Symbiodinium—Invertebrate Symbioses and the Role of Metabolomics
Abstract
:1. Introduction
2. Symbiodinium Symbiosis
3. Targeted Metabolite Analysis
3.1. Nutritional Roles of Symbiodinium
3.2. Diel Cycles
3.3. Biomarkers and Natural Products
4. Marine Metabolomics
5. Metabolomics in Systems Biology
6. Concluding Remarks
Acknowledgements
- Samples Availability: Available from the authors.
References
- Fiehn, O; Kopka, J; Dormann, P; Altmann, T; Trethewey, RN; Willmitzer, L. Metabolite Profiling for Plant Functional Genomics. Nat. Biotechnol 2000, 18, 1157–1161. [Google Scholar]
- Raamsdonk, LM; Teusink, B; Broadhurst, D; Zhang, NS; Hayes, A; Walsh, MC; Berden, JA; Brindle, KM; Kell, DB; Rowland, JJ; Westerhoff, HV; van Dam, K; Oliver, SG. A Functional Genomics Strategy that Uses Metabolome Data to Reveal the Phenotype of Silent Mutations. Nat. Biotechnol 2001, 19, 45–50. [Google Scholar]
- Oliver, SG; Winson, MK; Kell, DB; Baganz, F. Systematic Functional Analysis of the Yeast Genome. Trends Biotechnol 1998, 16, 373–378. [Google Scholar]
- Goodacre, R; Vaidyanathan, S; Dunn, WB; Harrigan, GG; Kell, DB. Metabolomics by Numbers: Acquiring and Understanding Global Metabolite Data. Trends Biotechnol 2004, 22, 245–252. [Google Scholar]
- Fiehn, O. Combining Genomics, Metabolome Analysis, and Biochemical Modelling to Understand Metabolic Networks. Comp. Funct. Genomics 2001, 2, 155–168. [Google Scholar]
- Rochfort, S. Metabolomics Reviewed: A New “Omics” Platform Technology for Systems Biology and Implications for Natural Products Research. J. Nat. Prod 2005, 68, 1813–1820. [Google Scholar]
- Viant, MR. Metabolomics of Aquatic Organisms: The New ‘Omics’ on the Block. Mar. Ecol. Prog. Ser 2007, 332, 301–306. [Google Scholar]
- Viant, MR; Bearden, DW; Bundy, JG; Burton, IW; Collette, TW; Ekman, DR; Ezernieks, V; Karakach, TK; Lin, CY; Rochfort, S; de Ropp, JS; Teng, Q; Tjeerdema, RS; Walter, JA; Wu, H. International NMR-Based Environmental Metabolomics Intercomparison Exercise. Environ. Sci. Technol 2009, 43, 219–225. [Google Scholar]
- Bundy, J; Davey, M; Viant, M. Environmental Metabolomics: A Critical Review and Future Perspectives. Metabolomics 2009, 5, 3–21. [Google Scholar]
- Kell, DB; Oliver, SG. Here Is the Evidence, Now What Is the Hypothesis? The Complementary Roles of Inductive and Hypothesis-Driven Science in the Post-Genomic Era. Bioessays 2004, 26, 99–105. [Google Scholar]
- Dixon, RA; Gang, DR; Charlton, AJ; Fiehn, O; Kuiper, HA; Reynolds, TL; Tjeerdema, RS; Jeffery, EH; German, JB; Ridley, WP; Seiber, JN. Applications of Metabolomics in Agriculture. J. Agric. Food Chem 2006, 54, 8984–8994. [Google Scholar]
