | [42] | Hong Kong | 418 patients | Hospitalized patients with community-acquired pneumonia | 12 months (Mar 2003 – Mar 2004) | 10 (2.4) | 8 (80) | Not performed | Not performed | Not performed | 2 died |
| 2 | [32] | Australia | 324 specimens | Children presented to Queensland hospitals or general practitioners with acute respiratory tract infections | 4 months (May 2004 – Aug 2004) | 10 (3.1) | Not mentioned | 11 (3.4) | 1 (0.3) | 0 (0) | Not mentioned |
| 3 | [35] | France | 135 patients | Hospitalized patients with respiratory symptoms | 2 months (Feb 2005 – Mar 2005) | 6 (4.4) | 3 (50) | 2 (1.5) | 0 (0) | 2 (1.5) | Not mentioned |
| 4 | [11] | USA | 1048 respiratory specimens from 851 children | Specimens from emergency department,inpatient wards, intensive care units, and hospital-affiliatedprimary care outpatient clinic | 12 months (Dec 2001 – Dec 2002) | 9 (1)patients | 5 (56) | Not performed | Not performed | Not performed | Not mentioned |
| 5 | [25] | Hong Kong | 4181 specimens | Hospitalized patients with acute respiratory tract infections | 12 months (April 2004 – Mar 2005) | 13 (0.3) | 8 (62) | 53 (1.3) | 4 (0.1) | 17 (0.4) | All survived |
| 6 | [14] | Switzerland | 540 specimens from 279 adults | Hospitalized patients with respiratory disease | 20 months | 4 (1.4) patients | Not mentioned | 12 (4.3)patients | 7 (2.5)patients | 6 (2.2)patients | Not mentioned |
| 7 | [12] | Italy | 2060 children | Children who attended the emergency department with an acute disease excluding trauma | 5 months (Nov 2003 – Mar 2004) | 0 (0) | - | 17 (0.8) | 42 (2) | 20 (1) | - |
| 8 | [29] | Italy | 227 children | Hospitalized children with acute respiratory infection or related conditions at the Pediatric Department | 12 months (Oct 2004 – Sep 2005) | 0 (0) | - | 6 (2.6) | 0 (0) | 1 (0.4) | - |
| 9 | [15] | Italy | 685 specimens from 426 patients | Hospitalized patients with acute respiratory tract infections | 7 months (Nov 2005 – May 2006) | 10 (2.3) patients | 8 (80) | 8 (1.9)patients | 20 (4.7)patients | 11 (2.6)patients | Not mentioned |
| 10 | [8] | Korea | 231 children | Hospitalized children with acute expiratory wheezing | 10 months (Feb 2006 – Nov 2006) | 0 (0) | - | 3 (1.3) | 2 (0.9) | 3 (1.3) | - |
| 11 | [9] | Thailand | 734 patients | Patients hospitalized with pneumonia | Study year 1: | Not mentioned | Not mentioned | ||||
| Sep 2003 – Aug 2004 | 3 (0.4) | 31 (4.2) | 3 (0.4) | 7 (1) | |||||||
| 1156 patients | Patients hospitalized with pneumonia | Study year 2: | |||||||||
| Sep 2004 – Oct 2005 | 9 (0.8) | 4 (0.4) | 7 (0.6) | 1 (0.1) | |||||||
| 513 patients | Outpatients with influenza-like illness | Study year 2: | |||||||||
| Sep 2004 – Oct 2005 | 1 (0.2) | 0 (0) | 2 (0.4) | 9 (1.8) | |||||||
| 12 | [21] | USA | 1043 children | Outpatients, emergency department patients and inpatients with acute respiratory illness | 12 months (Oct 2003 – Sep 2004) | 28 (2.7) | Not mentioned | 19 (1.8) | 8 (0.8) | 11 (1.1) | Not mentioned |
