Nanomaterials-Based Sensing Strategies for Electrochemical Detection of MicroRNAs
Abstract
:1. Introduction
2. Nanomaterials-Based Electrochemical Strategies for miRNAs Detection
2.1. Metallic Oxide
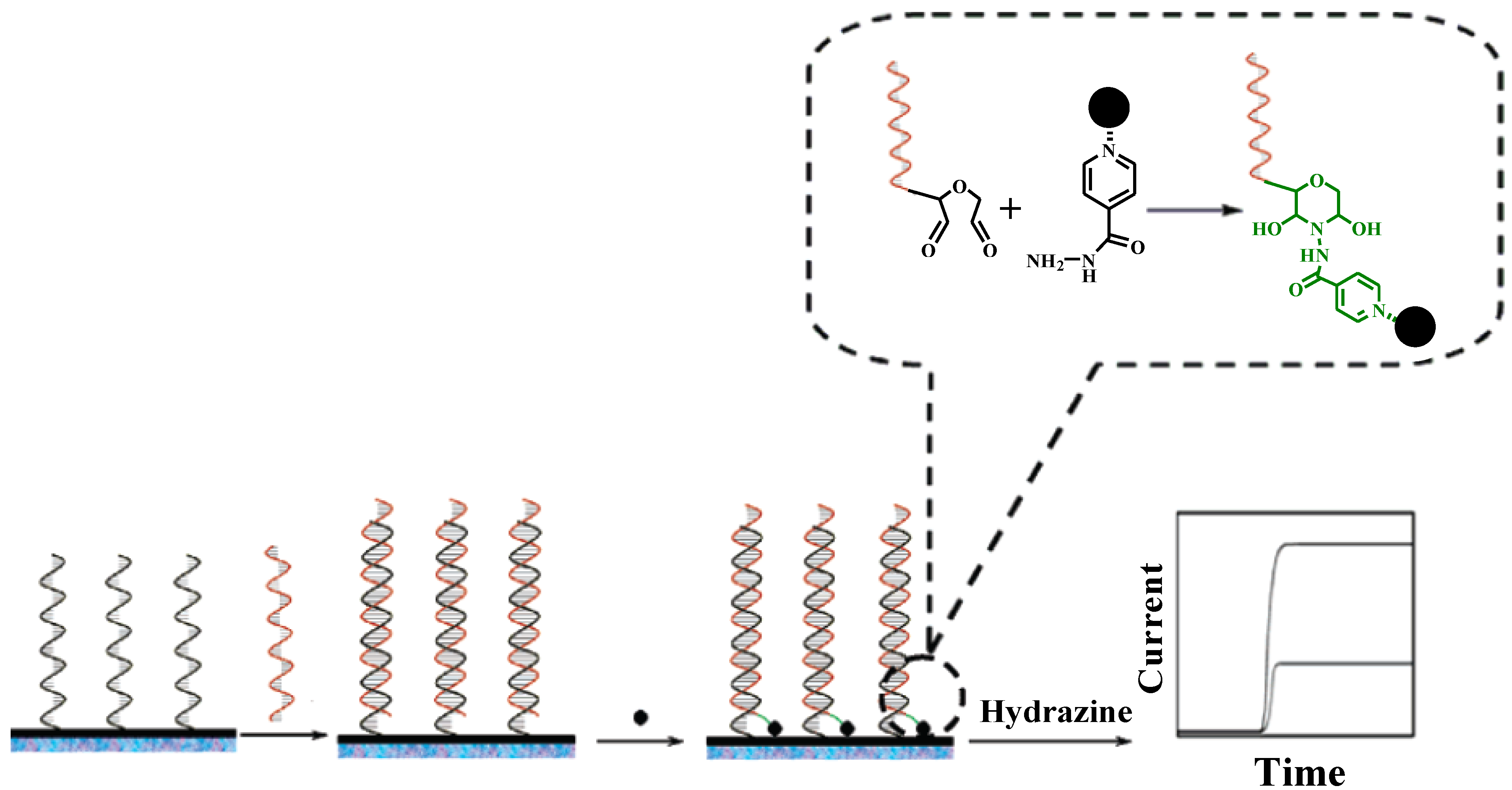
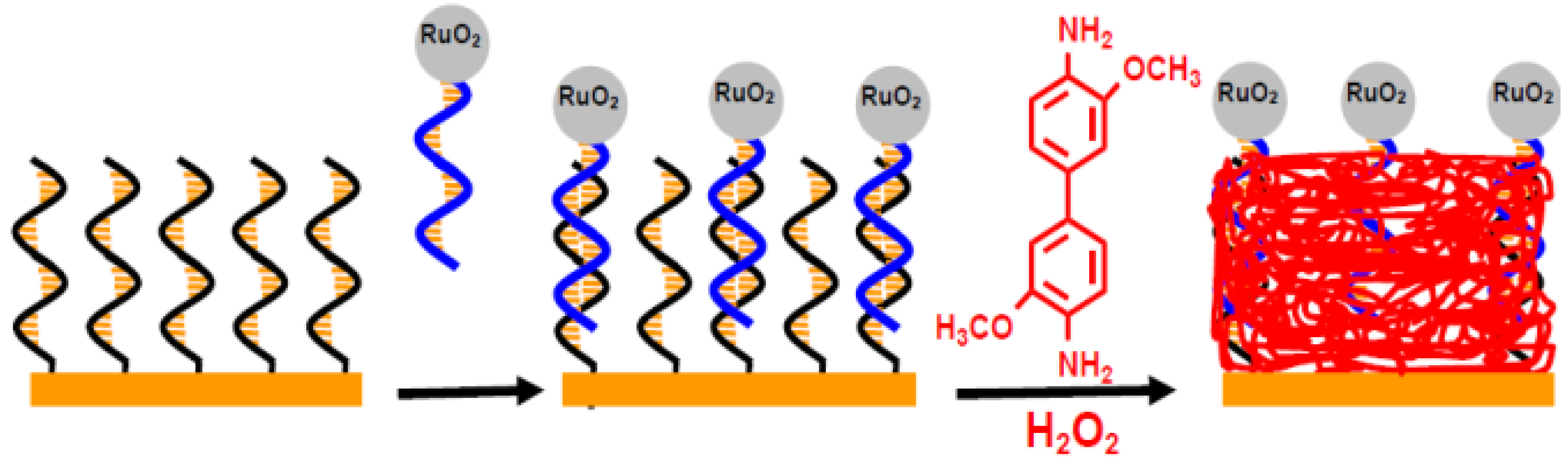
2.2. Gold Nanoparticles
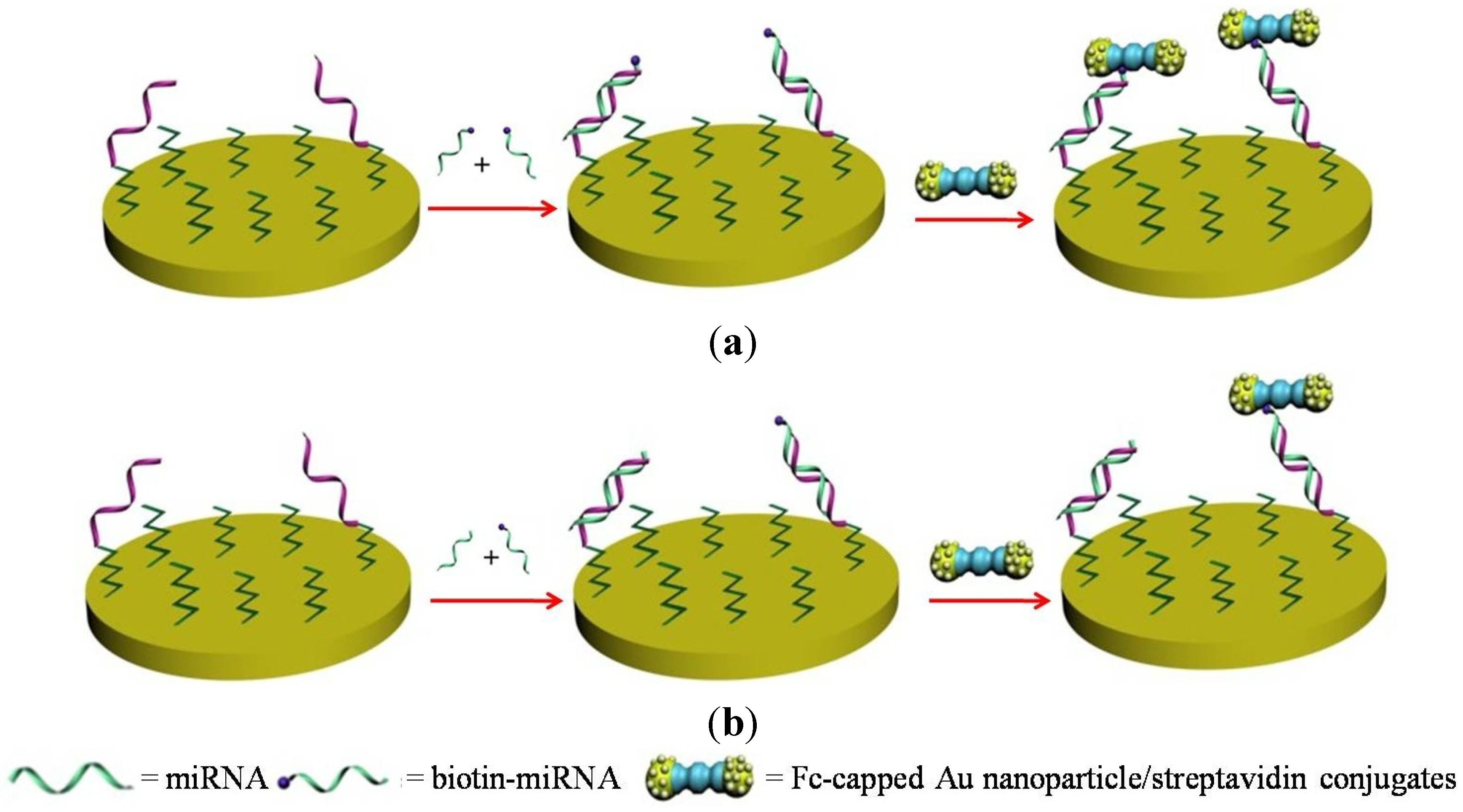

2.3. Silver Nanoclusters
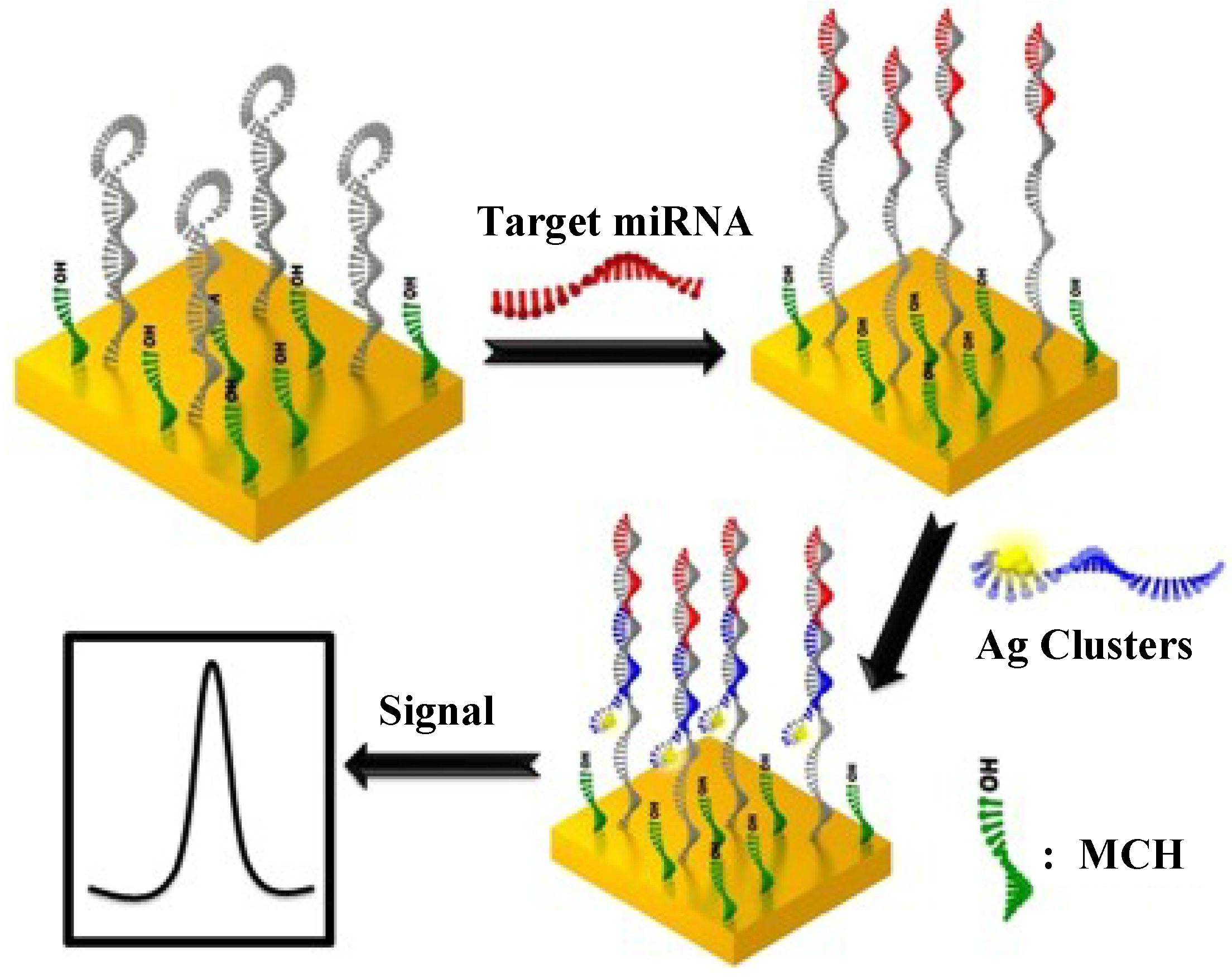
2.4. Carbon Nanomaterials
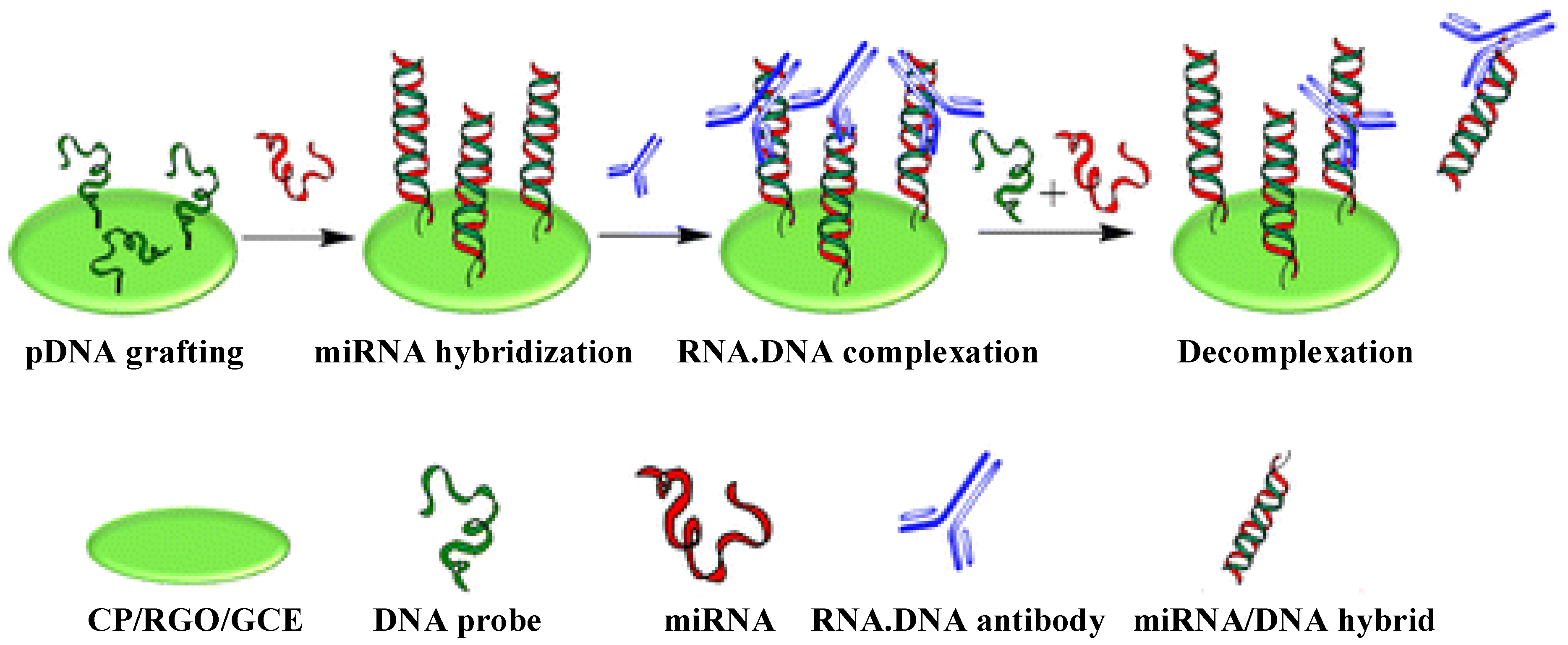
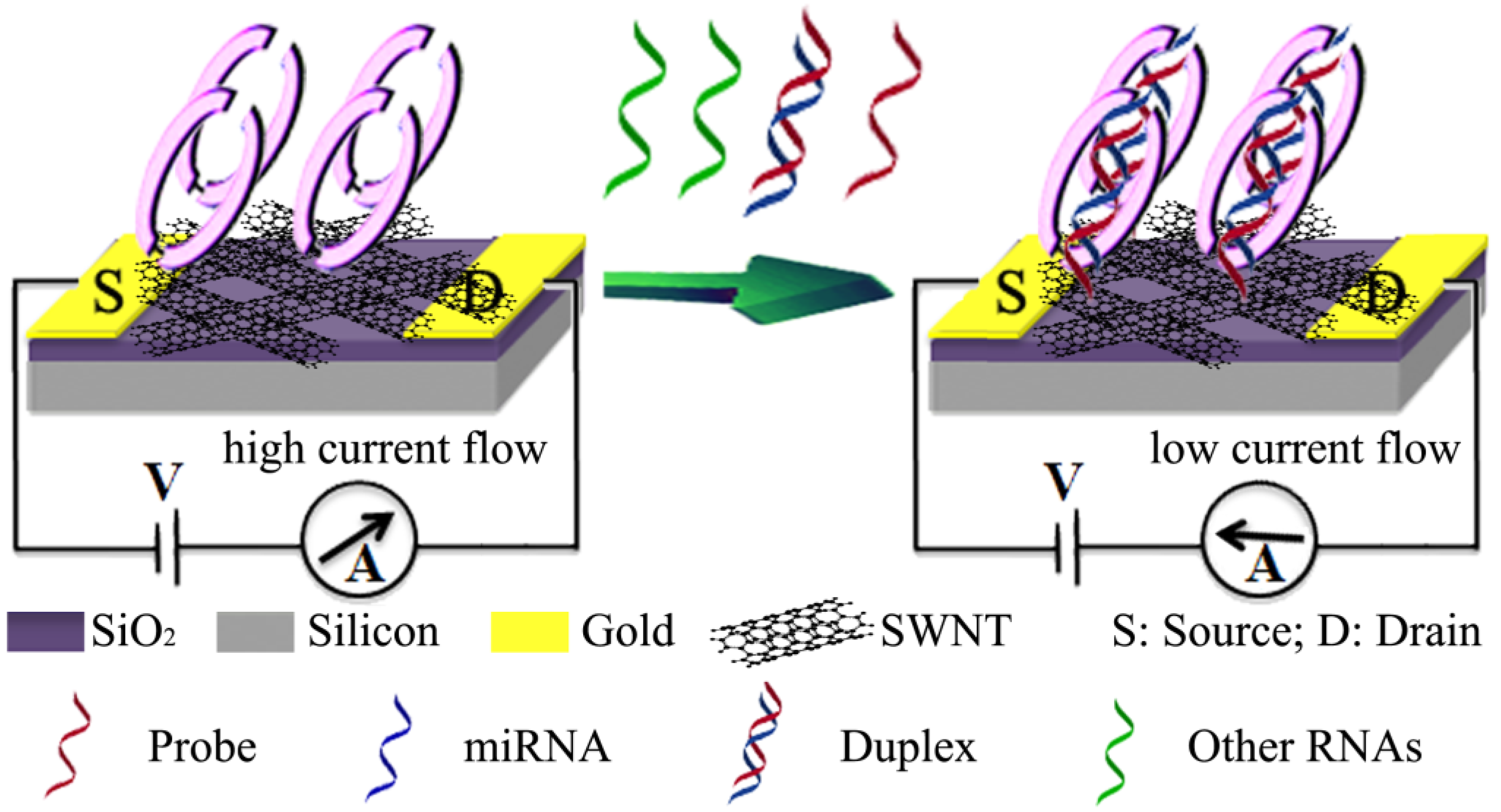
2.5. Quantum Dots
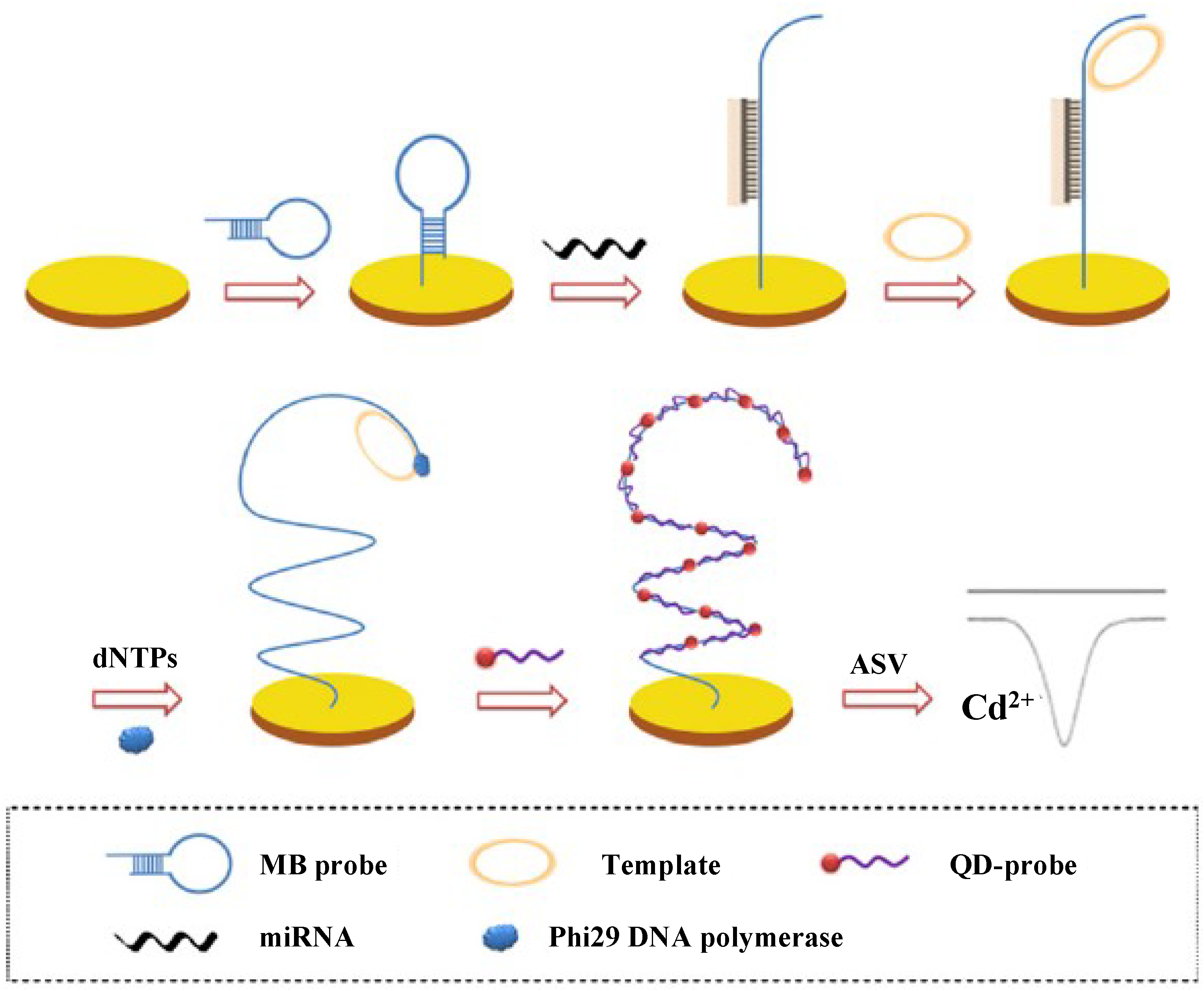
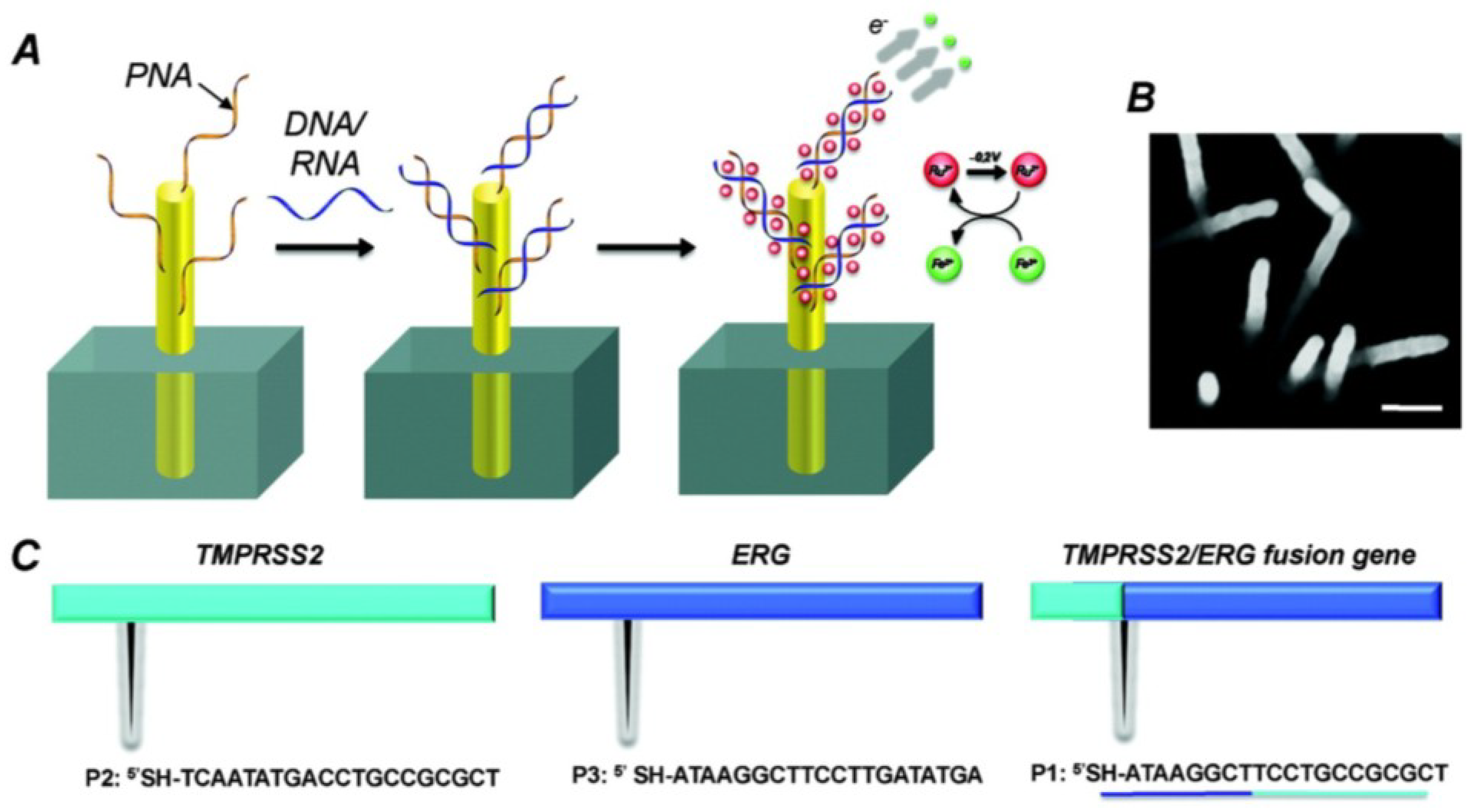
2.6. Nanostructured Electrodes
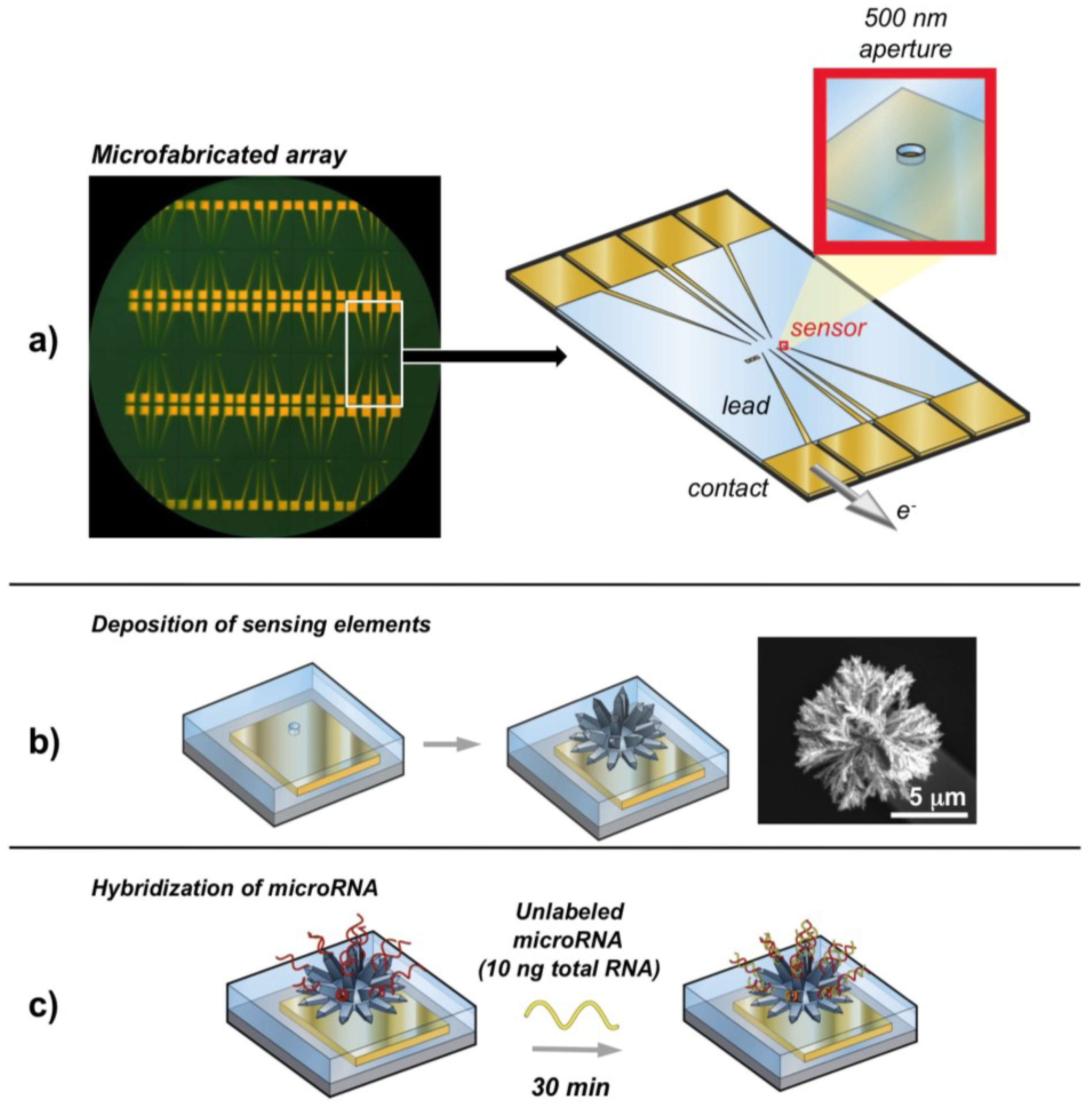
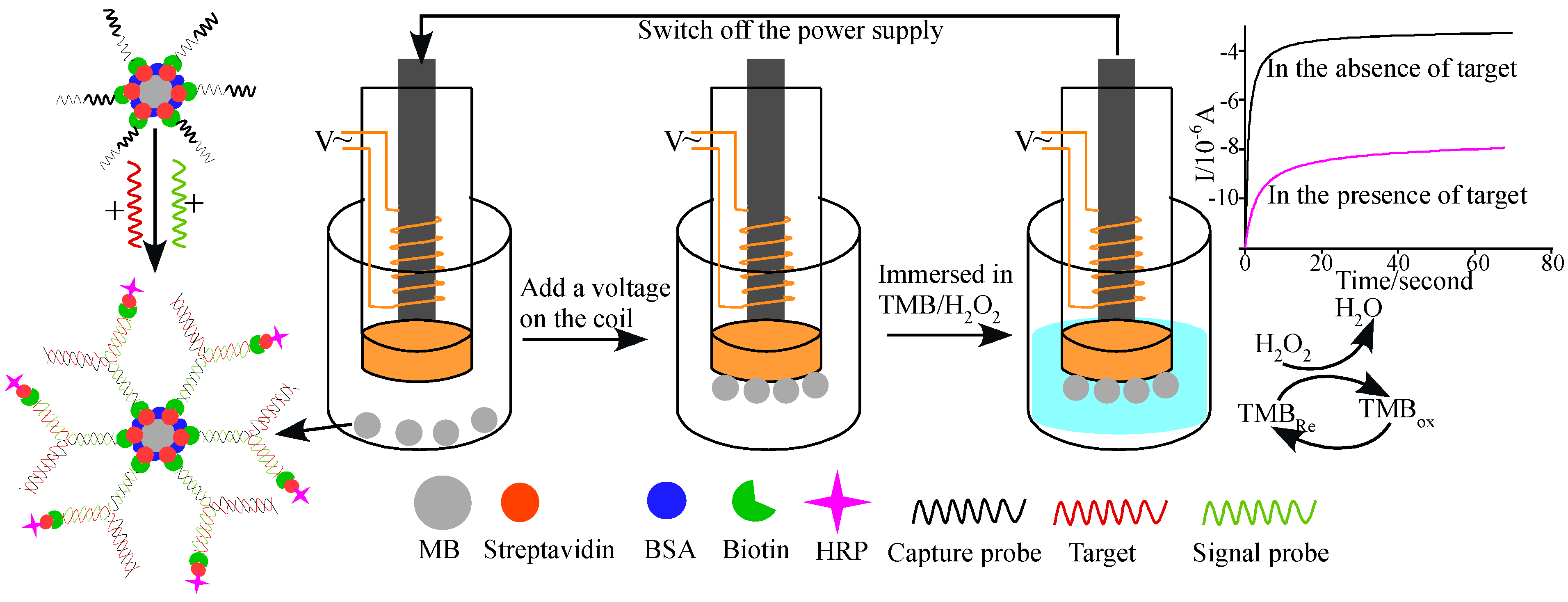
2.7. Magnetic Beads

3. Conclusions/Outlook
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Wienholds, E.; Kloosterman, W.P.; Miska, E.; Alvarez-Saavedra, E.; Berezikov, E.; de Bruijn, E.; Horvitz, H.R.; Kauppinen, S.; Plasterk, R.H. MicroRNA expression in zebrafish embryonic development. Science 2005, 309, 310–311. [Google Scholar] [CrossRef] [PubMed]
- Jovanovic, M.; Hengartner, M.O. MiRNAs and apoptosis: RNAs to die for. Oncogene 2006, 25, 6176–6187. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.D.; Esquela-Kerscher, A.; Stefani, G.; Byrom, M.; Kelnar, K.; Ovcharenko, D.; Wilson, M.; Wang, X.; Shelton, J.; Shingara, J.; et al. The let-7 microRNA represses cell proliferation pathways in human cells. Cancer Res. 2007, 67, 7713–7722. [Google Scholar] [CrossRef] [PubMed]
- Calin, G.A.; Croce, C.M. MicroRNA signatures in human cancers. Nat. Rev. Cancer 2006, 6, 857–866. [Google Scholar] [CrossRef] [PubMed]
- Roosbroeck, K.V.; Pollet, J.; Calin, G.A. miRNAs and long noncoding RNAs as biomarkers in human diseases. Expert Rev. Mol. Diagn. 2013, 13, 183–204. [Google Scholar] [CrossRef] [PubMed]
- Dong, H.; Lei, J.; Ding, L.; Wen, Y.; Ju, H.; Zhang, X. MicroRNA: Function, detection, and bioanalysis. Chem. Rev. 2013, 113, 6207–6233. [Google Scholar] [CrossRef] [PubMed]
- Qavi, A.J.; Bailey, R.C. Multiplexed detection and label-free quantitation of microRNAs using arrays of silicon photonic microring resonators. Angew. Chem. Int. Ed. 2010, 49, 4608–4611. [Google Scholar] [CrossRef]
- Cissell, K.A.; Shrestha, S.; Deo, S.K. MicroRNA detection: Challenge for analytical chemists. Anal. Chem. 2007, 79, 4754–4761. [Google Scholar] [CrossRef]