- Rochfort, S; Ezernieks, V; Bastian, SEP; Downey, MO. Sensory Attributes of Wine Influenced by Variety and Berry Shading Discriminated by NMR Metabolomics. Food Chem 2010, 121, 1296–1304. [Google Scholar]
- Ovenden, SPB; Gordon, BR; Bagas, CK; Muir, B; Rochfort, S; Bourne, DJ. A Study of the Metabolome of Ricinus communis for Forensic Applications. Aust. J. Chem 2010, 63, 8–21. [Google Scholar]
- Cienkowski, L. On the Production of Spores in the Radiolaria. Archiv. Fur. Mikroskop. Anatomie 1871, 7, 396–403. [Google Scholar]
- Muscatine, L; Porter, JW. Reef Corals: Mutualistic Symbioses Adapted to Nutrient-Poor Environments. Bioscience 1977, 27, 454–460. [Google Scholar]
- Smith, DC; Douglas, AE. The Biology of Symbiosis; Edward Arnold: London, UK, 1987. [Google Scholar]
- Trench, RK. The Cell Biology of Plant-Animal Symbiosis. Annu. Rev. Plant Physiol. Plant Mol. Biol 1979, 30, 485–531. [Google Scholar]
- Lewis, DH; Smith, DC. The Autotrophic Nutrition of Symbiotic Marine Coelenterates with Special Reference to Hermatypic Corals. I. Movement of Photosynthetic Products between the Symbionts. Proc. R. Soc. Lond. B Biol. Sci 1971, 178, 111–129. [Google Scholar]
- Hughes, TP; Baird, AH; Bellwood, DR; Card, M; Connolly, SR; Folke, C; Grosberg, R; Hoegh-Guldberg, O; Jackson, JBC; Kleypas, J; et al. Climate Change, Human Impacts, and the Resilience of Coral Reefs. Science 2003, 301, 929–933. [Google Scholar]
- Yellowlees, D; Rees, TAV; Leggat, W. Metabolic Interactions between Algal Symbionts and Invertebrate Hosts. Plant Cell Environ 2008, 31, 679–694. [Google Scholar]
- Stat, M; Carter, D; Hoegh-Guldberg, O. The Evolutionary History of Symbiodinium and Scleractinian Hosts—Symbiosis, Diversity, and the Effect of Climate Change. Perspect. Plant Ecol. Evol. Syst 2006, 8, 23–43. [Google Scholar]
- Roth, E; Jeon, K; Stacey, G. Palacios, RD, Verma, PS, Eds.; Molecular Genetics of Plant Microbe Interactions. In Homology in Endosymbiotic Systems: The Term “Symbiosome”; Proceedings of the 4th International Symposium of Molecular Genetics of Plant—Microbe Interactions, Acapulco, Mexico, 1988, APS Press: Acapulco, Mexico, 1988; pp. 220–225. [Google Scholar]
- Rands, ML; Loughman, BC; Douglas, AE. The Symbiotic Interface in an Alga Invertebrate Symbiosis. Proc. R. Soc. Lond. B Biol. Sci 1993, 253, 161–165. [Google Scholar]
- Muscatine, L; Blackburn, A; Gates, RD; Baghdasarian, G; Allemand, D. Cell-Specific Density of Symbiotic Dinoflagellates in Tropical Anthozoans. Coral Reefs 1998, 17, 329–337. [Google Scholar]
- Fitt, WK. Cellular Growth of Host and Symbiont in a Cnidarian-Zooxanthellar Symbiosis. Biol. Bull 2000, 198, 110–120. [Google Scholar]