| 13 | [30] | Switzerland | 112 infants | Infants followed prospectively during their first year of life for any respiratory or other disease symptoms and their treatment by weekly telephone interviews | 69 months(Apr 1999 – Dec 2004) | 1 (0.9) | Not mentioned | 7 (6.3) | 3 (2.7) | 9 (8) | Not mentioned |
| 14 | [36] | France | 1002 specimens from 928 children | Hospitalized children with respiratory or general symptoms | 9 months (Sep 2004 – May 2005) | 38 (3.8) specimens | 8 (24) of 34 patients | 27 (2.9)specimens | 2 (0.2)specimens | 33 (3.6)specimens | All survived |
| 15 | [20] | Jordan | 325 children | Children with acute respiratory infection admitted to the pediatric wards | 6 months (Dec 2003 – May 2004) | 0 (0) | - | Not performed | Not performed | 4 (1.2) | - |
| 16 | [3] | Italy | 322 infants | Hospitalized infants with acute respiratory disease | 24 months (Oct 2004 – Sep 2006) | 6 (1.9) | Not mentioned | 11 (3.4) | 1 (0.3) | 10 (3.1) | Not mentioned |
| 17 | [1] | Italy | 85 infants | Infants hospitalized for the first acute episode of wheezing | 6 months (Oct 2005 – Mar 2006) | 1 (1.2) | Not mentioned | 2 (2.4) | 0 (0) | 0 (0) | Not mentioned |
| 18 | [13] | France | 159 specimens | Specimens collected from mother-child couples admitted in labor to the Gynecology-Obstetrics Unit | 18 months (Jul 2003 – Jun 2004 and Mar 2005 – Aug 2005) | 1 (0.6) | Not mentioned | 0 (0) | 11 (6.9) | 0 (0) | Not mentioned |
| Studies | References | Place of study | Detection methods | Gene target | Target band size | Primer sequences |
|---|---|---|---|---|---|---|
| 1 | [42] | Hong Kong | RT-PCR | pol | 453 bp | Forward 5’-AAAGGATGTTGACAACCCTGTT-3’ |
| Reverse 5’-ATCATCATACTAAAATGCTTACA-3’ | ||||||
| 2 | [32] | Australia | RT-PCR | pol | 453 bp | Forward 5’-AAAGGATGTTGACAACCCTGTT-3’ |
| Reverse 5’-ATCATCATACTAAAATGCTTACA-3’ | ||||||
| 3 | [35] | France | RT-PCR | N | 443 bp | Forward 5’-ACCAATCTGAGCGAAATTACCAAAC-3’ |
| Reverse 5’-CGGAAACCTAGTAGGGATAGCTT-3’ | ||||||
| 4 | [11] | USA | RT-PCR | pol | 440 bp | Forward 5’-GGTTGGGATTATCCTAAATGTGA-3’ |
| Reverse 5’-CCATCATCACTCAAAATCATCATA-3’ | ||||||
| 5 | [25] | Hong Kong | RT-PCR | pol | 453 bp | Forward 5’-AAAGGATGTTGACAACCCTGTT-3’ |
| Reverse 5’-ATCATCATACTAAAATGCTTACA-3’ | ||||||
| 6 | [14] | Switzerland | Real-time | pol | 506 bp | Forward 5’-GAATTTTGTTGTTCACATGGTGATAGA-3’ |
| RT-PCR | Reverse 5’-GCAACCGCCACACATAACTATTT-3’ | |||||
| Probe 5’-FAM-TTTATCGCCTTGCGAATGAATGTGCTC-TAMRA-3’ | ||||||
| 7 | [12] | Italy | Real-time | N | 64 bp | Forward 5’-AGTTCCCATTGCTTTCGGAGTA-3’ |
| RT-PCR | Reverse 5’-CCGGCTGTGTCTATACCAATATCC-3’ | |||||
| Probe 5’-FAM-CCCCTTCTGAAGCAA-MGB-3’ | ||||||
| 8 | [29] | Italy | RT-PCR | pol | 440 bp | Forward 5’-GGTTGGGACTATCCTAAGTGTGA-3’ |
| Reverse 5’-CCATCATCAGATAGAATCATCATA-3’ | ||||||