- Liu, Y.-Q.; Zhang, M.; Yin, B.-C.; Ye, B.-C. Attomolar ultrasensitive microRNA detection by DNA-scaffolded silver-nanocluster probe based on isothermal amplification. Anal. Chem. 2012, 84, 5165–5169. [Google Scholar] [CrossRef] [PubMed]
- Paleček, E.; Bartošík, M. Electrochemistry of nucleic acids. Chem. Rev. 2012, 112, 3427–3481. [Google Scholar] [CrossRef] [PubMed]
- Katz, E.; Willner, I. Integrated nanoparticle-biomolecule hybrid systems: Synthesis, properties, and applications. Angew. Chem. Int. Ed. 2004, 43, 6042–6108. [Google Scholar] [CrossRef]
- Wang, J. Nanoparticle-based electrochemical DNA detection. Anal. Chim. Acta 2003, 500, 247–257. [Google Scholar] [CrossRef]
- Gao, Z.Q.; Yang, Z.C. Detection of microRNAs using electrocatalytic nanoparticle tags. Anal. Chem. 2006, 78, 1470–1477. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Zhang, J.; Zhang, G.; He, H. Polyaniline naowires on Si surfaces fabricated with DNA templates. J. Am. Chem. Soc. 2004, 126, 7097–7101. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Chen, X.T.; Trigg, A.D.; Tung, C.H.; Kong, J.M.; Gao, Z.Q. Detection of microRNAs using target-guided formation of conducting polymer nanowires in nanogaps. J. Am. Chem. Soc. 2007, 129, 5437–5443. [Google Scholar] [CrossRef] [PubMed]
- Gao, Z.; Deng, H.; Shen, W.; Ren, Y. A label-free biosensor for electrochemical detection of femtomolar microRNAs. Anal. Chem. 2013, 85, 1624–1630. [Google Scholar] [CrossRef] [PubMed]
- Shen, W.; Deng, H.; Ren, Y.; Gao, Z. A label-free microRNA biosensor based on DNAzyme-catalyzed and microRNA-guided formation of a thin insulating polymer film. Biosens. Bioelectron. 2013, 44, 171–176. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.; Yi, G.; Gao, Z. A highly sensitive microRNA biosensor based on ruthenium oxide nanoparticle-initiated polymerization of aniline. Chem. Commun. 2010, 46, 9131–9133. [Google Scholar] [CrossRef]
- Peng, Y.; Gao, Z. Amplified detection of microRNA based on ruthenium oxide nanoparticle-initiated deposition of an insulating film. Anal. Chem. 2011, 83, 820–827. [Google Scholar] [CrossRef] [PubMed]
- Rossi, L.M.; Dupont, J.; Machado, G.; Fichtner, P.F.P.; Radtke, C.; Baumvol, I.J.R.; Teixeire, R. Ruthenium dioxide nanoparticles in ionic liquids: Synthesis, characterization and catalytic properties in hydrogenation of olefins and arenes. J. Braz. Chem. Soc. 2004, 15, 904–910. [Google Scholar] [CrossRef]