- LaJeunesse, TC. Investigating the Biodiversity, Ecology, and Phylogeny of Endosymbiotic Dinoflagellates in the Genus Symbiodinium Using the ITS Region: In Search of A “Species” Level Marker. J. Phycol 2001, 37, 866–880. [Google Scholar]
- Zahl, PA; McLaughlin, JJA. Isolation and Cultivation of Zooxanthellae. Nature 1957, 180, 199–200. [Google Scholar]
- Muscatine, L; Hand, C. Direct Evidence for the Transfer of Materials from Symbiotic Algae to the Tissues of a Coelenterate. Proc. Natl. Acad. Sci. USA 1958, 44, 1259–1263. [Google Scholar]
- Muscatine, L; Karakashian, SJ; Karakashian, MW. Soluble Extracellular Products of Algae Symbiotic with a Ciliate, a Sponge and a Mutant Hydra. Comp. Biochem. Physiol 1967, 20, 1–6. [Google Scholar]
- Goreau, TF; Goreau, NI; Yonge, CM. On the Utilization of Photosynthetic Products from Zooxanthellae and of a Dissolved Amino Acid in Tridacna maxima f. Elongata (Mollusca: Bivalvia). J. Zool 1973, 169, 417–454. [Google Scholar]
- Von Holt, C; Von Holt, M. Transfer of Photosynthetic Products from Zooxanthellae to Coelenterate Hosts. Comp. Biochem. Physiol 1968, 24, 73–81. [Google Scholar]
- Cernichiari, E; Muscatine, L; Smith, DC. Maltose Excretion by the Symbiotic Algae of Hydra Viridis. Proc. R. Soc. Lond. B Biol. Sci 1969, 173, 557–576. [Google Scholar]
- Muscatine, L. Symbiosis of Hydra and Algae—III. Extracellular Products of the Algae. Comp. Biochem. Physiol 1965, 16, 77–92. [Google Scholar]
- Muscatine, L. Glycerol Excretion by Symbiotic Algae from Corals and Tridacna and Its Control by the Host. Science 1967, 156, 516–519. [Google Scholar]
- Bil, KY; Kolmakov, PV; Parnik, T; Titlyanov, EA; Muskatine, L. Photosynthetic Products in Zooxanthellae of the Symbiotic Corals Stylophora pistillata and Seriatopora coliendrum Situated at Various Depths. Fiziol. Rast. (Moscow) 1991, 38, 846–854. [Google Scholar]
- Von Holt, C. Uptake of Glycine and Release of Nucleoside Polyphosphates by Zooxanthellae. Comp. Biochem. Physiol 1968, 26, 1071–1079. [Google Scholar]
- Von Holt, C; Von Holt, M. Secretion of Organic Compounds by Zooxanthellae Isolated from Various Types of Zoanthus. Comp. Biochem. Physiol 1968, 24, 83–92. [Google Scholar]
- Patton, JS; Abraham, S; Benson, AA. Lipogenesis in the Intact Coral Pocillopora capitata and Its Isolated Zooxanthellae: Evidence for a Light-Driven Carbon Cycle between Symbiont and Host. Mar. Biol 1977, 44, 235–247. [Google Scholar]
- Kellogg, RB; Patton, JS. Lipid Droplets, Medium of Energy Exchange in the Symbiotic Anemone Condylactis gigantea: A Model Coral Polyp. Mar. Biol 1983, 75, 137–149. [Google Scholar]
