A New Hydroxylated Nonaprenylhydroquinone from the Mediterranean Marine Sponge Sarcotragus spinosulus
Abstract
:1. Introduction
2. Results and Discussion
3. Experimental Section
3.1. General
3.2. Sponge Material
3.3. Extraction and Isolation
3.4. Characterization Data
3.5. Biological Activity
4. Conclusions
Acknowledgments
- Samples Availability: Available from the authors.
References and Notes
- Cimino, G; De Stefano, S; Minale, L. Prenylated quinones in marine sponges Ircinia species. Experientia 1972, 28, 1401–1402. [Google Scholar]
- Pouchus, YF; Verbist, JF; Biard, JF; Boukef, K. Polyprenylhydroquinones de l’éponge Hippospongia communis: isolement et mise en évidence de formation d’artéfacts sur supports chromatographiques. J. Nat. Prod 1988, 51, 188–189. [Google Scholar]
- Venkateswarlu, Y; Venkata Rami Reddy, M. Three new heptaprenylhydroquinone derivatives from the sponge Ircinia fasciculata. J. Nat. Prod 1994, 57, 1286–1289. [Google Scholar]
- Waetjen, W; Putz, A; Chovolou, Y; Kampkoetter, A; Totzke, F; Kubbutat, MHG; Proksch, P; Konuklugil, B. Hexa-, hepta- and nonaprenylhydroquinones isolated from marine sponges Sarcotragus muscarum and Ircinia fasciculata inhibit NF-κB signalling in H4IIE cells. J. Pharm. Pharmacol 2009, 61, 919–924. [Google Scholar]
- Cimino, G; De Stefano, S; Minale, L. Polyprenyl derivatives from the sponge Ircinia spinosula: 2-Polyprenylbenzoquinones, 2-polyprenylbenzoquinols, prenylated furans and a C-31 difuranoterpene. Tetrahedron 1972, 28, 1315–1324. [Google Scholar]
- Tziveleka, LA; Abatis, D; Paulus, K; Bauer, R; Vagias, C; Roussis, V. Marine polyprenylated hydroquinones, quinones, and chromenols with inhibitory effects on leukotriene formation. Chem. Biodivers 2005, 2, 901–909. [Google Scholar]
- Fenical, W; McConnell, O. Simple antibiotics from the red seaweed Dasya pedicellata var Stanfordiana. Phytochemistry 1976, 15, 435–436. [Google Scholar]
- Sato, A; Shindo, T; Kasanuki, N; Hasegawa, K. Antioxidant metabolites from the tunicate Amaroucium multiplicatum. J. Nat. Prod 1989, 52, 975–981. [Google Scholar]
- Ochi, M; Kotsuki, H; Inoue, S; Taniguchi, M; Tokoroyama, T. Isolation of 2-(3,7,11-trimethyl-2,6,10-dodecatrienyl)hydroquinone from the brown seaweed Dictyopetris undulate. Chem. Lett 1979, 8, 831–832. [Google Scholar]
- Rochfort, SJ; Capon, RJ. New sesquiterpene/phenol from the Australian marine brown alga Perithalia caudata. J. Nat. Prod 1994, 57, 849–851. [Google Scholar]
- De Rosa, S; Crispino, A; De Guilio, A; Iodice, C. Biological effects of prenylated hydroquinones: in antimicrobial, brine shrimp, and fish lethality assays structure-activity relationship studies. J. Nat. Prod 1994, 57, 1711–1716. [Google Scholar]