- Saha, K.; Agasti, S.S.; Kim, C.; Li, X.; Rotello, V.M. Gold nanoparticles in chemical and biological sensing. Chem. Rev. 2012, 112, 2739–2779. [Google Scholar] [CrossRef] [PubMed]
- Guo, S.; Wang, E. Synthesis and electrochemical applications of gold nanoparticles. Anal. Chim. Acta 2007, 598, 181–192. [Google Scholar] [CrossRef] [PubMed]
- Pingarrón, J.M.; Yáñez-Sedeño, P.; González-Cortés, A. Gold nanoparticle-based electrochemical biosensors. Electrochim. Acta 2008, 53, 5848–5866. [Google Scholar] [CrossRef]
- Cao, X.; Ye, Y.; Liu, S. Gold nanoparticle-based signal amplification for biosensing. Anal. Biochem. 2011, 417, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Yi, X.; Tang, H.; Han, H.; Wu, M.; Zhou, F. Direct quantification of microRNA at low picomolar level in sera of glioma patients using a competitive hybridization followed by amplified voltammetric detection. Anal. Chem. 2012, 84, 6400–6406. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, S.; Abiko, N.; Haraguchi, N.; Fujita, H.; Seki, E.; Ono, T.; Yoshida, K.; Anza, J. Voltammetric response of ferroceneboronic acid to diol and phenolic compounds as possible pollutant. J. Environ. Sci. China 2011, 23, 1027–1032. [Google Scholar] [CrossRef] [PubMed]
- Park, J.-Y.; Chang, B.-Y.; Nam, H.; Park, S.-M. Selective electrochemical sensing of glycated hemoglobin (HbA1c) on thiophene-3-boronic acid self-assembled monolayer covered gold electrodes. Anal. Chem. 2008, 80, 8035–8044. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Du, J.; Li, S.; Yuan, B.; Han, H.; Jing, M.; Xia, N. Amplified voltammetric detection of dopamine using ferrocene-capped gold nanoparticle/streptavidin conjugates. Biosens. Bioelectron. 2013, 41, 730–735. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Liu, Y.; Liu, J.; Liu, Z. A Wulff-type boronate for boronate affinity capture of cis-diol compounds at medium acidic pH condition. Chem. Commun. 2011, 47, 8169–8171. [Google Scholar] [CrossRef]