- Cates, N; McLaughlin, JJA. Nutrient Availability for Zooxanthellae Derived from Physiological Activities of Condylactis spp. J. Exp. Mar. Biol. Ecol 1979, 37, 31–41. [Google Scholar]
- Muscatine, L; Lenhoff, HM. Symbiosis: On the Role of Algae Symbiotic with Hydra. Science 1963, 142, 956–958. [Google Scholar]
- Patton, JS; Battey, JF; Rigler, MW; Porter, JW; Black, CC; Burris, JE. A Comparison of the Metabolism of Bicarbonate 14C and Acetate 14C and the Variability of Species Lipid Composition in Reef Corals. Mar. Biol 1983, 75, 121–130. [Google Scholar]
- Dykens, JA; Shick, JM. Oxygen Production by Endosymbiotic Algae Controls Superoxide Dismutase Activity in Their Animal Host. Nature 1982, 297, 579–580. [Google Scholar]
- Onodera, K-i; Nakamura, H; Oba, Y; Ojika, M. Zooxanthellamide A, a Novel Polyhydroxy Metabolite from a Marine Dinoflagellate of Symbiodinium sp. Tetrahedron 2003, 59, 1067–1071. [Google Scholar]
- Onodera, K-i; Nakamura, H; Oba, Y; Ojika, M. Zooxanthellamide B, a Novel Large Polyhydroxy Metabolite from a Marine Dinoflagellate of Symbiodinium sp. Biosci. Biotechnol. Biochem 2004, 68, 955–958. [Google Scholar]
- Dunlap, WC; Yamamoto, Y. Small-Molecular Antioxidants in Marine Organisms: Antioxidant Activity of Mycosporine-Glycine. Comp. Biochem. Physiol 1995, 112, 106–114. [Google Scholar]
- Nakamura, H; Asari, T; Ohizumi, Y; Kobayashi, J; Yamasu, T; Murai, A. Isolation of Zooxanthellatoxins, Novel Vasoconstrictive Substances from the Zooxanthella Symbiodinium sp. Toxicon 1993, 31, 371–376. [Google Scholar]
- Nakamura, H; Kawase, Y; Maruyama, K; Murai, A. Studies on Polyketide Metabolites of a Symbiotic Dinoflagellate, Symbiodinium sp.: A New C30 Marine Alkaloid, Zooxanthellamine, a Plausible Precursor for Zoanthid Alkaloids. Bull. Chem. Soc. Jpn 1998, 71, 781–787. [Google Scholar]
- Trench, RK. The Physiology and Biochemistry of Zooxanthellae Symbiotic with Marine Coelenterates. I. The Assimilation of Photosynthetic Products of Zooxanthellae by Two Marine Coelenterates. Proc. R. Soc. Lond. B Biol. Sci 1971, 177, 225–235. [Google Scholar]
- Trench, RK. The Physiology and Biochemistry of Zooxanthellae Symbiotic with Marine Coelenterates. II. Liberation of Fixed 14C by Zooxanthellae in Vitro. Proc. R. Soc. Lond. B Biol. Sci 1971, 177, 237–250. [Google Scholar]
- Trench, RK. The Physiology and Biochemistry of Zooxanthellae Symbiotic with Marine Coelenterates. III. The Effect of Homogenates of Host Tissues on the Excretion of Photosynthetic Products in Vitro by Zooxanthellae from Two Marine Coelenterates. Proc. R. Soc. Lond. B Biol. Sci 1971, 177, 251–264. [Google Scholar]