- Mihopoulos, N; Vagias, C; Chinou, I; Roussakis, C; Scoullos, M; Harvala, C; Roussis, VZ. Antibacterial and cytotoxic natural and synthesized hydroquinones from sponge Ircinia spinosula. Z. Naturforsch 1999, 54c, 417–423. [Google Scholar]
- Loya, S; Rudi, A; Kashman, Y; Hizi, A. Mode of inhibition of HIV reverse transcriptase by 2-hexaprenylhydroquinone, a novel general inhibitor of RNA- and DNA-directed DNA polymerases. Biochem. J 1997, 324, 721–727. [Google Scholar]
- Terencio, MC; Ferrandiz, ML; Posadas, I; Roig, E; De Rosa, S; De Guilo, A; Paya, M; Alcaraz, MJ. Suppression of leukotriene B4 and tumour necrosis factor α release in acute inflammatory responses by novel prenylated hydroquinone derivatives. Arch. Pharmacol 1998, 357, 565–572. [Google Scholar]
- Gil, B; Sanz, MJ; Terencio, MC; De Guilio, A; De Rosa, S; Alcaraz, MJ; Paya, M. Effects of marine 2-polyprenyl-1,4-hydroquinones on phospholipase A2 activity and some inflammatory responses. Eur. J. Pharmacol 1995, 285, 281–288. [Google Scholar]
- Fusetani, N; Sugano, M; Matsunaga, S; Hashimoto, K; Shikama, H; Ohta, A; Nagano, H. Isolation of a hexaprenylhydroquinone sulfate from the marine sponge Dysidea sp. as a proton-potassium ATPase inhibitor. Experientia 1987, 43, 1233–1234. [Google Scholar]
- Stonik, VA; Makarieva, TN; Dmitrenok, AS. Sarcochromenol sulfates A–C and Sarcohydroquinone sulfates A–C, new natural products from the sponge Sarcotragus spinulosus. J. Nat. Prod 1992, 55, 1256–1260. [Google Scholar]
- De Rosa, S; Crispino, A; De Guilio, A; Iodice, C; Milone, A. Sulfated polyprenylhydroquinones from the sponge Ircinia spinosula. J. Nat. Prod 1995, 58, 1450–1454. [Google Scholar]
- Wakimoto, T; Maruyama, A; Matsunaga, S; Fusetani, N. Octa- and Nonaprenylhydroquinone Sulfates, Inhibitors of αl,3-Fucosyltransferase VII, from an Australian Marine Sponge Sarcotragus sp. Bioorg. Med. Chem. Lett 1999, 9, 727–730. [Google Scholar]
- Bifulco, G; Bruno, I; Minale, L; Riccio, R; Debitus, C; Bourdy, G; Vassas, A; Lavayre, J. Bioactive prenylhydroquinones sulfates and a novel C31 furanoterpene alcohol sulfate from the marine sponge, Ircinia sp. J. Nat. Prod 1995, 58, 1444–1449. [Google Scholar]
- Erdogan, I; Tanaka, J; Higa, T; Sener, B. Terpenoids from two sponge species of the Aegean Sea. Nat. Prod. Sci 1999, 5, 177–180. [Google Scholar]
- Tziveleka, LA; Kourounakis, AP; Kourounakis, PN; Roussis, V; Vagias, C. Antioxidant potential of natural and synthesised polyprenylated hydroquinones. Bioorg. Med. Chem 2002, 10, 935–939. [Google Scholar]
- Direct analysis of long polyprenylhydroquinones by electrospray ionization mass spectrometry (ESI-MS) or MALDITOF-MS often produces poor results requiring off-line time and sample-consuming derivatization techniques. Thus, silver trifluoroacetate was used as indicated in the experimental section to promote cationization of the isolated compounds by intense formation of [M + Ag]+ adducts.