- Potter, O.G.; Breadmore, M.C.; Hilder, E.F. Boronate functionalised polymer monoliths for microscale affinity chromatography. Analyst 2006, 131, 1094–1096. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.M.; Elaissari, A. Nucleic acid sample preparation for in vitro molecular diagnosis: From conventional techniques to biotechnology. Drug Discov. Today 2012, 17, 1199–1207. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Bryson, D.I.; Crumpton, J.B.; Wynn, J.; Santos, W.L. Branched peptide boronic acids (BPBAs): A novel mode of binding towards RNA. Chem. Commun. 2013, 49, 2436–2438. [Google Scholar] [CrossRef]
- Xia, N.; Zhang, L.; Wang, G.; Feng, Q.; Liu, L. Label-free and sensitive strategy for microRNAs detection based on the formation of boronate ester bonds and the dual-amplification of gold nanoparticles. Biosens. Bioelectron. 2013, 47, 461–466. [Google Scholar] [CrossRef] [PubMed]
- Tian, T.; Xiao, H.; Zhang, Z.; Long, Y.; Peng, S.; Wang, S.; Zhou, X.; Liu, S.; Zhou, X. Sensitive and convenient detection of microRNAs based on cascade amplification by catalytic DNAzymes. Chem. Eur. J. 2013, 19, 92–95. [Google Scholar] [CrossRef] [PubMed]
- Yang, N.; Cao, Y.; Han, P.; Zhu, X.; Sun, L.; Li, G. Tools for investigation of the RNA endonuclease activity of mammalian argonaute2 protein. Anal. Chem. 2012, 84, 2492–2497. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Zhao, F.; Ma, F.; Zhang, L.; Yang, S.; Xia, N. Electrochemical detection of β-amyloid peptides on electrode covered with N-terminus-specific antibody based on electrocatalytic O2 reduction by Aβ(1–16)-heme-modified gold nanoparticles. Biosens. Bioelectron. 2013, 49, 231–235. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Wang, M.; Meng, X.; Yin, H.; Ai, S. Amplified electrochemical microRNA biosensor using a hemin-G-quadruplex complex as the sensing element. RSC Adv. 2012, 2, 7140–7145. [Google Scholar] [CrossRef]
- Meng, X.; Zhou, Y.; Liang, Q.; Qu, X.; Yang, Q.; Yin, H.; Ai, S. Electrochemical determination of microRNA-21 based on bio bar code and hemin/G-quadruplet DNAenzyme. Analyst 2013, 138, 3409–3415. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Wang, M.; Xu, Z.; Ni, C.; Yin, H.; Ai, S. Investigation of the effect of phytohormone on the expression of microRNA-159a in Arabidopsis thaliana seedlings based on mimic enzyme catalysis systematic electrochemical biosensor. Biosens. Bioelectron. 2014, 54, 244–250. [Google Scholar] [CrossRef] [PubMed]
- Gao, Z. A highly sensitive electrochemical assay for microRNA expression profiling. Analyst 2012, 137, 1674–1679. [Google Scholar] [CrossRef] [PubMed]