- Gates, RD; Hoegh-Guldberg, O; McFall-Ngai, MJ; Bil, KY; Muscatine, L. Free Amino Acids Exhibit Anthozoan “Host Factor” Activity: They Induce the Release of Photosynthate from Symbiotic Dinoflagellates in Vitro. Proc. Natl. Acad. Sci. USA 1995, 92, 7430–7434. [Google Scholar]
- Withers, KJT; Grant, AJ; Hinde, R. Effects of Free Amino Acids on the Isolated Symbiotic Algae of the Coral Plesiastrea versipora (Lamarck): Absence of a Host Release Factor Response. Comp. Biochem. Physiol. Part A Mol. Integr. Physiol 1998, 120, 599–607. [Google Scholar]
- Cook, CB; Davy, SK. Are Free Amino Acids Responsible for the ‘Host Factor’ Effects on Symbiotic Zooxanthellae in Extracts of Host Tissue? Hydrobiologia 2001, 461, 71–78. [Google Scholar]
- Grant, AJ; Remond, M; Hinde, R. Low Molecular-Weight Factor from Plesiastrea versipora (Scleractinia) That Modifies Release and Glycerol Metabolism of Isolated Symbiotic Algae. Mar. Biol 1998, 130, 553–557. [Google Scholar]
- Grant, AJ; Remond, M; Withers, KJT; Hinde, R. Inhibition of Algal Photosynthesis by a Symbiotic Coral. Hydrobiologia 2001, 461, 63–69. [Google Scholar]
- Martin, S; Parton, R. Lipid Droplets: A Unified View of a Dynamic Organelle. Nat. Rev. Mol. Cell Biol 2006, 7, 373. [Google Scholar]
- Umlauf, E; Csaszar, E; Moertelmaier, M; Schuetz, GJ; Parton, RG; Prohaska, R. Association of Stomatin with Lipid Bodies. J. Biol. Chem 2004, 279, 23699–23709. [Google Scholar]
- Imanishi, Y; Gerke, V; Palczewski, K. Retinosomes: New Insights into Intracellular Managing of Hydrophobic Substances in Lipid Bodies. J. Cell Biol 2004, 166, 447–453. [Google Scholar]
- Luo, YJ; Wang, LH; Chen, WNU; Peng, SE; Tzen, JTC; Hsiao, YY; Huang, HJ; Fang, LS; Chen, CS. Ratiometric Imaging of Gastrodermal Lipid Bodies in Coral-Dinoflagellate Endosymbiosis. Coral Reefs 2009, 28, 289–301. [Google Scholar]
- Tchernov, D; Gorbunov, MY; de Vargas, C; Yadav, SN; Milligan, AJ; Haggblom, M; Falkowski, PG. Membrane Lipids of Symbiotic Algae Are Diagnostic of Sensitivity to Thermal Bleaching in Corals. Proc. Natl. Acad. Sci. USA 2004, 101, 13531–13535. [Google Scholar]
- Wang, JT; Douglas, AE. Essential Amino Acid Synthesis and Nitrogen Recycling in an Alga-Invertebrate Symbiosis. Mar. Biol 1999, 135, 219–222. [Google Scholar]
- Muscatine, L; D’Elia, CF. The Uptake, Retention, and Release of Ammonium by Reef Corals. Limnol. Oceanogr 1978, 23, 725–734. [Google Scholar]
- D’Elia, CF; Domotor, SL; Webb, KL. Nutrient Uptake Kinetics of Freshly Isolated Zooxanthellae. Mar. Biol 1983, 75, 157–167. [Google Scholar]
- Wang, JT; Douglas, AE. Nitrogen Recycling or Nitrogen Conservation in an Alga-Invertebrate Symbiosis? J. Exp. Biol 1998, 201, 2445–2453. [Google Scholar]