| 9 | [15] | Italy | RT-PCR | N | 516 bp | Forward 5’-CAGTGTTTTGGTAAAAGAGGACC-3’ |
| Reverse 5’-TACCACCTAGTGTCGAATTAGG-3’ | ||||||
| pol | 250 bp | Forward 5’-ACTCAAATGAATTTAAAATATGC-3’ | ||||
| Reverse 5’-TCACATTTAGGATAATCCCA-3’ | ||||||
| 10 | [8] | Korea | RT-PCR | pol | 921 bp | Forward 5’-GTTCAAGTGTCGCTGTTCA-3’ |
| Reverse 5’-CTATCATTATCACAATCCACAG-3’ | ||||||
| pol | 989 bp | Forward 5’-GGGTATGAAGTATCATCCTA-3’ | ||||
| Reverse 5’-GATAATCCCAACCCATAAGAAC-3’ | ||||||
| pol | 921 bp | Forward 5’-CATCTTATAAAGGATGTTGAC-3’ | ||||
| Reverse 5’-ACAAACAACACATGCACCTACAC-3’ | ||||||
| 11 | [9] | Thailand | Real-time | pol | 95 bp | Forward 5’-CCTTGCGAATGAATGTGCT-3’ |
| RT-PCR | Reverse 5’-TTGCATCACCACTGCTAGTACCAC-3’ | |||||
| Probe 5’-FAM-TGTGTGGCGGTTGCTATTATGTTAAGCCTG- Black Hole Quencher–1-3’ | ||||||
| 12 | [21] | USA | Real-time | pol | 96 bp | Forward 5’-TGGTGGCTGGGACGATATGT-3’ |
| RT-PCR | Reverse 5’-GGCATAGCACGATCACACTTAGG-3’ | |||||
| Probe 5’-6FAM-ATAATCCCAACCCATRAG-Quencher -3’ | ||||||
| 13 | [30] | Switzerland | Real-time | pol | 506 bp | Forward 5’-GAATTTTGTTGTTCACATGGTGATAGA-3’ |
| RT-PCR | Reverse 5’-GCAACCGCCACACATAACTATTT-3’ | |||||
| Probe 5’-FAM-TTTATCGCCTTGCGAATGAATGTGCTC-TAMRA-3’ | ||||||
| 14 | [36] | France | RT-PCR | N | 439 bp | Forward 5’-ATCTGARCGAAAYYAYCAAAC-3’ |
| Reverse 5’-CGYAAACCTAGTAGGGATAGCTT-3’ | ||||||
| 15 | [20] | Jordan | RT-PCR | pol | 453 bp | Forward 5’-AAAGGATGTTGACAACCCTGTT-3’ |
| Reverse 5’-ATCATCATACTAAAATGCTTACA-3’ | ||||||
| 16 | [3] | Italy | RT-PCR | pol | 220bp | Forward 5’-TTATGGGTTGGGATTATCCYAARTGTGAT-3’ |
| Reverse 5’-GTACTAGCRTCACCAGAAGTYGTACCACC-3’ | ||||||
| pol | 214bp | Forward2 5’-ATGGGATGGGACTATCCTAAGTGTGATAGAG-3’ | ||||
| Reverse2 5’-TTGCATCACCACTRCTAGTRCCACCAGGC-3’ | ||||||
| 17 | [1] | Italy | RT-PCR | pol | 440 bp | Forward 5’-GGTTGGGACTATCCTAAGTGTGA-3’ |
| Reverse 5’-CCATCATCAGATAGAATCATCATA-3’ | ||||||
| 18 | [13] | France | Real-time | pol | 96 bp | Forward 5’-TGGTGGCTGGGACGATATGT-3’ |
| RT-PCR | Reverse 5’-GGCATAGCACGATCACACTTAGG-3’ | |||||
| Probe 5’-6FAM-ATAATCCCAACCCATRAG-Quencher -3’ |
© 2009 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Woo, P.C.Y.; Lau, S.K.P.; Yip, C.C.Y.; Huang, Y.; Yuen, K.-Y. More and More Coronaviruses: Human Coronavirus HKU1. Viruses 2009, 1, 57-71. https://doi.org/10.3390/v1010057
Woo PCY, Lau SKP, Yip CCY, Huang Y, Yuen K-Y. More and More Coronaviruses: Human Coronavirus HKU1. Viruses. 2009; 1(1):57-71. https://doi.org/10.3390/v1010057
Chicago/Turabian StyleWoo, Patrick C. Y., Susanna K. P. Lau, Cyril C. Y. Yip, Yi Huang, and Kwok-Yung Yuen. 2009. "More and More Coronaviruses: Human Coronavirus HKU1" Viruses 1, no. 1: 57-71. https://doi.org/10.3390/v1010057
APA StyleWoo, P. C. Y., Lau, S. K. P., Yip, C. C. Y., Huang, Y., & Yuen, K.-Y. (2009). More and More Coronaviruses: Human Coronavirus HKU1. Viruses, 1(1), 57-71. https://doi.org/10.3390/v1010057