- Pöhlmann, C.; Sprinzl, M. Electrochemical detection of microRNAs via gap hybridization assay. Anal. Chem. 2010, 82, 4434–4440. [Google Scholar] [CrossRef] [PubMed]
- Kilic, T.; Topkaya, S.N.; Ariksoysal, D.O.; Ozsoz, M.; Ballar, P.; Erac, Y.; Gozen, O. Electrochemical based detection of microRNA, mir21 in breast cancer cells. Biosens. Bioelectron. 2012, 38, 195–201. [Google Scholar] [CrossRef] [PubMed]
- Xiang, G.; Jiang, D.; Luo, F.; Liu, F.; Liu, L.; Pu, X. Sensitive detection of microRNAs using hemin/G-quadruplexconcatamers as trace labels and RNA endonuclease-aided targetrecycling for amplification. Sens. Actuators B Chem. 2014, 195, 515–519. [Google Scholar] [CrossRef]
- Yin, H.; Zhou, Y.; Chen, C.; Zhu, L.; Ai, S. An electrochemical signal “off–on” sensing platform for microRNA detection. Analyst 2012, 137, 1389–1395. [Google Scholar] [CrossRef] [PubMed]
- Yin, H.; Zhou, Y.; Zhang, H.; Meng, X.; Ai, S. Electrochemical determination of microRNA-21 based on graphene, LNA integrated molecular beacon, AuNPs and biotin multifunctional bio bar codes and enzymatic assay system. Biosens. Bioelectron. 2012, 33, 247–253. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Wang, M.; Meng, X.; Yin, H.; Ai, S. MicroRNA-21 detection based on molecular switching by amperometry. New J. Chem. 2012, 36, 1985–1991. [Google Scholar] [CrossRef]
- Yang, H. Enzyme-based ultrasensitive electrochemical biosensors. Curr. Opin. Chem. Biol. 2012, 16, 422–428. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Xia, N.; Liu, H.; Kang, X.; Liu, X.; Xue, C.; He, X. Highly sensitive and label-free electrochemical detection of microRNAs based on triple signal amplification of multifunctional gold nanoparticles, enzymes and redox-cycling reaction. Biosens. Bioelectron. 2014, 53, 399–405. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Chai, Y.; Yuan, R.; Su, H.; Han, J. A novel label-free electrochemical microRNA biosensor using Pd nanoparticles as enhancer and linker. Analyst 2013, 138, 1060–1066. [Google Scholar] [CrossRef] [PubMed]
- Dong, H.; Jin, S.; Ju, H.; Hao, K.; Xu, L.-P.; Lu, H.; Zhang, X. Trace and label-free microRNA detection using oligonucleotide encapsulated silver nanoclusters as probes. Anal. Chem. 2012, 84, 8670–8674. [Google Scholar] [CrossRef] [PubMed]