- Franzisket, L. Uptake and Accumulation of Nitrate and Nitrite by Reef Corals. Naturwissenschaften 1973, 60, 552. [Google Scholar]
- Leggat, W; Hoegh-Guldberg, O; Dove, S; Yellowlees, D. Analysis of an EST Library from the Dinoflagellate (Symbiodinium sp.) Symbiont of Reef-Building Corals. J. Phycol 2007, 43, 1010–1021. [Google Scholar]
- Marubini, F; Davies, PS. Nitrate Increases Zooxanthellae Population Density and Reduces Skeletogenesis in Corals. Mar. Biol 1996, 127, 319–328. [Google Scholar]
- Kuhl, M; Cohen, Y; Dalsgaard, T; Jorgensen, BB; Revsbech, NP. Microenvironment and Photosynthesis of Zooxanthellae in Scleractinian Corals Studies with Microsensors for O2, pH and Light. Mar. Ecol. Prog. Ser 1995, 117, 159–172. [Google Scholar]
- Levy, O; Achituv, Y; Yacobi, YZ; Stambler, N; Dubinsky, Z. The Impact of Spectral Composition and Light Periodicity on the Activity of Two Antioxidant Enzymes (SOD and CAT) in the Coral Favia favus. J. Exp. Mar. Biol. Ecol 2006, 328, 35–46. [Google Scholar]
- Rees, TAV; Fitt, WK; Baillie, B; Yellowlees, D. A Method for Temporal Measurement of Haemolymph Composition in the Giant Clam Symbiosis and Its Application to Glucose and Glycerol Levels During a Diel Cycle. Limnol. Oceanogr 1993, 38, 213–217. [Google Scholar]
- Leggat, W; Marendy, EM; Baillie, B; Whitney, SM; Ludwig, M; Badger, MR; Yellowlees, D. Dinoflagellate Symbioses: Strategies and Adaptations for the Acquisition and Fixation of Inorganic Carbon. Funct. Plant Biol 2002, 29, 309–322. [Google Scholar]
- Nakamura, H; Kobayashi, J; Hirata, Y. Separation of Mycosporine-Like Amino-Acids in Marine Organisms Using Reversed-Phase High-Performance Liquid-Chromatography. J. Chromatogr 1982, 250, 113–118. [Google Scholar]
- Yakovleva, I; Hidaka, M. Diel Fluctuations of Mycosporine-Like Amino Acids in Shallow-Water Scleractinian Corals. Mar. Biol 2004, 145, 863–873. [Google Scholar]
- Banaszak, AT; LaJeunesse, TC; Trench, RK. The Synthesis of Mycosporine-Like Amino Acids (MAAs) by Cultured, Symbiotic Dinoflagellates. J. Exp. Mar. Biol. Ecol 2000, 249, 219–233. [Google Scholar]
- Korbee, N; Houvinen, P; Figueroa, FL; Aguilera, J; Karsten, U. Availability of Ammonium Influences Photosynthesis and the Accumulation of Mycosporine-Like Amino Acids in Two Porphyra Species (Bangiales, Rhodophyta). Mar. Biol 2005, 146, 645–654. [Google Scholar]
- Kashman, Y; Groweiss, A; Carmely, S; Kinamoni, Z; Czarkie, D; Rotem, M. Recent Research in Marine Natural Products from the Red Sea. Pure Appl. Chem 1982, 54, 1995–2010. [Google Scholar]
- Cuif, JP; Dauphin, Y; Freiwald, A; Gautret, P; Zibrowius, H. Biochemical Markers of Zooxanthellae Symbiosis in Soluble Matrices of Skeleton of 24 Scleractinia Species. Comp. Biochem. Physiol 1999, 123, 269–278. [Google Scholar]