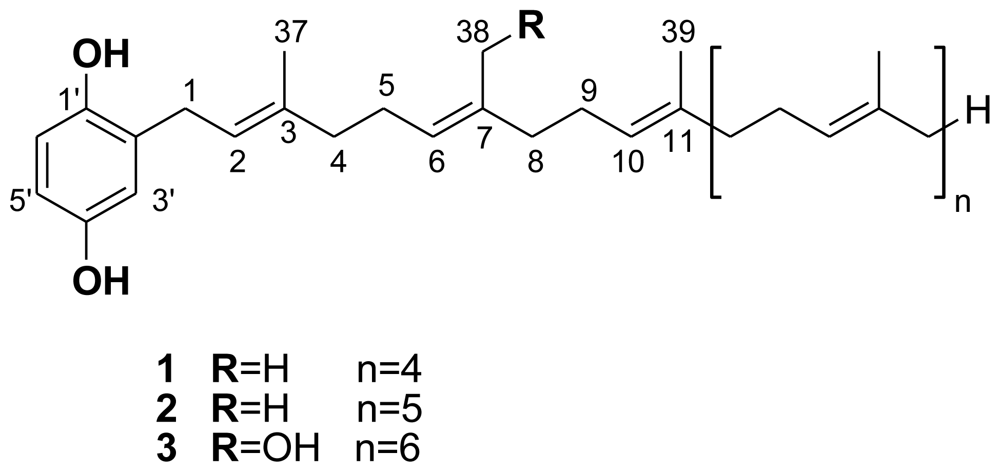
- Formally, the side-chain OH group gives rise to one (Z)-configured C=C bond, but the parent C=C arrangement remains (all-E).
- Jacquel, A; Herrant, M; Legros, L; Belhacene, N; Luciano, F; Pages, G; Hofman, P; Auberger, P. Imatinib induces mitochondria-dependent apoptosis of the Bcr-Abl positive K562 cell line and its differentiation towards the erythroid lineage. FASEB J 2003, 17, 2160–2162. [Google Scholar]
- Jacquel, A; Colosetti, P; Grosso, S; Belhacene, N; Puissant, A; Marchetti, S; Breittmayer, JP; Auberger, P. Apoptosis and erythroid differentiation triggered by Bcr-Abl inhibitors in CML cell lines are fully distinguishable processes that exhibit different sensitivity to caspase inhibition. Oncogene 2007, 26, 2445–2458. [Google Scholar]
- Puissant, A; Grosso, S; Jacquel, A; Belhacene, N; Colosetti, P; Cassuto, JP; Auberger, P. Imatinib mesylate-resistant human chronic myelogenous leukemia cell lines exhibit high sensitivity to the phytoalexin resveratrol. FASEB J 2008, 22, 1894–1904. [Google Scholar]

| No. | δC (ppm) | Mult. | δH (ppm) | J (Hz) | Mult. | COSY | HMBC |
|---|---|---|---|---|---|---|---|
| 1′ | 148.9 | qC | |||||
| 2′ | 130.2 | qC | |||||
| 3′ | 117.2 | CH | 6.52 | 3.00 | d | 1 | 1, 1′, 4′, 5′ |
| 4′ | 151.1 | qC | |||||
| 5′ | 113.8 | CH | 6.41 | 8.50, 3.00 | dd | 6′ | 3′, 4′ |
| 6′ | 116.5 | CH | 6.56 | 8.50 | d | 5′ | 1, 1′, 2, 5′ |
| 1 | 29.1 | CH2 | 3.22 | 7.33 | d | 2 | 1′, 2′, 3, 3′ |
| 2 | 124.0 | CH | 5.30 | m | 1 | 1, 4 | |
| 3 | 136.8 | qC | |||||
| 4 | 40.9 | CH2 | 1.98 | m | 5 | ||
| 5 | 27.7 | CH2 | 2.16 | m | 4, 6 | 3, 4, 6, 7 | |
| 6 | 128.8 | CH | 5.25 | m | 5 | ||
| 7 | 139.4 | qC | |||||
| 8 | 35.9 | CH2 | 2.11 | m | 9 | 9, 6, 10, 38 | |
| 9 | 27.7 | CH2 | 2.05 | m | 8, 10 | ||
| 10 | 125.6 | CH | 5.10 | m | 9 | ||
| 11 | 135.8 | qC | |||||
| 12 | 40.9 | CH2 | 1.96 | m | |||
| 13 | 27.7 | CH2 | 2.05 | m | |||
| 14 | 125.6 | CH | 5.10 | m | |||
| 15 | 135.8 | qC | |||||
| 16 | 40.9 | CH2 | 1.96 | m | |||
| 17 | 27.7 | CH2 | 2.05 | m | |||
| 18 | 125.6 | CH | 5.10 | m | |||
| 19 | 135.8 | qC | |||||
| 20 | 40.9 | CH2 | 1.96 | m | |||
| 21 | 27.7 | CH2 | 2.05 | m | |||
| 22 | 125.6 | CH | 5.10 | m | |||
| 23 | 135.8 | qC | |||||