- Chen, A.; Chatterjee, S. Nanomaterials based electrochemical sensors for biomedical applications. Chem. Soc. Rev. 2013, 42, 5425–5438. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Munge, B.; Patel, V.; Jensen, G.; Bhirde, A.; Gong, J.D.; Kim, S.N.; Gillespie, J.; Gutkind, J.S.; Papadimitrakopoulos, F.; et al. Carbon nanotube amplification strategies for highly sensitive immunodetection of cancer biomarkers. J. Am. Chem. Soc. 2006, 128, 11199–11205. [Google Scholar] [CrossRef] [PubMed]
- Zheng, D.; Vashist, S.K.; Dykas, M.M.; Saha, S.; Al-Rubeaan, K.; Lam, E.; Luong, J.H.T.; Sheu, F.S. Graphene versus multi-walled carbon nanotubes for electrochemical glucose biosensing. Materials 2013, 6, 1011–1027. [Google Scholar] [CrossRef]
- Tran, H.V.; Piro, B.; Reisberg, S.; Tran, L.D.; Duc, H.T.; Pham, M.C. Label-free and reagentless electrochemical detection of microRNAs using a conducting polymer nanostructured by carbon nanotubes: Application to prostate cancer biomarker miR-141. Biosens. Bioelectron. 2013, 49, 164–169. [Google Scholar] [CrossRef] [PubMed]
- Tran, H.V.; Piro, B.; Reisberg, S.; Duc, H.T.; Pham, M.C. Antibodies directed to RNA/DNA hybrids: An electrochemical immunosensor for microRNAs detection using graphene-composite electrodes. Anal. Chem. 2013, 85, 8469–8474. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.-I.; Li, B.-R.; Yit-Tsong, C. Silicon nanowire field-effect transistor-based biosensors for biomedical diagnosis and cellular recording investigation. Nano Today 2011, 6, 131–154. [Google Scholar] [CrossRef]
- Cheng, S.; Hotani, K.; Hideshima, S.; Kuroiwa, S.; Nakanishi, T.; Hashimoto, M.; Mori, Y.; Osaka, T. Field effect transistor biosensor using antigen binding fragment for detecting tumor marker in human serum. Materials 2014, 7, 2490–2500. [Google Scholar] [CrossRef]
- Labib, M.; Khan, N.; Ghobadloo, S.M.; Cheng, J.; Pezacki, J.P.; Berezovski, M.V. Three-mode electrochemical sensing of ultralow microRNA levels. J. Am. Chem. Soc. 2013, 135, 3027–3038. [Google Scholar] [CrossRef] [PubMed]
- Ramnani, P.; Gao, Y.; Ozsoz, M.; Mulchandani, A. Electronic detection of microRNA at attomolar level with high specificity. Anal. Chem. 2013, 85, 8061–8064. [Google Scholar] [CrossRef] [PubMed]