- Boroujerdi Arezue, FB; Vizcaino Maria, I; Meyers, A; Pollock Elizabeth, C; Huynh Sara, L; Schock Tracey, B; Morris Pamela, J; Bearden Daniel, W. NMR-Based Microbial Metabolomics and the Temperature-Dependent Coral Pathogen Vibrio coralliilyticus. Environ. Sci. Technol 2009, 43, 7658–7664. [Google Scholar]
- Sumner, LW; Amberg, A; Barrett, D; Beale, MH; Beger, R; Daykin, CA; Fan, TWM; Fiehn, O; Goodacre, R; Griffin, JL; et al. Proposed Minimum Reporting Standards for Chemical Analysis. Metabolomics 2007, 3, 211–221. [Google Scholar]
- Hines, A; Oladiran, GS; Bignell, JP; Stentiford, GD; Viant, MR. Direct Sampling of Organisms from the Field and Knowledge of Their Phenotype: Key Recommendations for Environmental Metabolomics. Environ. Sci. Technol 2007, 41, 3375–3381. [Google Scholar]
- Kluender, C; Sans-Piche, F; Riedl, J; Altenburger, R; Haertig, C; Laue, G; Schmitt-Jansen, M. A Metabolomics Approach to Assessing Phytotoxic Effects on the Green Alga Scenedesmus vacuolatus. Metabolomics 2009, 5, 59–71. [Google Scholar]
- Stat, M; Loh, WKW; LaJeunesse, TC; Hoegh-Guldberg, O; Carter, DA. Stability of Coral-Endosymbiont Associations During and after a Thermal Stress Event in the Southern Great Barrier Reef. Coral Reefs 2009, 28, 709–713. [Google Scholar]
- Rowan, R. Coral Bleaching: Thermal Adaptation in Reef Coral Symbionts. Nature 2004, 430, 742. [Google Scholar]
- Abrego, D; Ulstrup, KE; Willis, BL; van Oppen, MJH. Species-Specific Interactions between Algal Endosymbionts and Coral Hosts Define Their Bleaching Response to Heat and Light Stress. Proc. R. Soc. Lond. B Biol. Sci 2008, 275, 2273–2282. [Google Scholar]
- Cantin, N; van Oppen, M; Willis, B; Mieog, J; Negri, A. Juvenile Corals Can Acquire More Carbon from High-Performance Algal Symbionts. Coral Reefs 2009, 28, 405–414. [Google Scholar]
- Kim, JD; Lee, CG. Systemic Optimization of Microalgae for Bioactive Compound Production. Biotechnol. Bioprocess Eng 2005, 10, 418–424. [Google Scholar]
- Kortschak, RD; Samuel, G; Saint, R; Miller, DJ. Est Analysis of the Cnidarian Acropora Millepora Reveals Extensive Gene Loss and Rapid Sequence Divergence in the Model Invertebrates. Curr. Biol 2003, 13, 2190–2195. [Google Scholar]
- Meyer, E; Davies, S; Wang, S; Willis, BL; Abrego, D; Juenger, TE; Matz, MV. Genetic Variation in Responses to a Settlement Cue and Elevated Temperature in the Reef-Building Coral Acropora millepora. Mar. Ecol. Prog. Ser 2009, 392, 81–92. [Google Scholar]