| 24 | 40.9 | CH2 | 1.96 | m | |||
| 25 | 27.7 | CH2 | 2.05 | m | |||
| 26 | 125.6 | CH | 5.10 | m | |||
| 27 | 135.8 | qC | |||||
| 28 | 40.9 | CH2 | 1.96 | m | |||
| 29 | 27.7 | CH2 | 2.05 | m | |||
| 30 | 125.6 | CH | 5.10 | m | |||
| 31 | 135.8 | qC | |||||
| 32 | 40.9 | CH2 | 1.96 | m | |||
| 33 | 27.7 | CH2 | 2.05 | m | |||
| 34 | 125.6 | CH | 5.10 | m | |||
| 35 | 135.8 | qC | |||||
| 36 | 25.9 | CH3 | 1.65 | s | 34,45 | ||
| 37 | 16.2 | CH3 | 1.69 | s | 1,2 | 2,3,4 | |
| 38 | 60.0 | CH2 | 4.06 | s | 6,7,8 | ||
| 39 | 16.2 | CH3 | 1.58 | s | |||
| 40 | 16.2 | CH3 | 1.58 | s | |||
| 41 | 16.2 | CH3 | 1.58 | s | |||
| 42 | 16.2 | CH3 | 1.58 | s | |||
| 43 | 16.2 | CH3 | 1.58 | s | |||
| 44 | 16.2 | CH3 | 1.58 | s | |||
| 45 | 17.8 | CH3 | 1.58 | s |
| Compounds | Cell metabolism (IC50) | Cell number (IC50) |
|---|---|---|
| 1 | 8 | 7 |
| 2 | 10 | 12 |
| 3 | 193 | 191 |
| Imatinib | 0.4 | 0.5 |
© 2011 by the authors; licensee MDPI, Basel, Switzerland This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Abed, C.; Legrave, N.; Dufies, M.; Robert, G.; Guérineau, V.; Vacelet, J.; Auberger, P.; Amade, P.; Mehiri, M. A New Hydroxylated Nonaprenylhydroquinone from the Mediterranean Marine Sponge Sarcotragus spinosulus. Mar. Drugs 2011, 9, 1210-1219. https://doi.org/10.3390/md9071210
Abed C, Legrave N, Dufies M, Robert G, Guérineau V, Vacelet J, Auberger P, Amade P, Mehiri M. A New Hydroxylated Nonaprenylhydroquinone from the Mediterranean Marine Sponge Sarcotragus spinosulus. Marine Drugs. 2011; 9(7):1210-1219. https://doi.org/10.3390/md9071210
Chicago/Turabian StyleAbed, Charline, Nathalie Legrave, Maeva Dufies, Guillaume Robert, Vincent Guérineau, Jean Vacelet, Patrick Auberger, Philippe Amade, and Mohamed Mehiri. 2011. "A New Hydroxylated Nonaprenylhydroquinone from the Mediterranean Marine Sponge Sarcotragus spinosulus" Marine Drugs 9, no. 7: 1210-1219. https://doi.org/10.3390/md9071210
APA StyleAbed, C., Legrave, N., Dufies, M., Robert, G., Guérineau, V., Vacelet, J., Auberger, P., Amade, P., & Mehiri, M. (2011). A New Hydroxylated Nonaprenylhydroquinone from the Mediterranean Marine Sponge Sarcotragus spinosulus. Marine Drugs, 9(7), 1210-1219. https://doi.org/10.3390/md9071210