- Boon, E.M.; Ceres, D.M.; Drummond, T.G.; Hill, M.G.; Barton, J.K. Mutation detection by electrocatalysis at DNA-modified electrodes. Nat. Biotechnol. 2000, 18, 1096–1100. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Cai, X.; Wang, J.; Jonsson, C. Trace measurements of RNA by potentiometric stripping analysis at carbon paste electrodes. Anal. Chem. 1995, 67, 4065–4070. [Google Scholar] [CrossRef]
- Li, F.; Peng, J.; Wang, J.; Tang, H.; Tan, L.; Xie, Q.; Yao, S. Carbon nanotube-based label-free electrochemical biosensor for sensitive detection of miRNA-24. Biosens. Bioelectron. 2014, 54, 158–164. [Google Scholar] [CrossRef] [PubMed]
- Lusi, E.A.; Passamano, M.; Guarascio, P.; Scarpa, A.; Schiavo, L. Innovative electrochemical approach for an early detection of microRNAs. Anal. Chem. 2009, 81, 2819–2822. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Liu, G.; Merkoçi, A. Electrochemical coding technology for simultaneous detection of multiple DNA targets. J. Am. Chem. Soc. 2003, 125, 3214–3215. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Hu, L.; Zhou, H.; Abdel-Halim, E.S.; Zhu, J.-J. Molecular beacon structure mediated rolling circle amplification for ultrasensitive electrochemical detection of microRNA based on quantum dots tagging. Electrochem. Commun. 2013, 33, 80–83. [Google Scholar] [CrossRef]
- Zhu, W.; Su, X.; Gao, X.; Dai, Z.; Zou, X. A label-free and PCR-free electrochemical assay for multiplexed microRNA profiles by ligase chain reaction coupling with quantum dots barcodes. Biosens. Bioelectron. 2014, 53, 414–419. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Hui, A.; Pampalakis, G.; Soleymani, L.; Liu, F.F.; Sargent, E.H.; Kelley, S.O. Direct, electronic microRNA detection for the rapid determination of differential expression profiles. Angew. Chem. Int. Ed. 2009, 48, 8461–8464. [Google Scholar] [CrossRef]
- Vasilyeva, E.; Lam, B.; Fang, Z.; Minden, M.D.; Sargent, E.H.; Kelley, S.O. Direct genetic analysis of ten cancer cells: Tuning sensor structure and molecular probe design for efficient mRNA capture. Angew. Chem. Int. Ed. 2011, 50, 4137–4141. [Google Scholar] [CrossRef]
- Soleymani, L.; Fang, Z.; Lam, B.; Bin, X.; Vasilyeva, E.; Ross, A.J.; Sargent, E.H.; Kelley, S.O. Hierarchical nanotextured microelectrodes overcome the molecular transport barrier to achieve rapid, direct bacterial detection. ACS Nano 2011, 5, 3360–3366. [Google Scholar] [CrossRef] [PubMed]