- Voolstra, CR; Sunagawa, S; Schwarz, JA; Coffroth, MA; Yellowlees, D; Leggat, W; Medina, M. Evolutionary Analysis of Orthologous cDNA Sequences from Cultured and Symbiotic Dinoflagellate Symbionts of Reef-Building Corals (Dinophyceae: Symbiodinium). Comp. Biochem. Physiol. Part D Genomics Proteomics 2009, 4, 67–74. [Google Scholar]
- Santos, EM; Ball, JS; Williams, TD; Wu, HF; Ortega, F; Van Aerle, R; Katsiadaki, I; Falciani, F; Viant, MR; Chipman, JK; Tyler, CR. Identifying Health Impacts of Exposure to Copper Using Transcriptomics and Metabolomics in a Fish Model. Environ. Sci. Technol 2010, 44, 820–826. [Google Scholar]
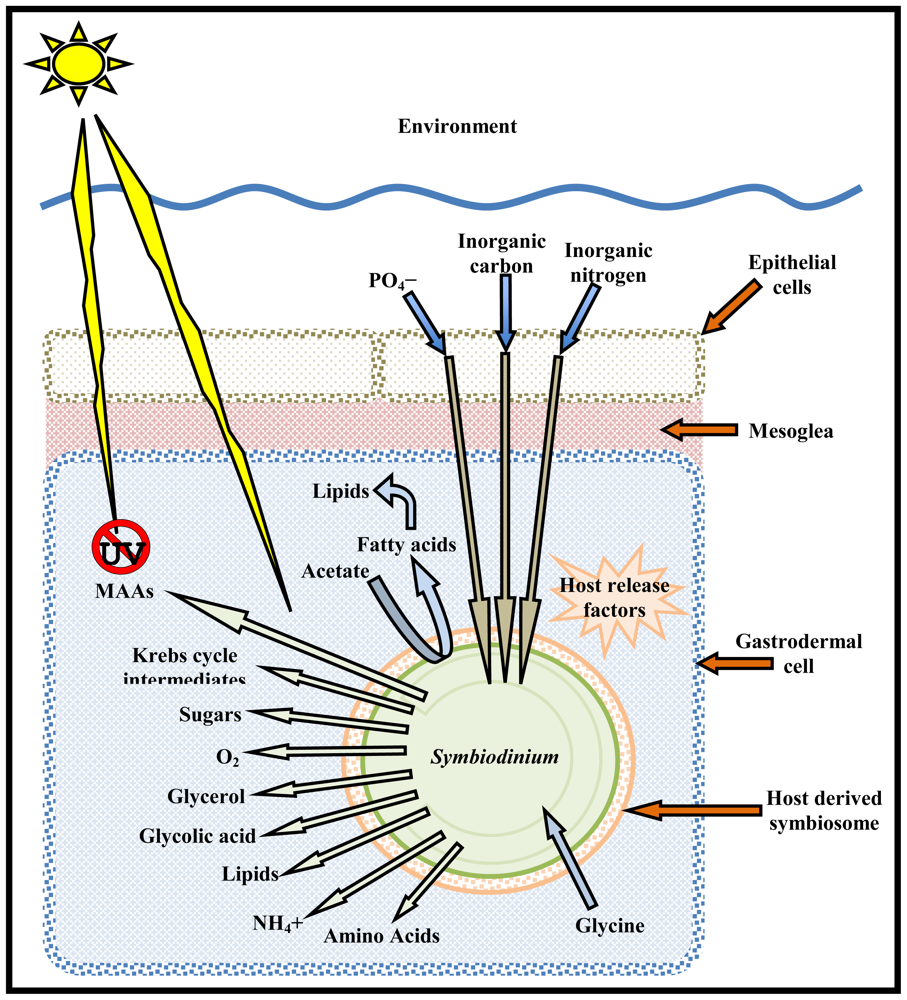
| Metabolite | Transport Direction | Symbiodinium specific | Function | Reference | |
|---|---|---|---|---|---|
| Host to symbiont | Symbiont to host | ||||
| Maltose | ✓ | Nutrition | [33] | ||
| Glycerol | ✓ | Nutrition | [34] | ||
| Glucose and other hexoses | ✓ | Nutrition | [29,35] | ||
| Glycolic acid | ✓ | Nutrition | [29] | ||
| Glycine | ✓ | Nucleotide synthesis and nutrition | [36] | ||
| Alanine | ✓ | Protein formation | [18] | ||
| Acetate | ✓ | Fatty acid synthesis | [37,38] | ||
| Fatty acids | ✓ | Lipid synthesis | [38] | ||
| Lipids | ✓ | Energy exchange | [39] | ||
| Lactate | ✓ | Metabolite | [37] | ||
| Succinate | ✓ | Krebs cycle | [37] | ||
| Citrate | ✓ | Krebs cycle | [37] | ||
| Ketoglutarate | ✓ | Krebs cycle | [37] | ||
| Malate | ✓ | Krebs cycle | [37] | ||
| Pyruvate | ✓ | Glycolysis product | [37] | ||
| PO43− | ✓ | Nutrition | [40] | ||
| NO2−, NO3−, NH4+ | ✓ | Amino acid synthesis | [40] | ||
| Inorganic carbon (i.e., CO2, HCO3−) | ✓ | Photosynthesis | [41,42] | ||
| O2 | ✓ | Photosynthesis | [43] | ||
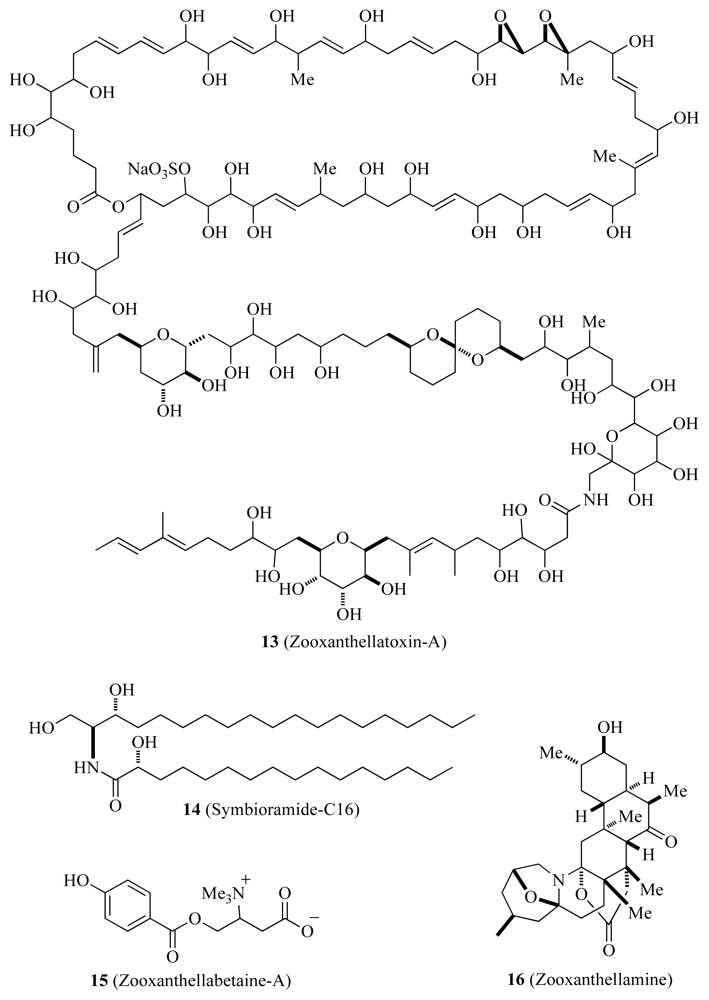
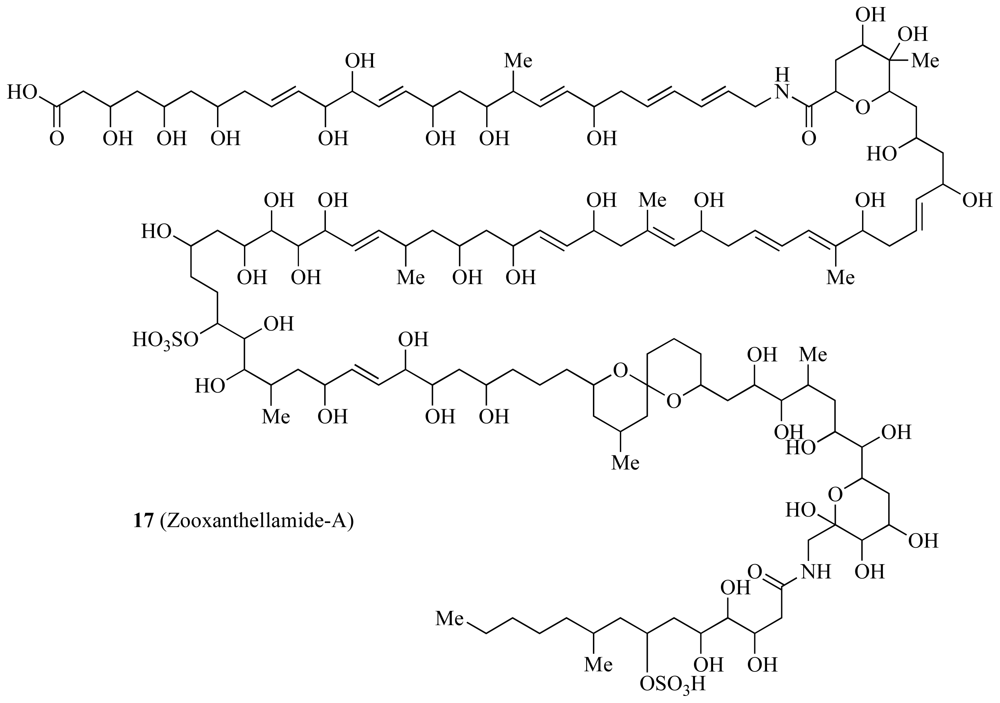
| Zooxanthellamide-A and B | ✓ | Unknown | [44,45] | ||
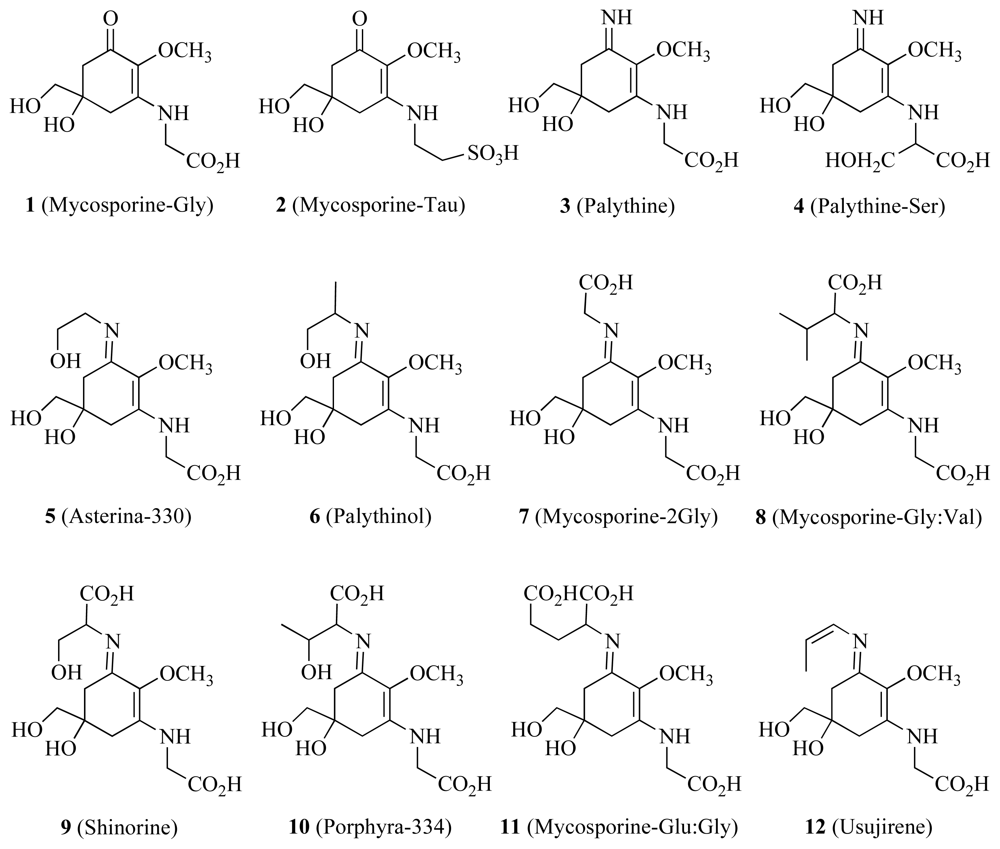
| Mycosporine-like amino acids | ✓ | UV light and free radical protection | [46] | ||
| Zooxanthellatoxins | ✓ | Unknown | [47] | ||
| Symbioramide-C16 | ✓ | Unknown | [48] | ||
| Zooxanthellabetaines | ✓ | Unknown | [48] | ||
| Zooxanthellamine | ✓ | Unknown | [48] | ||
© 2010 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Gordon, B.R.; Leggat, W. Symbiodinium—Invertebrate Symbioses and the Role of Metabolomics. Mar. Drugs 2010, 8, 2546-2568. https://doi.org/10.3390/md8102546
Gordon BR, Leggat W. Symbiodinium—Invertebrate Symbioses and the Role of Metabolomics. Marine Drugs. 2010; 8(10):2546-2568. https://doi.org/10.3390/md8102546
Chicago/Turabian StyleGordon, Benjamin R., and William Leggat. 2010. "Symbiodinium—Invertebrate Symbioses and the Role of Metabolomics" Marine Drugs 8, no. 10: 2546-2568. https://doi.org/10.3390/md8102546