- Fang, Z.; Kelley, S.O. Direct electrocatalytic mRNA detection using PNA-nanowire sensors. Anal. Chem. 2009, 81, 612–617. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Getz, G.; Miska, E.A.; Alvarez-Saavedra, E.; Lamb, J.; Peck, D.; Sweet-Cordero, A.; Ebet, B.L.; Mak, R.H.; Ferrando, A.A.; et al. MicroRNA expression profiles classify human cancers. Nature 2005, 435, 834–838. [Google Scholar] [CrossRef] [PubMed]
- Perez, J.M. Iron oxide nanoparticles: Hidden talent. Nat. Nanotechnol. 2007, 2, 535–536. [Google Scholar] [CrossRef] [PubMed]
- Bettazzi, F.; Hamid-Asl, E.; Lucia Esposito, C.; Quintavalle, C.; Formisano, N.; Laschi, S.; Catuogno, S.; Iaboni, M.; Marrazza, G.; Mascini, M.; et al. Electrochemical detection of miRNA-222 by use of a magnetic bead-based bioassay. Anal. Bioanal. Chem. 2013, 405, 1025–1034. [Google Scholar] [CrossRef] [PubMed]
- Erdem, A.; Congur, G.; Eksina, E. Multi channel screen printed array of electrodes for enzyme-linked voltammetric detection of microRNAs. Sens. Actuators B Chem. 2013, 188, 1089–1095. [Google Scholar] [CrossRef]
- Wang, Z.W.; Zhang, J.; Guo, Y.; Wu, X.Y.; Yang, W.J.; Xu, L.J.; Chen, J.H.; Fu, F.F. A novel electrically magnetic-controllable electrochemical biosensor for the ultra sensitive and specific detection of attomolar level oral cancer-related microRNA. Biosens. Bioelectron. 2013, 45, 108–113. [Google Scholar] [CrossRef] [PubMed]
- Bartosik, M.; Hrstka, R.; Palecek, E.; Vojtesek, B. Magnetic bead-based hybridization assay for electrochemical detection of microRNA. Anal. Chim. Acta 2014, 813, 35–40. [Google Scholar] [CrossRef] [PubMed]
- Campuzano, S.; Torrente-Rodríguez, R.M.; López-Hernández, E.; Conzuelo, F.; Granados, R.; Sánchez-Puelles, J.M.; Pingarrón, J.M. Magnetobiosensors based on viral protein p19 for microRNA determination in cancer cells and tissues. Angew. Chem. Int. Ed. 2014, 53, 6168–6171. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Xia, N.; Zhang, L. Nanomaterials-Based Sensing Strategies for Electrochemical Detection of MicroRNAs. Materials 2014, 7, 5366-5384. https://doi.org/10.3390/ma7075366
Xia N, Zhang L. Nanomaterials-Based Sensing Strategies for Electrochemical Detection of MicroRNAs. Materials. 2014; 7(7):5366-5384. https://doi.org/10.3390/ma7075366
Chicago/Turabian StyleXia, Ning, and Liping Zhang. 2014. "Nanomaterials-Based Sensing Strategies for Electrochemical Detection of MicroRNAs" Materials 7, no. 7: 5366-5384. https://doi.org/10.3390/ma7075366




